| UTHEALTH HOME ABOUT SBMI A-Z WEBMAIL INSIDE THE UNIVERSITY |

|
|||||||
|
Kinase Fusion Gene:PAN3_FLT3 |
Kinase Fusion Protein Summary |
 Kinase Fusion gene summary Kinase Fusion gene summary |
| Kinase Fusion partner gene information | Kinase Fusion gene name: PAN3_FLT3 | KinaseFusionDB ID: KFG4472 | FusionGDB2.0 ID: KFG4472 | Hgene | Tgene | Gene symbol | PAN3 | FLT3 | Gene ID | 255967 | 2322 | |
| Gene name | poly(A) specific ribonuclease subunit PAN3 | fms related receptor tyrosine kinase 3 | ||||||||||
| Synonyms | - | CD135|FLK-2|FLK2|STK1 | ||||||||||
| Cytomap | 13q12.2 | 13q12.2 | ||||||||||
| Type of gene | protein-coding | protein-coding | ||||||||||
| Description | PAN2-PAN3 deadenylation complex subunit PAN3PAB-dependent poly(A)-specific ribonuclease subunit 3PAB-dependent poly(A)-specific ribonuclease subunit PAN3PAB1P-dependent poly(A)-nucleasePAB1P-dependent poly(A)-specific ribonucleasePABP-dependent poly( | receptor-type tyrosine-protein kinase FLT3CD135 antigenFL cytokine receptorfetal liver kinase 2fms related tyrosine kinase 3fms-like tyrosine kinase 3growth factor receptor tyrosine kinase type IIIstem cell tyrosine kinase 1 | ||||||||||
| Modification date | 20240403 | 20240411 | ||||||||||
| UniProtAcc | Q58A45 | P36888 | ||||||||||
| Ensembl transtripts involved in fusion gene | ENST ids | ENST00000380958, ENST00000399613, ENST00000282391, ENST00000483842, | ENST00000469894, ENST00000241453, ENST00000380982, ENST00000537084, | |||||||||
| Context (manual curation of fusion genes in KinaseFusionDB) | PubMed: PAN3 [Title/Abstract] AND FLT3 [Title/Abstract] AND fusion [Title/Abstract] | |||||||||||
| Most frequent breakpoint (based on all fusion genes of FusionGDB 2.0) | PAN3(28752072)-FLT3(28644749), # samples:3 | |||||||||||
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | FLT3 | GO:0030097 | hemopoiesis | 7507245 |
 Kinase Fusion gene breakpoints across PAN3 (5'-gene) Kinase Fusion gene breakpoints across PAN3 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
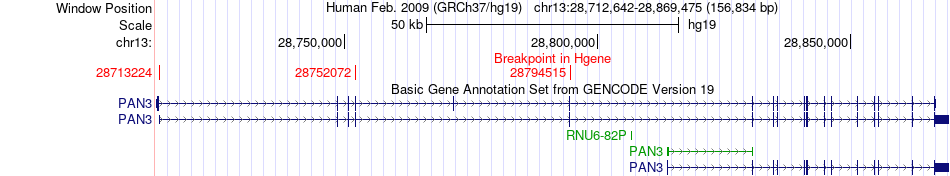 |
 Kinase Fusion gene breakpoints across FLT3 (3'-gene) Kinase Fusion gene breakpoints across FLT3 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
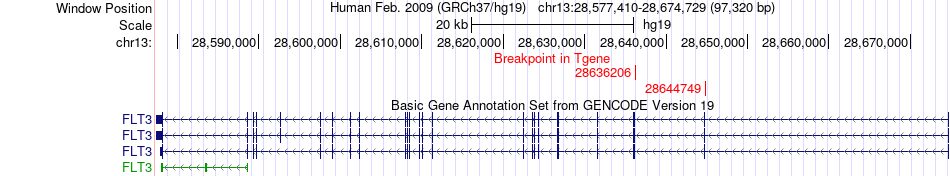 |
Top |
Kinase Fusion Gene Sample Information |
 Kinase Fusion gene information. Kinase Fusion gene information. |
 Kinase Fusion gene information from four resources (ChiTars 5.0, ChimerDB 4.0, COSMIC, and CCLE) Kinase Fusion gene information from four resources (ChiTars 5.0, ChimerDB 4.0, COSMIC, and CCLE)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Sample | Hgene | Hchr | Hbp | Tgene | Tchr | Tbp |
| ChimerDB4 | TCGA-AA-3712-01A | PAN3 | chr13 | 28752072 | FLT3 | chr13 | 28644749 |
