
|
|||||||
|
Fusion Protein:DDX3X-KRT13 |
Fusion Gene and Fusion Protein Summary |
 Fusion gene summary Fusion gene summary |
| Fusion partner gene information | Fusion gene name: DDX3X-KRT13 | FusionPDB ID: 21984 | FusionGDB2.0 ID: 21984 | Hgene | Tgene | Gene symbol | DDX3X | KRT13 | Gene ID | 1654 | 3860 |
| Gene name | DEAD-box helicase 3 X-linked | keratin 13 | |
| Synonyms | CAP-Rf|DBX|DDX14|DDX3|HLP2|MRX102 | CK13|K13|WSN2 | |
| Cytomap | Xp11.4 | 17q21.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | ATP-dependent RNA helicase DDX3XDEAD (Asp-Glu-Ala-Asp) box helicase 3, X-linkedDEAD (Asp-Glu-Ala-Asp) box polypeptide 3, X-linkedDEAD box protein 3, X-chromosomalDEAD box, X isoformDEAD/H (Asp-Glu-Ala-Asp/His) box polypeptide 3DEAD/H box-3helicase- | keratin, type I cytoskeletal 13CK-13cytokeratin 13keratin 13, type I | |
| Modification date | 20200329 | 20200313 | |
| UniProtAcc | O00571 Main function of 5'-partner protein: FUNCTION: Multifunctional ATP-dependent RNA helicase (PubMed:17357160, PubMed:21589879, PubMed:31575075). The ATPase activity can be stimulated by various ribo-and deoxynucleic acids indicative for a relaxed substrate specificity (PubMed:29222110). In vitro can unwind partially double-stranded DNA with a preference for 5'-single-stranded DNA overhangs (PubMed:17357160, PubMed:21589879). Binds RNA G-quadruplex (rG4s) structures, including those located in the 5'-UTR of NRAS mRNA (PubMed:30256975). Involved in many cellular processes, which do not necessarily require its ATPase/helicase catalytic activities (Probable). Involved in transcription regulation (PubMed:16818630, PubMed:18264132). Positively regulates CDKN1A/WAF1/CIP1 transcription in an SP1-dependent manner, hence inhibits cell growth. This function requires its ATPase, but not helicase activity (PubMed:16818630, PubMed:18264132). CDKN1A up-regulation may be cell-type specific (PubMed:18264132). Binds CDH1/E-cadherin promoter and represses its transcription (PubMed:18264132). Potentiates HNF4A-mediated MTTP transcriptional activation; this function requires ATPase, but not helicase activity. Facilitates HNF4A acetylation, possibly catalyzed by CREBBP/EP300, thereby increasing the DNA-binding affinity of HNF4 to its response element. In addition, disrupts the interaction between HNF4 and SHP that forms inactive heterodimers and enhances the formation of active HNF4 homodimers. By promoting HNF4A-induced MTTP expression, may play a role in lipid homeostasis (PubMed:28128295). May positively regulate TP53 transcription (PubMed:28842590). Associates with mRNPs, predominantly with spliced mRNAs carrying an exon junction complex (EJC) (PubMed:17095540, PubMed:18596238). Involved in the regulation of translation initiation (PubMed:18628297, PubMed:17667941, PubMed:22872150). Not involved in the general process of translation, but promotes efficient translation of selected complex mRNAs, containing highly structured 5'-untranslated regions (UTR) (PubMed:20837705, PubMed:22872150). This function depends on helicase activity (PubMed:20837705, PubMed:22872150). Might facilitate translation by resolving secondary structures of 5'-UTRs during ribosome scanning (PubMed:20837705). Alternatively, may act prior to 43S ribosomal scanning and promote 43S pre-initiation complex entry to mRNAs exhibiting specific RNA motifs, by performing local remodeling of transcript structures located close to the cap moiety (PubMed:22872150). Independently of its ATPase activity, promotes the assembly of functional 80S ribosomes and disassembles from ribosomes prior to the translation elongation process (PubMed:22323517). Positively regulates the translation of cyclin E1/CCNE1 mRNA and consequently promotes G1/S-phase transition during the cell cycle (PubMed:20837705). May activate TP53 translation (PubMed:28842590). Required for endoplasmic reticulum stress-induced ATF4 mRNA translation (PubMed:29062139). Independently of its ATPase/helicase activity, enhances IRES-mediated translation; this activity requires interaction with EIF4E (PubMed:17667941, PubMed:22323517). Independently of its ATPase/helicase activity, has also been shown specifically repress cap-dependent translation, possibly by acting on translation initiation factor EIF4E (PubMed:17667941). Involved in innate immunity, acting as a viral RNA sensor. Binds viral RNAs and promotes the production of type I interferon (IFN-alpha and IFN-beta) (PubMed:31575075, PubMed:20127681, PubMed:21170385). Potentiate MAVS/DDX58-mediated induction of IFNB in early stages of infection (PubMed:20127681, PubMed:21170385). Enhances IFNB1 expression via IRF3/IRF7 pathway and participates in NFKB activation in the presence of MAVS and TBK1 (PubMed:18583960, PubMed:18636090, PubMed:21170385, PubMed:27980081, PubMed:19913487). Involved in TBK1 and IKBKE-dependent IRF3 activation leading to IFNB induction, acts as a scaffolding adapter that links IKBKE and IRF3 and coordinates their activation (PubMed:23478265). Involved in the TLR7/TLR8 signaling pathway leading to type I interferon induction, including IFNA4 production. In this context, acts as an upstream regulator of IRF7 activation by MAP3K14/NIK and CHUK/IKKA. Stimulates CHUK autophosphorylation and activation following physiological activation of the TLR7 and TLR8 pathways, leading to MAP3K14/CHUK-mediated activatory phosphorylation of IRF7 (PubMed:30341167). Also stimulates MAP3K14/CHUK-dependent NF-kappa-B signaling (PubMed:30341167). Negatively regulates TNF-induced IL6 and IL8 expression, via the NF-kappa-B pathway. May act by interacting with RELA/p65 and trapping it in the cytoplasm (PubMed:27736973). May also bind IFNB promoter; the function is independent of IRF3 (PubMed:18583960). Involved in both stress and inflammatory responses (By similarity). Independently of its ATPase/helicase activity, required for efficient stress granule assembly through its interaction with EIF4E, hence