| UTHEALTH HOME ABOUT SBMI A-Z WEBMAIL INSIDE THE UNIVERSITY |

|
|||||||
|
Kinase Fusion Gene:CALML3_TYRO3 |
Kinase Fusion Protein Summary |
 Kinase Fusion gene summary Kinase Fusion gene summary |
| Kinase Fusion partner gene information | Kinase Fusion gene name: CALML3_TYRO3 | KinaseFusionDB ID: KFG838 | FusionGDB2.0 ID: KFG838 | Hgene | Tgene | Gene symbol | CALML3 | TYRO3 | Gene ID | 810 | 7301 | |
| Gene name | calmodulin like 3 | TYRO3 protein tyrosine kinase | ||||||||||
| Synonyms | CLP | BYK|Dtk|Etk-2|RSE|Rek|Sky|Tif | ||||||||||
| Cytomap | 10p15.1 | 15q15.1 | ||||||||||
| Type of gene | protein-coding | protein-coding | ||||||||||
| Description | calmodulin-like protein 3caM-like proteincalmodulin-related protein NB-1 | tyrosine-protein kinase receptor TYRO3tyrosine-protein kinase DTKtyrosine-protein kinase RSEtyrosine-protein kinase SKYtyrosine-protein kinase TIFtyrosine-protein kinase byk | ||||||||||
| Modification date | 20240411 | 20240411 | ||||||||||
| UniProtAcc | P27482 | Q06418 | ||||||||||
| Ensembl transtripts involved in fusion gene | ENST ids | ENST00000315238, | ENST00000263798, ENST00000559066, | |||||||||
| Context (manual curation of fusion genes in KinaseFusionDB) | PubMed: CALML3 [Title/Abstract] AND TYRO3 [Title/Abstract] AND fusion [Title/Abstract] | |||||||||||
| Most frequent breakpoint (based on all fusion genes of FusionGDB 2.0) | CALML3(5567458)-TYRO3(41854863), # samples:1 | |||||||||||
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | TYRO3 | GO:0046777 | protein autophosphorylation | 7511603|7537495 |
 Kinase Fusion gene breakpoints across CALML3 (5'-gene) Kinase Fusion gene breakpoints across CALML3 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
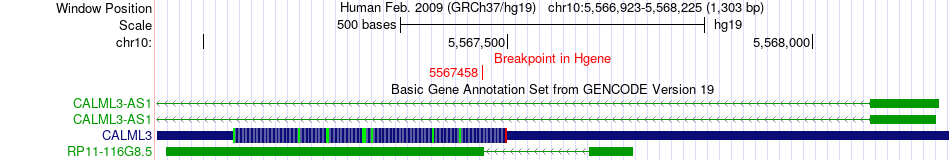 |
 Kinase Fusion gene breakpoints across TYRO3 (3'-gene) Kinase Fusion gene breakpoints across TYRO3 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
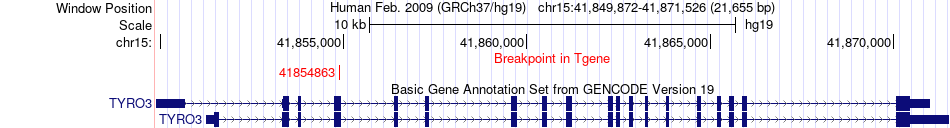 |
Top |
Kinase Fusion Gene Sample Information |
 Kinase Fusion gene information. Kinase Fusion gene information. |
 Kinase Fusion gene information from four resources (ChiTars 5.0, ChimerDB 4.0, COSMIC, and CCLE) Kinase Fusion gene information from four resources (ChiTars 5.0, ChimerDB 4.0, COSMIC, and CCLE)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Sample | Hgene | Hchr | Hbp | Tgene | Tchr | Tbp |
| ChimerDB4 | TCGA-D1-A1O8-01A | CALML3 | chr10 | 5567458 | TYRO3 | chr15 | 41854863 |
Top |
Kinase Fusion ORF Analysis |
 Kinase Fusion information from ORFfinder translation from full-length transcript sequence from KinaseFusionDB. Kinase Fusion information from ORFfinder translation from full-length transcript sequence from KinaseFusionDB. |