| ChimerDB4 | TCGA-AB-2853-03A | PAN3 | chr13 | 28794515 | FLT3 | chr13 | 28644749 |
| ChimerDB4 | TCGA-HS-A5NA-01A | PAN3 | chr13 | 28713224 | FLT3 | chr13 | 28636206 |
| ChimerDB4 | TCGA-HS-A5NA-01A | PAN3 | chr13 | 28713224 | FLT3 | chr13 | 28644749 |
| CCLE | SEMK2 | PAN3 | chr13 | 28835595 | FLT3 | chr13 | 28609810 |
Top |
Kinase Fusion ORF Analysis |
 Kinase Fusion information from ORFfinder translation from full-length transcript sequence from KinaseFusionDB. Kinase Fusion information from ORFfinder translation from full-length transcript sequence from KinaseFusionDB. |
| Henst | Tenst | Hgene | Hchr | Hbp | Tgene | Tchr | Tbp | Seq length (transcript) | Seq length (amino acids) |
| ENST00000380958 | ENST00000241453 | PAN3 | chr13 | 28794515 | FLT3 | chr13 | 28644749 | 4869 | 1312 |
Top |
Kinase Fusion Amino Acid Sequences |
 For individual full-length fusion transcript sequence from KinaseFusionDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. For individual full-length fusion transcript sequence from KinaseFusionDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >Henst_Tenst_Hgene_Hchr_Hbp_Tgene_Tchr_Tbp_length(fusion AA)_AAseq >ENST00000380958_ENST00000241453_PAN3_chr13_28794515_FLT3_chr13_28644749_length(amino acids)=1312 MNSGGGLPPPSAAASPSSSSLAAAVAVVAPPGVGGVPGGAAVGVKLKYCRYYAKDKTCFYGEECQFLHEDPAAGAAPGLGLHSNSVPLAL AGAPVAGFPPGAVAGGGAGPPPGPKKPDLGDPGTGAAAGGGGSSGGLDGPRLAIPGMDGGALTDTSLTDSYFSTSFIGVNGFGSPVETKY PLMQRMTNSSSSPSLLNDSAKPYSAHDPLTSPASSLFNDFGALNISQRRKPRKYRLGMLEERLVPMGSKARKAKNPIGCLADRCKSGVPI NMVWWNRVTENNLQTPNPTASEFIPKGGSTSRLSNVSQSNMSAFSQVFSHPSMGSPATAGLAPVVFSAMIFGTITNQDLPVIKCVLINHK NNDSSVGKSSSYPMVSESPEDLGCALRPQSSGTVYEAAAVEVDVSASITLQVLVDAPGNISCLWVFKHSSLNCQPHFDLQNRGVVSMVIL KMTETQAGEYLLFIQSEATNYTILFTVSIRNTLLYTLRRPYFRKMENQDALVCISESVPEPIVEWVLCDSQGESCKEESPAVVKKEEKVL HELFGTDIRCCARNELGRECTRLFTIDLNQTPQTTLPQLFLKVGEPLWIRCKAVHVNHGFGLTWELENKALEEGNYFEMSTYSTNRTMIR ILFAFVSSVARNDTGYYTCSSSKHPSQSALVTIVEKGFINATNSSEDYEIDQYEEFCFSVRFKAYPQIRCTWTFSRKSFPCEQKGLDNGY SISKFCNHKHQPGEYIFHAENDDAQFTKMFTLNIRRKPQVLAEASASQASCFSDGYPLPSWTWKKCSDKSPNCTEEITEGVWNRKANRKV FGQWVSSSTLNMSEAIKGFLVKCCAYNSLGTSCETILLNSPGPFPFIQDNISFYATIGVCLLFIVVLTLLICHKYKKQFRYESQLQMVQV TGSSDNEYFYVDFREYEYDLKWEFPRENLEFGKVLGSGAFGKVMNATAYGISKTGVSIQVAVKMLKEKADSSEREALMSELKMMTQLGSH ENIVNLLGACTLSGPIYLIFEYCCYGDLLNYLRSKREKFHRTWTEIFKEHNFSFYPTFQSHPNSSMPGSREVQIHPDSDQISGLHGNSFH SEDEIEYENQKRLEEEEDLNVLTFEDLLCFAYQVAKGMEFLEFKSCVHRDLAARNVLVTHGKVVKICDFGLARDIMSDSNYVVRGNARLP VKWMAPESLFEGIYTIKSDVWSYGILLWEIFSLGVNPYPGIPVDANFYKLIQNGFKMDQPFYATEEIYIIMQSCWAFDSRKRPSFPNLTS -------------------------------------------------------------- |
Multiple Sequence Alignment of All Fusion Protein Isoforms |
Top |
Kinase Fusion Protein Functional Features |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr13:28752072/chr13:28644749) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| PAN3 | FLT3 |
