| UTHEALTH HOME ABOUT SBMI A-Z WEBMAIL INSIDE THE UNIVERSITY |

|
|||||||
|
Kinase Fusion Gene:TP53BP1_PDGFRB |
Kinase Fusion Protein Summary |
 Kinase Fusion gene summary Kinase Fusion gene summary |
| Kinase Fusion partner gene information | Kinase Fusion gene name: TP53BP1_PDGFRB | KinaseFusionDB ID: KFG6690 | FusionGDB2.0 ID: KFG6690 | Hgene | Tgene | Gene symbol | TP53BP1 | PDGFRB | Gene ID | 7158 | 5159 | |
| Gene name | tumor protein p53 binding protein 1 | platelet derived growth factor receptor beta | ||||||||||
| Synonyms | 53BP1|TDRD30|p202|p53BP1 | CD140B|IBGC4|IMF1|JTK12|KOGS|PDGFR|PDGFR-1|PDGFR1|PENTT | ||||||||||
| Cytomap | 15q15.3 | 5q32 | ||||||||||
| Type of gene | protein-coding | protein-coding | ||||||||||
| Description | TP53-binding protein 1p53-binding protein 1tumor protein 53-binding protein, 1tumor suppressor p53-binding protein 1 | platelet-derived growth factor receptor betaActivated tyrosine kinase PDGFRBCD140 antigen-like family member BNDEL1-PDGFRBPDGF-R-betaPDGFR-betabeta-type platelet-derived growth factor receptorplatelet-derived growth factor receptor 1platelet-deriv | ||||||||||
| Modification date | 20240416 | 20240413 | ||||||||||
| UniProtAcc | Q12888 | P09619 | ||||||||||
| Ensembl transtripts involved in fusion gene | ENST ids | ENST00000263801, ENST00000382039, ENST00000382044, ENST00000450115, ENST00000605155, | ENST00000523456, ENST00000261799, | |||||||||
| Context (manual curation of fusion genes in KinaseFusionDB) | PubMed: TP53BP1 [Title/Abstract] AND PDGFRB [Title/Abstract] AND fusion [Title/Abstract] | |||||||||||
| Most frequent breakpoint (based on all fusion genes of FusionGDB 2.0) | ||||||||||||
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | TP53BP1 | GO:0006303 | double-strand break repair via nonhomologous end joining | 23333306|23760478|27153538|28241136|31135337 |
| Hgene | TP53BP1 | GO:0006974 | DNA damage response | 17500065 |
| Hgene | TP53BP1 | GO:0006974 | DNA damage response | 27153538|28241136 |
| Hgene | TP53BP1 | GO:0045830 | positive regulation of isotype switching | 23345425 |
| Hgene | TP53BP1 | GO:0051260 | protein homooligomerization | 23345425 |
| Hgene | TP53BP1 | GO:0097680 | double-strand break repair via classical nonhomologous end joining | 27153538 |
| Hgene | TP53BP1 | GO:1990166 | protein localization to site of double-strand break | 37696958 |
| Hgene | TP53BP1 | GO:2000042 | negative regulation of double-strand break repair via homologous recombination | 23333306|23345425 |
| Tgene | PDGFRB | GO:0007165 | signal transduction | 10821867 |
| Tgene | PDGFRB | GO:0010863 | positive regulation of phospholipase C activity | 1653029 |
| Tgene | PDGFRB | GO:0018108 | peptidyl-tyrosine phosphorylation | 1653029|2536956|2850496 |
| Tgene | PDGFRB | GO:0030335 | positive regulation of cell migration | 17470632 |
| Tgene | PDGFRB | GO:0032516 | positive regulation of phosphoprotein phosphatase activity | 7691811 |
| Tgene | PDGFRB | GO:0038091 | positive regulation of cell proliferation by VEGF-activated platelet derived growth factor receptor signaling pathway | 17470632 |
| Tgene | PDGFRB | GO:0046777 | protein autophosphorylation | 2536956|2850496 |
| Tgene | PDGFRB | GO:0048008 | platelet-derived growth factor receptor signaling pathway | 1314164|2536956 |
| Tgene | PDGFRB | GO:0051897 | positive regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction | 1314164 |
| Tgene | PDGFRB | GO:0060326 | cell chemotaxis | 2554309|17991872 |
 Kinase Fusion gene breakpoints across TP53BP1 (5'-gene) Kinase Fusion gene breakpoints across TP53BP1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 Kinase Fusion gene breakpoints across PDGFRB (3'-gene) Kinase Fusion gene breakpoints across PDGFRB (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
Top |
Kinase Fusion Gene Sample Information |
 Kinase Fusion gene information. Kinase Fusion gene information. |
 Kinase Fusion gene information from four resources (ChiTars 5.0, ChimerDB 4.0, COSMIC, and CCLE) Kinase Fusion gene information from four resources (ChiTars 5.0, ChimerDB 4.0, COSMIC, and CCLE)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Sample | Hgene | Hchr | Hbp | Tgene | Tchr | Tbp |