| Henst | Tenst | Hgene | Hchr | Hbp | Tgene | Tchr | Tbp | Seq length (transcript) | Seq length (amino acids) |
| ENST00000315238 | ENST00000559066 | CALML3 | chr10 | 5567458 | TYRO3 | chr15 | 41854863 | 3442 | 720 |
Top |
Kinase Fusion Amino Acid Sequences |
 For individual full-length fusion transcript sequence from KinaseFusionDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. For individual full-length fusion transcript sequence from KinaseFusionDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >Henst_Tenst_Hgene_Hchr_Hbp_Tgene_Tchr_Tbp_length(fusion AA)_AAseq >ENST00000315238_ENST00000559066_CALML3_chr10_5567458_TYRO3_chr15_41854863_length(amino acids)=720 MRYSSLTTKIGGPAPSPSVLNVTGVTQSTMFSCEAHNLKGLASSRTATVHLQALPAAPFNITVTKLSSSNASVAWMPGADGRALLQSCTV QVTQAPGGWEVLAVVVPVPPFTCLLRDLVPATNYSLRVRCANALGPSPYADWVPFQTKGLAPASAPQNLHAIRTDSGLILEWEEVIPEAP LEGPLGPYKLSWVQDNGTQDELTVEGTRANLTGWDPQKDLIVRVCVSNAVGCGPWSQPLVVSSHDRAGQQGPPHSRTSWVPVVLGVLTAL VTAAALALILLRKRRKETRFGQAFDSVMARGEPAVHFRAARSFNRERPERIEATLDSLGISDELKEKLEDVLIPEQQFTLGRMLGKGEFG SVREAQLKQEDGSFVKVAVKMLKADIIASSDIEEFLREAACMKEFDHPHVAKLVGVSLRSRAKGRLPIPMVILPFMKHGDLHAFLLASRI GENPFNLPLQTLIRFMVDIACGMEYLSSRNFIHRDLAARNCMLAEDMTVCVADFGLSRKIYSGDYYRQGCASKLPVKWLALESLADNLYT VQSDVWAFGVTMWEIMTRGQTPYAGIENAEIYNYLIGGNRLKQPPECMEDVYDLMYQCWSADPKQRPSFTCLRMELENILGQLSVLSASQ DPLYINIERAEEPTAGGSLELPGRDQPYSGAGDGSGMGAVGGTPSDCRYILTPGGLAEQPGQAEHQPESPLNETQRLLLLQQGLLPHSSC -------------------------------------------------------------- |
Multiple Sequence Alignment of All Fusion Protein Isoforms |
Top |
Kinase Fusion Protein Functional Features |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr10:5567458/chr15:41854863) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| CALML3 | TYRO3 |
| FUNCTION: May function as a specific light chain of unconventional myosin-10 (MYO10), also enhances MYO10 translation, possibly by acting as a chaperone for the emerging MYO10 heavy chain protein. May compete with calmodulin by binding, with different affinities, to cellular substrates. {ECO:0000269|PubMed:11278607, ECO:0000269|PubMed:18295593}. | FUNCTION: Receptor tyrosine kinase that transduces signals from the extracellular matrix into the cytoplasm by binding to several ligands including TULP1 or GAS6. Regulates many physiological processes including cell survival, migration and differentiation. Ligand binding at the cell surface induces dimerization and autophosphorylation of TYRO3 on its intracellular domain that provides docking sites for downstream signaling molecules. Following activation by ligand, interacts with PIK3R1 and thereby enhances PI3-kinase