| FUNCTION: Regulatory subunit of the poly(A)-nuclease (PAN) deadenylation complex, one of two cytoplasmic mRNA deadenylases involved in general and miRNA-mediated mRNA turnover. PAN specifically shortens poly(A) tails of RNA and the activity is stimulated by poly(A)-binding protein (PABP). PAN deadenylation is followed by rapid degradation of the shortened mRNA tails by the CCR4-NOT complex. Deadenylated mRNAs are then degraded by two alternative mechanisms, namely exosome-mediated 3'-5' exonucleolytic degradation, or deadenylation-dependent mRNA decapping and subsequent 5'-3' exonucleolytic degradation by XRN1. PAN3 acts as a regulator for PAN activity, recruiting the catalytic subunit PAN2 to mRNA via its interaction with RNA and PABP, and to miRNA targets via its interaction with GW182 family proteins. {ECO:0000255|HAMAP-Rule:MF_03181, ECO:0000269|PubMed:14583602, ECO:0000269|PubMed:23932717}.; FUNCTION: [Isoform 1]: Decreases PAN2-mediated deadenylation, possibly by preventing progression into the second CCR4-NOT mediated stage of biphasic deadenylation. Has a significant effect on mRNA stability, generally stabilizing a subset of the transcriptome. Stabilizes mRNAs degraded by the AU-rich element (ARE)-mediated mRNA decay pathway but promotes degradation of mRNAs by the microRNA-mediated pathway (PubMed:28559491). Its activity influences mRNP remodeling, specifically reducing formation of a subset of P-bodies containing GW220, an isoform of TNRC6A (PubMed:28559491). {ECO:0000269|PubMed:28559491}.; FUNCTION: [Isoform 3]: Enhances PAN2 deadenylase activity and has an extensive effect on mRNA stability, generally enhancing mRNA decay across the transcriptome by multiple pathways, including the AU-rich element (ARE)-mediated pathway, microRNA-mediated pathway and the nonsense-mediated pathway (NMD) (PubMed:28559491). Its activity is required for efficient P-body formation (PubMed:28559491). May be involved in regulating mRNAs of genes involved in cell cycle progression and cell proliferation (PubMed:28559491). {ECO:0000269|PubMed:28559491}. | FUNCTION: Tyrosine-protein kinase that acts as a cell-surface receptor for the cytokine FLT3LG and regulates differentiation, proliferation and survival of hematopoietic progenitor cells and of dendritic cells. Promotes phosphorylation of SHC1 and AKT1, and activation of the downstream effector MTOR. Promotes activation of RAS signaling and phosphorylation of downstream kinases, including MAPK1/ERK2 and/or MAPK3/ERK1. Promotes phosphorylation of FES, FER, PTPN6/SHP, PTPN11/SHP-2, PLCG1, and STAT5A and/or STAT5B. Activation of wild-type FLT3 causes only marginal activation of STAT5A or STAT5B. Mutations that cause constitutive kinase activity promote cell proliferation and