| ChimerKB3 | . | TP53BP1 | chr15 | 43707791 | PDGFRB | chr5 | 149506177 |
Top |
Kinase Fusion ORF Analysis |
 Kinase Fusion information from ORFfinder translation from full-length transcript sequence from KinaseFusionDB. Kinase Fusion information from ORFfinder translation from full-length transcript sequence from KinaseFusionDB. |
| Henst | Tenst | Hgene | Hchr | Hbp | Tgene | Tchr | Tbp | Seq length (transcript) | Seq length (amino acids) |
| ENST00000263801 | ENST00000261799 | TP53BP1 | chr15 | 43707791 | PDGFRB | chr5 | 149506177 | 8995 | 2271 |
Top |
Kinase Fusion Amino Acid Sequences |
 For individual full-length fusion transcript sequence from KinaseFusionDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. For individual full-length fusion transcript sequence from KinaseFusionDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >Henst_Tenst_Hgene_Hchr_Hbp_Tgene_Tchr_Tbp_length(fusion AA)_AAseq >ENST00000263801_ENST00000261799_TP53BP1_chr15_43707791_PDGFRB_chr5_149506177_length(amino acids)=2271 MDPTGSQLDSDFSQQDTPCLIIEDSQPESQVLEDDSGSHFSMLSRHLPNLQTHKENPVLDVVSNPEQTAGEERGDGNSGFNEHLKENKVA DPVDSSNLDTCGSISQVIEQLPQPNRTSSVLGMSVESAPAVEEEKGEELEQKEKEKEEDTSGNTTHSLGAEDTASSQLGFGVLELSQSQD VEENTVPYEVDKEQLQSVTTNSGYTRLSDVDANTAIKHEEQSNEDIPIAEQSSKDIPVTAQPSKDVHVVKEQNPPPARSEDMPFSPKASV AAMEAKEQLSAQELMESGLQIQKSPEPEVLSTQEDLFDQSNKTVSSDGCSTPSREEGGCSLASTPATTLHLLQLSGQRSLVQDSLSTNSS DLVAPSPDAFRSTPFIVPSSPTEQEGRQDKPMDTSVLSEEGGEPFQKKLQSGEPVELENPPLLPESTVSPQASTPISQSTPVFPPGSLPI PSQPQFSHDIFIPSPSLEEQSNDGKKDGDMHSSSLTVECSKTSEIEPKNSPEDLGLSLTGDSCKLMLSTSEYSQSPKMESLSSHRIDEDG ENTQIEDTEPMSPVLNSKFVPAENDSILMNPAQDGEVQLSQNDDKTKGDDTDTRDDISILATGCKGREETVAEDVCIDLTCDSGSQAVPS PATRSEALSSVLDQEEAMEIKEHHPEEGSSGSEVEEIPETPCESQGEELKEENMESVPLHLSLTETQSQGLCLQKEMPKKECSEAMEVET SVISIDSPQKLAILDQELEHKEQEAWEEATSEDSSVVIVDVKEPSPRVDVSCEPLEGVEKCSDSQSWEDIAPEIEPCAENRLDTKEEKSV EYEGDLKSGTAETEPVEQDSSQPSLPLVRADDPLRLDQELQQPQTQEKTSNSLTEDSKMANAKQLSSDAEAQKLGKPSAHASQSFCESSS ETPFHFTLPKEGDIIPPLTGATPPLIGHLKLEPKRHSTPIGISNYPESTIATSDVMSESMVETHDPILGSGKGDSGAAPDVDDKLCLRMK LVSPETEASEESLQFNLEKPATGERKNGSTAVAESVASPQKTMSVLSCICEARQENEARSEDPPTTPIRGNLLHFPSSQGEEEKEKLEGD HTIRQSQQPMKPISPVKDPVSPASQKMVIQGPSSPQGEAMVTDVLEDQKEGRSTNKENPSKALIERPSQNNIGIQTMECSLRVPETVSAA TQTIKNVCEQGTSTVDQNFGKQDATVQTERGSGEKPVSAPGDDTESLHSQGEEEFDMPQPPHGHVLHRHMRTIREVRTLVTRVITDVYYV DGTEVERKVTEETEEPIVECQECETEVSPSQTGGSSGDLGDISSFSSKASSLHRTSSGTSLSAMHSSGSSGKGAGPLRGKTSGTEPADFA LPSSRGGPGKLSPRKGVSQTGTPVCEEDGDAGLGIRQGGKAPVTPRGRGRRGRPPSRTTGTRETAVPGPLGIEDISPNLSPDDKSFSRVV PRVPDSTRRTDVGAGALRRSDSPEIPFQAAAGPSDGLDASSPGNSFVGLRVVAKWSSNGYFYSGKITRDVGAGKYKLLFDDGYECDVLGK DILLCDPIPLDTEVTALSEDEYFSAGVVKGHRKESGELYYSIEKEGQRKWYKRMAVILSLEQGNRLREQYGLGPYEAVTPLTKAADISLD NLVEGKRKRRSNVSSPATPTASSSSSTTPTRKITESPRASMGVLSGKRKLITSEEERSPAKRGRKSATVKPALPFKVVVISAILALVVLT IISLIILIMLWQKKPRYEIRWKVIESVSSDGHEYIYVDPMQLPYDSTWELPRDQLVLGRTLGSGAFGQVVEATAHGLSHSQATMKVAVKM LKSTARSSEKQALMSELKIMSHLGPHLNVVNLLGACTKGGPIYIITEYCRYGDLVDYLHRNKHTFLQHHSDKRRPPSAELYSNALPVGLP LPSHVSLTGESDGGYMDMSKDESVDYVPMLDMKGDVKYADIESSNYMAPYDNYVPSAPERTCRATLINESPVLSYMDLVGFSYQVANGME FLASKNCVHRDLAARNVLICEGKLVKICDFGLARDIMRDSNYISKGSTFLPLKWMAPESIFNSLYTTLSDVWSFGILLWEIFTLGGTPYP ELPMNEQFYNAIKRGYRMAQPAHASDEIYEIMQKCWEEKFEIRPPFSQLVLLLERLLGEGYKKKYQQVDEEFLRSDHPAILRSQARLPGF HGLRSPLDTSSVLYTAVQPNEGDNDYIIPLPDPKPEVADEGPLEGSPSLASSTLNEVNTSSTISCDSPLEPQDEPEPEPQLELQVEPEPE -------------------------------------------------------------- |
Multiple Sequence Alignment of All Fusion Protein Isoforms |
Top |
Kinase Fusion Protein Functional Features |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr15:/chr5:) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| TP53BP1 | PDGFRB |
