
|
|||||||
|
Fusion Protein:BCL2L11-BRAF |
Fusion Gene and Fusion Protein Summary |
 Fusion gene summary Fusion gene summary |
| Fusion partner gene information | Fusion gene name: BCL2L11-BRAF | FusionPDB ID: 9342 | FusionGDB2.0 ID: 9342 | Hgene | Tgene | Gene symbol | BCL2L11 | BRAF | Gene ID | 10018 | 673 |
| Gene name | BCL2 like 11 | B-Raf proto-oncogene, serine/threonine kinase | |
| Synonyms | BAM|BIM|BOD | B-RAF1|B-raf|BRAF1|NS7|RAFB1 | |
| Cytomap | 2q13 | 7q34 | |
| Type of gene | protein-coding | protein-coding | |
| Description | bcl-2-like protein 11BCL2-like 11 (apoptosis facilitator)bcl-2 interacting mediator of cell deathbcl-2 interacting protein Bimbcl-2-related ovarian death agonist | serine/threonine-protein kinase B-raf94 kDa B-raf proteinB-Raf proto-oncogene serine/threonine-protein kinase (p94)B-Raf serine/threonine-proteinmurine sarcoma viral (v-raf) oncogene homolog B1proto-oncogene B-Rafv-raf murine sarcoma viral oncogene | |
| Modification date | 20200313 | 20200329 | |
| UniProtAcc | O43521 Main function of 5'-partner protein: FUNCTION: Induces apoptosis and anoikis. Isoform BimL is more potent than isoform BimEL. Isoform Bim-alpha1, isoform Bim-alpha2 and isoform Bim-alpha3 induce apoptosis, although less potent than isoform BimEL, isoform BimL and isoform BimS. Isoform Bim-gamma induces apoptosis. Isoform Bim-alpha3 induces apoptosis possibly through a caspase-mediated pathway. Isoform BimAC and isoform BimABC lack the ability to induce apoptosis. {ECO:0000269|PubMed:11997495, ECO:0000269|PubMed:15486195, ECO:0000269|PubMed:9430630}. | P15056 Main function of 5'-partner protein: FUNCTION: Protein kinase involved in the transduction of mitogenic signals from the cell membrane to the nucleus (Probable). Phosphorylates MAP2K1, and thereby activates the MAP kinase signal transduction pathway (PubMed:21441910, PubMed:29433126). May play a role in the postsynaptic responses of hippocampal neurons (PubMed:1508179). {ECO:0000269|PubMed:1508179, ECO:0000269|PubMed:21441910, ECO:0000269|PubMed:29433126, ECO:0000305}. | |
| Ensembl transtripts involved in fusion gene | ENST ids | ENST00000308659, ENST00000337565, ENST00000357757, ENST00000393253, ENST00000393256, ENST00000405953, | ENST00000288602, |
| Fusion gene scores for assessment (based on all fusion genes of FusionGDB 2.0) | * DoF score | 7 X 4 X 6=168 | 48 X 58 X 16=44544 |
| # samples | 9 | 69 | |
| ** MAII score | log2(9/168*10)=-0.900464326449086 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(69/44544*10)=-6.0124909441832 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Fusion gene context | PubMed: BCL2L11 [Title/Abstract] AND BRAF [Title/Abstract] AND fusion [Title/Abstract] | ||
| Fusion neoantigen context | PubMed: BCL2L11 [Title/Abstract] AND BRAF [Title/Abstract] AND neoantigen [Title/Abstract] | ||
| Most frequent breakpoint (based on all fusion genes of FusionGDB 2.0) | BCL2L11(111881716)-BRAF(140482957), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | BCL2L11-BRAF seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. BCL2L11-BRAF seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. BCL2L11-BRAF seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. BCL2L11-BRAF seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | BCL2L11 | GO:0008630 | intrinsic apoptotic signaling pathway in response to DNA damage | 11546872 |
| Hgene | BCL2L11 | GO:0031334 | positive regulation of protein complex assembly | 21041309 |
| Hgene | BCL2L11 | GO:0034976 | response to endoplasmic reticulum stress | 22761832 |
| Hgene | BCL2L11 | GO:2000271 | positive regulation of fibroblast apoptotic process | 11997495 |
| Tgene | BRAF | GO:0000186 | activation of MAPKK activity | 29433126 |
| Tgene | BRAF | GO:0006468 | protein phosphorylation | 17563371 |
| Tgene | BRAF | GO:0010828 | positive regulation of glucose transmembrane transport | 23010278 |
| Tgene | BRAF | GO:0033138 | positive regulation of peptidyl-serine phosphorylation | 19667065 |
| Tgene | BRAF | GO:0043066 | negative regulation of apoptotic process | 19667065 |