activity. Activates the AKT survival pathway, including nuclear translocation of NF-kappa-B and up-regulation of transcription of NF-kappa-B-regulated genes. TYRO3 signaling plays a role in various processes such as neuron protection from excitotoxic injury, platelet aggregation and cytoskeleton reorganization. Also plays an important role in inhibition of Toll-like receptors (TLRs)-mediated innate immune response by activating STAT1, which selectively induces production of suppressors of cytokine signaling SOCS1 and SOCS3. {ECO:0000269|PubMed:20546121}.; FUNCTION: (Microbial infection) Acts as a receptor for lassa virus and lymphocytic choriomeningitis virus, possibly through GAS6 binding to phosphatidyl-serine at the surface of virion envelope. {ECO:0000269|PubMed:22156524, ECO:0000269|PubMed:22673088, ECO:0000269|PubMed:25277499}.; FUNCTION: (Microbial infection) Acts as a receptor for Ebolavirus, possibly through GAS6 binding to phosphatidyl-serine at the surface of virion envelope. {ECO:0000269|PubMed:17005688}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
 - Retained domain in the 5'-partner of fusion protein (protein functional feature from UniProt). - Retained domain in the 5'-partner of fusion protein (protein functional feature from UniProt). |
| Partner | Hgeneene | Hbp | Tgeneene | Tbp | ENST | BPexon | TotalExon | Protein feature loci | BPloci | TotalLen | Feature | Note |
 - Retained domain in the 3'-partner of fusion protein (protein functional feature from UniProt). - Retained domain in the 3'-partner of fusion protein (protein functional feature from UniProt). |
| Partner | Hgeneene | Hbp | Tgeneene | Tbp | ENST | BPexon | TotalExon | Protein feature loci | BPloci | TotalLen | Feature | Note |
| Tgene | CALML3 | 5567458 | TYRO3 | 41854863 | ENST00000315238 | 0 | 19 | 227_320 | 0 | 891 | Domain | Note=Fibronectin type-III 1;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00316 |
| Tgene | CALML3 | 5567458 | TYRO3 | 41854863 | ENST00000315238 | 0 | 19 | 325_416 | 0 | 891 | Domain | Note=Fibronectin type-III 2;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00316 |
| Tgene | CALML3 | 5567458 | TYRO3 | 41854863 | ENST00000315238 | 0 | 19 | 41_128 | 0 | 891 | Domain | Note=Ig-like C2-type 1 |
| Tgene | CALML3 | 5567458 | TYRO3 | 41854863 | ENST00000315238 | 0 | 19 | 139_220 | 0 | 891 | Domain | Note=Ig-like C2-type 2 |
| Tgene | CALML3 | 5567458 | TYRO3 | 41854863 | ENST00000315238 | 0 | 19 | 518_790 | 0 | 891 | Domain | Note=Protein kinase;Ontology_term=ECO:0000255;evidence=ECO:0000255|PROSITE-ProRule:PRU00159 |
Top |
Kinase Fusion Protein Structures |