| FUNCTION: Double-strand break (DSB) repair protein involved in response to DNA damage, telomere dynamics and class-switch recombination (CSR) during antibody genesis (PubMed:12364621, PubMed:22553214, PubMed:23333306, PubMed:17190600, PubMed:21144835, PubMed:27153538, PubMed:28241136, PubMed:31135337). Plays a key role in the repair of double-strand DNA breaks (DSBs) in response to DNA damage by promoting non-homologous end joining (NHEJ)-mediated repair of DSBs and specifically counteracting the function of the homologous recombination (HR) repair protein BRCA1 (PubMed:22553214, PubMed:23727112, PubMed:23333306, PubMed:27153538, PubMed:31135337). In response to DSBs, phosphorylation by ATM promotes interaction with RIF1 and dissociation from NUDT16L1/TIRR, leading to recruitment to DSBs sites (PubMed:28241136). Recruited to DSBs sites by recognizing and binding histone H2A monoubiquitinated at 'Lys-15' (H2AK15Ub) and histone H4 dimethylated at 'Lys-20' (H4K20me2), two histone marks that are present at DSBs sites (PubMed:23760478, PubMed:27153538, PubMed:28241136, PubMed:17190600). Required for immunoglobulin class-switch recombination (CSR) during antibody genesis, a process that involves the generation of DNA DSBs (PubMed:23345425). Participates in the repair and the orientation of the broken DNA ends during CSR (By similarity). In contrast, it is not required for classic NHEJ and V(D)J recombination (By similarity). Promotes NHEJ of dysfunctional telomeres via interaction with PAXIP1 (PubMed:23727112). {ECO:0000250|UniProtKB:P70399, ECO:0000269|PubMed:12364621, ECO:0000269|PubMed:17190600, ECO:0000269|PubMed:21144835, ECO:0000269|PubMed:22553214, ECO:0000269|PubMed:23333306, ECO:0000269|PubMed:23345425, ECO:0000269|PubMed:23727112, ECO:0000269|PubMed:23760478, ECO:0000269|PubMed:27153538, ECO:0000269|PubMed:28241136, ECO:0000269|PubMed:31135337}. | FUNCTION: Tyrosine-protein kinase that acts as a cell-surface receptor for homodimeric PDGFB and PDGFD and for heterodimers formed by PDGFA and PDGFB, and plays an essential role in the regulation of embryonic development, cell proliferation, survival, differentiation, chemotaxis and migration. Plays an essential role in blood vessel development by promoting proliferation, migration and recruitment of pericytes and smooth muscle cells to endothelial cells. Plays a role in the migration of vascular smooth muscle cells and the formation of neointima at vascular injury sites. Required for normal development of the cardiovascular system. Required for normal recruitment of pericytes (mesangial cells) in the kidney glomerulus, and for normal formation of a branched network of capillaries in kidney glomeruli. Promotes rearrangement of the actin cytoskeleton and the formation of membrane ruffles. Binding of its cognate ligands - homodimeric PDGFB, heterodimers formed by PDGFA and PDGFB or homodimeric PDGFD -leads to the activation of several signaling cascades; the response depends on the nature of the bound ligand and is modulated by the formation of heterodimers between PDGFRA and PDGFRB. Phosphorylates PLCG1, PIK3R1, PTPN11, RASA1/GAP, CBL, SHC1 and NCK1. Activation of PLCG1 leads to the production of the cellular signaling molecules diacylglycerol and inositol 1,4,5-trisphosphate, mobilization of cytosolic Ca(2+) and the activation of protein kinase C. Phosphorylation of PIK3R1, the regulatory subunit of phosphatidylinositol 3-kinase, leads to the activation of the AKT1 signaling pathway. Phosphorylation of SHC1, or of the C-terminus of PTPN11, creates a binding site for GRB2, resulting in the activation of HRAS, RAF1 and down-stream MAP kinases, including MAPK1/ERK2 and/or MAPK3/ERK1. Promotes phosphorylation and activation of SRC family kinases. Promotes phosphorylation of PDCD6IP/ALIX and STAM. Receptor signaling is down-regulated by protein phosphatases that dephosphorylate the receptor and its down-stream effectors, and by rapid internalization of the activated receptor. {ECO:0000269|PubMed:11297552, ECO:0000269|PubMed:11331881, ECO:0000269|PubMed:1314164, ECO:0000269|PubMed:1396585, ECO:0000269|PubMed:1653029, ECO:0000269|PubMed:1709159, ECO:0000269|PubMed:1846866, ECO:0000269|PubMed:20494825, ECO:0000269|PubMed:20529858, ECO:0000269|PubMed:21098708, ECO:0000269|PubMed:21679854, ECO:0000269|PubMed:21733313, ECO:0000269|PubMed:2554309, ECO:0000269|PubMed:26599395, ECO:0000269|PubMed:2835772, ECO:0000269|PubMed:2850496, ECO:0000269|PubMed:7685273, ECO:0000269|PubMed:7691811, ECO:0000269|PubMed:7692233, ECO:0000269|PubMed:8195171}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