| Tgene | BRAF | GO:0070374 | positive regulation of ERK1 and ERK2 cascade | 22065586 |
| Tgene | BRAF | GO:0071277 | cellular response to calcium ion | 18567582 |
| Tgene | BRAF | GO:0090150 | establishment of protein localization to membrane | 23010278 |
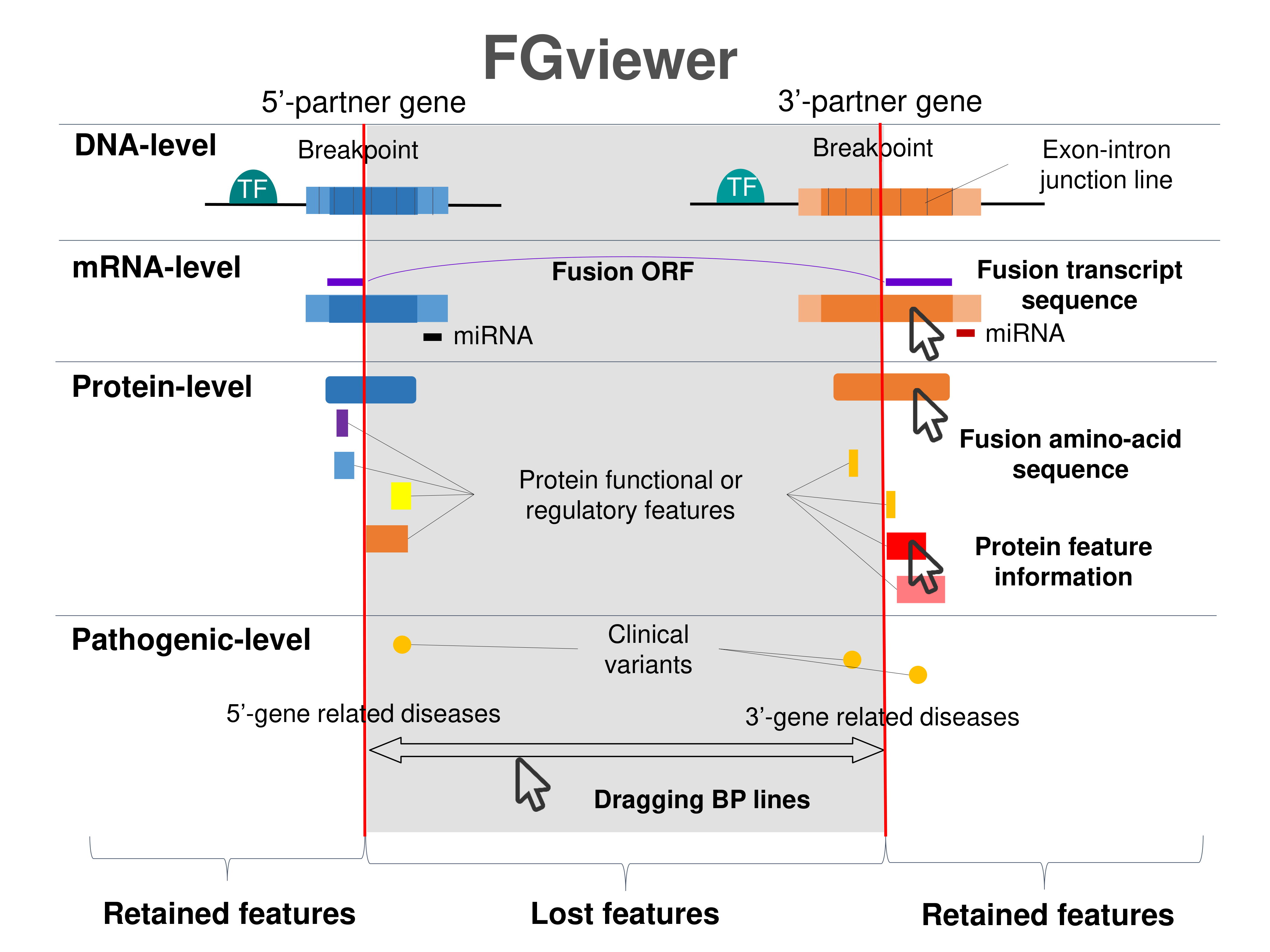
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr2:111881716/chr7:140482957) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Retention analysis results of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features, are available here. Retention analysis results of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features, are available here. |
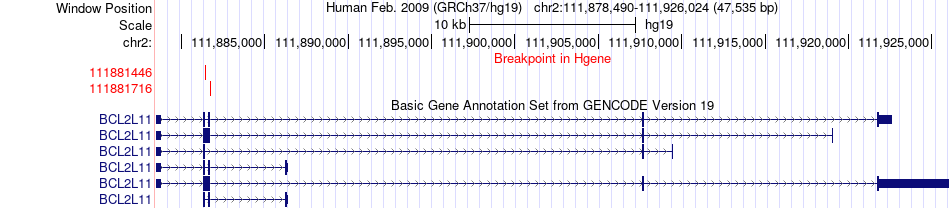
 Fusion gene breakpoints across BCL2L11 (5'-gene) Fusion gene breakpoints across BCL2L11 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
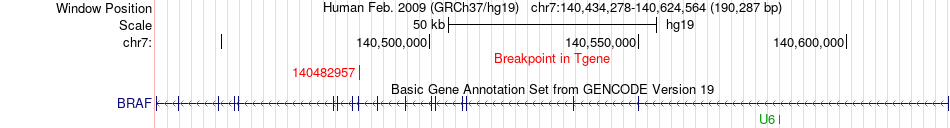
 Fusion gene breakpoints across BRAF (3'-gene) Fusion gene breakpoints across BRAF (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
Top |
Fusion Amino Acid Sequences |
 Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000337565 | BCL2L11 | chr2 | 111881716 | + | ENST00000288602 | BRAF | chr7 | 140482957 | - | 1744 | 502 | 126 | 1625 | 499 |
| ENST00000357757 | BCL2L11 | chr2 | 111881716 | + | ENST00000288602 | BRAF | chr7 | 140482957 | - | 1924 | 682 | 126 | 1805 | 559 |
| ENST00000308659 | BCL2L11 | chr2 | 111881716 | + | ENST00000288602 | BRAF | chr7 | 140482957 | - | 1744 | 502 | 126 | 1625 | 499 |
| ENST00000393256 | BCL2L11 | chr2 | 111881716 | + | ENST00000288602 | BRAF | chr7 | 140482957 | - | 1909 | 667 | 111 | 1790 | 559 |
| ENST00000405953 | BCL2L11 | chr2 | 111881716 | + | ENST00000288602 | BRAF | chr7 | 140482957 | - | 1456 | 214 | 0 | 1337 | 445 |
| ENST00000337565 | BCL2L11 | chr2 | 111881446 | + | ENST00000288602 | BRAF | chr7 | 140482957 | - | 1654 | 412 | 126 | 1535 | 469 |
| ENST00000393253 | BCL2L11 | chr2 | 111881446 | + | ENST00000288602 | BRAF | chr7 | 140482957 | - | 1654 | 412 | 126 | 1535 | 469 |
| ENST00000357757 | BCL2L11 | chr2 | 111881446 | + | ENST00000288602 | BRAF | chr7 | 140482957 | - | 1787 | 545 | 340 | 1668 | 442 |
| ENST00000308659 | BCL2L11 | chr2 | 111881446 | + | ENST00000288602 | BRAF | chr7 | 140482957 | - | 1654 | 412 | 126 | 1535 | 469 |
| ENST00000393256 | BCL2L11 | chr2 | 111881446 | + | ENST00000288602 | BRAF | chr7 | 140482957 | - | 1772 | 530 | 325 | 1653 | 442 |
| ENST00000405953 | BCL2L11 | chr2 | 111881446 | + | ENST00000288602 | BRAF | chr7 | 140482957 | - | 1366 | 124 | 0 | 1247 | 415 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000337565 | ENST00000288602 | BCL2L11 | chr2 | 111881716 | + | BRAF | chr7 | 140482957 | - | 0.013307793 | 0.98669213 |
| ENST00000357757 | ENST00000288602 | BCL2L11 | chr2 | 111881716 | + | BRAF | chr7 | 140482957 | - | 0.014716662 | 0.9852834 |
| ENST00000308659 | ENST00000288602 | BCL2L11 | chr2 | 111881716 | + | BRAF | chr7 | 140482957 | - | 0.013307793 | 0.98669213 |
| ENST00000393256 | ENST00000288602 | BCL2L11 | chr2 | 111881716 | + | BRAF | chr7 | 140482957 | - | 0.014069975 | 0.98593 |
| ENST00000405953 | ENST00000288602 | BCL2L11 | chr2 | 111881716 | + | BRAF | chr7 | 140482957 | - | 0.007845804 | 0.9921542 |
| ENST00000337565 | ENST00000288602 | BCL2L11 | chr2 | 111881446 | + | BRAF | chr7 | 140482957 | - | 0.006178714 | 0.99382126 |
| ENST00000393253 | ENST00000288602 | BCL2L11 | chr2 | 111881446 | + | BRAF | chr7 | 140482957 | - | 0.006178714 | 0.99382126 |
| ENST00000357757 | ENST00000288602 | BCL2L11 | chr2 | 111881446 | + | BRAF | chr7 | 140482957 | - | 0.003236262 | 0.9967637 |
| ENST00000308659 | ENST00000288602 | BCL2L11 | chr2 | 111881446 | + | BRAF | chr7 | 140482957 | - | 0.006178714 | 0.99382126 |
| ENST00000393256 | ENST00000288602 | BCL2L11 | chr2 | 111881446 | + | BRAF | chr7 | 140482957 | - | 0.00308201 | 0.99691796 |
| ENST00000405953 | ENST00000288602 | BCL2L11 | chr2 | 111881446 | + | BRAF | chr7 | 140482957 | - | 0.002835531 | 0.9971644 |
