| UTHEALTH HOME ABOUT SBMI A-Z WEBMAIL INSIDE THE UNIVERSITY |

|
|||||||
|
Kinase Fusion Gene:APPL2_CIT |
Kinase Fusion Protein Summary |
 Kinase Fusion gene summary Kinase Fusion gene summary |
| Kinase Fusion partner gene information | Kinase Fusion gene name: APPL2_CIT | KinaseFusionDB ID: KFG394 | FusionGDB2.0 ID: KFG394 | Hgene | Tgene | Gene symbol | APPL2 | CIT | Gene ID | 55198 | 79947 | |
| Gene name | adaptor protein, phosphotyrosine interacting with PH domain and leucine zipper 2 | |||||||||||
| Synonyms | DIP13B | |||||||||||
| Cytomap | 12q23.3 | |||||||||||
| Type of gene | protein-coding | |||||||||||
| Description | DCC-interacting protein 13-betaDIP13 betaadapter protein containing PH domain, PTB domain and leucine zipper motif 2adaptor protein, phosphotyrosine interaction, PH domain and leucine zipper containing 2 | |||||||||||
| Modification date | 20240403 | |||||||||||
| UniProtAcc | Q8NEU8 | Q96RK1 | ||||||||||
| Ensembl transtripts involved in fusion gene | ENST ids | ENST00000258530, ENST00000551662, ENST00000539978, ENST00000546731, ENST00000549573, | ENST00000537607, ENST00000261833, ENST00000392521, | |||||||||
| Context (manual curation of fusion genes in KinaseFusionDB) | PubMed: APPL2 [Title/Abstract] AND CIT [Title/Abstract] AND fusion [Title/Abstract] | |||||||||||
| Most frequent breakpoint (based on all fusion genes of FusionGDB 2.0) | APPL2(105629737)-CIT(120198859), # samples:3 | |||||||||||
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | APPL2 | GO:0006606 | protein import into nucleus | 26583432 |
| Hgene | APPL2 | GO:0051289 | protein homotetramerization | 23055524 |
| Hgene | APPL2 | GO:2000045 | regulation of G1/S transition of mitotic cell cycle | 15016378 |
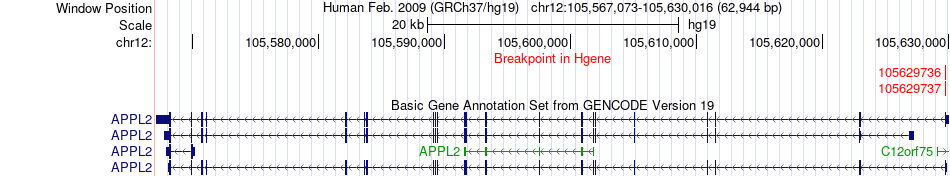
 Kinase Fusion gene breakpoints across APPL2 (5'-gene) Kinase Fusion gene breakpoints across APPL2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
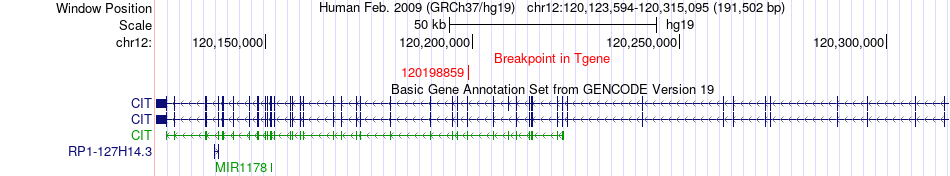
 Kinase Fusion gene breakpoints across CIT (3'-gene) Kinase Fusion gene breakpoints across CIT (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
Top |
Kinase Fusion Gene Sample Information |
 Kinase Fusion gene information. Kinase Fusion gene information. |
 Kinase Fusion gene information from four resources (ChiTars 5.0, ChimerDB 4.0, COSMIC, and CCLE) Kinase Fusion gene information from four resources (ChiTars 5.0, ChimerDB 4.0, COSMIC, and CCLE)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Sample | Hgene | Hchr | Hbp | Tgene | Tchr | Tbp |
| ChimerDB4 | TCGA-34-5232-01A | APPL2 | chr12 | 105629737 | CIT | chr12 | 120198859 |
| ChimerDB4 | TCGA-34-5232 | APPL2 | chr12 | 105629736 | CIT | chr12 | 120198859 |
Top |
Kinase Fusion ORF Analysis |
 Kinase Fusion information from ORFfinder translation from full-length transcript sequence from KinaseFusionDB. Kinase Fusion information from ORFfinder translation from full-length transcript sequence from KinaseFusionDB. |
| Henst | Tenst | Hgene | Hchr | Hbp | Tgene | Tchr | Tbp | Seq length (transcript) | Seq length (amino acids) |
| ENST00000258530 | ENST00000392521 | APPL2 | chr12 | 105629737 | CIT | chr12 | 120198859 | 6628 | 1368 |