 - Retained domain in the 5'-partner of fusion protein (protein functional feature from UniProt). - Retained domain in the 5'-partner of fusion protein (protein functional feature from UniProt). |
| Partner | Hgeneene | Hbp | Tgeneene | Tbp | ENST | BPexon | TotalExon | Protein feature loci | BPloci | TotalLen | Feature | Note |
 - Retained domain in the 3'-partner of fusion protein (protein functional feature from UniProt). - Retained domain in the 3'-partner of fusion protein (protein functional feature from UniProt). |
| Partner | Hgeneene | Hbp | Tgeneene | Tbp | ENST | BPexon | TotalExon | Protein feature loci | BPloci | TotalLen | Feature | Note |
| Tgene | TP53BP1 | 43707791 | PDGFRB | 149506177 | ENST00000263801 | 9 | 23 | 600_962 | 526 | 1107 | Domain | Note=Protein kinase;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00159 |
Top |
Kinase Fusion Protein Structures |
 CIF files of the predicted kinase fusion proteins CIF files of the predicted kinase fusion proteins * Here we show the 3D structure of the fusion proteins using Mol*. AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. Model confidence is shown from the pLDDT values per residue. pLDDT corresponds to the model’s prediction of its score on the local Distance Difference Test. It is a measure of local accuracy (from AlphfaFold website). To color code individual residues, we transformed individual PDB files into CIF format. |
| Kinase Fusion protein CIF link (fusion AA seq ID in KinaseFusionDB) | Henst | Tenst | Hgene | Hchr | Hbp | Tgene | Tchr | Tbp | AA seq | Len(AA seq) |
| PDB file >>>515_TP53BP1_PDGFRB | ENST00000263801 | ENST00000261799 | TP53BP1 | chr15 | 43707791 | PDGFRB | chr5 | 149506177 | MDPTGSQLDSDFSQQDTPCLIIEDSQPESQVLEDDSGSHFSMLSRHLPNLQTHKENPVLDVVSNPEQTAGEERGDGNSGFNEHLKENKVA DPVDSSNLDTCGSISQVIEQLPQPNRTSSVLGMSVESAPAVEEEKGEELEQKEKEKEEDTSGNTTHSLGAEDTASSQLGFGVLELSQSQD VEENTVPYEVDKEQLQSVTTNSGYTRLSDVDANTAIKHEEQSNEDIPIAEQSSKDIPVTAQPSKDVHVVKEQNPPPARSEDMPFSPKASV AAMEAKEQLSAQELMESGLQIQKSPEPEVLSTQEDLFDQSNKTVSSDGCSTPSREEGGCSLASTPATTLHLLQLSGQRSLVQDSLSTNSS DLVAPSPDAFRSTPFIVPSSPTEQEGRQDKPMDTSVLSEEGGEPFQKKLQSGEPVELENPPLLPESTVSPQASTPISQSTPVFPPGSLPI PSQPQFSHDIFIPSPSLEEQSNDGKKDGDMHSSSLTVECSKTSEIEPKNSPEDLGLSLTGDSCKLMLSTSEYSQSPKMESLSSHRIDEDG ENTQIEDTEPMSPVLNSKFVPAENDSILMNPAQDGEVQLSQNDDKTKGDDTDTRDDISILATGCKGREETVAEDVCIDLTCDSGSQAVPS PATRSEALSSVLDQEEAMEIKEHHPEEGSSGSEVEEIPETPCESQGEELKEENMESVPLHLSLTETQSQGLCLQKEMPKKECSEAMEVET SVISIDSPQKLAILDQELEHKEQEAWEEATSEDSSVVIVDVKEPSPRVDVSCEPLEGVEKCSDSQSWEDIAPEIEPCAENRLDTKEEKSV EYEGDLKSGTAETEPVEQDSSQPSLPLVRADDPLRLDQELQQPQTQEKTSNSLTEDSKMANAKQLSSDAEAQKLGKPSAHASQSFCESSS ETPFHFTLPKEGDIIPPLTGATPPLIGHLKLEPKRHSTPIGISNYPESTIATSDVMSESMVETHDPILGSGKGDSGAAPDVDDKLCLRMK LVSPETEASEESLQFNLEKPATGERKNGSTAVAESVASPQKTMSVLSCICEARQENEARSEDPPTTPIRGNLLHFPSSQGEEEKEKLEGD HTIRQSQQPMKPISPVKDPVSPASQKMVIQGPSSPQGEAMVTDVLEDQKEGRSTNKENPSKALIERPSQNNIGIQTMECSLRVPETVSAA TQTIKNVCEQGTSTVDQNFGKQDATVQTERGSGEKPVSAPGDDTESLHSQGEEEFDMPQPPHGHVLHRHMRTIREVRTLVTRVITDVYYV DGTEVERKVTEETEEPIVECQECETEVSPSQTGGSSGDLGDISSFSSKASSLHRTSSGTSLSAMHSSGSSGKGAGPLRGKTSGTEPADFA LPSSRGGPGKLSPRKGVSQTGTPVCEEDGDAGLGIRQGGKAPVTPRGRGRRGRPPSRTTGTRETAVPGPLGIEDISPNLSPDDKSFSRVV PRVPDSTRRTDVGAGALRRSDSPEIPFQAAAGPSDGLDASSPGNSFVGLRVVAKWSSNGYFYSGKITRDVGAGKYKLLFDDGYECDVLGK DILLCDPIPLDTEVTALSEDEYFSAGVVKGHRKESGELYYSIEKEGQRKWYKRMAVILSLEQGNRLREQYGLGPYEAVTPLTKAADISLD NLVEGKRKRRSNVSSPATPTASSSSSTTPTRKITESPRASMGVLSGKRKLITSEEERSPAKRGRKSATVKPALPFKVVVISAILALVVLT IISLIILIMLWQKKPRYEIRWKVIESVSSDGHEYIYVDPMQLPYDSTWELPRDQLVLGRTLGSGAFGQVVEATAHGLSHSQATMKVAVKM LKSTARSSEKQALMSELKIMSHLGPHLNVVNLLGACTKGGPIYIITEYCRYGDLVDYLHRNKHTFLQHHSDKRRPPSAELYSNALPVGLP LPSHVSLTGESDGGYMDMSKDESVDYVPMLDMKGDVKYADIESSNYMAPYDNYVPSAPERTCRATLINESPVLSYMDLVGFSYQVANGME FLASKNCVHRDLAARNVLICEGKLVKICDFGLARDIMRDSNYISKGSTFLPLKWMAPESIFNSLYTTLSDVWSFGILLWEIFTLGGTPYP ELPMNEQFYNAIKRGYRMAQPAHASDEIYEIMQKCWEEKFEIRPPFSQLVLLLERLLGEGYKKKYQQVDEEFLRSDHPAILRSQARLPGF HGLRSPLDTSSVLYTAVQPNEGDNDYIIPLPDPKPEVADEGPLEGSPSLASSTLNEVNTSSTISCDSPLEPQDEPEPEPQLELQVEPEPE | 2271 |