 CIF files of the predicted kinase fusion proteins CIF files of the predicted kinase fusion proteins * Here we show the 3D structure of the fusion proteins using Mol*. AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. Model confidence is shown from the pLDDT values per residue. pLDDT corresponds to the model’s prediction of its score on the local Distance Difference Test. It is a measure of local accuracy (from AlphfaFold website). To color code individual residues, we transformed individual PDB files into CIF format. |
| Kinase Fusion protein CIF link (fusion AA seq ID in KinaseFusionDB) | Henst | Tenst | Hgene | Hchr | Hbp | Tgene | Tchr | Tbp | AA seq | Len(AA seq) |
| PDB file >>>78_CALML3_TYRO3 | ENST00000315238 | ENST00000559066 | CALML3 | chr10 | 5567458 | TYRO3 | chr15 | 41854863 | MRYSSLTTKIGGPAPSPSVLNVTGVTQSTMFSCEAHNLKGLASSRTATVHLQALPAAPFNITVTKLSSSNASVAWMPGADGRALLQSCTV QVTQAPGGWEVLAVVVPVPPFTCLLRDLVPATNYSLRVRCANALGPSPYADWVPFQTKGLAPASAPQNLHAIRTDSGLILEWEEVIPEAP LEGPLGPYKLSWVQDNGTQDELTVEGTRANLTGWDPQKDLIVRVCVSNAVGCGPWSQPLVVSSHDRAGQQGPPHSRTSWVPVVLGVLTAL VTAAALALILLRKRRKETRFGQAFDSVMARGEPAVHFRAARSFNRERPERIEATLDSLGISDELKEKLEDVLIPEQQFTLGRMLGKGEFG SVREAQLKQEDGSFVKVAVKMLKADIIASSDIEEFLREAACMKEFDHPHVAKLVGVSLRSRAKGRLPIPMVILPFMKHGDLHAFLLASRI GENPFNLPLQTLIRFMVDIACGMEYLSSRNFIHRDLAARNCMLAEDMTVCVADFGLSRKIYSGDYYRQGCASKLPVKWLALESLADNLYT VQSDVWAFGVTMWEIMTRGQTPYAGIENAEIYNYLIGGNRLKQPPECMEDVYDLMYQCWSADPKQRPSFTCLRMELENILGQLSVLSASQ DPLYINIERAEEPTAGGSLELPGRDQPYSGAGDGSGMGAVGGTPSDCRYILTPGGLAEQPGQAEHQPESPLNETQRLLLLQQGLLPHSSC | 720 |
| 3D view using mol* of 78_CALML3_TYRO3 | ||||||||||
| PDB file >>>TKFP_128_CALML3_TYRO3 | ENST00000315238 | ENST00000559066 | CALML3 | chr10 | 5567458 | TYRO3 | chr15 | 41854863 | MRYSSLTTKIGGPAPSPSVLNVTGVTQSTMFSCEAHNLKGLASSRTATVHLQALPAAPFNITVTKLSSSNASVAWMPGADGRALLQSCTV QVTQAPGGWEVLAVVVPVPPFTCLLRDLVPATNYSLRVRCANALGPSPYADWVPFQTKGLAPASAPQNLHAIRTDSGLILEWEEVIPEAP LEGPLGPYKLSWVQDNGTQDELTVEGTRANLTGWDPQKDLIVRVCVSNAVGCGPWSQPLVVSSHDRAGQQGPPHSRTSWVPVVLGVLTAL VTAAALALILLRKRRKETRFGQAFDSVMARGEPAVHFRAARSFNRERPERIEATLDSLGISDELKEKLEDVLIPEQQFTLGRMLGKGEFG SVREAQLKQEDGSFVKVAVKMLKADIIASSDIEEFLREAACMKEFDHPHVAKLVGVSLRSRAKGRLPIPMVILPFMKHGDLHAFLLASRI GENPFNLPLQTLIRFMVDIACGMEYLSSRNFIHRDLAARNCMLAEDMTVCVADFGLSRKIYSGDYYRQGCASKLPVKWLALESLADNLYT VQSDVWAFGVTMWEIMTRGQTPYAGIENAEIYNYLIGGNRLKQPPECMEDVYDLMYQCWSADPKQRPSFTCLRMELENILGQLSVLSASQ DPLYINIERAEEPTAGGSLELPGRDQPYSGAGDGSGMGAVGGTPSDCRYILTPGGLAEQPGQAEHQPESPLNETQRLLLLQQGLLPHSSC | 720_CALML3_TYRO3 |
Top |
Comparison of Fusion Protein Isoforms |
 Superimpose the 3D Structures Among All Fusion Protein Isoforms Superimpose the 3D Structures Among All Fusion Protein Isoforms * Download the pdb file and open it from the molstar online viewer. |