promotes survival in stressed cells (PubMed:21883093). Independently of its helicase activity, regulates NLRP3 inflammasome assembly through interaction with NLRP3 and hence promotes cell death by pyroptosis during inflammation. This function is independent of helicase activity (By similarity). Therefore DDX3X availability may be used to interpret stress signals and choose between pro-survival stress granules and pyroptotic NLRP3 inflammasomes and serve as a live-or-die checkpoint in stressed cells (By similarity). In association with GSK3A/B, negatively regulates extrinsic apoptotic signaling pathway via death domain receptors, including TNFRSF10B, slowing down the rate of CASP3 activation following death receptor stimulation (PubMed:18846110). Cleavage by caspases may inactivate DDX3X and relieve the inhibition (PubMed:18846110). Independently of its ATPase/helicase activity, allosteric activator of CSNK1E. Stimulates CSNK1E-mediated phosphorylation of DVL2, thereby involved in the positive regulation of Wnt/beta-catenin signaling pathway. Also activates CSNK1A1 and CSNK1D in vitro, but it is uncertain if these targets are physiologically relevant (PubMed:23413191, PubMed:29222110). ATPase and casein kinase-activating functions are mutually exclusive (PubMed:29222110). May be involved in mitotic chromosome segregation (PubMed:21730191). {ECO:0000250|UniProtKB:Q62167, ECO:0000269|PubMed:16818630, ECO:0000269|PubMed:17095540, ECO:0000269|PubMed:17357160, ECO:0000269|PubMed:17667941, ECO:0000269|PubMed:18264132, ECO:0000269|PubMed:18583960, ECO:0000269|PubMed:18596238, ECO:0000269|PubMed:18628297, ECO:0000269|PubMed:18636090, ECO:0000269|PubMed:18846110, ECO:0000269|PubMed:19913487, ECO:0000269|PubMed:20127681, ECO:0000269|PubMed:20837705, ECO:0000269|PubMed:21170385, ECO:0000269|PubMed:21589879, ECO:0000269|PubMed:21730191, ECO:0000269|PubMed:21883093, ECO:0000269|PubMed:22323517, ECO:0000269|PubMed:22872150, ECO:0000269|PubMed:23413191, ECO:0000269|PubMed:23478265, ECO:0000269|PubMed:27736973, ECO:0000269|PubMed:27980081, ECO:0000269|PubMed:28128295, ECO:0000269|PubMed:28842590, ECO:0000269|PubMed:29062139, ECO:0000269|PubMed:29222110, ECO:0000269|PubMed:30256975, ECO:0000269|PubMed:30341167, ECO:0000269|PubMed:31575075, ECO:0000305}.; FUNCTION: (Microbial infection) Facilitates hepatitis C virus (HCV) replication (PubMed:29899501). During infection, HCV core protein inhibits the interaction between MAVS and DDX3X and therefore impairs MAVS-dependent INFB induction and might recruit DDX3X to HCV replication complex (PubMed:21170385). {ECO:0000269|PubMed:21170385, ECO:0000269|PubMed:29899501}.; FUNCTION: (Microbial infection) Facilitates HIV-1 replication (PubMed:15507209, PubMed:18583960, PubMed:21589879, PubMed:22872150, PubMed:29899501). Acts as a cofactor for XPO1-mediated nuclear export of HIV-1 Rev RNAs (PubMed:15507209, PubMed:18583960, PubMed:29899501). This function is strongly stimulated in the presence of TBK1 and