| 3D view using mol* of 515_TP53BP1_PDGFRB | ||||||||||
| PDB file >>>TKFP_876_TP53BP1_PDGFRB | ENST00000263801 | ENST00000261799 | TP53BP1 | chr15 | 43707791 | PDGFRB | chr5 | 149506177 | MDPTGSQLDSDFSQQDTPCLIIEDSQPESQVLEDDSGSHFSMLSRHLPNLQTHKENPVLDVVSNPEQTAGEERGDGNSGFNEHLKENKVA DPVDSSNLDTCGSISQVIEQLPQPNRTSSVLGMSVESAPAVEEEKGEELEQKEKEKEEDTSGNTTHSLGAEDTASSQLGFGVLELSQSQD VEENTVPYEVDKEQLQSVTTNSGYTRLSDVDANTAIKHEEQSNEDIPIAEQSSKDIPVTAQPSKDVHVVKEQNPPPARSEDMPFSPKASV AAMEAKEQLSAQELMESGLQIQKSPEPEVLSTQEDLFDQSNKTVSSDGCSTPSREEGGCSLASTPATTLHLLQLSGQRSLVQDSLSTNSS DLVAPSPDAFRSTPFIVPSSPTEQEGRQDKPMDTSVLSEEGGEPFQKKLQSGEPVELENPPLLPESTVSPQASTPISQSTPVFPPGSLPI PSQPQFSHDIFIPSPSLEEQSNDGKKDGDMHSSSLTVECSKTSEIEPKNSPEDLGLSLTGDSCKLMLSTSEYSQSPKMESLSSHRIDEDG ENTQIEDTEPMSPVLNSKFVPAENDSILMNPAQDGEVQLSQNDDKTKGDDTDTRDDISILATGCKGREETVAEDVCIDLTCDSGSQAVPS PATRSEALSSVLDQEEAMEIKEHHPEEGSSGSEVEEIPETPCESQGEELKEENMESVPLHLSLTETQSQGLCLQKEMPKKECSEAMEVET SVISIDSPQKLAILDQELEHKEQEAWEEATSEDSSVVIVDVKEPSPRVDVSCEPLEGVEKCSDSQSWEDIAPEIEPCAENRLDTKEEKSV EYEGDLKSGTAETEPVEQDSSQPSLPLVRADDPLRLDQELQQPQTQEKTSNSLTEDSKMANAKQLSSDAEAQKLGKPSAHASQSFCESSS ETPFHFTLPKEGDIIPPLTGATPPLIGHLKLEPKRHSTPIGISNYPESTIATSDVMSESMVETHDPILGSGKGDSGAAPDVDDKLCLRMK LVSPETEASEESLQFNLEKPATGERKNGSTAVAESVASPQKTMSVLSCICEARQENEARSEDPPTTPIRGNLLHFPSSQGEEEKEKLEGD HTIRQSQQPMKPISPVKDPVSPASQKMVIQGPSSPQGEAMVTDVLEDQKEGRSTNKENPSKALIERPSQNNIGIQTMECSLRVPETVSAA TQTIKNVCEQGTSTVDQNFGKQDATVQTERGSGEKPVSAPGDDTESLHSQGEEEFDMPQPPHGHVLHRHMRTIREVRTLVTRVITDVYYV DGTEVERKVTEETEEPIVECQECETEVSPSQTGGSSGDLGDISSFSSKASSLHRTSSGTSLSAMHSSGSSGKGAGPLRGKTSGTEPADFA LPSSRGGPGKLSPRKGVSQTGTPVCEEDGDAGLGIRQGGKAPVTPRGRGRRGRPPSRTTGTRETAVPGPLGIEDISPNLSPDDKSFSRVV PRVPDSTRRTDVGAGALRRSDSPEIPFQAAAGPSDGLDASSPGNSFVGLRVVAKWSSNGYFYSGKITRDVGAGKYKLLFDDGYECDVLGK DILLCDPIPLDTEVTALSEDEYFSAGVVKGHRKESGELYYSIEKEGQRKWYKRMAVILSLEQGNRLREQYGLGPYEAVTPLTKAADISLD NLVEGKRKRRSNVSSPATPTASSSSSTTPTRKITESPRASMGVLSGKRKLITSEEERSPAKRGRKSATVKPALPFKVVVISAILALVVLT IISLIILIMLWQKKPRYEIRWKVIESVSSDGHEYIYVDPMQLPYDSTWELPRDQLVLGRTLGSGAFGQVVEATAHGLSHSQATMKVAVKM LKSTARSSEKQALMSELKIMSHLGPHLNVVNLLGACTKGGPIYIITEYCRYGDLVDYLHRNKHTFLQHHSDKRRPPSAELYSNALPVGLP LPSHVSLTGESDGGYMDMSKDESVDYVPMLDMKGDVKYADIESSNYMAPYDNYVPSAPERTCRATLINESPVLSYMDLVGFSYQVANGME FLASKNCVHRDLAARNVLICEGKLVKICDFGLARDIMRDSNYISKGSTFLPLKWMAPESIFNSLYTTLSDVWSFGILLWEIFTLGGTPYP ELPMNEQFYNAIKRGYRMAQPAHASDEIYEIMQKCWEEKFEIRPPFSQLVLLLERLLGEGYKKKYQQVDEEFLRSDHPAILRSQARLPGF HGLRSPLDTSSVLYTAVQPNEGDNDYIIPLPDPKPEVADEGPLEGSPSLASSTLNEVNTSSTISCDSPLEPQDEPEPEPQLELQVEPEPE | 2271_TP53BP1_PDGFRB |
Top |
Comparison of Fusion Protein Isoforms |
 Superimpose the 3D Structures Among All Fusion Protein Isoforms Superimpose the 3D Structures Among All Fusion Protein Isoforms * Download the pdb file and open it from the molstar online viewer. |
 Comparison of the Secondary Structures of Fusion Protein Isoforms Comparison of the Secondary Structures of Fusion Protein Isoforms |
Top |
Comparison of Fusion Protein Sequences/Structures with Known Sequences/Structures from PDB |
Top |
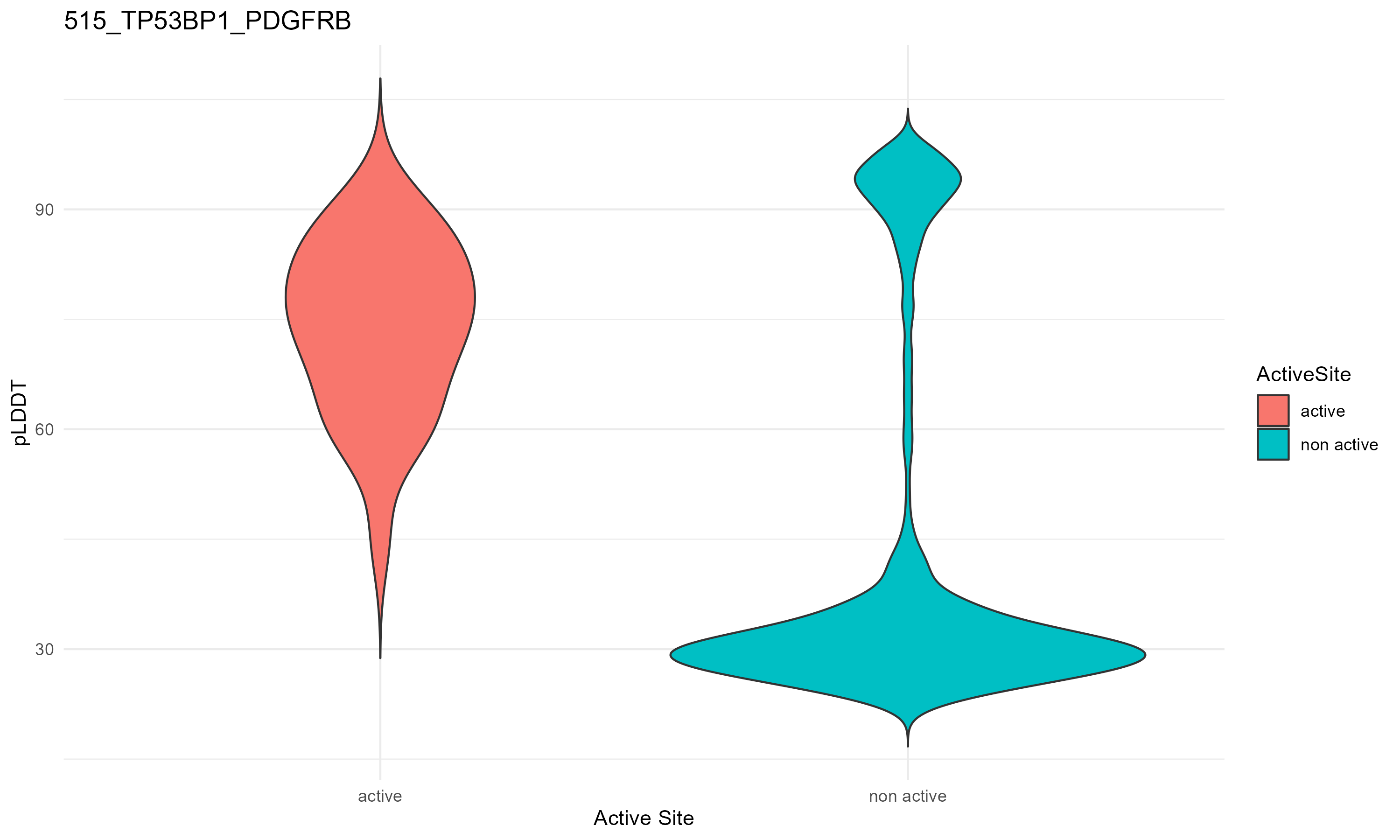
pLDDT score distribution |
 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2* AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. * The blue color at the bottom marks the best active site residues. |
| 515_TP53BP1_PDGFRB.png |
 |
| 515_TP53BP1_PDGFRB.png |
 |
Top |
Potential Active Site Information |
 The potential binding sites of these fusion proteins were identified using SiteMap, a module of the Schrodinger suite. The potential binding sites of these fusion proteins were identified using SiteMap, a module of the Schrodinger suite. |
| Kinase Fusion AA seq ID in KinaseFusionDB | Site score | Size | Dscore | Volume | Exposure | Enclosure | Contact | Phobic | Philic | Balance | Don/Acc | Residues |
Top |
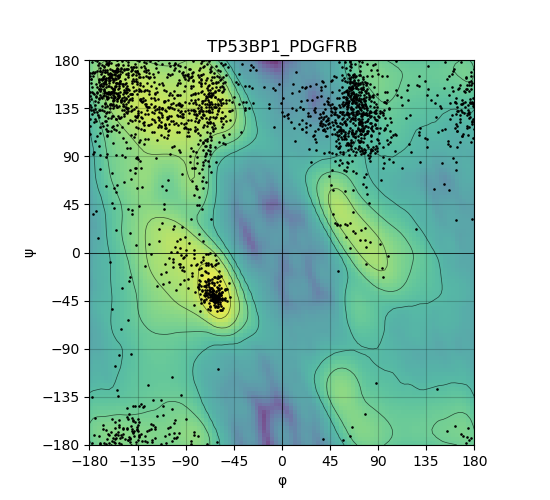
Ramachandran Plot of Kinase Fusion Protein Structure |
 Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. |
| 515_TP53BP1_PDGFRB_ramachandran.png |
 |
Top |
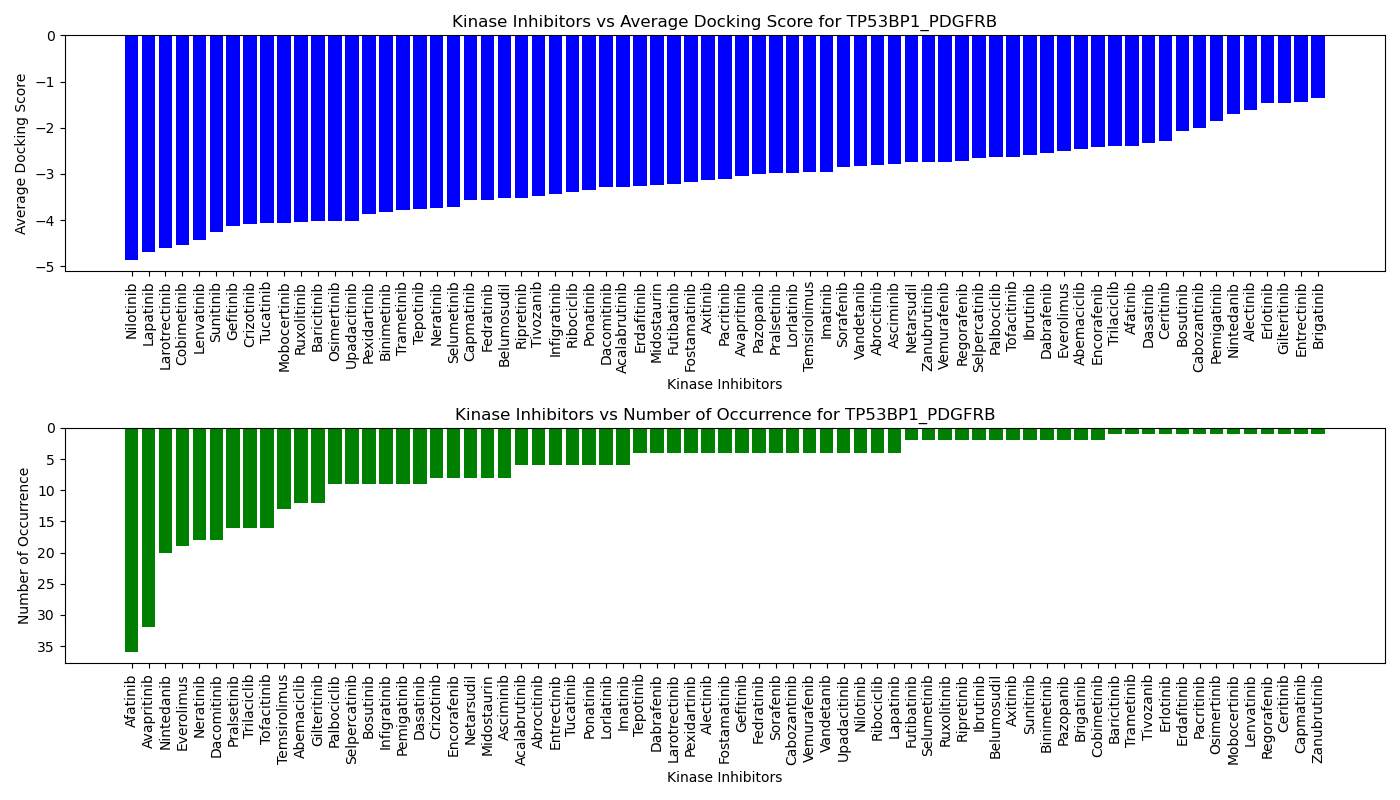
Virtual Screening Results |
 Distribution of the average docking score across all approved kinase inhibitors. Distribution of the average docking score across all approved kinase inhibitors.Distribution of the number of occurrence across all approved kinase inhibitors. |