| ENST00000258530 | ENST00000392521 | APPL2 | chr12 | 105629736 | CIT | chr12 | 120198859 | 6628 | 1368 |
Top |
Kinase Fusion Amino Acid Sequences |
 For individual full-length fusion transcript sequence from KinaseFusionDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. For individual full-length fusion transcript sequence from KinaseFusionDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >Henst_Tenst_Hgene_Hchr_Hbp_Tgene_Tchr_Tbp_length(fusion AA)_AAseq >ENST00000258530_ENST00000392521_APPL2_chr12_105629737_CIT_chr12_120198859_length(amino acids)=1368 MARRGRARGRRGHGVAAASPEPRAGECSRPPRLLPSVLPTQPSAEPRRTMPAVDKLLLEEALQDSPQVLDNQIKKDLADKETLENMMQRH EEEAHEKGKILSEQKAMINAMDSKIRSLEQRIVELSEANKLAANSSLFTQRNMKAQEEMISELRQQKFYLETQAGKLEAQNRKLEEQLEK ISHQDHSDKNRLLELETRLREVSLEHEEQKLELKRQLTELQLSLQERESQLTALQAARAALESQLRQAKTELEETTAEAEEEIQALTAHR DEIQRKFDALRNSCTVITDLEEQLNQLTEDNAELNNQNFYLSKQLDEASGANDEIVQLRSEVDHLRREITEREMQLTSQKQTMEALKTTC TMLEEQVMDLEALNDELLEKERQWEAWRSVLGDEKSQFECRVRELQRMLDTEKQSRARADQRITESRQVVELAVKEHKAEILALQQALKE QKLKAESLSDKLNDLEKKHAMLEMNARSLQQKLETERELKQRLLEEQAKLQQQMDLQKNHIFRLTQGLQEALDRADLLKTERSDLEYQLE NIQVLYSHEKVKMEGTISQQTKLIDFLQAKMDQPAKKKKGLFSRRKEDPALPTQVPLQYNELKLALEKEKARCAELEEALQKTRIELRSA REEAAHRKATDHPHPSTPATARQQIAMSAIVRSPEHQPSAMSLLAPPSSRRKESSTPEEFSRRLKERMHHNIPHRFNVGLNMRATKCAVC LDTVHFGRQASKCLECQVMCHPKCSTCLPATCGLPAEYATHFTEAFCRDKMNSPGLQTKEPSSSLHLEGWMKVPRNNKRGQQGWDRKYIV LEGSKVLIYDNEAREAGQRPVEEFELCLPDGDVSIHGAVGASELANTAKADVPYILKMESHPHTTCWPGRTLYLLAPSFPDKQRWVTALE SVVAGGRVSREKAEADAKLLGNSLLKLEGDDRLDMNCTLPFSDQVVLVGTEEGLYALNVLKNSLTHVPGIGAVFQIYIIKDLEKLLMIAG EERALCLVDVKKVKQSLAQSHLPAQPDISPNIFEAVKGCHLFGAGKIENGLCICAAMPSKVVILRYNENLSKYCIRKEIETSEPCSCIHF TNYSILIGTNKFYEIDMKQYTLEEFLDKNDHSLAPAVFAASSNSFPVSIVQVNSAGQREEYLLCFHEFGVFVDSYGRRSRTDDLKWSRLP LAFAYREPYLFVTHFNSLEVIEIQARSSAGTPARAYLDIPNPRYLGPAISSGAIYLASSYQDKLRVICCKGNLVKESGTEHHRGPSTSRS SPNKRGPPTYNEHITKRVASSPAPPEGPSHPREPSTPHRYREGRTELRRDKSPGRPLEREKSPGRMLSTRRERSPGRLFEDSSRGRLPAG -------------------------------------------------------------- >ENST00000258530_ENST00000392521_APPL2_chr12_105629736_CIT_chr12_120198859_length(amino acids)=1368 MARRGRARGRRGHGVAAASPEPRAGECSRPPRLLPSVLPTQPSAEPRRTMPAVDKLLLEEALQDSPQVLDNQIKKDLADKETLENMMQRH EEEAHEKGKILSEQKAMINAMDSKIRSLEQRIVELSEANKLAANSSLFTQRNMKAQEEMISELRQQKFYLETQAGKLEAQNRKLEEQLEK ISHQDHSDKNRLLELETRLREVSLEHEEQKLELKRQLTELQLSLQERESQLTALQAARAALESQLRQAKTELEETTAEAEEEIQALTAHR DEIQRKFDALRNSCTVITDLEEQLNQLTEDNAELNNQNFYLSKQLDEASGANDEIVQLRSEVDHLRREITEREMQLTSQKQTMEALKTTC TMLEEQVMDLEALNDELLEKERQWEAWRSVLGDEKSQFECRVRELQRMLDTEKQSRARADQRITESRQVVELAVKEHKAEILALQQALKE QKLKAESLSDKLNDLEKKHAMLEMNARSLQQKLETERELKQRLLEEQAKLQQQMDLQKNHIFRLTQGLQEALDRADLLKTERSDLEYQLE NIQVLYSHEKVKMEGTISQQTKLIDFLQAKMDQPAKKKKGLFSRRKEDPALPTQVPLQYNELKLALEKEKARCAELEEALQKTRIELRSA REEAAHRKATDHPHPSTPATARQQIAMSAIVRSPEHQPSAMSLLAPPSSRRKESSTPEEFSRRLKERMHHNIPHRFNVGLNMRATKCAVC LDTVHFGRQASKCLECQVMCHPKCSTCLPATCGLPAEYATHFTEAFCRDKMNSPGLQTKEPSSSLHLEGWMKVPRNNKRGQQGWDRKYIV LEGSKVLIYDNEAREAGQRPVEEFELCLPDGDVSIHGAVGASELANTAKADVPYILKMESHPHTTCWPGRTLYLLAPSFPDKQRWVTALE SVVAGGRVSREKAEADAKLLGNSLLKLEGDDRLDMNCTLPFSDQVVLVGTEEGLYALNVLKNSLTHVPGIGAVFQIYIIKDLEKLLMIAG EERALCLVDVKKVKQSLAQSHLPAQPDISPNIFEAVKGCHLFGAGKIENGLCICAAMPSKVVILRYNENLSKYCIRKEIETSEPCSCIHF TNYSILIGTNKFYEIDMKQYTLEEFLDKNDHSLAPAVFAASSNSFPVSIVQVNSAGQREEYLLCFHEFGVFVDSYGRRSRTDDLKWSRLP LAFAYREPYLFVTHFNSLEVIEIQARSSAGTPARAYLDIPNPRYLGPAISSGAIYLASSYQDKLRVICCKGNLVKESGTEHHRGPSTSRS SPNKRGPPTYNEHITKRVASSPAPPEGPSHPREPSTPHRYREGRTELRRDKSPGRPLEREKSPGRMLSTRRERSPGRLFEDSSRGRLPAG -------------------------------------------------------------- |
