
|
|||||||
|
Fusion Protein:TET1-CCAR1 |
Fusion Gene and Fusion Protein Summary |
 Fusion gene summary Fusion gene summary |
| Fusion partner gene information | Fusion gene name: TET1-CCAR1 | FusionPDB ID: 90156 | FusionGDB2.0 ID: 90156 | Hgene | Tgene | Gene symbol | TET1 | CCAR1 | Gene ID | 80312 | 55749 |
| Gene name | tet methylcytosine dioxygenase 1 | cell division cycle and apoptosis regulator 1 | |
| Synonyms | CXXC6|LCX|bA119F7.1 | - | |
| Cytomap | 10q21.3 | 10q21.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | methylcytosine dioxygenase TET1CXXC finger 6CXXC zinc finger 6CXXC-type zinc finger protein 6TET1 splice variant VP_DE4TET1 splice variant VP_DE456leukemia-associated protein with a CXXC domainten-eleven translocation 1 gene proteinten-eleven tran | cell division cycle and apoptosis regulator protein 1cell cycle and apoptosis regulatory protein 1death inducer with SAP domain | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | Q8NFU7 Main function of 5'-partner protein: FUNCTION: Dioxygenase that catalyzes the conversion of the modified genomic base 5-methylcytosine (5mC) into 5-hydroxymethylcytosine (5hmC) and plays a key role in active DNA demethylation (PubMed:19372391, PubMed:21496894). Also mediates subsequent conversion of 5hmC into 5-formylcytosine (5fC), and conversion of 5fC to 5-carboxylcytosine (5caC) (PubMed:21778364). In addition to its role in DNA demethylation, plays a more general role in chromatin regulation by recruiting histone modifying protein complexes to alter histone marks and chromatin accessibility, leading to both activation and repression of gene expression (PubMed:33833093). Plays therefore a role in many biological processes and diseases, including stem cell maintenance, T and B-cell development, inflammation regulation, genomic imprinting, neural activity or DNA repair (PubMed:31278917). Involved in the balance between pluripotency and lineage commitment of cells it plays a role in embryonic stem cells maintenance and inner cell mass cell specification. Plays an important role in the tumorigenicity of glioblastoma cells. TET1-mediated production of 5hmC acts as a recruitment signal for the CHTOP-methylosome complex to selective sites on the chromosome, where it methylates H4R3 and activates the transcription of genes involved in glioblastomagenesis (PubMed:25284789). Binds preferentially to DNA containing cytidine-phosphate-guanosine (CpG) dinucleotides over CpH (H=A, T, and C), hemimethylated-CpG and hemimethylated-hydroxymethyl-CpG (PubMed:29276034). Plays an essential role in the protection and maintenance of transcriptional and developmental programs together with QSER1 to inhibit the binding of DNMT3A/3B and therefore de novo methylation (PubMed:33833093). {ECO:0000269|PubMed:12124344, ECO:0000269|PubMed:19372391, ECO:0000269|PubMed:19372393, ECO:0000269|PubMed:21496894, ECO:0000269|PubMed:21778364, ECO:0000269|PubMed:25284789, ECO:0000269|PubMed:29276034, ECO:0000269|PubMed:31278917, ECO:0000269|PubMed:33833093}. | Q8IX12 Main function of 5'-partner protein: FUNCTION: Associates