 Predicted full-length fusion amino acid sequences. For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among all the predicted ones. Predicted full-length fusion amino acid sequences. For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among all the predicted ones. |
Get the fusion protein sequences from here. |
| Fusion protein sequence information is available in the fasta format. >FusionGDB ID_FusionGDB isoform ID_FGname_Hgene_Hchr_Hbp_Henst_Tgene_Tchr_Tbp_Tenst_length(fusion AA) seq_BP |
Top |
Fusion Protein Breakpoint Sequences for BCL2L11-BRAF |
 +/-13 AA sequence from the breakpoints of the fusion protein sequences. +/-13 AA sequence from the breakpoints of the fusion protein sequences. |
| Hgene | Hchr | Hbp | Tgene | Tchr | Tbp | Length(fusion protein) | BP in fusion protein | Peptide |
| BCL2L11 | chr2 | 111881446 | BRAF | chr7 | 140482957 | 124 | 41 | RPGAPTSLQTEPQGSTTGLSATPPAS |
| BCL2L11 | chr2 | 111881446 | BRAF | chr7 | 140482957 | 412 | 95 | RPGAPTSLQTEPQGSTTGLSATPPAS |
| BCL2L11 | chr2 | 111881446 | BRAF | chr7 | 140482957 | 530 | 68 | PPCQAFNHYLSAMGSTTGLSATPPAS |
| BCL2L11 | chr2 | 111881446 | BRAF | chr7 | 140482957 | 545 | 68 | PPCQAFNHYLSAMGSTTGLSATPPAS |
| BCL2L11 | chr2 | 111881716 | BRAF | chr7 | 140482957 | 214 | 71 | PPCQAFNHYLSAMGSTTGLSATPPAS |
| BCL2L11 | chr2 | 111881716 | BRAF | chr7 | 140482957 | 502 | 125 | PPCQAFNHYLSAMGSTTGLSATPPAS |
| BCL2L11 | chr2 | 111881716 | BRAF | chr7 | 140482957 | 667 | 185 | PPCQAFNHYLSAMGSTTGLSATPPAS |
| BCL2L11 | chr2 | 111881716 | BRAF | chr7 | 140482957 | 682 | 185 | PPCQAFNHYLSAMGSTTGLSATPPAS |
Top |
Potential FusionNeoAntigen Information of BCL2L11-BRAF in HLA I |
 Multiple sequence alignments of the potential FusionNeoAntigens per fusion breakpoints. If the MSA is empty, then it means that there were predicted fusion neoantigens in this fusion breakpoint, but those predicted fusion neoantigens were not across the breakpoint, which is not fusion-specific. Multiple sequence alignments of the potential FusionNeoAntigens per fusion breakpoints. If the MSA is empty, then it means that there were predicted fusion neoantigens in this fusion breakpoint, but those predicted fusion neoantigens were not across the breakpoint, which is not fusion-specific. |
| BCL2L11-BRAF_111881446_140482957.msa |
 Potential FusionNeoAntigen Information Potential FusionNeoAntigen Information* We used NetMHCpan v4.1 (%rank<0.5) and deepHLApan v1.1 (immunogenic score>0.5) |
| Fusion gene | Hchr | Hbp | Tgene | Tchr | Tbp | HLA I | FusionNeoAntigen peptide | Binding score | Immunogenic score | Neoantigen start (at BP 13) | Neoantigen end (at BP 13) |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-B39:06 | NHYLSAMGS | 0.9645 | 0.7372 | 6 | 15 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-B35:03 | SAMGSTTGL | 0.9079 | 0.9303 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B35:08 | EPQGSTTGL | 0.8251 | 0.632 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-B07:10 | SAMGSTTGL | 0.8185 | 0.5316 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-B46:01 | SAMGSTTGL | 0.794 | 0.6253 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-B35:02 | SAMGSTTGL | 0.7928 | 0.9873 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-B35:04 | SAMGSTTGL | 0.7928 | 0.9873 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B35:03 | EPQGSTTGL | 0.7476 | 0.6773 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B35:02 | EPQGSTTGL | 0.5208 | 0.8178 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B35:04 | EPQGSTTGL | 0.5208 | 0.8178 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-B81:01 | SAMGSTTGL | 0.4624 | 0.7435 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B81:01 | TEPQGSTTGL | 0.8438 | 0.6105 | 9 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B82:01 | TEPQGSTTGL | 0.5728 | 0.5471 | 9 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B07:05 | TEPQGSTTGL | 0.5473 | 0.5112 | 9 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-A02:22 | YLSAMGSTTGL | 0.987 | 0.7641 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-A02:24 | YLSAMGSTTGL | 0.9573 | 0.7315 