Multiple Sequence Alignment of All Fusion Protein Isoforms |
Top |
Kinase Fusion Protein Functional Features |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr12:105629737/chr12:120198859) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| APPL2 | CIT |
| FUNCTION: Multifunctional adapter protein that binds to various membrane receptors, nuclear factors and signaling proteins to regulate many processes, such as cell proliferation, immune response, endosomal trafficking and cell metabolism (PubMed:26583432, PubMed:15016378, PubMed:24879834). Regulates signaling pathway leading to cell proliferation through interaction with RAB5A and subunits of the NuRD/MeCP1 complex (PubMed:15016378). Plays a role in immune response by modulating phagocytosis, inflammatory and innate immune responses. In macrophages, enhances Fc-gamma receptor-mediated phagocytosis through interaction with RAB31 leading to activation of PI3K/Akt signaling. In response to LPS, modulates inflammatory responses by playing a key role on the regulation of TLR4 signaling and in the nuclear translocation of RELA/NF-kappa-B p65 and the secretion of pro- and anti-inflammatory cytokines. Also functions as a negative regulator of innate immune response via inhibition of AKT1 signaling pathway by forming a complex with APPL1 and PIK3R1 (By similarity). Plays a role in endosomal trafficking of TGFBR1 from the endosomes to the nucleus (PubMed:26583432). Plays a role in cell metabolism by regulating adiponecting ans insulin signaling pathways and adaptative thermogenesis (PubMed:24879834) (By similarity). In muscle, negatively regulates adiponectin-simulated glucose uptake and fatty acid oxidation by inhibiting adiponectin signaling pathway through APPL1 sequestration thereby antagonizing APPL1 action (By similarity). In muscles, negativeliy regulates insulin-induced plasma membrane recruitment of GLUT4 and glucose uptake through interaction with TBC1D1 (PubMed:24879834). Plays a role in cold and diet-induced adaptive thermogenesis by activating ventromedial hypothalamus (VMH) neurons throught AMPK inhibition which enhances sympathetic outflow to subcutaneous white adipose tissue (sWAT), sWAT beiging and cold tolerance (By similarity). Also plays a role in other signaling pathways namely Wnt/beta-catenin, HGF and glucocorticoid receptor signaling (PubMed:19433865) (By similarity). Positive regulator of beta-catenin/TCF-dependent transcription through direct interaction with RUVBL2/reptin resulting in the relief of RUVBL2-mediated repression of beta-catenin/TCF target genes by modulating the interactions within the beta-catenin-reptin-HDAC complex (PubMed:19433865). May affect adult neurogenesis in hippocampus and olfactory system via regulating the sensitivity of glucocorticoid