with components of the Mediator and p160 coactivator complexes that play a role as intermediaries transducing regulatory signals from upstream transcriptional activator proteins to basal transcription machinery at the core promoter. Recruited to endogenous nuclear receptor target genes in response to the appropriate hormone. Also functions as a p53 coactivator. May thus play an important role in transcriptional regulation (By similarity). May be involved in apoptosis signaling in the presence of the reinoid CD437. Apoptosis induction involves sequestration of 14-3-3 protein(s) and mediated altered expression of multiple cell cycle regulatory genes including MYC, CCNB1 and CDKN1A. Plays a role in cell cycle progression and/or cell proliferation (PubMed:12816952). In association with CALCOCO1 enhances GATA1- and MED1-mediated transcriptional activation from the gamma-globin promoter during erythroid differentiation of K562 erythroleukemia cells (PubMed:24245781). Can act as a both a coactivator and corepressor of AR-mediated transcription. Contributes to chromatin looping and AR transcription complex assembly by stabilizing AR-GATA2 association on chromatin and facilitating MED1 and RNA polymerase II recruitment to AR-binding sites. May play an important role in the growth and tumorigenesis of prostate cancer cells (PubMed:23887938). {ECO:0000250|UniProtKB:Q8CH18, ECO:0000269|PubMed:12816952, ECO:0000269|PubMed:23887938, ECO:0000269|PubMed:24245781}. | |
| Ensembl transtripts involved in fusion gene | ENST ids | ENST00000373644, | ENST00000483264, ENST00000265872, ENST00000535016, ENST00000543719, |
| Fusion gene scores for assessment (based on all fusion genes of FusionGDB 2.0) | * DoF score | 10 X 14 X 7=980 | 5 X 6 X 3=90 |
| # samples | 14 | 6 | |
| ** MAII score | log2(14/980*10)=-2.8073549220576 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(6/90*10)=-0.584962500721156 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Fusion gene context | PubMed: TET1 [Title/Abstract] AND CCAR1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Fusion neoantigen context | PubMed: TET1 [Title/Abstract] AND CCAR1 [Title/Abstract] AND neoantigen [Title/Abstract] | ||
| Most frequent breakpoint (based on all fusion genes of FusionGDB 2.0) | CCAR1(70520949)-TET1(70432652), # samples:2 CCAR1(70517134)-TET1(70432652), # samples:2 TET1(70446464)-CCAR1(70549467), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | TET1-CCAR1 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. TET1-CCAR1 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. TET1-CCAR1 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. TET1-CCAR1 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. CCAR1-TET1 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. CCAR1-TET1 seems lost the major protein functional domain in Hgene partner, which is a tumor suppressor due to the frame-shifted ORF. CCAR1-TET1 seems lost the major protein functional domain in Tgene partner, which is a CGC due to the frame-shifted ORF. CCAR1-TET1 seems lost the major protein functional domain in Tgene