resistance to apoptosis via the activation of multiple signaling pathways. {ECO:0000269|PubMed:10080542, ECO:0000269|PubMed:11090077, ECO:0000269|PubMed:14504097, ECO:0000269|PubMed:16266983, ECO:0000269|PubMed:16627759, ECO:0000269|PubMed:18490735, ECO:0000269|PubMed:20111072, ECO:0000269|PubMed:21067588, ECO:0000269|PubMed:21262971, ECO:0000269|PubMed:21516120, ECO:0000269|PubMed:7507245}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
 - Retained domain in the 5'-partner of fusion protein (protein functional feature from UniProt). - Retained domain in the 5'-partner of fusion protein (protein functional feature from UniProt). |
| Partner | Hgeneene | Hbp | Tgeneene | Tbp | ENST | BPexon | TotalExon | Protein feature loci | BPloci | TotalLen | Feature | Note |
 - Retained domain in the 3'-partner of fusion protein (protein functional feature from UniProt). - Retained domain in the 3'-partner of fusion protein (protein functional feature from UniProt). |
| Partner | Hgeneene | Hbp | Tgeneene | Tbp | ENST | BPexon | TotalExon | Protein feature loci | BPloci | TotalLen | Feature | Note |
| Tgene | PAN3 | 28794515 | FLT3 | 28644749 | ENST00000380958 | 0 | 23 | 253_343 | 14 | 953 | Domain | Note=Ig-like C2-type |
| Tgene | PAN3 | 28794515 | FLT3 | 28644749 | ENST00000380958 | 0 | 24 | 253_343 | 14 | 994 | Domain | Note=Ig-like C2-type |
| Tgene | PAN3 | 28794515 | FLT3 | 28644749 | ENST00000380958 | 0 | 23 | 610_943 | 14 | 953 | Domain | Note=Protein kinase;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00159 |
| Tgene | PAN3 | 28794515 | FLT3 | 28644749 | ENST00000380958 | 0 | 24 | 610_943 | 14 | 994 | Domain | Note=Protein kinase;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00159 |
Top |
Kinase Fusion Protein Structures |
 CIF files of the predicted kinase fusion proteins CIF files of the predicted kinase fusion proteins * Here we show the 3D structure of the fusion proteins using Mol*. AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. Model confidence is shown from the pLDDT values per residue. pLDDT corresponds to the model’s prediction of its score on the local Distance Difference Test. It is a measure of local accuracy (from AlphfaFold website). To color code individual residues, we transformed individual PDB files into CIF format. |
| Kinase Fusion protein CIF link (fusion AA seq ID in KinaseFusionDB) | Henst | Tenst | Hgene | Hchr | Hbp | Tgene | Tchr | Tbp | AA seq | Len(AA seq) |