requires DDX3X ATPase activity (PubMed:18583960). {ECO:0000269|PubMed:15507209, ECO:0000269|PubMed:18583960, ECO:0000269|PubMed:21589879, ECO:0000269|PubMed:22872150, ECO:0000269|PubMed:29899501}.; FUNCTION: (Microbial infection) Facilitates Zika virus (ZIKV) replication. {ECO:0000269|PubMed:29899501}.; FUNCTION: (Microbial infection) Facilitates Dengue virus (DENV) replication. {ECO:0000269|PubMed:29899501}.; FUNCTION: (Microbial infection) Facilitates Venezuelan equine encephalitis virus (VEEV) replication. {ECO:0000269|PubMed:27105836}. | P13646 Main function of 5'-partner protein: | |
| Ensembl transtripts involved in fusion gene | ENST ids | ENST00000478993, ENST00000399959, ENST00000441189, ENST00000457138, ENST00000542215, | ENST00000246635, ENST00000336861, ENST00000587544, ENST00000587118, |
| Fusion gene scores for assessment (based on all fusion genes of FusionGDB 2.0) | * DoF score | 8 X 9 X 6=432 | 19 X 33 X 6=3762 |
| # samples | 8 | 24 | |
| ** MAII score | log2(8/432*10)=-2.43295940727611 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(24/3762*10)=-3.97039353791468 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Fusion gene context | PubMed: DDX3X [Title/Abstract] AND KRT13 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Fusion neoantigen context | PubMed: DDX3X [Title/Abstract] AND KRT13 [Title/Abstract] AND neoantigen [Title/Abstract] | ||
| Most frequent breakpoint (based on all fusion genes of FusionGDB 2.0) | DDX3X(41206905)-KRT13(39661579), # samples:1 DDX3X(41206907)-KRT13(39661625), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | DDX3X-KRT13 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. DDX3X-KRT13 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. DDX3X-KRT13 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. DDX3X-KRT13 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. DDX3X-KRT13 seems lost the major protein functional domain in Hgene partner, which is a CGC due to the frame-shifted ORF. DDX3X-KRT13 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. DDX3X-KRT13 seems lost the major protein functional domain in Hgene partner, which is a tumor suppressor due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | DDX3X | GO:0009615 | response to virus | 18636090 |
| Hgene | DDX3X | GO:0010501 | RNA secondary structure unwinding | 22872150 |
| Hgene | DDX3X | GO:0010628 | positive regulation of gene expression | 10074132 |
| Hgene | DDX3X | GO:0030308 | negative regulation of cell growth | 16818630 |
| Hgene | DDX3X | GO:0031333 | negative regulation of protein complex assembly | 17667941 |
| Hgene | DDX3X | GO:0031954 | positive regulation of protein autophosphorylation | 30341167 |
| Hgene | DDX3X | GO:0032727 | positive regulation of interferon-alpha production | 30341167 |
| Hgene | DDX3X | GO:0032728 | positive regulation of interferon-beta production | 27980081 |
| Hgene | DDX3X | GO:0034063 | stress granule assembly | 21883093 |