 Comparison of the Secondary Structures of Fusion Protein Isoforms Comparison of the Secondary Structures of Fusion Protein Isoforms |
Top |
Comparison of Fusion Protein Sequences/Structures with Known Sequences/Structures from PDB |
Top |
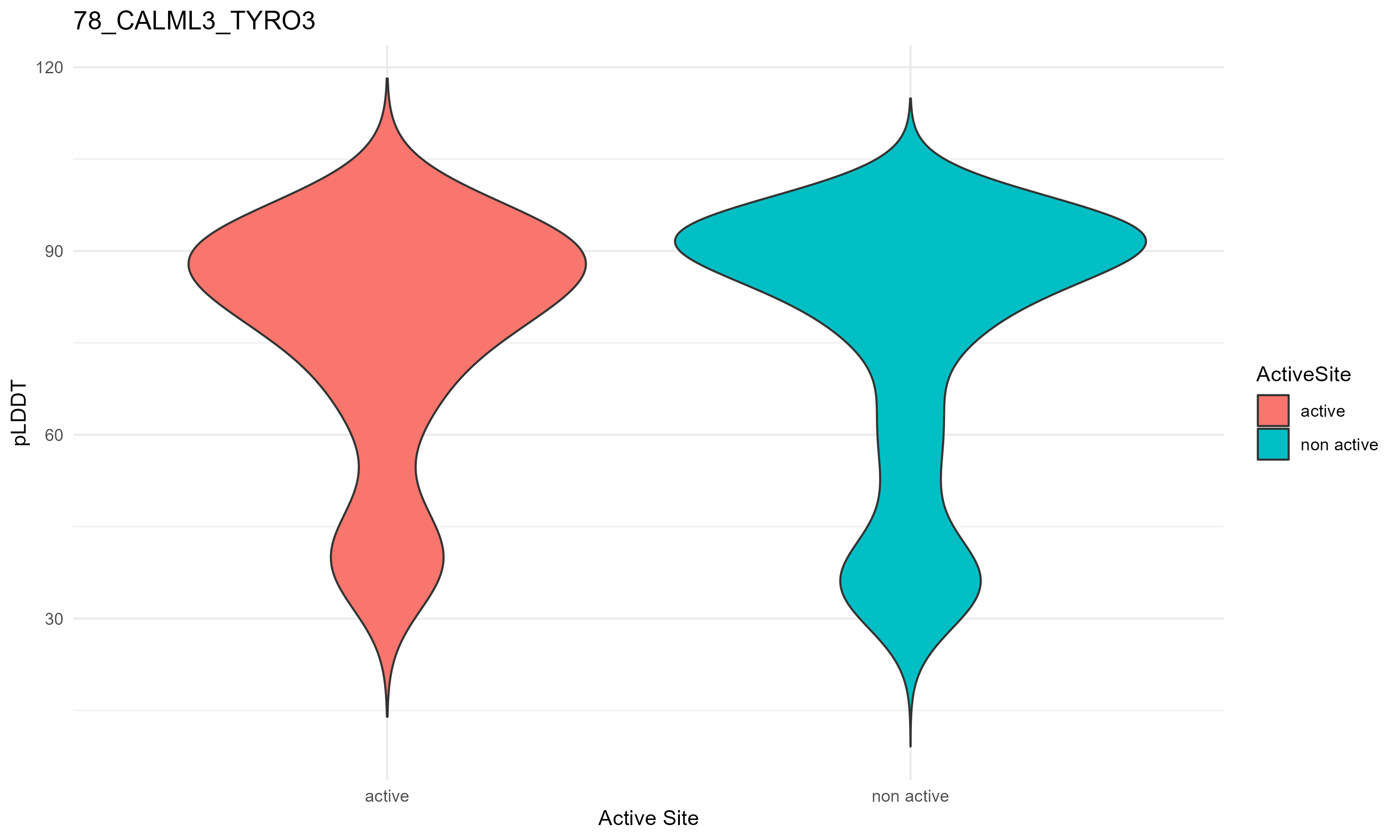
pLDDT score distribution |
 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2* AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. * The blue color at the bottom marks the best active site residues. |
| 78_CALML3_TYRO3.png |
 |
| 78_CALML3_TYRO3.png |
 |
Top |
Potential Active Site Information |
 The potential binding sites of these fusion proteins were identified using SiteMap, a module of the Schrodinger suite. The potential binding sites of these fusion proteins were identified using SiteMap, a module of the Schrodinger suite. |
| Kinase Fusion AA seq ID in KinaseFusionDB | Site score | Size | Dscore | Volume | Exposure | Enclosure | Contact | Phobic | Philic | Balance | Don/Acc | Residues |
Top |
Ramachandran Plot of Kinase Fusion Protein Structure |
 Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. |
| 78_CALML3_TYRO3_ramachandran.png |
 |
Top |
Virtual Screening Results |
 Distribution of the average docking score across all approved kinase inhibitors. Distribution of the average docking score across all approved kinase inhibitors.Distribution of the number of occurrence across all approved kinase inhibitors. |
| 5'-kinase fusion protein case |
| 3'-kinase fusion protein case |
Top |
 Drug information from DrugBank of the top 20 interacting small molecules. Drug information from DrugBank of the top 20 interacting small molecules.* The detailed information of individual kinase inhibitors are available in the download page. |
| Fusion gene name info | Drug | Docking score | Glide g score | Glide energy |
Top |
Kinase-Substrate Information of CALML3_TYRO3 |
 Phosphorylation target of the kinase Phosphorylation target of the kinase(phosphosite, 03-17-2024) |
| Kinase | Kinase UniProt Acc | Kinase species | Substrate | Substrate UniProt Acc | Substrate phosphorylated residues | Substrate phosphorylated sites (+/-7AA) | Domain |
| TYRO3 | Q06418 | human | MLKL | Q8NB16 | Y376 | DRVkstAyLSPQELE | PK_Tyr_Ser-Thr |
| TYRO3 | Q06418 | human | AKT1 | P31749 | T308 | kDGAtMKtFCGtPEy | Pkinase |
 Biological Network Integration of This Kinase and Substrates Biological Network Integration of This Kinase and Substrates (GeneMANIA website) |
 Enriched GO biological processes of the phosphorylation target genes of the kinase Enriched GO biological processes of the phosphorylation target genes of the kinase |
| Kinase | GOID | GO term | P.adjust |