| PDB file >>>360_PAN3_FLT3 | ENST00000380958 | ENST00000241453 | PAN3 | chr13 | 28794515 | FLT3 | chr13 | 28644749 | MNSGGGLPPPSAAASPSSSSLAAAVAVVAPPGVGGVPGGAAVGVKLKYCRYYAKDKTCFYGEECQFLHEDPAAGAAPGLGLHSNSVPLAL AGAPVAGFPPGAVAGGGAGPPPGPKKPDLGDPGTGAAAGGGGSSGGLDGPRLAIPGMDGGALTDTSLTDSYFSTSFIGVNGFGSPVETKY PLMQRMTNSSSSPSLLNDSAKPYSAHDPLTSPASSLFNDFGALNISQRRKPRKYRLGMLEERLVPMGSKARKAKNPIGCLADRCKSGVPI NMVWWNRVTENNLQTPNPTASEFIPKGGSTSRLSNVSQSNMSAFSQVFSHPSMGSPATAGLAPVVFSAMIFGTITNQDLPVIKCVLINHK NNDSSVGKSSSYPMVSESPEDLGCALRPQSSGTVYEAAAVEVDVSASITLQVLVDAPGNISCLWVFKHSSLNCQPHFDLQNRGVVSMVIL KMTETQAGEYLLFIQSEATNYTILFTVSIRNTLLYTLRRPYFRKMENQDALVCISESVPEPIVEWVLCDSQGESCKEESPAVVKKEEKVL HELFGTDIRCCARNELGRECTRLFTIDLNQTPQTTLPQLFLKVGEPLWIRCKAVHVNHGFGLTWELENKALEEGNYFEMSTYSTNRTMIR ILFAFVSSVARNDTGYYTCSSSKHPSQSALVTIVEKGFINATNSSEDYEIDQYEEFCFSVRFKAYPQIRCTWTFSRKSFPCEQKGLDNGY SISKFCNHKHQPGEYIFHAENDDAQFTKMFTLNIRRKPQVLAEASASQASCFSDGYPLPSWTWKKCSDKSPNCTEEITEGVWNRKANRKV FGQWVSSSTLNMSEAIKGFLVKCCAYNSLGTSCETILLNSPGPFPFIQDNISFYATIGVCLLFIVVLTLLICHKYKKQFRYESQLQMVQV TGSSDNEYFYVDFREYEYDLKWEFPRENLEFGKVLGSGAFGKVMNATAYGISKTGVSIQVAVKMLKEKADSSEREALMSELKMMTQLGSH ENIVNLLGACTLSGPIYLIFEYCCYGDLLNYLRSKREKFHRTWTEIFKEHNFSFYPTFQSHPNSSMPGSREVQIHPDSDQISGLHGNSFH SEDEIEYENQKRLEEEEDLNVLTFEDLLCFAYQVAKGMEFLEFKSCVHRDLAARNVLVTHGKVVKICDFGLARDIMSDSNYVVRGNARLP VKWMAPESLFEGIYTIKSDVWSYGILLWEIFSLGVNPYPGIPVDANFYKLIQNGFKMDQPFYATEEIYIIMQSCWAFDSRKRPSFPNLTS | 1312 |
| 3D view using mol* of 360_PAN3_FLT3 | ||||||||||
| PDB file >>>TKFP_615_PAN3_FLT3 | ENST00000380958 | ENST00000241453 | PAN3 | chr13 | 28794515 | FLT3 | chr13 | 28644749 | MNSGGGLPPPSAAASPSSSSLAAAVAVVAPPGVGGVPGGAAVGVKLKYCRYYAKDKTCFYGEECQFLHEDPAAGAAPGLGLHSNSVPLAL AGAPVAGFPPGAVAGGGAGPPPGPKKPDLGDPGTGAAAGGGGSSGGLDGPRLAIPGMDGGALTDTSLTDSYFSTSFIGVNGFGSPVETKY PLMQRMTNSSSSPSLLNDSAKPYSAHDPLTSPASSLFNDFGALNISQRRKPRKYRLGMLEERLVPMGSKARKAKNPIGCLADRCKSGVPI NMVWWNRVTENNLQTPNPTASEFIPKGGSTSRLSNVSQSNMSAFSQVFSHPSMGSPATAGLAPVVFSAMIFGTITNQDLPVIKCVLINHK NNDSSVGKSSSYPMVSESPEDLGCALRPQSSGTVYEAAAVEVDVSASITLQVLVDAPGNISCLWVFKHSSLNCQPHFDLQNRGVVSMVIL KMTETQAGEYLLFIQSEATNYTILFTVSIRNTLLYTLRRPYFRKMENQDALVCISESVPEPIVEWVLCDSQGESCKEESPAVVKKEEKVL HELFGTDIRCCARNELGRECTRLFTIDLNQTPQTTLPQLFLKVGEPLWIRCKAVHVNHGFGLTWELENKALEEGNYFEMSTYSTNRTMIR ILFAFVSSVARNDTGYYTCSSSKHPSQSALVTIVEKGFINATNSSEDYEIDQYEEFCFSVRFKAYPQIRCTWTFSRKSFPCEQKGLDNGY SISKFCNHKHQPGEYIFHAENDDAQFTKMFTLNIRRKPQVLAEASASQASCFSDGYPLPSWTWKKCSDKSPNCTEEITEGVWNRKANRKV FGQWVSSSTLNMSEAIKGFLVKCCAYNSLGTSCETILLNSPGPFPFIQDNISFYATIGVCLLFIVVLTLLICHKYKKQFRYESQLQMVQV TGSSDNEYFYVDFREYEYDLKWEFPRENLEFGKVLGSGAFGKVMNATAYGISKTGVSIQVAVKMLKEKADSSEREALMSELKMMTQLGSH ENIVNLLGACTLSGPIYLIFEYCCYGDLLNYLRSKREKFHRTWTEIFKEHNFSFYPTFQSHPNSSMPGSREVQIHPDSDQISGLHGNSFH SEDEIEYENQKRLEEEEDLNVLTFEDLLCFAYQVAKGMEFLEFKSCVHRDLAARNVLVTHGKVVKICDFGLARDIMSDSNYVVRGNARLP VKWMAPESLFEGIYTIKSDVWSYGILLWEIFSLGVNPYPGIPVDANFYKLIQNGFKMDQPFYATEEIYIIMQSCWAFDSRKRPSFPNLTS | 1312_PAN3_FLT3 |
Top |
Comparison of Fusion Protein Isoforms |
 Superimpose the 3D Structures Among All Fusion Protein Isoforms Superimpose the 3D Structures Among All Fusion Protein Isoforms * Download the pdb file and open it from the molstar online viewer. |