receptor. Required for fibroblast migration through HGF cell signaling (By similarity). {ECO:0000250|UniProtKB:Q8K3G9, ECO:0000269|PubMed:15016378, ECO:0000269|PubMed:19433865, ECO:0000269|PubMed:24879834, ECO:0000269|PubMed:26583432}. | FUNCTION: Acts as a transcriptional coactivator for TFAP2/AP-2. Enhances estrogen-dependent transactivation mediated by estrogen receptors. May function as an inhibitor of transactivation by HIF1A by disrupting HIF1A interaction with CREBBP. May be involved in regulation of gene expression during development and differentiation of blood cells, endothelial cells and mammary epithelial cells. {ECO:0000269|PubMed:11744733, ECO:0000269|PubMed:15342390}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
 - Retained domain in the 5'-partner of fusion protein (protein functional feature from UniProt). - Retained domain in the 5'-partner of fusion protein (protein functional feature from UniProt). |
| Partner | Hgeneene | Hbp | Tgeneene | Tbp | ENST | BPexon | TotalExon | Protein feature loci | BPloci | TotalLen | Feature | Note |
 - Retained domain in the 3'-partner of fusion protein (protein functional feature from UniProt). - Retained domain in the 3'-partner of fusion protein (protein functional feature from UniProt). |
| Partner | Hgeneene | Hbp | Tgeneene | Tbp | ENST | BPexon | TotalExon | Protein feature loci | BPloci | TotalLen | Feature | Note |
| Tgene | APPL2 | 105629736 | CIT | 120198859 | ENST00000258530 | 17 | 47 | 1591_1881 | 726 | 2028 | Domain | Note=CNH;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00795 |
| Tgene | APPL2 | 105629736 | CIT | 120198859 | ENST00000258530 | 18 | 48 | 1591_1881 | 768 | 2070 | Domain | Note=CNH;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00795 |
| Tgene | APPL2 | 105629737 | CIT | 120198859 | ENST00000258530 | 17 | 47 | 1591_1881 | 726 | 2028 | Domain | Note=CNH;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00795 |
| Tgene | APPL2 | 105629737 | CIT | 120198859 | ENST00000258530 | 18 | 48 | 1591_1881 | 768 | 2070 | Domain | Note=CNH;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00795 |
| Tgene | APPL2 | 105629736 | CIT | 120198859 | ENST00000258530 | 17 | 47 | 1443_1563 | 726 | 2028 | Domain | Note=PH;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00145 |
| Tgene | APPL2 | 105629736 | CIT | 120198859 | ENST00000258530 | 18 | 48 | 1443_1563 | 768 | 2070 | Domain | Note=PH;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00145 |
| Tgene | APPL2 | 105629737 | CIT | 120198859 | ENST00000258530 | 17 | 47 | 1443_1563 | 726 | 2028 | Domain | Note=PH;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00145 |
| Tgene | APPL2 | 105629737 | CIT | 120198859 | ENST00000258530 | 18 | 48 | 1443_1563 | 768 | 2070 | Domain | Note=PH;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00145 |
Top |
Kinase Fusion Protein Structures |
 CIF files of the predicted kinase fusion proteins CIF files of the predicted kinase fusion proteins * Here we show the 3D structure of the fusion proteins using Mol*. AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. Model confidence is shown from the pLDDT values per residue. pLDDT corresponds to the model’s prediction of its score on the local Distance Difference Test. It is a measure of local accuracy (from AlphfaFold website). To color code individual residues, we transformed individual PDB files into CIF format. |
