| UTHEALTH HOME ABOUT SBMI A-Z WEBMAIL INSIDE THE UNIVERSITY |

|
|||||||
|
Kinase Fusion Gene:AHI1_SGK1 |
Kinase Fusion Protein Summary |
 Kinase Fusion gene summary Kinase Fusion gene summary |
| Kinase Fusion partner gene information | Kinase Fusion gene name: AHI1_SGK1 | KinaseFusionDB ID: KFG235 | FusionGDB2.0 ID: KFG235 | Hgene | Tgene | Gene symbol | AHI1 | SGK1 | Gene ID | 54806 | 6446 | |
| Gene name | Abelson helper integration site 1 | serum/glucocorticoid regulated kinase 1 | ||||||||||
| Synonyms | AHI-1|JBTS3|ORF1|dJ71N10.1 | SGK | ||||||||||
| Cytomap | 6q23.3 | 6q23.2 | ||||||||||
| Type of gene | protein-coding | protein-coding | ||||||||||
| Description | jouberinabelson helper integration site 1 protein homologcontatins SH3 and WD40 domains | serine/threonine-protein kinase Sgk1serine/threonine protein kinase SGK | ||||||||||
| Modification date | 20240411 | 20240305 | ||||||||||
| UniProtAcc | Q8N157 | O00141 | ||||||||||
| Ensembl transtripts involved in fusion gene | ENST ids | ENST00000367800, ENST00000417892, ENST00000457866, ENST00000327035, ENST00000367798, ENST00000488690, ENST00000528103, ENST00000531527, ENST00000534469, | ENST00000237305, ENST00000367857, ENST00000413996, ENST00000475719, ENST00000489458, ENST00000524929, ENST00000528577, ENST00000367858, | |||||||||
| Context (manual curation of fusion genes in KinaseFusionDB) | PubMed: AHI1 [Title/Abstract] AND SGK1 [Title/Abstract] AND fusion [Title/Abstract] | |||||||||||
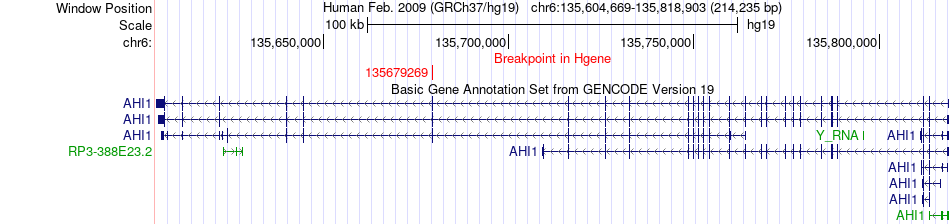
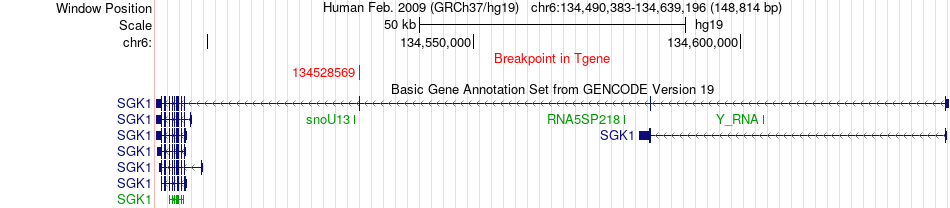
| Most frequent breakpoint (based on all fusion genes of FusionGDB 2.0) | AHI1(135679269)-SGK1(134528569), # samples:1 | |||||||||||
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Kinase Fusion gene breakpoints across AHI1 (5'-gene) Kinase Fusion gene breakpoints across AHI1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
 Kinase Fusion gene breakpoints across SGK1 (3'-gene) Kinase Fusion gene breakpoints across SGK1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
Top |
Kinase Fusion Gene Sample Information |
 Kinase Fusion gene information. Kinase Fusion gene information. |
 Kinase Fusion gene information from four resources (ChiTars 5.0, ChimerDB 4.0, COSMIC, and CCLE) Kinase Fusion gene information from four resources (ChiTars 5.0, ChimerDB 4.0, COSMIC, and CCLE)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Sample | Hgene | Hchr | Hbp | Tgene | Tchr | Tbp |
| ChimerDB4 | TCGA-R6-A6DN | AHI1 | chr6 | 135679269 | SGK1 | chr6 | 134528569 |
Top |
Kinase Fusion ORF Analysis |
 Kinase Fusion information from ORFfinder translation from full-length transcript sequence from KinaseFusionDB. Kinase Fusion information from ORFfinder translation from full-length transcript sequence from KinaseFusionDB. |
| Henst | Tenst | Hgene | Hchr | Hbp | Tgene | Tchr | Tbp | Seq length (transcript) | Seq length (amino acids) |
| ENST00000457866 | ENST00000367858 | AHI1 | chr6 | 135679269 | SGK1 | chr6 | 134528569 | 5782 | 1486 |
Top |
Kinase Fusion Amino Acid Sequences |
 For individual full-length fusion transcript sequence from KinaseFusionDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. For individual full-length fusion transcript sequence from KinaseFusionDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >Henst_Tenst_Hgene_Hchr_Hbp_Tgene_Tchr_Tbp_length(fusion AA)_AAseq >ENST00000457866_ENST00000367858_AHI1_chr6_135679269_SGK1_chr6_134528569_length(amino acids)=1486 MPTAESEAKVKTKVRFEELLKTHSDLMREKKKLKKKLVRSEENISPDTIRSNLHYMKETTSDDPDTIRSNLPHIKETTSDDVSAANTNNL KKSTRVTKNKLRNTQLATENPNGDASVEEDKQGKPNKKVIKTVPQLTTQDLKPETPENKVDSTHQKTHTKPQPGVDHQKSEKANEGREET DLEEDEELMQAYQCHVTEEMAKEIKRKIRKKLKEQLTYFPSDTLFHDDKLSSEKRKKKKEVPVFSKAETSTLTISGDTVEGEQKKESSVR