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-A02:67 | YLSAMGSTTGL | 0.9573 | 0.7315 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-A02:30 | YLSAMGSTTGL | 0.9573 | 0.7315 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-A02:13 | YLSAMGSTTGL | 0.9549 | 0.8862 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-A02:11 | YLSAMGSTTGL | 0.9506 | 0.7626 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-A02:27 | YLSAMGSTTGL | 0.9482 | 0.8051 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-A02:16 | YLSAMGSTTGL | 0.9337 | 0.6422 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-A02:29 | YLSAMGSTTGL | 0.9059 | 0.7351 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B07:05 | QTEPQGSTTGL | 0.8967 | 0.527 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-A02:38 | YLSAMGSTTGL | 0.8957 | 0.8545 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-A02:17 | YLSAMGSTTGL | 0.8842 | 0.8179 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-A02:19 | YLSAMGSTTGL | 0.88 | 0.7204 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B82:01 | QTEPQGSTTGL | 0.7204 | 0.7437 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C03:19 | SAMGSTTGL | 0.9991 | 0.991 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C03:07 | SAMGSTTGL | 0.9991 | 0.967 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C03:08 | SAMGSTTGL | 0.9987 | 0.8622 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C15:06 | SAMGSTTGL | 0.9982 | 0.974 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C04:06 | SAMGSTTGL | 0.9916 | 0.9788 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C06:03 | SAMGSTTGL | 0.9834 | 0.9972 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C12:04 | SAMGSTTGL | 0.9826 | 0.9977 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C12:12 | SAMGSTTGL | 0.9812 | 0.9631 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C08:13 | SAMGSTTGL | 0.9808 | 0.9863 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C08:04 | SAMGSTTGL | 0.9808 | 0.9863 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C02:06 | SAMGSTTGL | 0.9409 | 0.9834 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C08:03 | SAMGSTTGL | 0.9182 | 0.9941 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C03:14 | SAMGSTTGL | 0.8188 | 0.9914 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-B35:12 | SAMGSTTGL | 0.7928 | 0.9873 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-B07:12 | SAMGSTTGL | 0.7463 | 0.6687 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C01:30 | SAMGSTTGL | 0.5607 | 0.9816 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C01:17 | SAMGSTTGL | 0.5472 | 0.9806 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B35:12 | EPQGSTTGL | 0.5208 | 0.8178 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B39:10 | EPQGSTTGL | 0.3113 | 0.8483 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B42:02 | EPQGSTTGL | 0.2878 | 0.6993 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B42:01 | EPQGSTTGL | 0.219 | 0.6904 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B39:08 | TEPQGSTTGL | 0.833 | 0.8134 | 9 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B07:12 | TEPQGSTTGL | 0.6469 | 0.5385 | 9 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B39:10 | TEPQGSTTGL | 0.6338 | 0.8423 | 9 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B42:01 | TEPQGSTTGL | 0.4611 | 0.7445 | 9 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-A02:02 | YLSAMGSTTGL | 0.9887 | 0.691 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-A02:01 | YLSAMGSTTGL | 0.9573 | 0.7315 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B07:12 | QTEPQGSTTGL | 0.9349 | 0.5032 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C03:04 | SAMGSTTGL | 0.9992 | 0.9861 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C03:03 | SAMGSTTGL | 0.9992 | 0.9861 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C03:67 | SAMGSTTGL | 0.9985 | 0.9756 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C03:05 | SAMGSTTGL | 0.998 | 0.927 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C03:17 | SAMGSTTGL | 0.998 | 0.985 