| Kinase Fusion protein CIF link (fusion AA seq ID in KinaseFusionDB) | Henst | Tenst | Hgene | Hchr | Hbp | Tgene | Tchr | Tbp | AA seq | Len(AA seq) |
| PDB file >>>31_APPL2_CIT | ENST00000258530 | ENST00000392521 | APPL2 | chr12 | 105629736 | CIT | chr12 | 120198859 | MARRGRARGRRGHGVAAASPEPRAGECSRPPRLLPSVLPTQPSAEPRRTMPAVDKLLLEEALQDSPQVLDNQIKKDLADKETLENMMQRH EEEAHEKGKILSEQKAMINAMDSKIRSLEQRIVELSEANKLAANSSLFTQRNMKAQEEMISELRQQKFYLETQAGKLEAQNRKLEEQLEK ISHQDHSDKNRLLELETRLREVSLEHEEQKLELKRQLTELQLSLQERESQLTALQAARAALESQLRQAKTELEETTAEAEEEIQALTAHR DEIQRKFDALRNSCTVITDLEEQLNQLTEDNAELNNQNFYLSKQLDEASGANDEIVQLRSEVDHLRREITEREMQLTSQKQTMEALKTTC TMLEEQVMDLEALNDELLEKERQWEAWRSVLGDEKSQFECRVRELQRMLDTEKQSRARADQRITESRQVVELAVKEHKAEILALQQALKE QKLKAESLSDKLNDLEKKHAMLEMNARSLQQKLETERELKQRLLEEQAKLQQQMDLQKNHIFRLTQGLQEALDRADLLKTERSDLEYQLE NIQVLYSHEKVKMEGTISQQTKLIDFLQAKMDQPAKKKKGLFSRRKEDPALPTQVPLQYNELKLALEKEKARCAELEEALQKTRIELRSA REEAAHRKATDHPHPSTPATARQQIAMSAIVRSPEHQPSAMSLLAPPSSRRKESSTPEEFSRRLKERMHHNIPHRFNVGLNMRATKCAVC LDTVHFGRQASKCLECQVMCHPKCSTCLPATCGLPAEYATHFTEAFCRDKMNSPGLQTKEPSSSLHLEGWMKVPRNNKRGQQGWDRKYIV LEGSKVLIYDNEAREAGQRPVEEFELCLPDGDVSIHGAVGASELANTAKADVPYILKMESHPHTTCWPGRTLYLLAPSFPDKQRWVTALE SVVAGGRVSREKAEADAKLLGNSLLKLEGDDRLDMNCTLPFSDQVVLVGTEEGLYALNVLKNSLTHVPGIGAVFQIYIIKDLEKLLMIAG EERALCLVDVKKVKQSLAQSHLPAQPDISPNIFEAVKGCHLFGAGKIENGLCICAAMPSKVVILRYNENLSKYCIRKEIETSEPCSCIHF TNYSILIGTNKFYEIDMKQYTLEEFLDKNDHSLAPAVFAASSNSFPVSIVQVNSAGQREEYLLCFHEFGVFVDSYGRRSRTDDLKWSRLP LAFAYREPYLFVTHFNSLEVIEIQARSSAGTPARAYLDIPNPRYLGPAISSGAIYLASSYQDKLRVICCKGNLVKESGTEHHRGPSTSRS SPNKRGPPTYNEHITKRVASSPAPPEGPSHPREPSTPHRYREGRTELRRDKSPGRPLEREKSPGRMLSTRRERSPGRLFEDSSRGRLPAG | 1368 |
| 3D view using mol* of 31_APPL2_CIT | ||||||||||
| PDB file >>>TKFP_47_APPL2_CIT | ENST00000258530 | ENST00000392521 | APPL2 | chr12 | 105629737 | CIT | chr12 | 120198859 | MARRGRARGRRGHGVAAASPEPRAGECSRPPRLLPSVLPTQPSAEPRRTMPAVDKLLLEEALQDSPQVLDNQIKKDLADKETLENMMQRH EEEAHEKGKILSEQKAMINAMDSKIRSLEQRIVELSEANKLAANSSLFTQRNMKAQEEMISELRQQKFYLETQAGKLEAQNRKLEEQLEK ISHQDHSDKNRLLELETRLREVSLEHEEQKLELKRQLTELQLSLQERESQLTALQAARAALESQLRQAKTELEETTAEAEEEIQALTAHR DEIQRKFDALRNSCTVITDLEEQLNQLTEDNAELNNQNFYLSKQLDEASGANDEIVQLRSEVDHLRREITEREMQLTSQKQTMEALKTTC TMLEEQVMDLEALNDELLEKERQWEAWRSVLGDEKSQFECRVRELQRMLDTEKQSRARADQRITESRQVVELAVKEHKAEILALQQALKE QKLKAESLSDKLNDLEKKHAMLEMNARSLQQKLETERELKQRLLEEQAKLQQQMDLQKNHIFRLTQGLQEALDRADLLKTERSDLEYQLE NIQVLYSHEKVKMEGTISQQTKLIDFLQAKMDQPAKKKKGLFSRRKEDPALPTQVPLQYNELKLALEKEKARCAELEEALQKTRIELRSA REEAAHRKATDHPHPSTPATARQQIAMSAIVRSPEHQPSAMSLLAPPSSRRKESSTPEEFSRRLKERMHHNIPHRFNVGLNMRATKCAVC LDTVHFGRQASKCLECQVMCHPKCSTCLPATCGLPAEYATHFTEAFCRDKMNSPGLQTKEPSSSLHLEGWMKVPRNNKRGQQGWDRKYIV LEGSKVLIYDNEAREAGQRPVEEFELCLPDGDVSIHGAVGASELANTAKADVPYILKMESHPHTTCWPGRTLYLLAPSFPDKQRWVTALE SVVAGGRVSREKAEADAKLLGNSLLKLEGDDRLDMNCTLPFSDQVVLVGTEEGLYALNVLKNSLTHVPGIGAVFQIYIIKDLEKLLMIAG EERALCLVDVKKVKQSLAQSHLPAQPDISPNIFEAVKGCHLFGAGKIENGLCICAAMPSKVVILRYNENLSKYCIRKEIETSEPCSCIHF TNYSILIGTNKFYEIDMKQYTLEEFLDKNDHSLAPAVFAASSNSFPVSIVQVNSAGQREEYLLCFHEFGVFVDSYGRRSRTDDLKWSRLP LAFAYREPYLFVTHFNSLEVIEIQARSSAGTPARAYLDIPNPRYLGPAISSGAIYLASSYQDKLRVICCKGNLVKESGTEHHRGPSTSRS SPNKRGPPTYNEHITKRVASSPAPPEGPSHPREPSTPHRYREGRTELRRDKSPGRPLEREKSPGRMLSTRRERSPGRLFEDSSRGRLPAG | 1368_APPL2_CIT |