SVSSDSHQDDEISSMEQSTEDSMQDDTKPKPKKTKKKTKAVADNNEDVDGDGVHEITSRDSPVYPKCLLDDDLVLGVYIHRTDRLKSDFM ISHPMVKIHVVDEHTGQYVKKDDSGRPVSSYYEKENVDYILPIMTQPYDFKQLKSRLPEWEEQIVFNENFPYLLRGSDESPKVILFFEIL DFLSVDEIKNNSEVQNQECGFRKIAWAFLKLLGANGNANINSKLRLQLYYPPTKPRSPLSVVEAFEWWSKCPRNHYPSTLYVTVRGLKVP DCIKPSYRSMMALQEEKGKPVHCERHHESSSVDTEPGLEESKEVIKWKRLPGQACRIPNKHLFSLNAGERGCFCLDFSHNGRILAAACAS RDGYPIILYEIPSGRFMRELCGHLNIIYDLSWSKDDHYILTSSSDGTARIWKNEINNTNTFRVLPHPSFVYTAKFHPAVRELVVTGCYDS MIRIWKVEMREDSAILVRQFDVHKSFINSLCFDTEGHHMYSGDCTGVIVVWNTYVKINDLEHSVHHWTINKEIKETEFKGIPISYLEIHP NGKRLLIHTKDSTLRIMDLRILVARKFVGAANYREKIHSTLTPCGTFLFAGSEDGIVYVWNPETGEQVAMYSDLPFKSPIRDISYHPFEN MVAFCAFGQNEPILLYIYDFHVAQQEAEMFKRYNGTFPLPGIHQSQDALCTCPKLPHQGSFQIDEFVHTESSSTKMQLVKQRLETVTEVI RSCAAKVNKNLSFTSPPAVSSQQSKLKQSNMLTAQEILHQFGFTQTGIISIERKPCNHQVDTAPTVREPCNHANILTKPDPRTFWTNDDP AFMKQRRMGLNDFIQKIANNSYACKHPEVQSILKISQPQEPELMNANPSPPPSPSQQINLGPSSNPHAKPSDFHFLKVIGKGSFGKVLLA RHKAEEVFYAVKVLQKKAILKKKEEKHIMSERNVLLKNVKHPFLVGLHFSFQTADKLYFVLDYINGGELFYHLQRERCFLEPRARFYAAE IASALGYLHSLNIVYRDLKPENILLDSQGHIVLTDFGLCKENIEHNSTTSTFCGTPEYLAPEVLHKQPYDRTVDWWCLGAVLYEMLYGLP PFYSRNTAEMYDNILNKPLQLKPNITNSARHLLEGLLQKDRTKRLGAKDDFMEIKSHVFFSLINWDDLINKKITPPFNPNVSGPNDLRHF -------------------------------------------------------------- |
Multiple Sequence Alignment of All Fusion Protein Isoforms |
Top |
Kinase Fusion Protein Functional Features |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr6:135679269/chr6:134528569) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| AHI1 | SGK1 |
| FUNCTION: Involved in vesicle trafficking and required for ciliogenesis, formation of primary non-motile cilium, and recruitment of RAB8A to the basal body of primary cilium. Component of the tectonic-like complex, a complex localized at the transition zone of primary cilia and acting as a barrier that prevents diffusion of transmembrane proteins between the cilia and plasma membranes. Involved in neuronal differentiation. As a positive modulator of classical Wnt signaling, may play a crucial role in ciliary signaling during cerebellum embryonic development (PubMed:21623382). {ECO:0000250|UniProtKB:Q8K3E5, ECO:0000269|PubMed:21623382}. | FUNCTION: Serine/threonine-protein kinase which is involved in the regulation of a wide variety of ion channels, membrane transporters, cellular enzymes, transcription factors, neuronal excitability, cell growth, proliferation, survival, migration and apoptosis. Plays an important role in cellular stress response. Contributes to regulation of renal Na(+) retention, renal K(+) elimination, salt appetite, gastric acid secretion, intestinal Na(+)/H(+) exchange and nutrient transport, insulin-dependent salt sensitivity of blood pressure, salt sensitivity of peripheral glucose uptake, cardiac repolarization and memory consolidation. Up-regulates Na(+) channels: SCNN1A/ENAC, SCN5A and ASIC1/ACCN2, K(+) channels: KCNJ1/ROMK1, KCNA1-5, KCNQ1-5 and KCNE1, epithelial Ca(2+) channels: TRPV5 and TRPV6, chloride channels: BSND, CLCN2 and CFTR, glutamate transporters: SLC1A3/EAAT1, SLC1A2 /EAAT2, SLC1A1/EAAT3, SLC1A6/EAAT4 and SLC1A7/EAAT5, amino acid transporters: SLC1A5/ASCT2, SLC38A1/SN1 and SLC6A19, creatine transporter: SLC6A8, Na(+)/dicarboxylate cotransporter: SLC13A2/NADC1, Na(+)-dependent phosphate cotransporter: SLC34A2/NAPI-2B, glutamate receptor: GRIK2/GLUR6. Up-regulates carriers: SLC9A3/NHE3, SLC12A1/NKCC2, SLC12A3/NCC, SLC5A3/SMIT, SLC2A1/GLUT1, SLC5A1/SGLT1 and SLC15A2/PEPT2. Regulates enzymes: GSK3A/B, PMM2 and Na(+)/K(+) ATPase, and transcription factors: CTNNB1 and nuclear factor NF-kappa-B. Stimulates sodium transport into epithelial cells by enhancing the stability and expression of SCNN1A/ENAC. This is achieved by phosphorylating the NEDD4L ubiquitin E3 