| TYRO3 | ID | Description | 0.00e+00 |
| TYRO3 | GO:0010918 | positive regulation of mitochondrial membrane potential | 2.48e-02 |
| TYRO3 | GO:0015911 | long-chain fatty acid import across plasma membrane | 2.48e-02 |
| TYRO3 | GO:0032070 | regulation of deoxyribonuclease activity | 2.48e-02 |
| TYRO3 | GO:0070673 | response to interleukin-18 | 2.48e-02 |
| TYRO3 | GO:0022605 | mammalian oogenesis stage | 2.48e-02 |
| TYRO3 | GO:0046322 | negative regulation of fatty acid oxidation | 2.48e-02 |
| TYRO3 | GO:0070141 | response to UV-A | 2.48e-02 |
| TYRO3 | GO:0140052 | cellular response to oxidised low-density lipoprotein particle stimulus | 2.48e-02 |
| TYRO3 | GO:0003376 | sphingosine-1-phosphate receptor signaling pathway | 2.48e-02 |
| TYRO3 | GO:0010763 | positive regulation of fibroblast migration | 2.48e-02 |
| TYRO3 | GO:0045725 | positive regulation of glycogen biosynthetic process | 2.48e-02 |
| TYRO3 | GO:0045737 | positive regulation of cyclin-dependent protein serine/threonine kinase activity | 2.48e-02 |
| TYRO3 | GO:0045838 | positive regulation of membrane potential | 2.48e-02 |
| TYRO3 | GO:0070207 | protein homotrimerization | 2.48e-02 |
| TYRO3 | GO:2000402 | negative regulation of lymphocyte migration | 2.48e-02 |
| TYRO3 | GO:0071380 | cellular response to prostaglandin E stimulus | 2.48e-02 |
| TYRO3 | GO:0007252 | I-kappaB phosphorylation | 2.48e-02 |
| TYRO3 | GO:0044539 | long-chain fatty acid import into cell | 2.48e-02 |
| TYRO3 | GO:0060644 | mammary gland epithelial cell differentiation | 2.48e-02 |
| TYRO3 | GO:0070206 | protein trimerization | 2.48e-02 |
| TYRO3 | GO:0070875 | positive regulation of glycogen metabolic process | 2.48e-02 |
| TYRO3 | GO:0043217 | myelin maintenance | 2.48e-02 |
| TYRO3 | GO:0051000 | positive regulation of nitric-oxide synthase activity | 2.48e-02 |
| TYRO3 | GO:0090520 | sphingolipid mediated signaling pathway | 2.48e-02 |
| TYRO3 | GO:1904031 | positive regulation of cyclin-dependent protein kinase activity | 2.48e-02 |
| TYRO3 | GO:0035821 | modulation of process of another organism | 2.48e-02 |
| TYRO3 | GO:0048266 | behavioral response to pain | 2.48e-02 |
| TYRO3 | GO:0060716 | labyrinthine layer blood vessel development | 2.48e-02 |
| TYRO3 | GO:0031998 | regulation of fatty acid beta-oxidation | 2.48e-02 |
| TYRO3 | GO:0090201 | negative regulation of release of cytochrome c from mitochondria | 2.48e-02 |
| TYRO3 | GO:0140354 | lipid import into cell | 2.48e-02 |
| TYRO3 | GO:1902001 | fatty acid transmembrane transport | 2.48e-02 |
| TYRO3 | GO:1902018 | negative regulation of cilium assembly | 2.48e-02 |