| PDB file >>>TKFP_48_APPL2_CIT | ENST00000258530 | ENST00000392521 | APPL2 | chr12 | 105629736 | CIT | chr12 | 120198859 | MARRGRARGRRGHGVAAASPEPRAGECSRPPRLLPSVLPTQPSAEPRRTMPAVDKLLLEEALQDSPQVLDNQIKKDLADKETLENMMQRH EEEAHEKGKILSEQKAMINAMDSKIRSLEQRIVELSEANKLAANSSLFTQRNMKAQEEMISELRQQKFYLETQAGKLEAQNRKLEEQLEK ISHQDHSDKNRLLELETRLREVSLEHEEQKLELKRQLTELQLSLQERESQLTALQAARAALESQLRQAKTELEETTAEAEEEIQALTAHR DEIQRKFDALRNSCTVITDLEEQLNQLTEDNAELNNQNFYLSKQLDEASGANDEIVQLRSEVDHLRREITEREMQLTSQKQTMEALKTTC TMLEEQVMDLEALNDELLEKERQWEAWRSVLGDEKSQFECRVRELQRMLDTEKQSRARADQRITESRQVVELAVKEHKAEILALQQALKE QKLKAESLSDKLNDLEKKHAMLEMNARSLQQKLETERELKQRLLEEQAKLQQQMDLQKNHIFRLTQGLQEALDRADLLKTERSDLEYQLE NIQVLYSHEKVKMEGTISQQTKLIDFLQAKMDQPAKKKKGLFSRRKEDPALPTQVPLQYNELKLALEKEKARCAELEEALQKTRIELRSA REEAAHRKATDHPHPSTPATARQQIAMSAIVRSPEHQPSAMSLLAPPSSRRKESSTPEEFSRRLKERMHHNIPHRFNVGLNMRATKCAVC LDTVHFGRQASKCLECQVMCHPKCSTCLPATCGLPAEYATHFTEAFCRDKMNSPGLQTKEPSSSLHLEGWMKVPRNNKRGQQGWDRKYIV LEGSKVLIYDNEAREAGQRPVEEFELCLPDGDVSIHGAVGASELANTAKADVPYILKMESHPHTTCWPGRTLYLLAPSFPDKQRWVTALE SVVAGGRVSREKAEADAKLLGNSLLKLEGDDRLDMNCTLPFSDQVVLVGTEEGLYALNVLKNSLTHVPGIGAVFQIYIIKDLEKLLMIAG EERALCLVDVKKVKQSLAQSHLPAQPDISPNIFEAVKGCHLFGAGKIENGLCICAAMPSKVVILRYNENLSKYCIRKEIETSEPCSCIHF TNYSILIGTNKFYEIDMKQYTLEEFLDKNDHSLAPAVFAASSNSFPVSIVQVNSAGQREEYLLCFHEFGVFVDSYGRRSRTDDLKWSRLP LAFAYREPYLFVTHFNSLEVIEIQARSSAGTPARAYLDIPNPRYLGPAISSGAIYLASSYQDKLRVICCKGNLVKESGTEHHRGPSTSRS SPNKRGPPTYNEHITKRVASSPAPPEGPSHPREPSTPHRYREGRTELRRDKSPGRPLEREKSPGRMLSTRRERSPGRLFEDSSRGRLPAG | 1368_APPL2_CIT |
Top |
Comparison of Fusion Protein Isoforms |
 Superimpose the 3D Structures Among All Fusion Protein Isoforms Superimpose the 3D Structures Among All Fusion Protein Isoforms * Download the pdb file and open it from the molstar online viewer. |
 Comparison of the Secondary Structures of Fusion Protein Isoforms Comparison of the Secondary Structures of Fusion Protein Isoforms |
Top |
Comparison of Fusion Protein Sequences/Structures with Known Sequences/Structures from PDB |
Top |
pLDDT score distribution |
 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2* AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. * The blue color at the bottom marks the best active site residues. |
| 31_APPL2_CIT.png |
 |
| 31_APPL2_CIT.png |
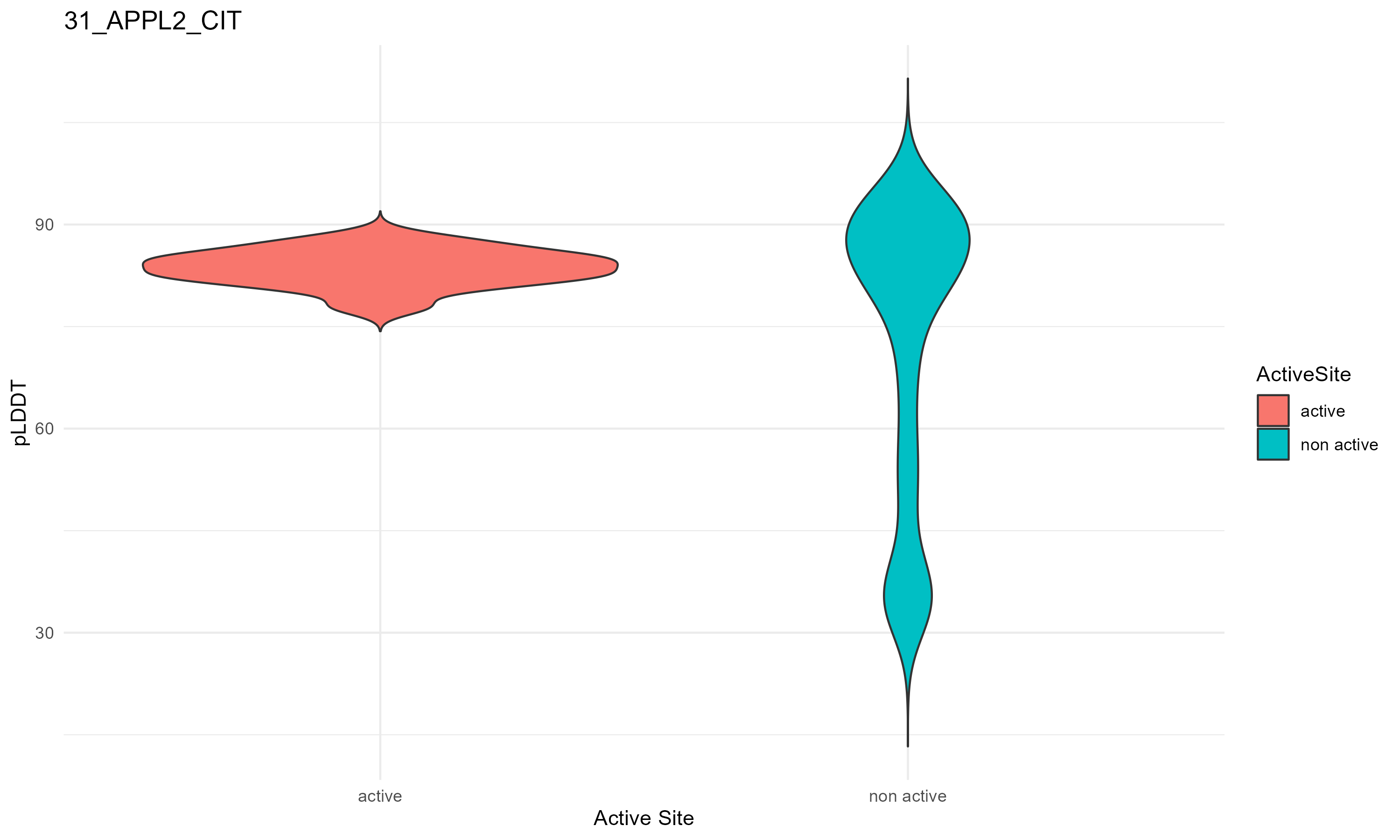 |
Top |
Potential Active Site Information |
 The potential binding sites of these fusion proteins were identified using SiteMap, a module of the Schrodinger suite. The potential binding sites of these fusion proteins were identified using SiteMap, a module of the Schrodinger suite. |
| Kinase Fusion AA seq ID in KinaseFusionDB | Site score | Size | Dscore | Volume | Exposure | Enclosure | Contact | Phobic | Philic | Balance | Don/Acc | Residues |
Top |
Ramachandran Plot of Kinase Fusion Protein Structure |
 Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. |
| 31_APPL2_CIT_ramachandran.png |