ligase, promoting its interaction with 14-3-3 proteins, thereby preventing it from binding to SCNN1A/ENAC and targeting it for degradation. Regulates store-operated Ca(+2) entry (SOCE) by stimulating ORAI1 and STIM1. Regulates KCNJ1/ROMK1 directly via its phosphorylation or indirectly via increased interaction with SLC9A3R2/NHERF2. Phosphorylates MDM2 and activates MDM2-dependent ubiquitination of p53/TP53. Phosphorylates MAPT/TAU and mediates microtubule depolymerization and neurite formation in hippocampal neurons. Phosphorylates SLC2A4/GLUT4 and up-regulates its activity. Phosphorylates APBB1/FE65 and promotes its localization to the nucleus. Phosphorylates MAPK1/ERK2 and activates it by enhancing its interaction with MAP2K1/MEK1 and MAP2K2/MEK2. Phosphorylates FBXW7 and plays an inhibitory role in the NOTCH1 signaling. Phosphorylates FOXO1 resulting in its relocalization from the nucleus to the cytoplasm. Phosphorylates FOXO3, promoting its exit from the nucleus and interference with FOXO3-dependent transcription. Phosphorylates BRAF and MAP3K3/MEKK3 and inhibits their activity. Phosphorylates SLC9A3/NHE3 in response to dexamethasone, resulting in its activation and increased localization at the cell membrane. Phosphorylates CREB1. Necessary for vascular remodeling during angiogenesis. Sustained high levels and activity may contribute to conditions such as hypertension and diabetic nephropathy. Isoform 2 exhibited a greater effect on cell plasma membrane expression of SCNN1A/ENAC and Na(+) transport than isoform 1. {ECO:0000269|PubMed:11154281, ECO:0000269|PubMed:11410590, ECO:0000269|PubMed:11696533, ECO:0000269|PubMed:12397388, ECO:0000269|PubMed:12590200, ECO:0000269|PubMed:12634932, ECO:0000269|PubMed:12650886, ECO:0000269|PubMed:12761204, ECO:0000269|PubMed:12911626, ECO:0000269|PubMed:14623317, ECO:0000269|PubMed:14706641, ECO:0000269|PubMed:15040001, ECO:0000269|PubMed:15044175, ECO:0000269|PubMed:15234985, ECO:0000269|PubMed:15319523, ECO:0000269|PubMed:15496163, ECO:0000269|PubMed:15733869, ECO:0000269|PubMed:15737648, ECO:0000269|PubMed:15845389, ECO:0000269|PubMed:15888551, ECO:0000269|PubMed:16036218, ECO:0000269|PubMed:16443776, ECO:0000269|PubMed:16982696, ECO:0000269|PubMed:17382906, ECO:0000269|PubMed:18005662, ECO:0000269|PubMed:18304449, ECO:0000269|PubMed:18753299, ECO:0000269|PubMed:19447520, ECO:0000269|PubMed:19756449, ECO:0000269|PubMed:20511718, ECO:0000269|PubMed:20730100, ECO:0000269|PubMed:21865597}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
 - Retained domain in the 5'-partner of fusion protein (protein functional feature from UniProt). - Retained domain in the 5'-partner of fusion protein (protein functional feature from UniProt). |
| Partner | Hgeneene | Hbp | Tgeneene | Tbp | ENST | BPexon | TotalExon | Protein feature loci | BPloci | TotalLen | Feature | Note |
 - Retained domain in the 3'-partner of fusion protein (protein functional feature from UniProt). - Retained domain in the 3'-partner of fusion protein (protein functional feature from UniProt). |
| Partner | Hgeneene | Hbp | Tgeneene | Tbp | ENST | BPexon | TotalExon | Protein feature loci | BPloci | TotalLen | Feature | Note |
| Tgene | AHI1 | 135679269 | SGK1 | 134528569 | ENST00000457866 | 0 | 11 | 356_431 | 0 | 422 | Domain | Note=AGC-kinase C-terminal;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00618 |
| Tgene | AHI1 | 135679269 | SGK1 | 134528569 | ENST00000457866 | 0 | 12 | 356_431 | 0 | 432 | Domain | Note=AGC-kinase C-terminal;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00618 |
| Tgene | AHI1 | 135679269 | SGK1 | 134528569 | ENST00000457866 | 0 | 12 | 356_431 | 0 | 446 | Domain | Note=AGC-kinase C-terminal;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00618 |