| Hgene | DDX3X | GO:0034157 | positive regulation of toll-like receptor 7 signaling pathway | 30341167 |
| Hgene | DDX3X | GO:0034161 | positive regulation of toll-like receptor 8 signaling pathway | 30341167 |
| Hgene | DDX3X | GO:0035556 | intracellular signal transduction | 18636090 |
| Hgene | DDX3X | GO:0045727 | positive regulation of translation | 18596238|22872150 |
| Hgene | DDX3X | GO:0045944 | positive regulation of transcription by RNA polymerase II | 16818630|18636090|28128295 |
| Hgene | DDX3X | GO:0071243 | cellular response to arsenic-containing substance | 21883093 |
| Hgene | DDX3X | GO:0071470 | cellular response to osmotic stress | 21883093 |
| Hgene | DDX3X | GO:0071902 | positive regulation of protein serine/threonine kinase activity | 23413191 |
| Hgene | DDX3X | GO:1902523 | positive regulation of protein K63-linked ubiquitination | 27980081 |
| Tgene | KRT13 | GO:0007010 | cytoskeleton organization | 21371075 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chrX:41206905/chr17:39661579) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
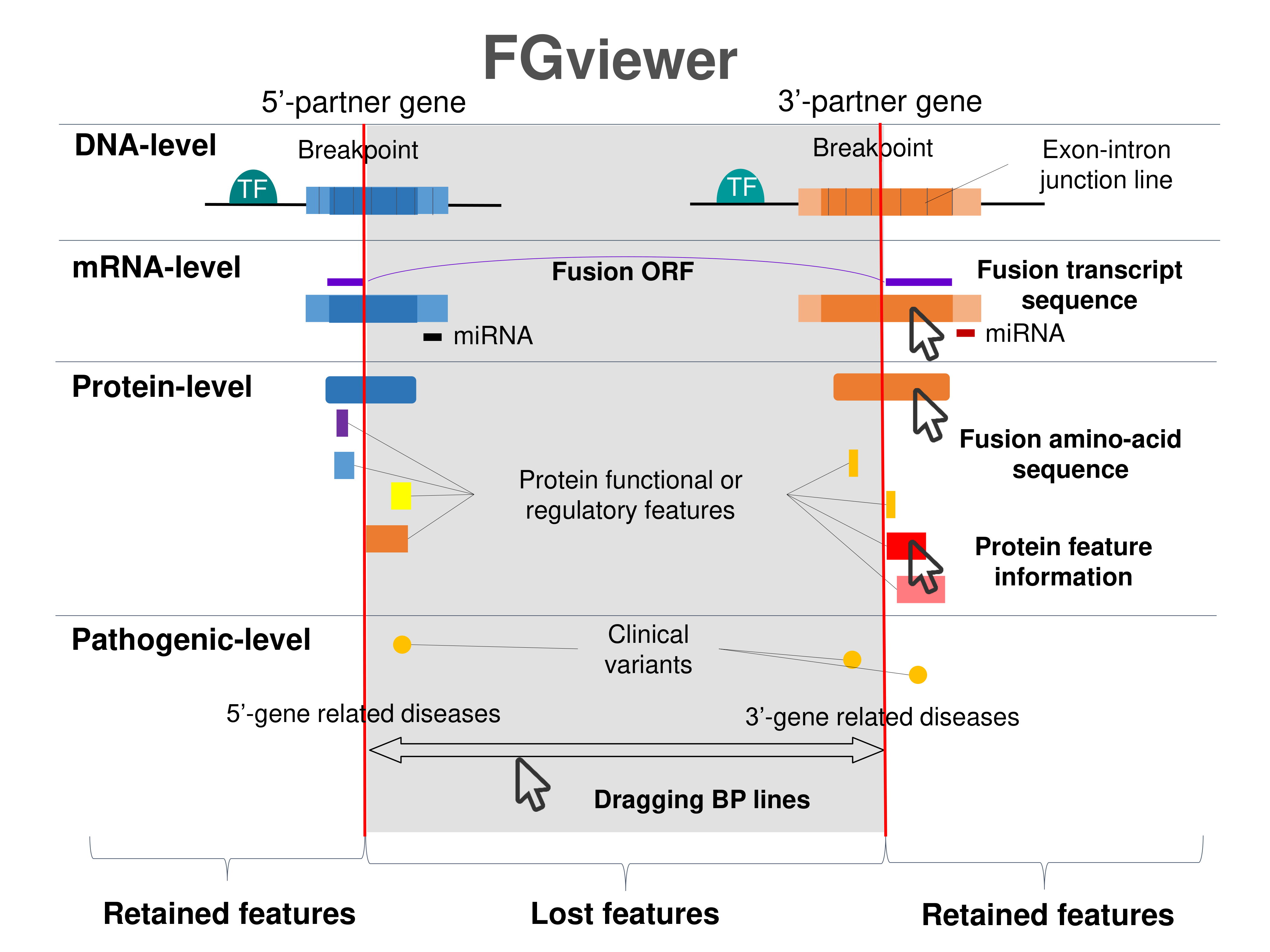 |
 Retention analysis results of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features, are available here. Retention analysis results of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features, are available here. |
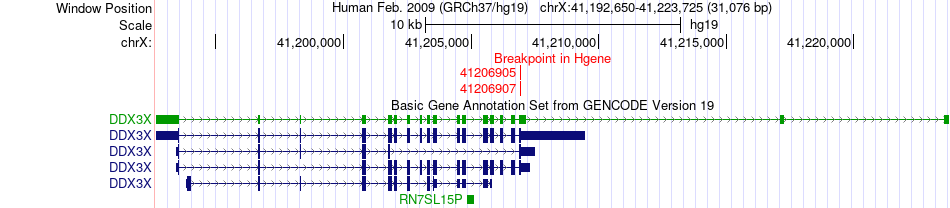
 Fusion gene breakpoints across DDX3X (5'-gene) Fusion gene breakpoints across DDX3X (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
 Fusion gene breakpoints across KRT13 (3'-gene) Fusion gene breakpoints across KRT13 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
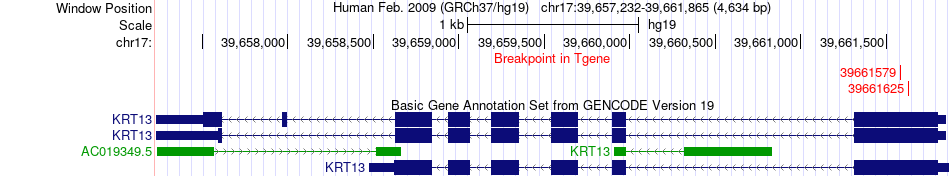 |
Top |
Fusion Amino Acid Sequences |
 Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000399959 | DDX3X | chrX | 41206905 | + | ENST00000336861 | KRT13 | chr17 | 39661579 | - | 6724 | 5321 | 855 | 2846 | 663 |
| ENST00000399959 | DDX3X | chrX | 41206905 | + | ENST00000246635 | KRT13 | chr17 | 39661579 | - | 6750 | 5321 | 855 | 2846 | 663 |
| ENST00000399959 | DDX3X | chrX | 41206905 | + | ENST00000587544 | KRT13 | chr17 | 39661579 | - | 6491 | 5321 | 855 | 2846 | 663 |
| ENST00000441189 | DDX3X | chrX | 41206905 | + | ENST00000336861 | KRT13 | chr17 | 39661579 | - | 2441 | 1038 | 91 | 513 | 140 |
| ENST00000441189 | DDX3X | chrX | 41206905 | + | ENST00000246635 | KRT13 | chr17 | 39661579 | - | 2467 | 1038 | 91 | 513 | 140 |
| ENST00000441189 | DDX3X | chrX | 41206905 | + | ENST00000587544 | KRT13 | chr17 | 39661579 | - | 2208 | 1038 | 91 | 513 | 140 |
| ENST00000399959 | DDX3X | chrX | 41206907 | + | ENST00000336861 | KRT13 | chr17 | 39661625 | - | 6768 | 5319 | 855 | 2828 | 657 |
| ENST00000399959 | DDX3X | chrX | 41206907 | + | ENST00000246635 | KRT13 | chr17 | 39661625 | - | 6794 | 5319 | 855 | 2828 | 657 |
| ENST00000399959 | DDX3X | chrX | 41206907 | + | ENST00000587544 | KRT13 | chr17 | 39661625 | - | 6535 | 5319 | 855 | 2828 | 657 |