partner, which is a epigenetic factor due to the frame-shifted ORF. CCAR1-TET1 seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. CCAR1-TET1 seems lost the major protein functional domain in Tgene partner, which is a transcription factor due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | CCAR1 | GO:0043065 | positive regulation of apoptotic process | 12816952 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr10:70520949/chr10:70432652) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
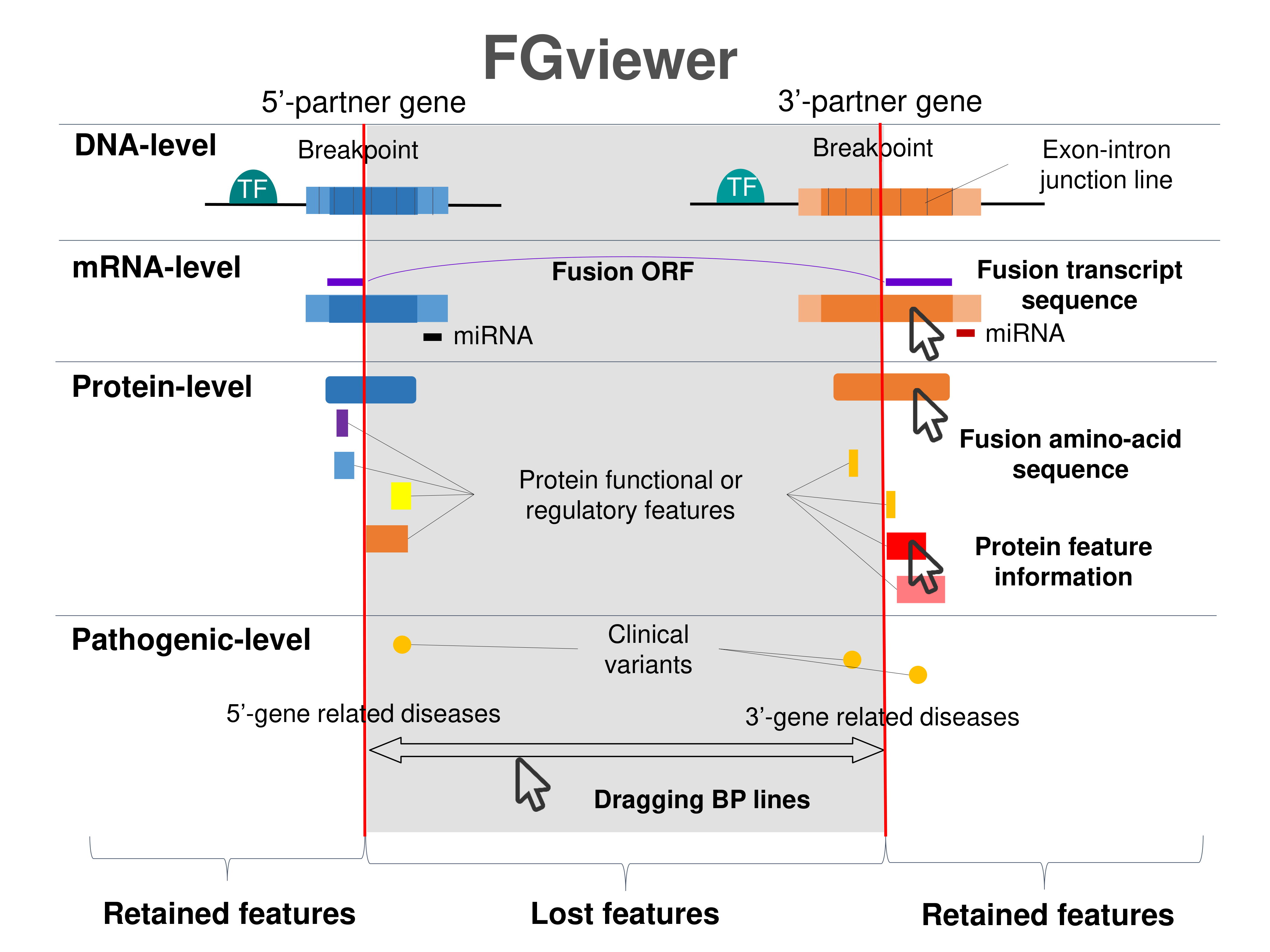 |
 Retention analysis results of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features, are available here. Retention analysis results of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features, are available here. |
 Fusion gene breakpoints across TET1 (5'-gene) Fusion gene breakpoints across TET1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
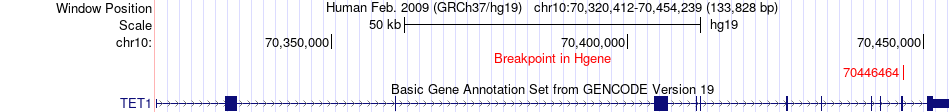 |
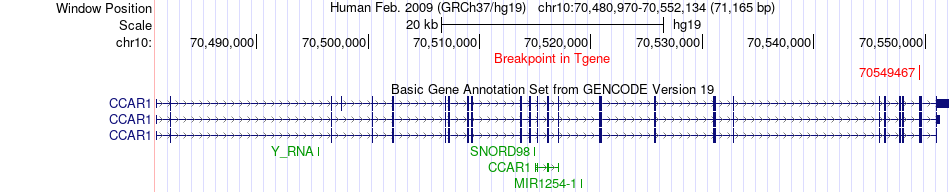
 Fusion gene breakpoints across CCAR1 (3'-gene) Fusion gene breakpoints across CCAR1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
Top |
Fusion Amino Acid Sequences |
 Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000373644 | TET1 | chr10 | 70446464 | + | ENST00000535016 | CCAR1 | chr10 | 70549467 | + | 6165 | 5613 | 176 | 5878 | 1900 |
| ENST00000373644 | TET1 | chr10 | 70446464 | + | ENST00000265872 | CCAR1 | chr10 | 70549467 | + | 6990 | 5613 | 176 | 5878 | 1900 |
| ENST00000373644 | TET1 | chr10 | 70446464 | + | ENST00000543719 | CCAR1 | chr10 | 70549467 | + | 5879 | 5613 | 176 | 5878 | 1901 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000373644 | ENST00000535016 | TET1 | chr10 | 70446464 | + | CCAR1 | chr10 | 70549467 | + | 0.000253077 | 0.99974686 |
| ENST00000373644 | ENST00000265872 | TET1 | chr10 | 70446464 | + | CCAR1 | chr10 | 70549467 | + | 0.000200823 | 0.9997992 |
| ENST00000373644 | ENST00000543719 | TET1 | chr10 | 70446464 | + | CCAR1 | chr10 | 70549467 | + | 0.000308784 | 0.99969125 |
 Predicted full-length fusion amino acid sequences. For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among all the predicted ones. Predicted full-length fusion amino acid sequences. For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among all the predicted ones. |
Get the fusion protein sequences from here. |
| Fusion protein sequence information is available in the fasta format. >FusionGDB ID_FusionGDB isoform ID_FGname_Hgene_Hchr_Hbp_Henst_Tgene_Tchr_Tbp_Tenst_length(fusion AA) seq_BP |
Top |
Fusion Protein Breakpoint Sequences for TET1-CCAR1 |
 +/-13 AA sequence from the breakpoints of the fusion protein sequences. +/-13 AA sequence from the breakpoints of the fusion protein sequences. |
| Hgene | Hchr | Hbp | Tgene | Tchr | Tbp | Length(fusion protein) | BP in fusion protein | Peptide |
Top |
Potential FusionNeoAntigen Information of TET1-CCAR1 in HLA I |