| Tgene | AHI1 | 135679269 | SGK1 | 134528569 | ENST00000457866 | 0 | 12 | 356_431 | 0 | 460 | Domain | Note=AGC-kinase C-terminal;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00618 |
| Tgene | AHI1 | 135679269 | SGK1 | 134528569 | ENST00000457866 | 1 | 14 | 356_431 | 95 | 527 | Domain | Note=AGC-kinase C-terminal;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00618 |
| Tgene | AHI1 | 135679269 | SGK1 | 134528569 | ENST00000457866 | 0 | 11 | 98_355 | 0 | 422 | Domain | Note=Protein kinase;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00159 |
| Tgene | AHI1 | 135679269 | SGK1 | 134528569 | ENST00000457866 | 0 | 12 | 98_355 | 0 | 432 | Domain | Note=Protein kinase;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00159 |
| Tgene | AHI1 | 135679269 | SGK1 | 134528569 | ENST00000457866 | 0 | 12 | 98_355 | 0 | 446 | Domain | Note=Protein kinase;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00159 |
| Tgene | AHI1 | 135679269 | SGK1 | 134528569 | ENST00000457866 | 0 | 12 | 98_355 | 0 | 460 | Domain | Note=Protein kinase;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00159 |
| Tgene | AHI1 | 135679269 | SGK1 | 134528569 | ENST00000457866 | 1 | 14 | 98_355 | 95 | 527 | Domain | Note=Protein kinase;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00159 |
Top |
Kinase Fusion Protein Structures |
 CIF files of the predicted kinase fusion proteins CIF files of the predicted kinase fusion proteins * Here we show the 3D structure of the fusion proteins using Mol*. AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. Model confidence is shown from the pLDDT values per residue. pLDDT corresponds to the model’s prediction of its score on the local Distance Difference Test. It is a measure of local accuracy (from AlphfaFold website). To color code individual residues, we transformed individual PDB files into CIF format. |
| Kinase Fusion protein CIF link (fusion AA seq ID in KinaseFusionDB) | Henst | Tenst | Hgene | Hchr | Hbp | Tgene | Tchr | Tbp | AA seq | Len(AA seq) |
| PDB file >>>14_AHI1_SGK1 | ENST00000457866 | ENST00000367858 | AHI1 | chr6 | 135679269 | SGK1 | chr6 | 134528569 | MPTAESEAKVKTKVRFEELLKTHSDLMREKKKLKKKLVRSEENISPDTIRSNLHYMKETTSDDPDTIRSNLPHIKETTSDDVSAANTNNL KKSTRVTKNKLRNTQLATENPNGDASVEEDKQGKPNKKVIKTVPQLTTQDLKPETPENKVDSTHQKTHTKPQPGVDHQKSEKANEGREET DLEEDEELMQAYQCHVTEEMAKEIKRKIRKKLKEQLTYFPSDTLFHDDKLSSEKRKKKKEVPVFSKAETSTLTISGDTVEGEQKKESSVR SVSSDSHQDDEISSMEQSTEDSMQDDTKPKPKKTKKKTKAVADNNEDVDGDGVHEITSRDSPVYPKCLLDDDLVLGVYIHRTDRLKSDFM ISHPMVKIHVVDEHTGQYVKKDDSGRPVSSYYEKENVDYILPIMTQPYDFKQLKSRLPEWEEQIVFNENFPYLLRGSDESPKVILFFEIL DFLSVDEIKNNSEVQNQECGFRKIAWAFLKLLGANGNANINSKLRLQLYYPPTKPRSPLSVVEAFEWWSKCPRNHYPSTLYVTVRGLKVP DCIKPSYRSMMALQEEKGKPVHCERHHESSSVDTEPGLEESKEVIKWKRLPGQACRIPNKHLFSLNAGERGCFCLDFSHNGRILAAACAS RDGYPIILYEIPSGRFMRELCGHLNIIYDLSWSKDDHYILTSSSDGTARIWKNEINNTNTFRVLPHPSFVYTAKFHPAVRELVVTGCYDS MIRIWKVEMREDSAILVRQFDVHKSFINSLCFDTEGHHMYSGDCTGVIVVWNTYVKINDLEHSVHHWTINKEIKETEFKGIPISYLEIHP NGKRLLIHTKDSTLRIMDLRILVARKFVGAANYREKIHSTLTPCGTFLFAGSEDGIVYVWNPETGEQVAMYSDLPFKSPIRDISYHPFEN MVAFCAFGQNEPILLYIYDFHVAQQEAEMFKRYNGTFPLPGIHQSQDALCTCPKLPHQGSFQIDEFVHTESSSTKMQLVKQRLETVTEVI RSCAAKVNKNLSFTSPPAVSSQQSKLKQSNMLTAQEILHQFGFTQTGIISIERKPCNHQVDTAPTVREPCNHANILTKPDPRTFWTNDDP AFMKQRRMGLNDFIQKIANNSYACKHPEVQSILKISQPQEPELMNANPSPPPSPSQQINLGPSSNPHAKPSDFHFLKVIGKGSFGKVLLA RHKAEEVFYAVKVLQKKAILKKKEEKHIMSERNVLLKNVKHPFLVGLHFSFQTADKLYFVLDYINGGELFYHLQRERCFLEPRARFYAAE IASALGYLHSLNIVYRDLKPENILLDSQGHIVLTDFGLCKENIEHNSTTSTFCGTPEYLAPEVLHKQPYDRTVDWWCLGAVLYEMLYGLP PFYSRNTAEMYDNILNKPLQLKPNITNSARHLLEGLLQKDRTKRLGAKDDFMEIKSHVFFSLINWDDLINKKITPPFNPNVSGPNDLRHF | 1486 |
| 3D view using mol* of 14_AHI1_SGK1 | ||||||||||