 Comparison of the Secondary Structures of Fusion Protein Isoforms Comparison of the Secondary Structures of Fusion Protein Isoforms |
Top |
Comparison of Fusion Protein Sequences/Structures with Known Sequences/Structures from PDB |
Top |
pLDDT score distribution |
 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2* AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. * The blue color at the bottom marks the best active site residues. |
| 360_PAN3_FLT3.png |
 |
| 360_PAN3_FLT3.png |
 |
Top |
Potential Active Site Information |
 The potential binding sites of these fusion proteins were identified using SiteMap, a module of the Schrodinger suite. The potential binding sites of these fusion proteins were identified using SiteMap, a module of the Schrodinger suite. |
| Kinase Fusion AA seq ID in KinaseFusionDB | Site score | Size | Dscore | Volume | Exposure | Enclosure | Contact | Phobic | Philic | Balance | Don/Acc | Residues |
Top |
Ramachandran Plot of Kinase Fusion Protein Structure |
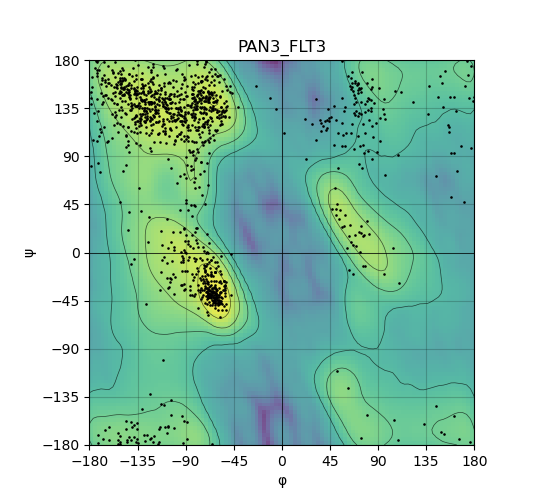
 Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. |
| 360_PAN3_FLT3_ramachandran.png |
 |
Top |
Virtual Screening Results |
 Distribution of the average docking score across all approved kinase inhibitors. Distribution of the average docking score across all approved kinase inhibitors.Distribution of the number of occurrence across all approved kinase inhibitors. |
| 5'-kinase fusion protein case |
| 3'-kinase fusion protein case |
Top |
 Drug information from DrugBank of the top 20 interacting small molecules. Drug information from DrugBank of the top 20 interacting small molecules.* The detailed information of individual kinase inhibitors are available in the download page. |
| Fusion gene name info | Drug | Docking score | Glide g score | Glide energy |
Top |
Kinase-Substrate Information of PAN3_FLT3 |
 Phosphorylation target of the kinase Phosphorylation target of the kinase(phosphosite, 03-17-2024) |
| Kinase | Kinase UniProt Acc | Kinase species | Substrate | Substrate UniProt Acc | Substrate phosphorylated residues | Substrate phosphorylated sites (+/-7AA) | Domain |
| FLT3 | P36888 | human | FLT3 | P36888 | Y955 | ADAEEAMyQNVDGRV | |
| FLT3 | P36888 | human | FLT3 | P36888 | Y726 | kEHNFSFyPtFQSHP | PK_Tyr_Ser-Thr |
| FLT3 | P36888 | human | FLT3 | P36888 | Y591 | SSDNEyFyVDFREyE | |
| FLT3 | P36888 | human | SIRT3 | Q9NTG7 | Y226 | KGLLLRLyTQNIDGL | |
| FLT3 | P36888 | human | IDH1 | O75874 | Y391 | PNVQRsDyLNTFEFM | Iso_dh |
| FLT3 | P36888 | human | IDH2 | P48735 | Y107 | sALATQkysVAVkCA | Iso_dh |
| FLT3 | P36888 | human | FLT3 | P36888 | Y969 | VSECPHtyQNrrPFS | |
| FLT3 | P36888 | human | PTPN11 | Q06124 | Y62 | KIQNtGDyyDLyGGE | SH2 |
| FLT3 | P36888 | human | FLT3 | P36888 | Y842 | DIMSDsNyVVRGNAR | PK_Tyr_Ser-Thr |