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C15:02 | SAMGSTTGL | 0.9979 | 0.9678 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C15:05 | SAMGSTTGL | 0.9977 | 0.9758 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C03:02 | SAMGSTTGL | 0.9972 | 0.9798 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C16:04 | SAMGSTTGL | 0.9925 | 0.9911 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C12:03 | SAMGSTTGL | 0.9887 | 0.9922 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C12:02 | SAMGSTTGL | 0.987 | 0.9907 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C03:06 | SAMGSTTGL | 0.9858 | 0.9931 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C08:01 | SAMGSTTGL | 0.9182 | 0.9941 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C02:02 | SAMGSTTGL | 0.9169 | 0.9878 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C02:10 | SAMGSTTGL | 0.9169 | 0.9878 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-A68:02 | SAMGSTTGL | 0.9139 | 0.8139 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C16:02 | SAMGSTTGL | 0.8994 | 0.9959 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-B35:13 | SAMGSTTGL | 0.8994 | 0.9349 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-B07:13 | SAMGSTTGL | 0.8978 | 0.8798 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C16:01 | SAMGSTTGL | 0.8862 | 0.9903 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-A69:01 | SAMGSTTGL | 0.8339 | 0.7922 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-B35:09 | SAMGSTTGL | 0.7928 | 0.9873 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C17:01 | SAMGSTTGL | 0.7803 | 0.9821 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-B35:22 | SAMGSTTGL | 0.7442 | 0.6304 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-B15:30 | SAMGSTTGL | 0.7089 | 0.953 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C01:02 | SAMGSTTGL | 0.6028 | 0.9803 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C01:03 | SAMGSTTGL | 0.5904 | 0.9672 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B35:09 | EPQGSTTGL | 0.5208 | 0.8178 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B67:01 | EPQGSTTGL | 0.3425 | 0.7917 | 10 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B40:49 | TEPQGSTTGL | 0.8021 | 0.5282 | 9 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B67:01 | TEPQGSTTGL | 0.6138 | 0.8157 | 9 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B82:02 | TEPQGSTTGL | 0.5728 | 0.5471 | 9 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B41:03 | TEPQGSTTGL | 0.4044 | 0.6125 | 9 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C03:04 | YLSAMGSTTGL | 0.9871 | 0.9954 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 | HLA-C03:03 | YLSAMGSTTGL | 0.9871 | 0.9954 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B67:01 | QTEPQGSTTGL | 0.7537 | 0.8292 | 8 | 19 |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 | HLA-B82:02 | QTEPQGSTTGL | 0.7204 | 0.7437 | 8 | 19 |
Top |
Potential FusionNeoAntigen Information of BCL2L11-BRAF in HLA II |
 Multiple sequence alignments of the potential FusionNeoAntigens per fusion breakpoints. If the MSA is empty, then it means that there were predicted fusion neoantigens in this fusion breakpoint, but those predicted fusion neoantigens were not across the breakpoint, which is not fusion-specific. Multiple sequence alignments of the potential FusionNeoAntigens per fusion breakpoints. If the MSA is empty, then it means that there were predicted fusion neoantigens in this fusion breakpoint, but those predicted fusion neoantigens were not across the breakpoint, which is not fusion-specific. |
 Potential FusionNeoAntigen Information Potential FusionNeoAntigen Information * We used NetMHCIIpan v4.1 (%rank<0.5). |
| Fusion gene | Hchr | Hbp | Tgene | Tchr | Tbp | HLA II | FusionNeoAntigen peptide | Neoantigen start (at BP 13) | Neoantigen end (at BP 13) |
Top |
Fusion breakpoint peptide structures of BCL2L11-BRAF |