| PDB file >>>TKFP_22_AHI1_SGK1 | ENST00000457866 | ENST00000367858 | AHI1 | chr6 | 135679269 | SGK1 | chr6 | 134528569 | MPTAESEAKVKTKVRFEELLKTHSDLMREKKKLKKKLVRSEENISPDTIRSNLHYMKETTSDDPDTIRSNLPHIKETTSDDVSAANTNNL KKSTRVTKNKLRNTQLATENPNGDASVEEDKQGKPNKKVIKTVPQLTTQDLKPETPENKVDSTHQKTHTKPQPGVDHQKSEKANEGREET DLEEDEELMQAYQCHVTEEMAKEIKRKIRKKLKEQLTYFPSDTLFHDDKLSSEKRKKKKEVPVFSKAETSTLTISGDTVEGEQKKESSVR SVSSDSHQDDEISSMEQSTEDSMQDDTKPKPKKTKKKTKAVADNNEDVDGDGVHEITSRDSPVYPKCLLDDDLVLGVYIHRTDRLKSDFM ISHPMVKIHVVDEHTGQYVKKDDSGRPVSSYYEKENVDYILPIMTQPYDFKQLKSRLPEWEEQIVFNENFPYLLRGSDESPKVILFFEIL DFLSVDEIKNNSEVQNQECGFRKIAWAFLKLLGANGNANINSKLRLQLYYPPTKPRSPLSVVEAFEWWSKCPRNHYPSTLYVTVRGLKVP DCIKPSYRSMMALQEEKGKPVHCERHHESSSVDTEPGLEESKEVIKWKRLPGQACRIPNKHLFSLNAGERGCFCLDFSHNGRILAAACAS RDGYPIILYEIPSGRFMRELCGHLNIIYDLSWSKDDHYILTSSSDGTARIWKNEINNTNTFRVLPHPSFVYTAKFHPAVRELVVTGCYDS MIRIWKVEMREDSAILVRQFDVHKSFINSLCFDTEGHHMYSGDCTGVIVVWNTYVKINDLEHSVHHWTINKEIKETEFKGIPISYLEIHP NGKRLLIHTKDSTLRIMDLRILVARKFVGAANYREKIHSTLTPCGTFLFAGSEDGIVYVWNPETGEQVAMYSDLPFKSPIRDISYHPFEN MVAFCAFGQNEPILLYIYDFHVAQQEAEMFKRYNGTFPLPGIHQSQDALCTCPKLPHQGSFQIDEFVHTESSSTKMQLVKQRLETVTEVI RSCAAKVNKNLSFTSPPAVSSQQSKLKQSNMLTAQEILHQFGFTQTGIISIERKPCNHQVDTAPTVREPCNHANILTKPDPRTFWTNDDP AFMKQRRMGLNDFIQKIANNSYACKHPEVQSILKISQPQEPELMNANPSPPPSPSQQINLGPSSNPHAKPSDFHFLKVIGKGSFGKVLLA RHKAEEVFYAVKVLQKKAILKKKEEKHIMSERNVLLKNVKHPFLVGLHFSFQTADKLYFVLDYINGGELFYHLQRERCFLEPRARFYAAE IASALGYLHSLNIVYRDLKPENILLDSQGHIVLTDFGLCKENIEHNSTTSTFCGTPEYLAPEVLHKQPYDRTVDWWCLGAVLYEMLYGLP PFYSRNTAEMYDNILNKPLQLKPNITNSARHLLEGLLQKDRTKRLGAKDDFMEIKSHVFFSLINWDDLINKKITPPFNPNVSGPNDLRHF | 1486_AHI1_SGK1 |
Top |
Comparison of Fusion Protein Isoforms |
 Superimpose the 3D Structures Among All Fusion Protein Isoforms Superimpose the 3D Structures Among All Fusion Protein Isoforms * Download the pdb file and open it from the molstar online viewer. |
 Comparison of the Secondary Structures of Fusion Protein Isoforms Comparison of the Secondary Structures of Fusion Protein Isoforms |
Top |
Comparison of Fusion Protein Sequences/Structures with Known Sequences/Structures from PDB |
Top |
pLDDT score distribution |
 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2* AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. * The blue color at the bottom marks the best active site residues. |
| 14_AHI1_SGK1.png |
 |
| 14_AHI1_SGK1.png |
 |
Top |
Potential Active Site Information |
 The potential binding sites of these fusion proteins were identified using SiteMap, a module of the Schrodinger suite. The potential binding sites of these fusion proteins were identified using SiteMap, a module of the Schrodinger suite. |
| Kinase Fusion AA seq ID in KinaseFusionDB | Site score | Size | Dscore | Volume | Exposure | Enclosure | Contact | Phobic | Philic | Balance | Don/Acc | Residues |
Top |
Ramachandran Plot of Kinase Fusion Protein Structure |
 Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. |
| 14_AHI1_SGK1_ramachandran.png |
 |
Top |
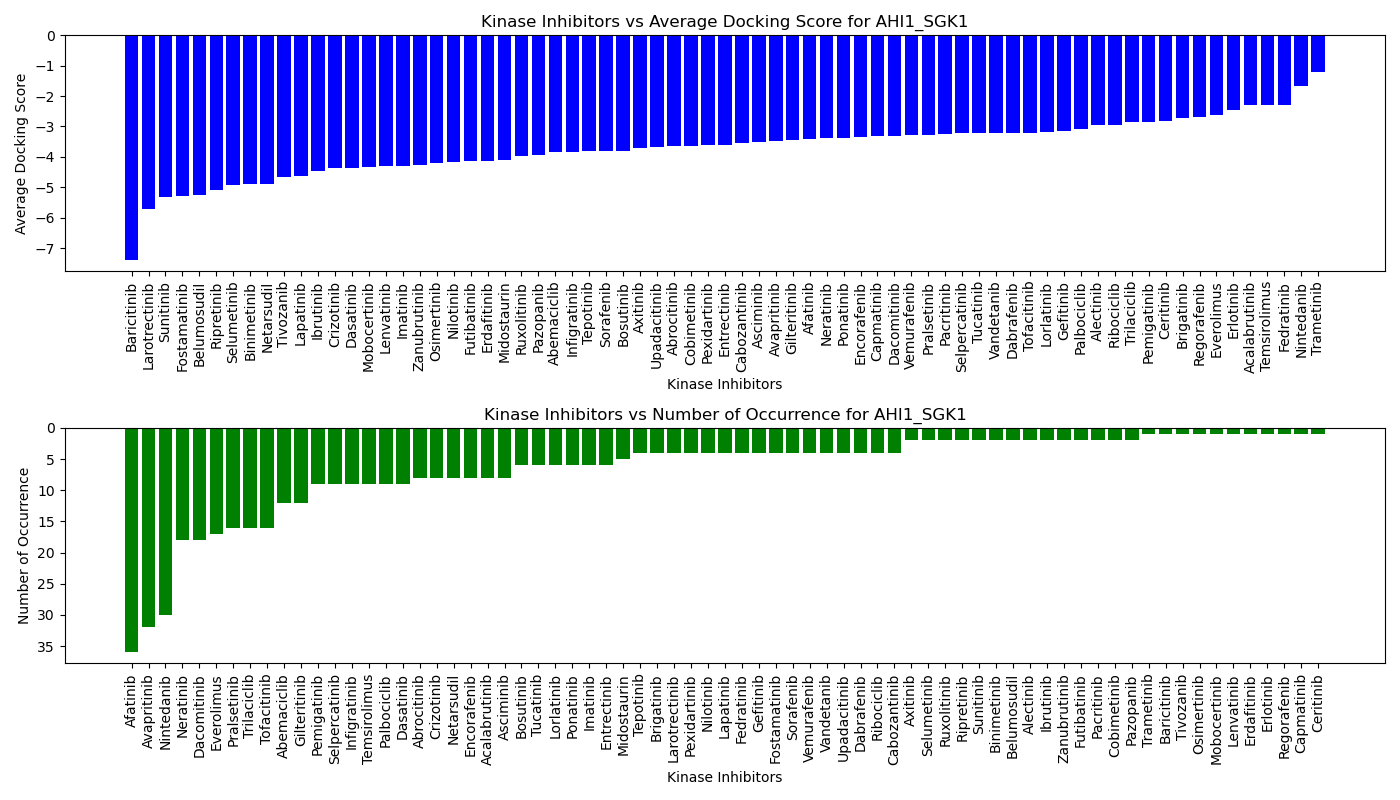
Virtual Screening Results |
 Distribution of the average docking score across all approved kinase inhibitors. Distribution of the average docking score across all approved kinase inhibitors.Distribution of the number of occurrence across all approved kinase inhibitors. |
| 5'-kinase fusion protein case |
| 3'-kinase fusion protein case |
 |
Top |
 Drug information from DrugBank of the top 20 interacting small molecules. Drug information from DrugBank of the top 20 interacting small molecules.* The detailed information of individual kinase inhibitors are available in the download page. |