| ENST00000441189 | DDX3X | chrX | 41206907 | + | ENST00000336861 | KRT13 | chr17 | 39661625 | - | 2485 | 1036 | 1901 | 2392 | 163 |
| ENST00000441189 | DDX3X | chrX | 41206907 | + | ENST00000246635 | KRT13 | chr17 | 39661625 | - | 2511 | 1036 | 1927 | 2418 | 163 |
| ENST00000441189 | DDX3X | chrX | 41206907 | + | ENST00000587544 | KRT13 | chr17 | 39661625 | - | 2252 | 1036 | 1668 | 2159 | 163 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000399959 | ENST00000336861 | DDX3X | chrX | 41206905 | + | KRT13 | chr17 | 39661579 | - | 0.000884035 | 0.999116 |
| ENST00000399959 | ENST00000246635 | DDX3X | chrX | 41206905 | + | KRT13 | chr17 | 39661579 | - | 0.000922997 | 0.99907696 |
| ENST00000399959 | ENST00000587544 | DDX3X | chrX | 41206905 | + | KRT13 | chr17 | 39661579 | - | 0.00091004 | 0.99908996 |
| ENST00000441189 | ENST00000336861 | DDX3X | chrX | 41206905 | + | KRT13 | chr17 | 39661579 | - | 0.009790508 | 0.99020946 |
| ENST00000441189 | ENST00000246635 | DDX3X | chrX | 41206905 | + | KRT13 | chr17 | 39661579 | - | 0.015351403 | 0.9846485 |
| ENST00000441189 | ENST00000587544 | DDX3X | chrX | 41206905 | + | KRT13 | chr17 | 39661579 | - | 0.008597597 | 0.99140245 |
| ENST00000399959 | ENST00000336861 | DDX3X | chrX | 41206907 | + | KRT13 | chr17 | 39661625 | - | 0.000933795 | 0.9990662 |
| ENST00000399959 | ENST00000246635 | DDX3X | chrX | 41206907 | + | KRT13 | chr17 | 39661625 | - | 0.000976246 | 0.9990238 |
| ENST00000399959 | ENST00000587544 | DDX3X | chrX | 41206907 | + | KRT13 | chr17 | 39661625 | - | 0.000971992 | 0.99902797 |
| ENST00000441189 | ENST00000336861 | DDX3X | chrX | 41206907 | + | KRT13 | chr17 | 39661625 | - | 0.00835389 | 0.9916461 |
| ENST00000441189 | ENST00000246635 | DDX3X | chrX | 41206907 | + | KRT13 | chr17 | 39661625 | - | 0.009178149 | 0.99082184 |
| ENST00000441189 | ENST00000587544 | DDX3X | chrX | 41206907 | + | KRT13 | chr17 | 39661625 | - | 0.007682975 | 0.9923171 |
 Predicted full-length fusion amino acid sequences. For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among all the predicted ones. Predicted full-length fusion amino acid sequences. For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among all the predicted ones. |
Get the fusion protein sequences from here. |
| Fusion protein sequence information is available in the fasta format. >FusionGDB ID_FusionGDB isoform ID_FGname_Hgene_Hchr_Hbp_Henst_Tgene_Tchr_Tbp_Tenst_length(fusion AA) seq_BP |
Top |
Fusion Protein Breakpoint Sequences for DDX3X-KRT13 |
 +/-13 AA sequence from the breakpoints of the fusion protein sequences. +/-13 AA sequence from the breakpoints of the fusion protein sequences. |
| Hgene | Hchr | Hbp | Tgene | Tchr | Tbp | Length(fusion protein) | BP in fusion protein | Peptide |
Top |
Potential FusionNeoAntigen Information of DDX3X-KRT13 in HLA I |