 3D structures of the fusion breakpoint peptide of 14AA sequence that have potential fusion neoantigens 3D structures of the fusion breakpoint peptide of 14AA sequence that have potential fusion neoantigens* The minimum length of the amino acid sequence in RoseTTAFold is 14AA. Here, we predicted the 14AA fusion protein breakpoint sequence not the fusion neoantigen peptide, which is shorter than 14 AA. |
| File name | BPseq | Hgene | Tgene | Hchr | Hbp | Tchr | Tbp | AAlen |
| 6189 | NHYLSAMGSTTGLS | BCL2L11 | BRAF | chr2 | 111881446 | chr7 | 140482957 | 545 |
| File name | BPseq | Hgene | Tgene | Hchr | Hbp | Tchr | Tbp | AAlen |
| 8780 | SLQTEPQGSTTGLS | BCL2L11 | BRAF | chr2 | 111881446 | chr7 | 140482957 | 412 |
Top |
Filtering FusionNeoAntigens Through Checking the Interaction with HLAs in 3D of BCL2L11-BRAF |
 Virtual screening between 25 HLAs (from PDB) and FusionNeoAntigens Virtual screening between 25 HLAs (from PDB) and FusionNeoAntigens* We used Glide to predict the interaction between HLAs and neoantigens. |
| HLA allele | PDB ID | File name | BPseq | Docking score | Glide score |
| HLA-B14:02 | 3BVN | 6189 | NHYLSAMGSTTGLS | -7.15543 | -7.26883 |
| HLA-B14:02 | 3BVN | 6189 | NHYLSAMGSTTGLS | -4.77435 | -5.80965 |
| HLA-B52:01 | 3W39 | 6189 | NHYLSAMGSTTGLS | -6.80875 | -6.92215 |
| HLA-B52:01 | 3W39 | 6189 | NHYLSAMGSTTGLS | -4.20386 | -5.23916 |
| HLA-A11:01 | 4UQ2 | 6189 | NHYLSAMGSTTGLS | -7.5194 | -8.5547 |
| HLA-A11:01 | 4UQ2 | 6189 | NHYLSAMGSTTGLS | -6.9601 | -7.0735 |
| HLA-A24:02 | 5HGA | 6189 | NHYLSAMGSTTGLS | -7.52403 | -7.63743 |
| HLA-A24:02 | 5HGA | 6189 | NHYLSAMGSTTGLS | -5.82433 | -6.85963 |
| HLA-B27:05 | 6PYJ | 6189 | NHYLSAMGSTTGLS | -3.28285 | -4.31815 |
| HLA-B44:05 | 3DX8 | 6189 | NHYLSAMGSTTGLS | -5.91172 | -6.94702 |
| HLA-B44:05 | 3DX8 | 6189 | NHYLSAMGSTTGLS | -4.24346 | -4.35686 |
| HLA-B14:02 | 3BVN | 8780 | SLQTEPQGSTTGLS | -7.15543 | -7.26883 |
| HLA-B14:02 | 3BVN | 8780 | SLQTEPQGSTTGLS | -4.77435 | -5.80965 |
| HLA-B52:01 | 3W39 | 8780 | SLQTEPQGSTTGLS | -6.80875 | -6.92215 |
| HLA-B52:01 | 3W39 | 8780 | SLQTEPQGSTTGLS | -4.20386 | -5.23916 |
| HLA-A11:01 | 4UQ2 | 8780 | SLQTEPQGSTTGLS | -7.5194 | -8.5547 |
| HLA-A11:01 | 4UQ2 | 8780 | SLQTEPQGSTTGLS | -6.9601 | -7.0735 |
| HLA-A24:02 | 5HGA | 8780 | SLQTEPQGSTTGLS | -7.52403 | -7.63743 |
| HLA-A24:02 | 5HGA | 8780 | SLQTEPQGSTTGLS | -5.82433 | -6.85963 |
| HLA-B27:05 | 6PYJ | 8780 | SLQTEPQGSTTGLS | -3.28285 | -4.31815 |
| HLA-B44:05 | 3DX8 | 8780 | SLQTEPQGSTTGLS | -5.91172 | -6.94702 |
| HLA-B44:05 | 3DX8 | 8780 | SLQTEPQGSTTGLS | -4.24346 | -4.35686 |
Top |
Vaccine Design for the FusionNeoAntigens of BCL2L11-BRAF |
 mRNA and peptide sequences of FusionNeoAntigens that have potential interaction with HLA-Is. mRNA and peptide sequences of FusionNeoAntigens that have potential interaction with HLA-Is. |
| Fusion gene | Hchr | Hbp | Tchr | Tbp | Start in +/-13AA | End in +/-13AA | FusionNeoAntigen peptide sequence | FusionNeoAntigen RNA sequence |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 10 | 19 | EPQGSTTGL | AGCCACAAGGATCAACCACAGGTTTGT |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 10 | 19 | SAMGSTTGL | GTGCAATGGGATCAACCACAGGTTTGT |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 6 | 15 | NHYLSAMGS | ACCACTATCTCAGTGCAATGGGATCAA |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 8 | 19 | QTEPQGSTTGL | AGACAGAGCCACAAGGATCAACCACAGGTTTGT |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 8 | 19 | YLSAMGSTTGL | ATCTCAGTGCAATGGGATCAACCACAGGTTTGT |
| BCL2L11-BRAF | chr2 | 111881446 | chr7 | 140482957 | 9 | 19 | TEPQGSTTGL | CAGAGCCACAAGGATCAACCACAGGTTTGT |
 mRNA and peptide sequences of FusionNeoAntigens that have potential interaction with HLA-IIs. mRNA and peptide sequences of FusionNeoAntigens that have potential interaction with HLA-IIs. |
| Fusion gene | Hchr | Hbp | Tchr | Tbp | Start in +/-13AA | End in +/-13AA | FusionNeoAntigen peptide | FusionNEoAntigen RNA sequence |
Top |
Information of the samples that have these potential fusion neoantigens of BCL2L11-BRAF |