| Fusion gene name info | Drug | Docking score | Glide g score | Glide energy |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Baricitinib | -7.38605 | -7.38605 | -40.6563 |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Netarsudil | -6.76213 | -6.77323 | -54.1894 |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Netarsudil | -6.76213 | -6.77323 | -54.1894 |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Larotrectinib | -6.51813 | -6.51813 | -45.6358 |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Abrocitinib | -6.2384900000000005 | -6.24959 | -38.9322 |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Abrocitinib | -6.2384900000000005 | -6.24959 | -38.9322 |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Larotrectinib | -6.17695 | -6.17695 | -43.951 |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Fostamatinib | -6.01936 | -6.08296 | -56.5353 |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Fostamatinib | -6.01936 | -6.08296 | -56.5353 |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Lapatinib | -5.93823 | -6.02703 | -55.06100000000001 |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Larotrectinib | -5.87326 | -5.87326 | -42.2495 |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Dasatinib | -5.80729 | -6.29319 | -55.6657 |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Dasatinib | -5.80729 | -6.29319 | -55.6657 |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Dasatinib | -5.80729 | -6.29319 | -55.6657 |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Sunitinib | -5.76839 | -5.77259 | -39.6084 |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Belumosudil | -5.48123 | -5.48893 | -48.6795 |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Selpercatinib | -5.4621699999999995 | -5.4926699999999995 | -55.8775 |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Selpercatinib | -5.4621699999999995 | -5.4926699999999995 | -55.8775 |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Neratinib | -5.39503 | -5.57733 | -57.0824 |
| 14_AHI1_SGK1-DOCK_HTVS_1-001 | Neratinib | -5.39503 | -5.57733 | -57.0824 |
Top |
Kinase-Substrate Information of AHI1_SGK1 |
 Phosphorylation target of the kinase Phosphorylation target of the kinase(phosphosite, 03-17-2024) |
| Kinase | Kinase UniProt Acc | Kinase species | Substrate | Substrate UniProt Acc | Substrate phosphorylated residues | Substrate phosphorylated sites (+/-7AA) | Domain |
| SGK1 | O00141 | human | NEDD4L | Q96PU5 | S342 | ssRLRsCsVtDAVAE | |
| SGK1 | O00141 | human | TBC1D4 | O60343 | S318 | EFRsRCssVtGVQRR | PID |
| SGK1 | O00141 | human | NDRG2 | Q9UN36 | T330 | tRLsRsRtAsLtsAA | |
| SGK1 | O00141 | human | NCSTN | Q92542 | S437 | FLRArNIsGVVLADH | Nicastrin |
| SGK1 | O00141 | human | NDRG2 | Q9UN36 | S346 | VDGNRsRsRtLsQss | |
| SGK1 | O00141 | human | CDKN1B | P46527 | T157 | GIRkrPAtDDSSTQN | |
| SGK1 | O00141 | human | KMT2D | O14686-3 | S1331 | RGRARLKsTASSIET | |
| SGK1 | O00141 | human | MKNK1 | Q9BUB5-2 | S353 | QVLQRNSsTMDLTLF | |
| SGK1 | O00141 | human | GSK3B | P49841 | Y216 | RGEPNVsyICsRyyR | Pkinase |
| SGK1 | O00141 | human | FOXO3 | O43524 | S315 | DFRsRTNsNAstVsG | |
| SGK1 | O00141 | human | NEDD4L | Q96PU5 | S448 | IRRPRsLssPtVtLS | |
| SGK1 | O00141 | human | MAPK1 | P28482 | S29 | GPRyTNLsYIGEGAy | Pkinase |
| SGK1 | O00141 | human | TSC2 | P49815 | S981 | AFRCRSIsVSEHVVR | |
| SGK1 | O00141 | human | TBC1D4 | O60343 | S570 | AKRsLtssLENIFsr | |
| SGK1 | O00141 | human | MAP2K4 | P45985 | S80 | IERLRtHsIESSGkL | |
| SGK1 | O00141 | human | TSC2 | P49815 | S1130 | GARDRVRsMsGGHGL | |
| SGK1 | O00141 | human | WNK4 | Q96J92 | S1190 | SSRQRRLsKGSFPTS | |
| SGK1 | O00141 | human | NDRG1 | Q92597 | T366 | GtRsRsHtsEGAHLD | |
| SGK1 | O00141 | human | HTT | P42858 | S419 | GGRsRsGsIVELIAG | |
| SGK1 | O00141 | human | FBXW7 | Q969H0 | S227 | QQRRRITsVQPPTGL | |
| SGK1 | O00141 | human | NFKBIA | P25963 | S32 | LLDDRHDsGLDsMkD | |
| SGK1 | O00141 | human | APBB1 | O00213 | S610 | ECRVRFLsFLAVGRD | PID |