| FLT3 | P36888 | human | FLT3 | P36888 | Y599 | VDFREyEyDLkWEFP | |
| FLT3 | P36888 | human | FLT3 | P36888 | Y589 | TGSSDNEyFyVDFRE | |
| FLT3 | P36888 | human | PDHA1 | P08559 | Y301 | MsDPGVsyRtREEIQ | E1_dh |
| FLT3 | P36888 | human | PDK1 | Q15118 | Y243 | ARRLCDLyyINSPEL |
 Biological Network Integration of This Kinase and Substrates Biological Network Integration of This Kinase and Substrates (GeneMANIA website) |
 Enriched GO biological processes of the phosphorylation target genes of the kinase Enriched GO biological processes of the phosphorylation target genes of the kinase |
| Kinase | GOID | GO term | P.adjust |
| FLT3 | ID | Description | 0.00e+00 |
| FLT3 | GO:0006099 | tricarboxylic acid cycle | 6.25e-05 |
| FLT3 | GO:0009060 | aerobic respiration | 7.24e-05 |
| FLT3 | GO:0045333 | cellular respiration | 1.12e-04 |
| FLT3 | GO:0015980 | energy derivation by oxidation of organic compounds | 2.62e-04 |
| FLT3 | GO:0019216 | regulation of lipid metabolic process | 2.62e-04 |
| FLT3 | GO:0006086 | acetyl-CoA biosynthetic process from pyruvate | 3.97e-04 |
| FLT3 | GO:0006163 | purine nucleotide metabolic process | 5.57e-04 |
| FLT3 | GO:0072350 | tricarboxylic acid metabolic process | 5.57e-04 |
| FLT3 | GO:0006103 | 2-oxoglutarate metabolic process | 5.57e-04 |
| FLT3 | GO:0072521 | purine-containing compound metabolic process | 5.57e-04 |
| FLT3 | GO:0006085 | acetyl-CoA biosynthetic process | 6.02e-04 |
| FLT3 | GO:0006084 | acetyl-CoA metabolic process | 1.90e-03 |
| FLT3 | GO:0006739 | NADP metabolic process | 3.10e-03 |
| FLT3 | GO:0035384 | thioester biosynthetic process | 3.10e-03 |
| FLT3 | GO:0071616 | acyl-CoA biosynthetic process | 3.10e-03 |
| FLT3 | GO:0033866 | nucleoside bisphosphate biosynthetic process | 3.95e-03 |
| FLT3 | GO:0034030 | ribonucleoside bisphosphate biosynthetic process | 3.95e-03 |
| FLT3 | GO:0034033 | purine nucleoside bisphosphate biosynthetic process | 3.95e-03 |
| FLT3 | GO:0006081 | cellular aldehyde metabolic process | 5.45e-03 |
| FLT3 | GO:0019362 | pyridine nucleotide metabolic process | 6.28e-03 |
| FLT3 | GO:0046496 | nicotinamide nucleotide metabolic process | 6.28e-03 |
| FLT3 | GO:0072524 | pyridine-containing compound metabolic process | 6.94e-03 |
| FLT3 | GO:0043648 | dicarboxylic acid metabolic process | 7.79e-03 |
| FLT3 | GO:0006637 | acyl-CoA metabolic process | 7.79e-03 |
| FLT3 | GO:0035383 | thioester metabolic process | 7.79e-03 |
| FLT3 | GO:0006090 | pyruvate metabolic process | 1.13e-02 |
| FLT3 | GO:0033865 | nucleoside bisphosphate metabolic process | 1.14e-02 |
| FLT3 | GO:0033875 | ribonucleoside bisphosphate metabolic process | 1.14e-02 |
| FLT3 | GO:0034032 | purine nucleoside bisphosphate metabolic process | 1.14e-02 |
| FLT3 | GO:0046887 | positive regulation of hormone secretion | 1.39e-02 |