 Multiple sequence alignments of the potential FusionNeoAntigens per fusion breakpoints. If the MSA is empty, then it means that there were predicted fusion neoantigens in this fusion breakpoint, but those predicted fusion neoantigens were not across the breakpoint, which is not fusion-specific. Multiple sequence alignments of the potential FusionNeoAntigens per fusion breakpoints. If the MSA is empty, then it means that there were predicted fusion neoantigens in this fusion breakpoint, but those predicted fusion neoantigens were not across the breakpoint, which is not fusion-specific. |
 Potential FusionNeoAntigen Information Potential FusionNeoAntigen Information* We used NetMHCpan v4.1 (%rank<0.5) and deepHLApan v1.1 (immunogenic score>0.5) |
| Fusion gene | Hchr | Hbp | Tgene | Tchr | Tbp | HLA I | FusionNeoAntigen peptide | Binding score | Immunogenic score | Neoantigen start (at BP 13) | Neoantigen end (at BP 13) |
Top |
Potential FusionNeoAntigen Information of TET1-CCAR1 in HLA II |
 Multiple sequence alignments of the potential FusionNeoAntigens per fusion breakpoints. If the MSA is empty, then it means that there were predicted fusion neoantigens in this fusion breakpoint, but those predicted fusion neoantigens were not across the breakpoint, which is not fusion-specific. Multiple sequence alignments of the potential FusionNeoAntigens per fusion breakpoints. If the MSA is empty, then it means that there were predicted fusion neoantigens in this fusion breakpoint, but those predicted fusion neoantigens were not across the breakpoint, which is not fusion-specific. |
 Potential FusionNeoAntigen Information Potential FusionNeoAntigen Information * We used NetMHCIIpan v4.1 (%rank<0.5). |
| Fusion gene | Hchr | Hbp | Tgene | Tchr | Tbp | HLA II | FusionNeoAntigen peptide | Neoantigen start (at BP 13) | Neoantigen end (at BP 13) |
Top |
Fusion breakpoint peptide structures of TET1-CCAR1 |
 3D structures of the fusion breakpoint peptide of 14AA sequence that have potential fusion neoantigens 3D structures of the fusion breakpoint peptide of 14AA sequence that have potential fusion neoantigens* The minimum length of the amino acid sequence in RoseTTAFold is 14AA. Here, we predicted the 14AA fusion protein breakpoint sequence not the fusion neoantigen peptide, which is shorter than 14 AA. |
Top |
Filtering FusionNeoAntigens Through Checking the Interaction with HLAs in 3D of TET1-CCAR1 |
 Virtual screening between 25 HLAs (from PDB) and FusionNeoAntigens Virtual screening between 25 HLAs (from PDB) and FusionNeoAntigens* We used Glide to predict the interaction between HLAs and neoantigens. |
| HLA allele | PDB ID | File name | BPseq | Docking score | Glide score |
Top |
Vaccine Design for the FusionNeoAntigens of TET1-CCAR1 |
 mRNA and peptide sequences of FusionNeoAntigens that have potential interaction with HLA-Is. mRNA and peptide sequences of FusionNeoAntigens that have potential interaction with HLA-Is. |
| Fusion gene | Hchr | Hbp | Tchr | Tbp | Start in +/-13AA | End in +/-13AA | FusionNeoAntigen peptide sequence | FusionNeoAntigen RNA sequence |
 mRNA and peptide sequences of FusionNeoAntigens that have potential interaction with HLA-IIs. mRNA and peptide sequences of FusionNeoAntigens that have potential interaction with HLA-IIs. |