| SGK1 | O00141 | human | NDRG1 | Q92597 | T346 | GtRsRsHtsEGtRsR | |
| SGK1 | O00141 | human | NDRG1 | Q92597 | T356 | GtRsRsHtsEGtRsR | |
| SGK1 | O00141 | human | TSC2 | P49815 | T1462 | GLrPrGytISDsAPs | |
| SGK1 | O00141 | human | NDRG2 | Q9UN36 | T348 | GNRsRsRtLsQssEs | |
| SGK1 | O00141 | human | CREB1 | P16220 | S119 | EILsRRPsyRkILND | pKID |
| SGK1 | O00141 | human | NEDD4L | Q96PU5 | T367 | AGRARsstVtGGEEP | |
| SGK1 | O00141 | human | TSC2 | P49815 | S939 | sFRARstsLNERPKs | |
| SGK1 | O00141 | human | GSK3B | P49841 | S9 | SGRPRttsFAEsCkP | |
| SGK1 | O00141 | human | ZBTB33 | Q86T24 | T606 | yLSDRSStIPAMkDD | |
| SGK1 | O00141 | human | NDRG2 | Q9UN36 | S332 | LsRsRtAsLtsAAsV | |
| SGK1 | O00141 | human | TBC1D4 | O60343 | S751 | EGRKRtsstCsNEsL | |
| SGK1 | O00141 | human | MDM2 | Q00987 | S166 | SsRRRAIsEtEENsD | |
| SGK1 | O00141 | human | MAPT | P10636-8 | S214 | GsRsRtPsLPtPPtR | |
| SGK1 | O00141 | human | TBC1D4 | O60343 | S341 | QPRRRHAsAPsHVQP | PID |
| SGK1 | O00141 | human | CDKN1B | P46527 | T198 | PGLRRRQt_______ | |
| SGK1 | O00141 | human | TSC2 | P49815 | S1798 | GQRKRLIssVEDFTE | |
| SGK1 | O00141 | human | SGK1 | O00141 | S422 | AEAFLGFsYAPPTDS | Pkinase_C |
| SGK1 | O00141 | human | CARHSP1 | Q9Y2V2 | S52 | tRRtrtFsAtVRAsQ | |
| SGK1 | O00141 | human | WNK1 | Q9H4A3 | T60 | EYRRRRHtMDKDSrG | |
| SGK1 | O00141 | human | TBC1D4 | O60343 | T642 | QFRRRAHtFsHPPss | |
| SGK1 | O00141 | human | SLC2A4 | P14672 | S274 | LERERPLsLLQLLGs | Sugar_tr |
| SGK1 | O00141 | human | NDRG1 | Q92597 | S330 | LMRsRtAsGssVtsL | |
| SGK1 | O00141 | human | TBC1D4 | O60343 | T568 | NKAKRsLtssLENIF | |
| SGK1 | O00141 | human | TBC1D4 | O60343 | S588 | RMRGRLGsVDsFERs | |
| SGK1 | O00141 | human | TSC2 | P49815 | S1132 | RDRVRsMsGGHGLRV | |
| SGK1 | O00141 | human | TBC1D4 | O60343 | S666 | GRAQGVRsPLLRQSS | |
| SGK1 | O00141 | human | WNK4 | Q96J92 | S1201 | FPTSRRNsLQRSEPP | |
| SGK1 | O00141 | human | PIKFYVE | Q9Y2I7 | S318 | LsLDRsGsPMVPSyE | |
| SGK1 | O00141 | human | RICTOR | Q6R327 | T1135 | NRRIRtLtEPsVDFN | RICTOR_phospho |
| SGK1 | O00141 | human | IKBKB | O14920 | S181 | DQGsLCtsFVGTLQy | Pkinase |
| SGK1 | O00141 | human | NDRG1 | Q92597 | T328 | TRLMRsRtAsGssVt | |
| SGK1 | O00141 | human | WNK4 | Q96J92 | S1217 | PGIMRRNsLSGSStG |
 Biological Network Integration of This Kinase and Substrates Biological Network Integration of This Kinase and Substrates (GeneMANIA website) |
 Enriched GO biological processes of the phosphorylation target genes of the kinase Enriched GO biological processes of the phosphorylation target genes of the kinase |
| Kinase | GOID | GO term | P.adjust |
| SGK1 | ID | Description | 0.00e+00 |
| SGK1 | GO:0018105 | peptidyl-serine phosphorylation | 1.12e-05 |
| SGK1 | GO:0018209 | peptidyl-serine modification | 1.12e-05 |
| SGK1 | GO:0007611 | learning or memory | 7.35e-05 |
| SGK1 | GO:0043434 | response to peptide hormone | 7.35e-05 |
| SGK1 | GO:0050890 | cognition | 1.02e-04 |
| SGK1 | GO:0062197 | cellular response to chemical stress | 1.02e-04 |
| SGK1 | GO:0016049 | cell growth | 1.26e-04 |
| SGK1 | GO:0010506 | regulation of autophagy | 1.57e-04 |
| SGK1 | GO:0010038 | response to metal ion | 1.57e-04 |
| SGK1 | GO:0010821 | regulation of mitochondrion organization | 4.93e-04 |
| SGK1 | GO:0071375 | cellular response to peptide hormone stimulus | 9.00e-04 |
| SGK1 | GO:0031647 | regulation of protein stability | 1.01e-03 |
| SGK1 | GO:0006913 | nucleocytoplasmic transport | 1.01e-03 |
| SGK1 | GO:0051169 | nuclear transport | 1.01e-03 |
| SGK1 | GO:0071496 | cellular response to external stimulus | 1.23e-03 |
| SGK1 | GO:0032869 | cellular response to insulin stimulus | 1.36e-03 |