 Multiple sequence alignments of the potential FusionNeoAntigens per fusion breakpoints. If the MSA is empty, then it means that there were predicted fusion neoantigens in this fusion breakpoint, but those predicted fusion neoantigens were not across the breakpoint, which is not fusion-specific. Multiple sequence alignments of the potential FusionNeoAntigens per fusion breakpoints. If the MSA is empty, then it means that there were predicted fusion neoantigens in this fusion breakpoint, but those predicted fusion neoantigens were not across the breakpoint, which is not fusion-specific. |
 Potential FusionNeoAntigen Information Potential FusionNeoAntigen Information* We used NetMHCpan v4.1 (%rank<0.5) and deepHLApan v1.1 (immunogenic score>0.5) |
| Fusion gene | Hchr | Hbp | Tgene | Tchr | Tbp | HLA I | FusionNeoAntigen peptide | Binding score | Immunogenic score | Neoantigen start (at BP 13) | Neoantigen end (at BP 13) |
Top |
Potential FusionNeoAntigen Information of DDX3X-KRT13 in HLA II |
 Multiple sequence alignments of the potential FusionNeoAntigens per fusion breakpoints. If the MSA is empty, then it means that there were predicted fusion neoantigens in this fusion breakpoint, but those predicted fusion neoantigens were not across the breakpoint, which is not fusion-specific. Multiple sequence alignments of the potential FusionNeoAntigens per fusion breakpoints. If the MSA is empty, then it means that there were predicted fusion neoantigens in this fusion breakpoint, but those predicted fusion neoantigens were not across the breakpoint, which is not fusion-specific. |
 Potential FusionNeoAntigen Information Potential FusionNeoAntigen Information * We used NetMHCIIpan v4.1 (%rank<0.5). |
| Fusion gene | Hchr | Hbp | Tgene | Tchr | Tbp | HLA II | FusionNeoAntigen peptide | Neoantigen start (at BP 13) | Neoantigen end (at BP 13) |
Top |
Fusion breakpoint peptide structures of DDX3X-KRT13 |
 3D structures of the fusion breakpoint peptide of 14AA sequence that have potential fusion neoantigens 3D structures of the fusion breakpoint peptide of 14AA sequence that have potential fusion neoantigens* The minimum length of the amino acid sequence in RoseTTAFold is 14AA. Here, we predicted the 14AA fusion protein breakpoint sequence not the fusion neoantigen peptide, which is shorter than 14 AA. |
Top |
Filtering FusionNeoAntigens Through Checking the Interaction with HLAs in 3D of DDX3X-KRT13 |
 Virtual screening between 25 HLAs (from PDB) and FusionNeoAntigens Virtual screening between 25 HLAs (from PDB) and FusionNeoAntigens* We used Glide to predict the interaction between HLAs and neoantigens. |
| HLA allele | PDB ID | File name | BPseq | Docking score | Glide score |
Top |
Vaccine Design for the FusionNeoAntigens of DDX3X-KRT13 |
 mRNA and peptide sequences of FusionNeoAntigens that have potential interaction with HLA-Is. mRNA and peptide sequences of FusionNeoAntigens that have potential interaction with HLA-Is. |
| Fusion gene | Hchr | Hbp | Tchr | Tbp | Start in +/-13AA | End in +/-13AA | FusionNeoAntigen peptide sequence | FusionNeoAntigen RNA sequence |