 These samples were reported as having these fusion breakpoints. For individual breakpoints, we checked the open reading frames considering multiple gene isoforms and chose the in-frame fusion genes only. Then, we made fusion protein sequences and predicted the fusion neoantigens. These fusion-positive samples may have these potential fusion neoantigens. These samples were reported as having these fusion breakpoints. For individual breakpoints, we checked the open reading frames considering multiple gene isoforms and chose the in-frame fusion genes only. Then, we made fusion protein sequences and predicted the fusion neoantigens. These fusion-positive samples may have these potential fusion neoantigens. |
| Cancer type | Fusion gene | Hchr | Hbp | Henst | Tchr | Tbp | Tenst | Sample |
| THCA | BCL2L11-BRAF | chr2 | 111881446 | ENST00000308659 | chr7 | 140482957 | ENST00000288602 | TCGA-ET-A2MX-01A |
| THCA | BCL2L11-BRAF | chr2 | 111881446 | ENST00000357757 | chr7 | 140482957 | ENST00000288602 | TCGA-ET-A2MX-01A |
Top |
Potential target of CAR-T therapy development for BCL2L11-BRAF |
 Predicted 3D structure. We used RoseTTAFold. Predicted 3D structure. We used RoseTTAFold. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, to provide the retention of the transmembrane domain, we only show the protein feature retention information of those transmembrane features Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, to provide the retention of the transmembrane domain, we only show the protein feature retention information of those transmembrane features* Minus value of BPloci means that the break point is located before the CDS. |
| - In-frame and retained 'Transmembrane'. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
 Subcellular localization prediction of the transmembrane domain retained fusion proteins Subcellular localization prediction of the transmembrane domain retained fusion proteins* We used DeepLoc 1.0. The order of the X-axis of the barplot is as follows: Entry_ID, Localization, Type, Nucleus, Cytoplasm, Extracellular, Mitochondrion, Cell_membrane, Endoplasmic_reticulum, Plastid, Golgi.apparatus, Lysosome.Vacuole, Peroxisome. Y-axis is the output score of DeepLoc. Clicking the image will open a new tab with a large image. |
| Hgene | Hchr | Hbp | Henst | Tgene | Tchr | Tbp | Tenst | DeepLoc result |
Top |
Related Drugs to BCL2L11-BRAF |
 Drugs used for this fusion-positive patient. Drugs used for this fusion-positive patient. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Drug | Source | PMID |
Top |
Related Diseases to BCL2L11-BRAF |
 Diseases that have this fusion gene. Diseases that have this fusion gene. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Disease | Source | PMID |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
| Tgene | BRAF | C0025202 | melanoma | 24 | CGI;CTD_human;UNIPROT |
| Tgene | BRAF | C1275081 | Cardio-facio-cutaneous syndrome | 14 | CLINGEN;CTD_human;GENOMICS_ENGLAND;ORPHANET;UNIPROT |
| Tgene | BRAF | C0009402 | Colorectal Carcinoma | 8 | CTD_human;UNIPROT |
| Tgene | BRAF | C0028326 | Noonan Syndrome | 8 | CLINGEN;CTD_human;GENOMICS_ENGLAND;ORPHANET |
| Tgene | BRAF | C0238463 | Papillary thyroid carcinoma | 8 | CTD_human;ORPHANET |
| Tgene | BRAF | C0040136 | Thyroid Neoplasm | 6 | CGI;CTD_human |
| Tgene | BRAF | C0151468 | Thyroid Gland Follicular Adenoma | 6 | CTD_human |
| Tgene | BRAF | C0175704 | LEOPARD Syndrome | 6 | CLINGEN;GENOMICS_ENGLAND |
| Tgene | BRAF | C0549473 | Thyroid carcinoma | 6 | CGI;CTD_human |
| Tgene | BRAF | C3150970 | NOONAN SYNDROME 7 | 5 | CTD_human;GENOMICS_ENGLAND;UNIPROT |
| Tgene | BRAF | C0009404 | Colorectal Neoplasms | 4 | CTD_human |
| Tgene | BRAF | C3150971 | LEOPARD SYNDROME 3 | 4 | CTD_human;GENOMICS_ENGLAND;UNIPROT |
| Tgene | BRAF | C1519086 | Pilomyxoid astrocytoma | 3 | ORPHANET |
| Tgene | BRAF | C0004565 | Melanoma, B16 | 2 | CTD_human |
| Tgene | BRAF | C0009075 | Melanoma, Cloudman S91 | 2 | CTD_human |
| Tgene | BRAF | C0018598 | Melanoma, Harding-Passey | 2 | CTD_human |
| Tgene | BRAF | C0023443 | Hairy Cell Leukemia | 2 | CGI;ORPHANET |
| Tgene | BRAF | C0025205 | Melanoma, Experimental | 2 | CTD_human |
| Tgene | BRAF | C0033578 | Prostatic Neoplasms | 2 | CTD_human |
| Tgene | BRAF | C0152013 | Adenocarcinoma of lung (disorder) | 2 | CGI;CTD_human |
| Tgene | BRAF | C0376358 | Malignant neoplasm of prostate | 2 | CTD_human |
| Tgene | BRAF | C0587248 | Costello syndrome (disorder) | 2 | CLINGEN;CTD_human |
| Tgene | BRAF | C3501843 | Nonmedullary Thyroid Carcinoma | 2 | CTD_human |
| Tgene | BRAF | C3501844 | Familial Nonmedullary Thyroid Cancer | 2 | CTD_human |
| Tgene | BRAF | C0002448 | Ameloblastoma | 1 | CTD_human |
| Tgene | BRAF | C0004114 | Astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0010276 | Craniopharyngioma | 1 | CTD_human;ORPHANET |
| Tgene | BRAF | C0011860 | Diabetes Mellitus, Non-Insulin-Dependent | 1 | CTD_human |
| Tgene | BRAF | C0017638 | Glioma | 1 | CGI;CTD_human |
| Tgene | BRAF | C0019621 | Histiocytosis, Langerhans-Cell | 1 | CGI;ORPHANET |
| Tgene | BRAF | C0022665 | Kidney Neoplasm | 1 | CTD_human |
| Tgene | BRAF | C0023903 | Liver neoplasms | 1 | CTD_human |
| Tgene | BRAF | C0024232 | Lymphatic Metastasis | 1 | CTD_human |
| Tgene | BRAF | C0024694 | Mandibular Neoplasms | 1 | CTD_human |
| Tgene | BRAF | C0027659 | Neoplasms, Experimental | 1 | CTD_human |
| Tgene | BRAF | C0027962 | Melanocytic nevus | 1 | GENOMICS_ENGLAND |
| Tgene | BRAF | C0036920 | Sezary Syndrome | 1 | CTD_human |
| Tgene | BRAF | C0041409 | Turner Syndrome, Male | 1 | CTD_human |
| Tgene | BRAF | C0079773 | Lymphoma, T-Cell, Cutaneous | 1 | CTD_human |
| Tgene | BRAF | C0205768 | Subependymal Giant Cell Astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0206686 | Adrenocortical carcinoma | 1 | CTD_human |
| Tgene | BRAF | C0206754 | Neuroendocrine Tumors | 1 | CTD_human |
| Tgene | BRAF | C0259783 | mixed gliomas | 1 | CTD_human |
| Tgene | BRAF | C0278875 | Adult Craniopharyngioma | 1 | CTD_human |
| Tgene | BRAF | C0280783 | Juvenile Pilocytic Astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0280785 | Diffuse Astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0334579 | Anaplastic astrocytoma | 1 | CGI;CTD_human |
| Tgene | BRAF | C0334580 | Protoplasmic astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0334581 | Gemistocytic astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0334582 | Fibrillary Astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0334583 | Pilocytic Astrocytoma | 1 | CGI;CTD_human |
| Tgene | BRAF | C0338070 | Childhood Cerebral Astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0345904 | Malignant neoplasm of liver | 1 | CTD_human |
| Tgene | BRAF | C0376407 | Granulomatous Slack Skin | 1 | CTD_human |
| Tgene | BRAF | C0406803 | Syringocystadenoma Papilliferum | 1 | GENOMICS_ENGLAND |
| Tgene | BRAF | C0431128 | Papillary craniopharyngioma | 1 | CTD_human |
| Tgene | BRAF | C0431129 | Adamantinous Craniopharyngioma | 1 | CTD_human |
| Tgene | BRAF | C0547065 | Mixed oligoastrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0555198 | Malignant Glioma | 1 | CTD_human |
| Tgene | BRAF | C0596263 | Carcinogenesis | 1 | CTD_human |
| Tgene | BRAF | C0684249 | Carcinoma of lung | 1 | CGI;UNIPROT |
| Tgene | BRAF | C0740457 | Malignant neoplasm of kidney | 1 | CTD_human |
| Tgene | BRAF | C0750935 | Cerebral Astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0750936 | Intracranial Astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0751061 | Craniopharyngioma, Child | 1 | CTD_human |
| Tgene | BRAF | C0920269 | Microsatellite Instability | 1 | CTD_human |
| Tgene | BRAF | C1527404 | Female Pseudo-Turner Syndrome | 1 | CTD_human |
| Tgene | BRAF | C1704230 | Grade I Astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C1721098 | Replication Error Phenotype | 1 | CTD_human |
| Tgene | BRAF | C2239176 | Liver carcinoma | 1 | CTD_human |
| Tgene | BRAF | C4551484 | Leopard Syndrome 1 | 1 | GENOMICS_ENGLAND |
| Tgene | BRAF | C4551602 | Noonan Syndrome 1 | 1 | CTD_human |
| Tgene | BRAF | C4721532 | Lymphoma, Non-Hodgkin, Familial | 1 | UNIPROT |
| Tgene | BRAF | C4733333 | familial non-medullary thyroid cancer | 1 | GENOMICS_ENGLAND |