| FLT3 | GO:0045834 | positive regulation of lipid metabolic process | 1.43e-02 |
| FLT3 | GO:1902652 | secondary alcohol metabolic process | 1.48e-02 |
| FLT3 | GO:0044272 | sulfur compound biosynthetic process | 1.65e-02 |
| FLT3 | GO:0050796 | regulation of insulin secretion | 1.68e-02 |
| FLT3 | GO:0050731 | positive regulation of peptidyl-tyrosine phosphorylation | 1.69e-02 |
| FLT3 | GO:0046890 | regulation of lipid biosynthetic process | 2.03e-02 |
| FLT3 | GO:0006006 | glucose metabolic process | 2.03e-02 |
| FLT3 | GO:0090276 | regulation of peptide hormone secretion | 2.03e-02 |
| FLT3 | GO:0002791 | regulation of peptide secretion | 2.03e-02 |
| FLT3 | GO:0030073 | insulin secretion | 2.03e-02 |
| FLT3 | GO:0090087 | regulation of peptide transport | 2.03e-02 |
| FLT3 | GO:0009152 | purine ribonucleotide biosynthetic process | 2.25e-02 |
| FLT3 | GO:0009260 | ribonucleotide biosynthetic process | 2.49e-02 |
| FLT3 | GO:0050730 | regulation of peptidyl-tyrosine phosphorylation | 2.49e-02 |
| FLT3 | GO:0019318 | hexose metabolic process | 2.49e-02 |
| FLT3 | GO:0046390 | ribose phosphate biosynthetic process | 2.49e-02 |
| FLT3 | GO:0030072 | peptide hormone secretion | 2.56e-02 |
| FLT3 | GO:0002790 | peptide secretion | 2.59e-02 |
| FLT3 | GO:0046883 | regulation of hormone secretion | 2.59e-02 |
Top |
Related Drugs to PAN3_FLT3 |
 Drugs used for this fusion-positive patient. Drugs used for this fusion-positive patient. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Drug | Source | PMID |
 Distribution of the number of studies mentioning PAN3-FLT3 and kinase inhibitors the PubMed Abstract (04-01-2024) Distribution of the number of studies mentioning PAN3-FLT3 and kinase inhibitors the PubMed Abstract (04-01-2024) |
| Fusion gene - drug pair 1 | Fusion gene - drug pair 2 | PMID | Publication date | DOI | Study title |
Top |
Related Diseases to PAN3_FLT3 |
 Diseases that have this fusion gene. Diseases that have this fusion gene. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Disease | Source | PMID |
 Related diseases from the literature mentioned this fusion gene and drug. Related diseases from the literature mentioned this fusion gene and drug. (PubMed, 04-01-2024) |
| MeSH ID | MeSH term |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
Top |
Clinical Trials of the Found Drugs/Small Molecules |
 Statistics of the Clinical Trials of the Found Kinase Inibitors from clinicaltrials.gov (06-17-2024) Statistics of the Clinical Trials of the Found Kinase Inibitors from clinicaltrials.gov (06-17-2024) |
 Clinical Trials from clinicaltrials.gov (06-17-2024) Clinical Trials from clinicaltrials.gov (06-17-2024) |
| Fusion Gene | Kinase Inhibitor | NCT ID | Study Status | Phases | Disease | # Enrolment | Date |