| SGK1 | GO:0045732 | positive regulation of protein catabolic process | 1.40e-03 |
| SGK1 | GO:1903828 | negative regulation of protein localization | 1.51e-03 |
| SGK1 | GO:1901653 | cellular response to peptide | 1.51e-03 |
| SGK1 | GO:0044058 | regulation of digestive system process | 1.87e-03 |
| SGK1 | GO:0071356 | cellular response to tumor necrosis factor | 2.01e-03 |
| SGK1 | GO:0048588 | developmental cell growth | 2.01e-03 |
| SGK1 | GO:1903146 | regulation of autophagy of mitochondrion | 2.06e-03 |
| SGK1 | GO:0001558 | regulation of cell growth | 2.15e-03 |
| SGK1 | GO:0007613 | memory | 2.15e-03 |
| SGK1 | GO:0031669 | cellular response to nutrient levels | 2.15e-03 |
| SGK1 | GO:0034599 | cellular response to oxidative stress | 2.24e-03 |
| SGK1 | GO:0034612 | response to tumor necrosis factor | 2.32e-03 |
| SGK1 | GO:0031400 | negative regulation of protein modification process | 2.38e-03 |
| SGK1 | GO:0043666 | regulation of phosphoprotein phosphatase activity | 2.51e-03 |
| SGK1 | GO:0032868 | response to insulin | 2.51e-03 |
| SGK1 | GO:0031648 | protein destabilization | 2.74e-03 |
| SGK1 | GO:0031668 | cellular response to extracellular stimulus | 2.90e-03 |
| SGK1 | GO:0071456 | cellular response to hypoxia | 3.25e-03 |
| SGK1 | GO:0043086 | negative regulation of catalytic activity | 3.31e-03 |
| SGK1 | GO:0010508 | positive regulation of autophagy | 3.32e-03 |
| SGK1 | GO:0043525 | positive regulation of neuron apoptotic process | 3.48e-03 |
| SGK1 | GO:0034614 | cellular response to reactive oxygen species | 3.48e-03 |
| SGK1 | GO:0031667 | response to nutrient levels | 3.52e-03 |
| SGK1 | GO:0036294 | cellular response to decreased oxygen levels | 3.62e-03 |
| SGK1 | GO:0034504 | protein localization to nucleus | 3.62e-03 |
| SGK1 | GO:0030002 | intracellular monoatomic anion homeostasis | 3.62e-03 |
| SGK1 | GO:0030644 | intracellular chloride ion homeostasis | 3.62e-03 |
| SGK1 | GO:0010921 | regulation of phosphatase activity | 3.85e-03 |
| SGK1 | GO:0051168 | nuclear export | 4.05e-03 |
| SGK1 | GO:0030157 | pancreatic juice secretion | 4.05e-03 |
| SGK1 | GO:0072584 | caveolin-mediated endocytosis | 4.05e-03 |
| SGK1 | GO:0048638 | regulation of developmental growth | 4.13e-03 |
| SGK1 | GO:0071453 | cellular response to oxygen levels | 4.33e-03 |
Top |
Related Drugs to AHI1_SGK1 |
 Drugs used for this fusion-positive patient. Drugs used for this fusion-positive patient. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Drug | Source | PMID |
 Distribution of the number of studies mentioning AHI1-SGK1 and kinase inhibitors the PubMed Abstract (04-01-2024) Distribution of the number of studies mentioning AHI1-SGK1 and kinase inhibitors the PubMed Abstract (04-01-2024) |
| Fusion gene - drug pair 1 | Fusion gene - drug pair 2 | PMID | Publication date | DOI | Study title |
Top |
Related Diseases to AHI1_SGK1 |
 Diseases that have this fusion gene. Diseases that have this fusion gene. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Disease | Source | PMID |
 Related diseases from the literature mentioned this fusion gene and drug. Related diseases from the literature mentioned this fusion gene and drug. (PubMed, 04-01-2024) |
| MeSH ID | MeSH term |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
Top |
Clinical Trials of the Found Drugs/Small Molecules |
 Statistics of the Clinical Trials of the Found Kinase Inibitors from clinicaltrials.gov (06-17-2024) Statistics of the Clinical Trials of the Found Kinase Inibitors from clinicaltrials.gov (06-17-2024) |
 Clinical Trials from clinicaltrials.gov (06-17-2024) Clinical Trials from clinicaltrials.gov (06-17-2024) |
| Fusion Gene | Kinase Inhibitor | NCT ID | Study Status | Phases | Disease | # Enrolment | Date |