| Fusion gene | Hchr | Hbp | Tchr | Tbp | Start in +/-13AA | End in +/-13AA | FusionNeoAntigen peptide | FusionNEoAntigen RNA sequence |
Top |
Information of the samples that have these potential fusion neoantigens of TET1-CCAR1 |
 These samples were reported as having these fusion breakpoints. For individual breakpoints, we checked the open reading frames considering multiple gene isoforms and chose the in-frame fusion genes only. Then, we made fusion protein sequences and predicted the fusion neoantigens. These fusion-positive samples may have these potential fusion neoantigens. These samples were reported as having these fusion breakpoints. For individual breakpoints, we checked the open reading frames considering multiple gene isoforms and chose the in-frame fusion genes only. Then, we made fusion protein sequences and predicted the fusion neoantigens. These fusion-positive samples may have these potential fusion neoantigens. |
| Cancer type | Fusion gene | Hchr | Hbp | Henst | Tchr | Tbp | Tenst | Sample |
Top |
Potential target of CAR-T therapy development for TET1-CCAR1 |
 Predicted 3D structure. We used RoseTTAFold. Predicted 3D structure. We used RoseTTAFold. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, to provide the retention of the transmembrane domain, we only show the protein feature retention information of those transmembrane features Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, to provide the retention of the transmembrane domain, we only show the protein feature retention information of those transmembrane features* Minus value of BPloci means that the break point is located before the CDS. |
| - In-frame and retained 'Transmembrane'. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
 Subcellular localization prediction of the transmembrane domain retained fusion proteins Subcellular localization prediction of the transmembrane domain retained fusion proteins* We used DeepLoc 1.0. The order of the X-axis of the barplot is as follows: Entry_ID, Localization, Type, Nucleus, Cytoplasm, Extracellular, Mitochondrion, Cell_membrane, Endoplasmic_reticulum, Plastid, Golgi.apparatus, Lysosome.Vacuole, Peroxisome. Y-axis is the output score of DeepLoc. Clicking the image will open a new tab with a large image. |
| Hgene | Hchr | Hbp | Henst | Tgene | Tchr | Tbp | Tenst | DeepLoc result |
Top |
Related Drugs to TET1-CCAR1 |
 Drugs used for this fusion-positive patient. Drugs used for this fusion-positive patient. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Drug | Source | PMID |
Top |
Related Diseases to TET1-CCAR1 |
 Diseases that have this fusion gene. Diseases that have this fusion gene. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Disease | Source | PMID |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
| Hgene | TET1 | C0033975 | Psychotic Disorders | 1 | PSYGENET |
| Hgene | TET1 | C0036341 | Schizophrenia | 1 | PSYGENET |
| Hgene | TET1 | C0349204 | Nonorganic psychosis | 1 | PSYGENET |
| Hgene | TET1 | C1510586 | Autism Spectrum Disorders | 1 | CTD_human |