| 5'-kinase fusion protein case |
| 3'-kinase fusion protein case |
 |
Top |
 Drug information from DrugBank of the top 20 interacting small molecules. Drug information from DrugBank of the top 20 interacting small molecules.* The detailed information of individual kinase inhibitors are available in the download page. |
| Fusion gene name info | Drug | Docking score | Glide g score | Glide energy |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Nilotinib | -5.79663 | -5.93623 | -44.9815 |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Nilotinib | -5.79663 | -5.93623 | -44.9815 |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Larotrectinib | -5.29758 | -5.29758 | -38.3285 |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Lapatinib | -5.2759 | -5.3647 | -55.2762 |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Neratinib | -5.19335 | -5.37925 | -47.544 |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Neratinib | -5.16309 | -5.34539 | -48.6644 |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Neratinib | -5.16309 | -5.34539 | -48.6644 |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Larotrectinib | -4.9281 | -4.9281 | -34.957 |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Larotrectinib | -4.86841 | -4.86841 | -38.8524 |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Lapatinib | -4.8621 | -4.9509 | -48.2889 |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Ruxolitinib | -4.76628 | -4.76628 | -26.9669 |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Upadacitinib | -4.758319999999999 | -4.75932 | -31.1345 |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Afatinib | -4.6663 | -4.8486 | -32.0839 |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Afatinib | -4.6663 | -4.8486 | -32.0839 |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Afatinib | -4.6649 | -4.8486 | -32.0839 |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Dacomitinib | -4.63754 | -4.74654 | -36.5442 |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Crizotinib | -4.58047 | -4.91667 | -41.6677 |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Crizotinib | -4.58047 | -4.91667 | -41.6677 |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Tucatinib | -4.5724 | -5.0352 | -37.9893 |
| 515_TP53BP1_PDGFRB-DOCK_HTVS_1-001 | Tucatinib | -4.56989 | -5.0326900000000006 | -37.881 |
Top |
Kinase-Substrate Information of TP53BP1_PDGFRB |
 Phosphorylation target of the kinase Phosphorylation target of the kinase(phosphosite, 03-17-2024) |
| Kinase | Kinase UniProt Acc | Kinase species | Substrate | Substrate UniProt Acc | Substrate phosphorylated residues | Substrate phosphorylated sites (+/-7AA) | Domain |
| PDGFRB | P09619 | human | PDGFRB | P09619 | Y1021 | PNEGDNDyIIPLPDP | |
| PDGFRB | P09619 | human | PDGFRA | P16234 | Y754 | ERKEVsKysDIQRsL | PK_Tyr_Ser-Thr |
| PDGFRB | P09619 | human | PDGFRB | P09619 | Y1009 | LDTSSVLyTAVQPNE | |
| PDGFRB | P09619 | human | PTK2 | Q05397 | Y5 | ___MAAAyLDPNLNH | |
| PDGFRB | P09619 | human | ABL2 | P42684 | Y272 | KCNKPTVyGVsPIHD | |
| PDGFRB | P09619 | human | PDGFRB | P09619 | Y579 | VSSDGHEyIyVDPMQ | |
| PDGFRB | P09619 | human | PDGFRB | P09619 | Y562 | LWQKKPRyEIRWKVI | |
| PDGFRB | P09619 | human | THOC5 | Q13769 | Y225 | EIEVKKEyLSSLQPR | FmiP_Thoc5 |
| PDGFRB | P09619 | human | ABL2 | P42684 | Y139 | EKLRVLGyNQNGEWS | SH3_1 |
| PDGFRB | P09619 | human | PLCG1 | P19174 | Y783 | EGRNPGFyVEANPMP | |
| PDGFRB | P09619 | human | PDGFRB | P09619 | Y740 | TGESDGGyMDMSKDE | PK_Tyr_Ser-Thr |
| PDGFRB | P09619 | human | ABL2 | P42684 | Y439 | RLMtGDtytAHAGAk | PK_Tyr_Ser-Thr |
| PDGFRB | P09619 | human | FYN | P06241 | Y28 | sLNQssGyRyGTDPT | |
| PDGFRB | P09619 | human | ETV6 | P41212 | Y17 | IkQERIsytPPEsPV | |
| PDGFRB | P09619 | human | PTK2 | Q05397 | Y194 | ALEKKSNyEVLEkDV | FERM_M |
| PDGFRB | P09619 | human | PDGFRB | P09619 | Y763 | DMKGDVKyADIESSN | PK_Tyr_Ser-Thr |
| PDGFRB | P09619 | human | TNK2 | Q07912 | Y635 | PLPPPPAyDDVAQDE | |
| PDGFRB | P09619 | human | PDGFRB | P09619 | Y581 | SDGHEyIyVDPMQLP | |
| PDGFRB | P09619 | human | ABL2 | P42684 | Y116 | PNLFVALyDFVAsGD | SH3_1 |
| PDGFRB | P09619 | human | ABL2 | P42684 | Y303 | GGQyGEVyVGVWKKy | PK_Tyr_Ser-Thr |
| PDGFRB | P09619 | human | MUC1 | P15941 | Y1218 | tyHtHGRyVPPsstD | |
| PDGFRB | P09619 | human | ABL2 | P42684 | Y310 | yVGVWKKysLtVAVK | PK_Tyr_Ser-Thr |
| PDGFRB | P09619 | human | PDGFRB | P09619 | Y857 | DIMRDSNyISKGSTF | PK_Tyr_Ser-Thr |
| PDGFRB | P09619 | human | PDGFRB | P09619 | Y775 | SSNyMAPyDNyVPSA | PK_Tyr_Ser-Thr |
| PDGFRB | P09619 | human | MUC1 | P15941 | Y1203 | IFPARDtyHPMsEyP | |
| PDGFRB | P09619 | human | PDGFRB | P09619 | Y771 | ADIESSNyMAPyDNy | PK_Tyr_Ser-Thr |
| PDGFRB | P09619 | human | ABL2 | P42684 | Y161 | QGWVPsNyItPVNSL | |
| PDGFRB | P09619 | human | ETV6 | P41212 | Y27 | PEsPVPsyAsstPLH | |
| PDGFRB | P09619 | human | SRC | P12931 | Y419 | RLIEDNEytARQGAk | PK_Tyr_Ser-Thr |
| PDGFRB | P09619 | human | PDGFRB | P09619 | Y751 | SKDESVDyVPMLDMK | PK_Tyr_Ser-Thr |
| PDGFRB | P09619 | human | ABL2 | P42684 | Y299 | HKLGGGQyGEVyVGV | PK_Tyr_Ser-Thr |
| PDGFRB | P09619 | human | PDGFRB | P09619 | Y778 | yMAPyDNyVPSAPER | PK_Tyr_Ser-Thr |
 Biological Network Integration of This Kinase and Substrates Biological Network Integration of This Kinase and Substrates (GeneMANIA website) |
 Enriched GO biological processes of the phosphorylation target genes of the kinase Enriched GO biological processes of the phosphorylation target genes of the kinase |
| Kinase | GOID | GO term | P.adjust |
| PDGFRB | ID | Description | 0.00e+00 |
| PDGFRB | GO:0018108 | peptidyl-tyrosine phosphorylation | 1.87e-08 |
| PDGFRB | GO:0018212 | peptidyl-tyrosine modification | 1.87e-08 |
| PDGFRB | GO:0038093 | Fc receptor signaling pathway | 6.42e-06 |
| PDGFRB | GO:0036120 | cellular response to platelet-derived growth factor stimulus | 4.63e-05 |
| PDGFRB | GO:0036119 | response to platelet-derived growth factor | 4.63e-05 |
| PDGFRB | GO:0002433 | immune response-regulating cell surface receptor signaling pathway involved in phagocytosis | 4.63e-05 |
| PDGFRB | GO:0038096 | Fc-gamma receptor signaling pathway involved in phagocytosis | 4.63e-05 |
| PDGFRB | GO:0002431 | Fc receptor mediated stimulatory signaling pathway | 6.79e-05 |
| PDGFRB | GO:0002862 | negative regulation of inflammatory response to antigenic stimulus | 6.79e-05 |
| PDGFRB | GO:0038094 | Fc-gamma receptor signaling pathway | 6.79e-05 |
| PDGFRB | GO:0010863 | positive regulation of phospholipase C activity | 8.77e-05 |
| PDGFRB | GO:1900274 | regulation of phospholipase C activity | 9.45e-05 |
| PDGFRB | GO:0051897 | positive regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction | 1.34e-04 |
| PDGFRB | GO:0002861 | regulation of inflammatory response to antigenic stimulus | 1.52e-04 |
| PDGFRB | GO:0010518 | positive regulation of phospholipase activity | 1.52e-04 |
| PDGFRB | GO:0048013 | ephrin receptor signaling pathway | 1.61e-04 |
| PDGFRB | GO:0046777 | protein autophosphorylation | 1.87e-04 |
| PDGFRB | GO:0048008 | platelet-derived growth factor receptor signaling pathway | 2.11e-04 |
| PDGFRB | GO:0010517 | regulation of phospholipase activity | 2.11e-04 |
| PDGFRB | GO:0060193 | positive regulation of lipase activity | 2.11e-04 |
| PDGFRB | GO:0048010 | vascular endothelial growth factor receptor signaling pathway | 2.11e-04 |
| PDGFRB | GO:0051896 | regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction | 3.46e-04 |
| PDGFRB | GO:0060191 | regulation of lipase activity | 4.05e-04 |
| PDGFRB | GO:0002437 | inflammatory response to antigenic stimulus | 4.19e-04 |
| PDGFRB | GO:0043491 | phosphatidylinositol 3-kinase/protein kinase B signal transduction | 5.64e-04 |
| PDGFRB | GO:0002429 | immune response-activating cell surface receptor signaling pathway | 7.80e-04 |
| PDGFRB | GO:0035791 | platelet-derived growth factor receptor-beta signaling pathway | 8.61e-04 |
| PDGFRB | GO:0007173 | epidermal growth factor receptor signaling pathway | 8.61e-04 |
| PDGFRB | GO:0033674 | positive regulation of kinase activity | 8.77e-04 |