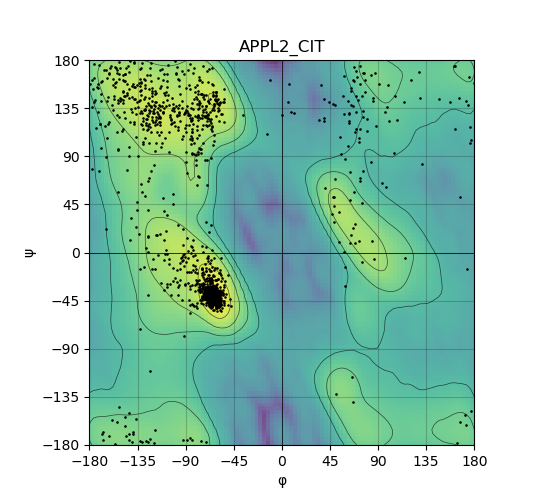 |
Top |
Virtual Screening Results |
 Distribution of the average docking score across all approved kinase inhibitors. Distribution of the average docking score across all approved kinase inhibitors.Distribution of the number of occurrence across all approved kinase inhibitors. |
| 5'-kinase fusion protein case |
| 3'-kinase fusion protein case |
Top |
 Drug information from DrugBank of the top 20 interacting small molecules. Drug information from DrugBank of the top 20 interacting small molecules.* The detailed information of individual kinase inhibitors are available in the download page. |
| Fusion gene name info | Drug | Docking score | Glide g score | Glide energy |
Top |
Kinase-Substrate Information of APPL2_CIT |
 Phosphorylation target of the kinase Phosphorylation target of the kinase(phosphosite, 03-17-2024) |
| Kinase | Kinase UniProt Acc | Kinase species | Substrate | Substrate UniProt Acc | Substrate phosphorylated residues | Substrate phosphorylated sites (+/-7AA) | Domain |
| CIT | O14578 | human | GLI2 | P10070 | S149 | IsAArGLsPADVAQE | |
| CIT | O14578 | human | MYL9 | P24844 | T19 | KKRPQRAtsNVFAMF | |
| CIT | O14578 | human | MYL9 | P24844 | S20 | KRPQRAtsNVFAMFD |
 Biological Network Integration of This Kinase and Substrates Biological Network Integration of This Kinase and Substrates (GeneMANIA website) |
 Enriched GO biological processes of the phosphorylation target genes of the kinase Enriched GO biological processes of the phosphorylation target genes of the kinase |
| Kinase | GOID | GO term | P.adjust |
| CIT | ID | Description | 0.00e+00 |
| CIT | GO:0033089 | positive regulation of T cell differentiation in thymus | 2.55e-02 |
| CIT | GO:0033504 | floor plate development | 2.55e-02 |
| CIT | GO:0060579 | ventral spinal cord interneuron fate commitment | 2.55e-02 |
| CIT | GO:0060581 | cell fate commitment involved in pattern specification | 2.55e-02 |
| CIT | GO:0021514 | ventral spinal cord interneuron differentiation | 2.55e-02 |
| CIT | GO:0002076 | osteoblast development | 2.55e-02 |
| CIT | GO:0021513 | spinal cord dorsal/ventral patterning | 2.55e-02 |
| CIT | GO:0021511 | spinal cord patterning | 2.55e-02 |
| CIT | GO:0033081 | regulation of T cell differentiation in thymus | 2.55e-02 |
| CIT | GO:0021952 | central nervous system projection neuron axonogenesis | 2.55e-02 |
| CIT | GO:0031069 | hair follicle morphogenesis | 2.55e-02 |
| CIT | GO:0048566 | embryonic digestive tract development | 2.55e-02 |
| CIT | GO:0009954 | proximal/distal pattern formation | 2.55e-02 |
| CIT | GO:0048665 | neuron fate specification | 2.55e-02 |
| CIT | GO:0021696 | cerebellar cortex morphogenesis | 2.55e-02 |
| CIT | GO:0048730 | epidermis morphogenesis | 2.55e-02 |
| CIT | GO:0045740 | positive regulation of DNA replication | 2.55e-02 |
| CIT | GO:0021955 | central nervous system neuron axonogenesis | 2.55e-02 |
| CIT | GO:0021983 | pituitary gland development | 2.55e-02 |
| CIT | GO:0021587 | cerebellum morphogenesis | 2.55e-02 |
| CIT | GO:0021517 | ventral spinal cord development | 2.55e-02 |
| CIT | GO:0021575 | hindbrain morphogenesis | 2.55e-02 |
| CIT | GO:0048546 | digestive tract morphogenesis | 2.55e-02 |
| CIT | GO:0021515 | cell differentiation in spinal cord | 2.55e-02 |
| CIT | GO:0021695 | cerebellar cortex development | 2.55e-02 |
| CIT | GO:0030239 | myofibril assembly | 3.14e-02 |
| CIT | GO:0070527 | platelet aggregation | 3.14e-02 |
| CIT | GO:0048663 | neuron fate commitment | 3.14e-02 |
| CIT | GO:0055002 | striated muscle cell development | 3.14e-02 |
| CIT | GO:0021536 | diencephalon development | 3.15e-02 |
| CIT | GO:0033077 | T cell differentiation in thymus | 3.15e-02 |
| CIT | GO:0021954 | central nervous system neuron development | 3.15e-02 |
| CIT | GO:0001942 | hair follicle development | 3.15e-02 |
| CIT | GO:0042475 | odontogenesis of dentin-containing tooth | 3.15e-02 |
| CIT | GO:0022404 | molting cycle process | 3.15e-02 |
| CIT | GO:0022405 | hair cycle process | 3.15e-02 |
| CIT | GO:0034109 | homotypic cell-cell adhesion | 3.15e-02 |
| CIT | GO:0009953 | dorsal/ventral pattern formation | 3.15e-02 |
| CIT | GO:0021510 | spinal cord development | 3.15e-02 |
| CIT | GO:0021549 | cerebellum development | 3.15e-02 |
| CIT | GO:0001708 | cell fate specification | 3.15e-02 |
| CIT | GO:0022037 | metencephalon development | 3.15e-02 |
| CIT | GO:0045582 | positive regulation of T cell differentiation | 3.15e-02 |
| CIT | GO:0042303 | molting cycle | 3.15e-02 |
| CIT | GO:0042633 | hair cycle | 3.15e-02 |
| CIT | GO:0006275 | regulation of DNA replication | 3.15e-02 |
| CIT | GO:0010927 | cellular component assembly involved in morphogenesis | 3.15e-02 |
| CIT | GO:0098773 | skin epidermis development | 3.15e-02 |
| CIT | GO:0045621 | positive regulation of lymphocyte differentiation | 3.15e-02 |
Top |
Related Drugs to APPL2_CIT |
 Drugs used for this fusion-positive patient. Drugs used for this fusion-positive patient. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Drug | Source | PMID |
 Distribution of the number of studies mentioning APPL2-CIT and kinase inhibitors the PubMed Abstract (04-01-2024) Distribution of the number of studies mentioning APPL2-CIT and kinase inhibitors the PubMed Abstract (04-01-2024) |
| Fusion gene - drug pair 1 | Fusion gene - drug pair 2 | PMID | Publication date | DOI | Study title |
Top |
Related Diseases to APPL2_CIT |
 Diseases that have this fusion gene. Diseases that have this fusion gene. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Disease | Source | PMID |
 Related diseases from the literature mentioned this fusion gene and drug. Related diseases from the literature mentioned this fusion gene and drug. (PubMed, 04-01-2024) |
| MeSH ID | MeSH term |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
Top |
Clinical Trials of the Found Drugs/Small Molecules |
 Statistics of the Clinical Trials of the Found Kinase Inibitors from clinicaltrials.gov (06-17-2024) Statistics of the Clinical Trials of the Found Kinase Inibitors from clinicaltrials.gov (06-17-2024) |
 Clinical Trials from clinicaltrials.gov (06-17-2024) Clinical Trials from clinicaltrials.gov (06-17-2024) |
| Fusion Gene | Kinase Inhibitor | NCT ID | Study Status | Phases | Disease | # Enrolment | Date |