| TYRO3 | GO:0071379 | cellular response to prostaglandin stimulus | 2.48e-02 |
| TYRO3 | GO:2000010 | positive regulation of protein localization to cell surface | 2.48e-02 |
| TYRO3 | GO:0032069 | regulation of nuclease activity | 2.48e-02 |
| TYRO3 | GO:0003323 | type B pancreatic cell development | 2.48e-02 |
| TYRO3 | GO:0032891 | negative regulation of organic acid transport | 2.48e-02 |
| TYRO3 | GO:0034695 | response to prostaglandin E | 2.48e-02 |
| TYRO3 | GO:0071676 | negative regulation of mononuclear cell migration | 2.48e-02 |
| TYRO3 | GO:0050995 | negative regulation of lipid catabolic process | 2.48e-02 |
| TYRO3 | GO:0022011 | myelination in peripheral nervous system | 2.48e-02 |
| TYRO3 | GO:0032292 | peripheral nervous system axon ensheathment | 2.48e-02 |
| TYRO3 | GO:0032770 | positive regulation of monooxygenase activity | 2.48e-02 |
| TYRO3 | GO:0003309 | type B pancreatic cell differentiation | 2.48e-02 |
| TYRO3 | GO:0005979 | regulation of glycogen biosynthetic process | 2.48e-02 |
| TYRO3 | GO:0010962 | regulation of glucan biosynthetic process | 2.48e-02 |
| TYRO3 | GO:0014044 | Schwann cell development | 2.48e-02 |
| TYRO3 | GO:0050999 | regulation of nitric-oxide synthase activity | 2.48e-02 |
Top |
Related Drugs to CALML3_TYRO3 |
 Drugs used for this fusion-positive patient. Drugs used for this fusion-positive patient. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Drug | Source | PMID |
 Distribution of the number of studies mentioning CALML3-TYRO3 and kinase inhibitors the PubMed Abstract (04-01-2024) Distribution of the number of studies mentioning CALML3-TYRO3 and kinase inhibitors the PubMed Abstract (04-01-2024) |
| Fusion gene - drug pair 1 | Fusion gene - drug pair 2 | PMID | Publication date | DOI | Study title |
Top |
Related Diseases to CALML3_TYRO3 |
 Diseases that have this fusion gene. Diseases that have this fusion gene. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Disease | Source | PMID |
 Related diseases from the literature mentioned this fusion gene and drug. Related diseases from the literature mentioned this fusion gene and drug. (PubMed, 04-01-2024) |
| MeSH ID | MeSH term |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
Top |
Clinical Trials of the Found Drugs/Small Molecules |
 Statistics of the Clinical Trials of the Found Kinase Inibitors from clinicaltrials.gov (06-17-2024) Statistics of the Clinical Trials of the Found Kinase Inibitors from clinicaltrials.gov (06-17-2024) |
 Clinical Trials from clinicaltrials.gov (06-17-2024) Clinical Trials from clinicaltrials.gov (06-17-2024) |
| Fusion Gene | Kinase Inhibitor | NCT ID | Study Status | Phases | Disease | # Enrolment | Date |