| PDGFRB | GO:0002768 | immune response-regulating cell surface receptor signaling pathway | 9.40e-04 |
| PDGFRB | GO:2001243 | negative regulation of intrinsic apoptotic signaling pathway | 9.40e-04 |
| PDGFRB | GO:0072224 | metanephric glomerulus development | 1.11e-03 |
| PDGFRB | GO:0038127 | ERBB signaling pathway | 1.14e-03 |
| PDGFRB | GO:2000811 | negative regulation of anoikis | 1.18e-03 |
| PDGFRB | GO:0055003 | cardiac myofibril assembly | 1.24e-03 |
| PDGFRB | GO:0061298 | retina vasculature development in camera-type eye | 1.24e-03 |
| PDGFRB | GO:0050900 | leukocyte migration | 1.29e-03 |
| PDGFRB | GO:0043552 | positive regulation of phosphatidylinositol 3-kinase activity | 1.45e-03 |
| PDGFRB | GO:0051347 | positive regulation of transferase activity | 1.46e-03 |
| PDGFRB | GO:0050921 | positive regulation of chemotaxis | 1.49e-03 |
| PDGFRB | GO:0002223 | stimulatory C-type lectin receptor signaling pathway | 1.54e-03 |
| PDGFRB | GO:1990840 | response to lectin | 1.54e-03 |
| PDGFRB | GO:1990858 | cellular response to lectin | 1.54e-03 |
| PDGFRB | GO:0034614 | cellular response to reactive oxygen species | 1.60e-03 |
| PDGFRB | GO:2000209 | regulation of anoikis | 1.60e-03 |
| PDGFRB | GO:0090218 | positive regulation of lipid kinase activity | 1.85e-03 |
| PDGFRB | GO:0050731 | positive regulation of peptidyl-tyrosine phosphorylation | 1.97e-03 |
| PDGFRB | GO:0002757 | immune response-activating signaling pathway | 1.99e-03 |
| PDGFRB | GO:0032102 | negative regulation of response to external stimulus | 2.07e-03 |
Top |
Related Drugs to TP53BP1_PDGFRB |
 Drugs used for this fusion-positive patient. Drugs used for this fusion-positive patient. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Drug | Source | PMID |
 Distribution of the number of studies mentioning TP53BP1-PDGFRB and kinase inhibitors the PubMed Abstract (04-01-2024) Distribution of the number of studies mentioning TP53BP1-PDGFRB and kinase inhibitors the PubMed Abstract (04-01-2024) |
 |
| Fusion gene - drug pair 1 | Fusion gene - drug pair 2 | PMID | Publication date | DOI | Study title |
| TP53BP1-PDGFRB AND Imatinib | 15492236 | 2004-10-15 | 10.1158/0008-5472.can-04-2005 | p53-Binding protein 1 is fused to the platelet-derived growth factor receptor beta in a patient with a t(5;15)(q33;q22) and an imatinib-responsive eosinophilic myeloproliferative disorder. |
Top |
Related Diseases to TP53BP1_PDGFRB |
 Diseases that have this fusion gene. Diseases that have this fusion gene. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Disease | Source | PMID |
 Related diseases from the literature mentioned this fusion gene and drug. Related diseases from the literature mentioned this fusion gene and drug. (PubMed, 04-01-2024) |
| MeSH ID | MeSH term |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
| Tgene | PDGFRB | C3554321 | BASAL GANGLIA CALCIFICATION, IDIOPATHIC, 4 | 6 | CTD_human;GENOMICS_ENGLAND;UNIPROT |
| Tgene | PDGFRB | C0393590 | Fahr's syndrome (disorder) | 3 | GENOMICS_ENGLAND;ORPHANET |
| Tgene | PDGFRB | C4225270 | Kosaki overgrowth syndrome | 3 | CTD_human;GENOMICS_ENGLAND;ORPHANET;UNIPROT |
| Tgene | PDGFRB | C4551572 | MYOFIBROMATOSIS, INFANTILE, 1 | 3 | GENOMICS_ENGLAND;UNIPROT |
| Tgene | PDGFRB | C0013421 | Dystonia | 2 | GENOMICS_ENGLAND |
| Tgene | PDGFRB | C0023480 | Leukemia, Myelomonocytic, Chronic | 2 | ORPHANET |
| Tgene | PDGFRB | C0023893 | Liver Cirrhosis, Experimental | 2 | CTD_human |
| Tgene | PDGFRB | C0036341 | Schizophrenia | 2 | PSYGENET |
| Tgene | PDGFRB | C0432284 | Infantile myofibromatosis | 2 | CTD_human;GENOMICS_ENGLAND;ORPHANET |
Top |
Clinical Trials of the Found Drugs/Small Molecules |
 Statistics of the Clinical Trials of the Found Kinase Inibitors from clinicaltrials.gov (06-17-2024) Statistics of the Clinical Trials of the Found Kinase Inibitors from clinicaltrials.gov (06-17-2024) |
 Clinical Trials from clinicaltrials.gov (06-17-2024) Clinical Trials from clinicaltrials.gov (06-17-2024) |
| Fusion Gene | Kinase Inhibitor | NCT ID | Study Status | Phases | Disease | # Enrolment | Date |