 mRNA and peptide sequences of FusionNeoAntigens that have potential interaction with HLA-IIs. mRNA and peptide sequences of FusionNeoAntigens that have potential interaction with HLA-IIs. |
| Fusion gene | Hchr | Hbp | Tchr | Tbp | Start in +/-13AA | End in +/-13AA | FusionNeoAntigen peptide | FusionNEoAntigen RNA sequence |
Top |
Information of the samples that have these potential fusion neoantigens of DDX3X-KRT13 |
 These samples were reported as having these fusion breakpoints. For individual breakpoints, we checked the open reading frames considering multiple gene isoforms and chose the in-frame fusion genes only. Then, we made fusion protein sequences and predicted the fusion neoantigens. These fusion-positive samples may have these potential fusion neoantigens. These samples were reported as having these fusion breakpoints. For individual breakpoints, we checked the open reading frames considering multiple gene isoforms and chose the in-frame fusion genes only. Then, we made fusion protein sequences and predicted the fusion neoantigens. These fusion-positive samples may have these potential fusion neoantigens. |
| Cancer type | Fusion gene | Hchr | Hbp | Henst | Tchr | Tbp | Tenst | Sample |
Top |
Potential target of CAR-T therapy development for DDX3X-KRT13 |
 Predicted 3D structure. We used RoseTTAFold. Predicted 3D structure. We used RoseTTAFold. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, to provide the retention of the transmembrane domain, we only show the protein feature retention information of those transmembrane features Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, to provide the retention of the transmembrane domain, we only show the protein feature retention information of those transmembrane features* Minus value of BPloci means that the break point is located before the CDS. |
| - In-frame and retained 'Transmembrane'. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
 Subcellular localization prediction of the transmembrane domain retained fusion proteins Subcellular localization prediction of the transmembrane domain retained fusion proteins* We used DeepLoc 1.0. The order of the X-axis of the barplot is as follows: Entry_ID, Localization, Type, Nucleus, Cytoplasm, Extracellular, Mitochondrion, Cell_membrane, Endoplasmic_reticulum, Plastid, Golgi.apparatus, Lysosome.Vacuole, Peroxisome. Y-axis is the output score of DeepLoc. Clicking the image will open a new tab with a large image. |
| Hgene | Hchr | Hbp | Henst | Tgene | Tchr | Tbp | Tenst | DeepLoc result |
Top |
Related Drugs to DDX3X-KRT13 |
 Drugs used for this fusion-positive patient. Drugs used for this fusion-positive patient. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Drug | Source | PMID |
Top |
Related Diseases to DDX3X-KRT13 |
 Diseases that have this fusion gene. Diseases that have this fusion gene. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Disease | Source | PMID |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |

