
|
|||||||
|
Fusion Protein:SETD2-NISCH |
Fusion Gene and Fusion Protein Summary |
 Fusion gene summary Fusion gene summary |
| Fusion partner gene information | Fusion gene name: SETD2-NISCH | FusionPDB ID: 80877 | FusionGDB2.0 ID: 80877 | Hgene | Tgene | Gene symbol | SETD2 | NISCH | Gene ID | 29072 | 11188 |
| Gene name | SET domain containing 2, histone lysine methyltransferase | nischarin | |
| Synonyms | HBP231|HIF-1|HIP-1|HSPC069|HYPB|KMT3A|LLS|SET2|p231HBP | I-1|IR1|IRAS|hIRAS | |
| Cytomap | 3p21.31 | 3p21.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | histone-lysine N-methyltransferase SETD2SET domain containing 2huntingtin interacting protein 1huntingtin yeast partner Bhuntingtin-interacting protein Blysine N-methyltransferase 3Aprotein-lysine N-methyltransferase SETD2 | nischarinI-1 receptor candidate proteinI1R candidate proteinimidazoline receptor 1imidazoline receptor antisera selected | |
| Modification date | 20200315 | 20200320 | |
| UniProtAcc | Q9BYW2 Main function of 5'-partner protein: FUNCTION: Histone methyltransferase that specifically trimethylates 'Lys-36' of histone H3 (H3K36me3) using dimethylated 'Lys-36' (H3K36me2) as substrate (PubMed:16118227, PubMed:19141475, PubMed:21526191, PubMed:21792193, PubMed:23043551, PubMed:27474439). It is capable of trimethylating unmethylated H3K36 (H3K36me0) in vitro (PubMed:19332550). Represents the main enzyme generating H3K36me3, a specific tag for epigenetic transcriptional activation (By similarity). Plays a role in chromatin structure modulation during elongation by coordinating recruitment of the FACT complex and by interacting with hyperphosphorylated POLR2A (PubMed:23325844). Acts as a key regulator of DNA mismatch repair in G1 and early S phase by generating H3K36me3, a mark required to recruit MSH6 subunit of the MutS alpha complex: early recruitment of the MutS alpha complex to chromatin to be replicated allows a quick identification of mismatch DNA to initiate the mismatch repair reaction (PubMed:23622243). Required for DNA double-strand break repair in response to DNA damage: acts by mediating formation of H3K36me3, promoting recruitment of RAD51 and DNA repair via homologous recombination (HR) (PubMed:24843002). Acts as a tumor suppressor (PubMed:24509477). H3K36me3 also plays an essential role in the maintenance of a heterochromatic state, by recruiting DNA methyltransferase DNMT3A (PubMed:27317772). H3K36me3 is also enhanced in intron-containing genes, suggesting that SETD2 recruitment is enhanced by splicing and that splicing is coupled to recruitment of elongating RNA polymerase (PubMed:21792193). Required during angiogenesis (By similarity). Required for endoderm development by promoting embryonic stem cell differentiation toward endoderm: acts by mediating formation of H3K36me3 in distal promoter regions of FGFR3, leading to regulate transcription initiation of FGFR3 (By similarity). In addition to histones, also mediates methylation of other proteins, such as tubulins and STAT1 (PubMed:27518565, PubMed:28753426). Trimethylates 'Lys-40' of alpha-tubulins such as TUBA1B (alpha-TubK40me3); alpha-TubK40me3 is required for normal mitosis and cytokinesis and may be a specific tag in cytoskeletal remodeling (PubMed:27518565). Involved in interferon-alpha-induced antiviral defense by mediating both monomethylation of STAT1 at 'Lys-525' and catalyzing H3K36me3 on promoters of some interferon-stimulated genes (ISGs) to activate gene transcription (PubMed:28753426). {ECO:0000250|UniProtKB:E9Q5F9, ECO:0000269|PubMed:16118227, ECO:0000269|PubMed:19141475, ECO:0000269|PubMed:21526191, ECO:0000269|PubMed:21792193, ECO:0000269|PubMed:23043551, ECO:0000269|PubMed:23325844, ECO:0000269|PubMed:23622243, ECO:0000269|PubMed:24509477, ECO:0000269|PubMed:24843002, ECO:0000269|PubMed:27317772, ECO:0000269|PubMed:27474439, ECO:0000269|PubMed:27518565, ECO:0000269|PubMed:28753426}.; FUNCTION: (Microbial infection) Recruited to the promoters of adenovirus 12 E1A gene in case of infection, possibly leading to regulate its expression. {ECO:0000269|PubMed:11461154}. | Q9Y2I1 Main function of 5'-partner protein: FUNCTION: Acts either as the functional imidazoline-1 receptor (I1R) candidate or as a membrane-associated mediator of the I1R signaling. Binds numerous imidazoline ligands that induces initiation of cell-signaling cascades triggering to cell survival, growth and migration. Its activation by the agonist rilmenidine induces an increase in phosphorylation of mitogen-activated protein kinases MAPK1 and MAPK3 in rostral ventrolateral medulla (RVLM) neurons that exhibited rilmenidine-evoked hypotension (By similarity). Blocking its activation with efaroxan abolished rilmenidine-induced mitogen-activated protein kinase phosphorylation in RVLM neurons (By similarity). Acts as a modulator of Rac-regulated signal transduction pathways (By similarity). Suppresses Rac1-stimulated cell migration by interacting with PAK1 and inhibiting its kinase activity (By similarity). Also blocks Pak-independent Rac signaling by interacting with RAC1 and inhibiting Rac1-stimulated NF-kB response element and cyclin D1 promoter activation (By similarity). Inhibits also LIMK1 kinase activity by reducing LIMK1 'Tyr-508' phosphorylation (By similarity). Inhibits Rac-induced cell migration and invasion in breast and colon epithelial cells (By similarity). Inhibits lamellipodia formation, when overexpressed (By similarity). Plays a role in protection against apoptosis. Involved in association with IRS4 in the enhancement of insulin activation of MAPK1 and MAPK3. When overexpressed, induces a redistribution of cell surface ITGA5 integrin to intracellular endosomal structures. {ECO:0000250, ECO:0000269|PubMed:10882231, ECO:0000269|PubMed:12868002, ECO:0000269|PubMed:15028619, ECO:0000269|PubMed:15028621, ECO:0000269|PubMed:15475348}. | |
| Ensembl transtripts involved in fusion gene | ENST ids | ENST00000409792, ENST00000492397, | ENST00000464280, ENST00000345716, ENST00000420808, ENST00000479054, ENST00000488380, |
| Fusion gene scores for assessment (based on all fusion genes of FusionGDB 2.0) | * DoF score | 27 X 19 X 16=8208 | 6 X 5 X 4=120 |
| # samples | 41 | 6 | |
| ** MAII score | log2(41/8208*10)=-4.32333491610161 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(6/120*10)=-1 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Fusion gene context | PubMed: SETD2 [Title/Abstract] AND NISCH [Title/Abstract] AND fusion [Title/Abstract] | ||
| Fusion neoantigen context | PubMed: SETD2 [Title/Abstract] AND NISCH [Title/Abstract] AND neoantigen [Title/Abstract] | ||
| Most frequent breakpoint (based on all fusion genes of FusionGDB 2.0) | SETD2(47125210)-NISCH(52504875), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | SETD2-NISCH seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. SETD2-NISCH seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. SETD2-NISCH seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. SETD2-NISCH seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | SETD2 | GO:0010569 | regulation of double-strand break repair via homologous recombination | 24843002 |
| Hgene | SETD2 | GO:0018023 | peptidyl-lysine trimethylation | 27518565 |
| Hgene | SETD2 | GO:0018026 | peptidyl-lysine monomethylation | 28753426 |
| Hgene | SETD2 | GO:0032465 | regulation of cytokinesis | 27518565 |
| Hgene | SETD2 | GO:0032727 | positive regulation of interferon-alpha production | 28753426 |
| Hgene | SETD2 | GO:0034340 | response to type I interferon | 28753426 |
| Hgene | SETD2 | GO:0051607 | defense response to virus | 28753426 |
| Hgene | SETD2 | GO:0097198 | histone H3-K36 trimethylation | 23043551|24843002|26002201|27474439|28753426 |
| Hgene | SETD2 | GO:0097676 | histone H3-K36 dimethylation | 26002201 |
| Hgene | SETD2 | GO:1902850 | microtubule cytoskeleton organization involved in mitosis | 27518565 |
| Hgene | SETD2 | GO:1905634 | regulation of protein localization to chromatin | 24843002 |
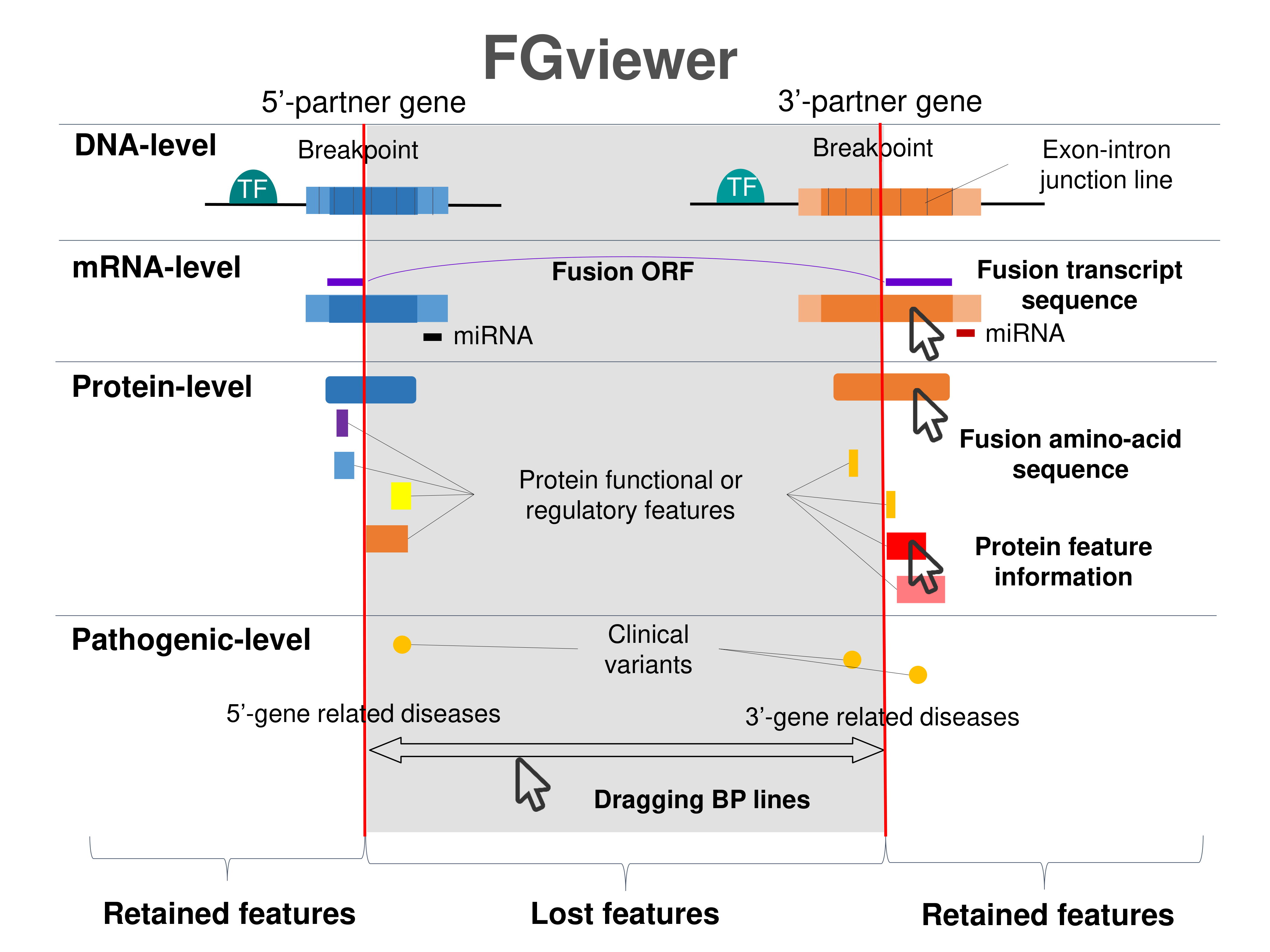
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr3:47125210/chr3:52504875) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Retention analysis results of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features, are available here. Retention analysis results of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features, are available here. |
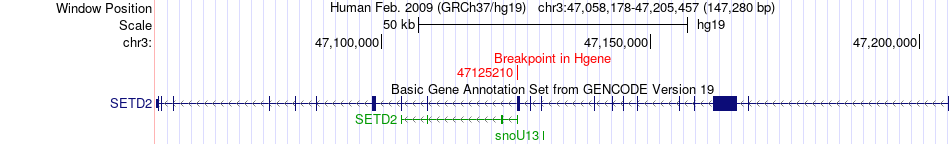
 Fusion gene breakpoints across SETD2 (5'-gene) Fusion gene breakpoints across SETD2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
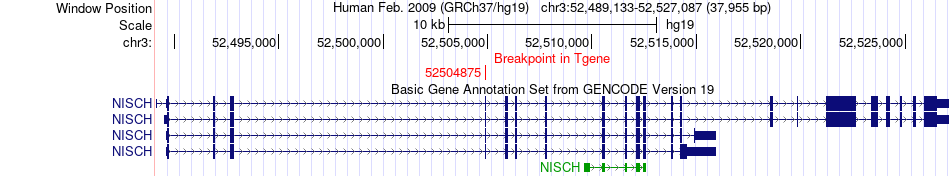
 Fusion gene breakpoints across NISCH (3'-gene) Fusion gene breakpoints across NISCH (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
Top |
Fusion Amino Acid Sequences |
 Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000409792 | SETD2 | chr3 | 47125210 | - | ENST00000479054 | NISCH | chr3 | 52504875 | + | 10844 | 6103 | 1 | 10257 | 3418 |
| ENST00000409792 | SETD2 | chr3 | 47125210 | - | ENST00000345716 | NISCH | chr3 | 52504875 | + | 10847 | 6103 | 1 | 10257 | 3418 |
| ENST00000409792 | SETD2 | chr3 | 47125210 | - | ENST00000488380 | NISCH | chr3 | 52504875 | + | 8886 | 6103 | 1 | 7494 | 2497 |
| ENST00000409792 | SETD2 | chr3 | 47125210 | - | ENST00000420808 | NISCH | chr3 | 52504875 | + | 8307 | 6103 | 1 | 7290 | 2429 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000409792 | ENST00000479054 | SETD2 | chr3 | 47125210 | - | NISCH | chr3 | 52504875 | + | 0.000898658 | 0.9991014 |
| ENST00000409792 | ENST00000345716 | SETD2 | chr3 | 47125210 | - | NISCH | chr3 | 52504875 | + | 0.000901776 | 0.9990983 |
| ENST00000409792 | ENST00000488380 | SETD2 | chr3 | 47125210 | - | NISCH | chr3 | 52504875 | + | 0.000463816 | 0.99953616 |
| ENST00000409792 | ENST00000420808 | SETD2 | chr3 | 47125210 | - | NISCH | chr3 | 52504875 | + | 0.000557386 | 0.99944264 |
 Predicted full-length fusion amino acid sequences. For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among all the predicted ones. Predicted full-length fusion amino acid sequences. For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among all the predicted ones. |
Get the fusion protein sequences from here. |
| Fusion protein sequence information is available in the fasta format. >FusionGDB ID_FusionGDB isoform ID_FGname_Hgene_Hchr_Hbp_Henst_Tgene_Tchr_Tbp_Tenst_length(fusion AA) seq_BP |
Top |
Fusion Protein Breakpoint Sequences for SETD2-NISCH |
 +/-13 AA sequence from the breakpoints of the fusion protein sequences. +/-13 AA sequence from the breakpoints of the fusion protein sequences. |
| Hgene | Hchr | Hbp | Tgene | Tchr | Tbp | Length(fusion protein) | BP in fusion protein | Peptide |
| SETD2 | chr3 | 47125210 | NISCH | chr3 | 52504875 | 6103 | 2032 | SDLATKLLDSWKDLKEINGITAALAE |
Top |
Potential FusionNeoAntigen Information of SETD2-NISCH in HLA I |
 Multiple sequence alignments of the potential FusionNeoAntigens per fusion breakpoints. If the MSA is empty, then it means that there were predicted fusion neoantigens in this fusion breakpoint, but those predicted fusion neoantigens were not across the breakpoint, which is not fusion-specific. Multiple sequence alignments of the potential FusionNeoAntigens per fusion breakpoints. If the MSA is empty, then it means that there were predicted fusion neoantigens in this fusion breakpoint, but those predicted fusion neoantigens were not across the breakpoint, which is not fusion-specific. |
| SETD2-NISCH_47125210_52504875.msa |
 Potential FusionNeoAntigen Information Potential FusionNeoAntigen Information* We used NetMHCpan v4.1 (%rank<0.5) and deepHLApan v1.1 (immunogenic score>0.5) |
| Fusion gene | Hchr | Hbp | Tgene | Tchr | Tbp | HLA I | FusionNeoAntigen peptide | Binding score | Immunogenic score | Neoantigen start (at BP 13) | Neoantigen end (at BP 13) |
| SETD2-NISCH | chr3 | 47125210 | chr3 | 52504875 | 6103 | HLA-B50:02 | DLKEINGITAA | 0.9739 | 0.5262 | 12 | 23 |
| SETD2-NISCH | chr3 | 47125210 | chr3 | 52504875 | 6103 | HLA-B51:07 | DSWKDLKEI | 0.9956 | 0.6819 | 8 | 17 |
Top |
Potential FusionNeoAntigen Information of SETD2-NISCH in HLA II |
 Multiple sequence alignments of the potential FusionNeoAntigens per fusion breakpoints. If the MSA is empty, then it means that there were predicted fusion neoantigens in this fusion breakpoint, but those predicted fusion neoantigens were not across the breakpoint, which is not fusion-specific. Multiple sequence alignments of the potential FusionNeoAntigens per fusion breakpoints. If the MSA is empty, then it means that there were predicted fusion neoantigens in this fusion breakpoint, but those predicted fusion neoantigens were not across the breakpoint, which is not fusion-specific. |
| SETD2-NISCH_47125210_52504875.msa |
 Potential FusionNeoAntigen Information Potential FusionNeoAntigen Information * We used NetMHCIIpan v4.1 (%rank<0.5). |
| Fusion gene | Hchr | Hbp | Tgene | Tchr | Tbp | HLA II | FusionNeoAntigen peptide | Neoantigen start (at BP 13) | Neoantigen end (at BP 13) |
| SETD2-NISCH | chr3 | 47125210 | chr3 | 52504875 | 6103 | DRB1-1457 | TKLLDSWKDLKEING | 4 | 19 |
Top |
Fusion breakpoint peptide structures of SETD2-NISCH |
 3D structures of the fusion breakpoint peptide of 14AA sequence that have potential fusion neoantigens 3D structures of the fusion breakpoint peptide of 14AA sequence that have potential fusion neoantigens* The minimum length of the amino acid sequence in RoseTTAFold is 14AA. Here, we predicted the 14AA fusion protein breakpoint sequence not the fusion neoantigen peptide, which is shorter than 14 AA. |
| File name | BPseq | Hgene | Tgene | Hchr | Hbp | Tchr | Tbp | AAlen |
| 5206 | LLDSWKDLKEINGI | SETD2 | NISCH | chr3 | 47125210 | chr3 | 52504875 | 6103 |
Top |
Filtering FusionNeoAntigens Through Checking the Interaction with HLAs in 3D of SETD2-NISCH |
 Virtual screening between 25 HLAs (from PDB) and FusionNeoAntigens Virtual screening between 25 HLAs (from PDB) and FusionNeoAntigens* We used Glide to predict the interaction between HLAs and neoantigens. |
| HLA allele | PDB ID | File name | BPseq | Docking score | Glide score |
| HLA-B14:02 | 3BVN | 5206 | LLDSWKDLKEINGI | -6.39555 | -6.50735 |
| HLA-B14:02 | 3BVN | 5206 | LLDSWKDLKEINGI | -3.70119 | -4.74429 |
| HLA-B52:01 | 3W39 | 5206 | LLDSWKDLKEINGI | -7.02029 | -7.13209 |
| HLA-B52:01 | 3W39 | 5206 | LLDSWKDLKEINGI | -6.19524 | -7.23834 |
| HLA-B18:01 | 4JQV | 5206 | LLDSWKDLKEINGI | -4.46522 | -4.57702 |
| HLA-A11:01 | 4UQ2 | 5206 | LLDSWKDLKEINGI | -9.23254 | -10.2756 |
| HLA-A11:01 | 4UQ2 | 5206 | LLDSWKDLKEINGI | -8.60916 | -8.72096 |
| HLA-A24:02 | 5HGA | 5206 | LLDSWKDLKEINGI | -9.68299 | -9.79479 |
| HLA-A24:02 | 5HGA | 5206 | LLDSWKDLKEINGI | -5.25555 | -6.29865 |
| HLA-B27:05 | 6PYJ | 5206 | LLDSWKDLKEINGI | -5.8374 | -5.9492 |
| HLA-B44:05 | 3DX8 | 5206 | LLDSWKDLKEINGI | -7.84132 | -7.95312 |
| HLA-B44:05 | 3DX8 | 5206 | LLDSWKDLKEINGI | -2.56171 | -3.60481 |
Top |
Vaccine Design for the FusionNeoAntigens of SETD2-NISCH |
 mRNA and peptide sequences of FusionNeoAntigens that have potential interaction with HLA-Is. mRNA and peptide sequences of FusionNeoAntigens that have potential interaction with HLA-Is. |
| Fusion gene | Hchr | Hbp | Tchr | Tbp | Start in +/-13AA | End in +/-13AA | FusionNeoAntigen peptide sequence | FusionNeoAntigen RNA sequence |
| SETD2-NISCH | chr3 | 47125210 | chr3 | 52504875 | 12 | 23 | DLKEINGITAA | AAGGAGATAAATGGCATCACCGCGGCACTGGCT |
| SETD2-NISCH | chr3 | 47125210 | chr3 | 52504875 | 8 | 17 | DSWKDLKEI | TGGAAAGACCTAAAGGAGATAAATGGC |
 mRNA and peptide sequences of FusionNeoAntigens that have potential interaction with HLA-IIs. mRNA and peptide sequences of FusionNeoAntigens that have potential interaction with HLA-IIs. |
| Fusion gene | Hchr | Hbp | Tchr | Tbp | Start in +/-13AA | End in +/-13AA | FusionNeoAntigen peptide | FusionNEoAntigen RNA sequence |
| SETD2-NISCH | chr3 | 47125210 | chr3 | 52504875 | 4 | 19 | TKLLDSWKDLKEING | CTCCTGGACAGTTGGAAAGACCTAAAGGAGATAAATGGCATCACC |
Top |
Information of the samples that have these potential fusion neoantigens of SETD2-NISCH |
 These samples were reported as having these fusion breakpoints. For individual breakpoints, we checked the open reading frames considering multiple gene isoforms and chose the in-frame fusion genes only. Then, we made fusion protein sequences and predicted the fusion neoantigens. These fusion-positive samples may have these potential fusion neoantigens. These samples were reported as having these fusion breakpoints. For individual breakpoints, we checked the open reading frames considering multiple gene isoforms and chose the in-frame fusion genes only. Then, we made fusion protein sequences and predicted the fusion neoantigens. These fusion-positive samples may have these potential fusion neoantigens. |
| Cancer type | Fusion gene | Hchr | Hbp | Henst | Tchr | Tbp | Tenst | Sample |
| MESO | SETD2-NISCH | chr3 | 47125210 | ENST00000409792 | chr3 | 52504875 | ENST00000345716 | TCGA-ZN-A9VO-01A |
Top |
Potential target of CAR-T therapy development for SETD2-NISCH |
 Predicted 3D structure. We used RoseTTAFold. Predicted 3D structure. We used RoseTTAFold. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, to provide the retention of the transmembrane domain, we only show the protein feature retention information of those transmembrane features Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, to provide the retention of the transmembrane domain, we only show the protein feature retention information of those transmembrane features* Minus value of BPloci means that the break point is located before the CDS. |
| - In-frame and retained 'Transmembrane'. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
 Subcellular localization prediction of the transmembrane domain retained fusion proteins Subcellular localization prediction of the transmembrane domain retained fusion proteins* We used DeepLoc 1.0. The order of the X-axis of the barplot is as follows: Entry_ID, Localization, Type, Nucleus, Cytoplasm, Extracellular, Mitochondrion, Cell_membrane, Endoplasmic_reticulum, Plastid, Golgi.apparatus, Lysosome.Vacuole, Peroxisome. Y-axis is the output score of DeepLoc. Clicking the image will open a new tab with a large image. |
| Hgene | Hchr | Hbp | Henst | Tgene | Tchr | Tbp | Tenst | DeepLoc result |
Top |
Related Drugs to SETD2-NISCH |
 Drugs used for this fusion-positive patient. Drugs used for this fusion-positive patient. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Drug | Source | PMID |
Top |
Related Diseases to SETD2-NISCH |
 Diseases that have this fusion gene. Diseases that have this fusion gene. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Disease | Source | PMID |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
| Hgene | SETD2 | C0279702 | Conventional (Clear Cell) Renal Cell Carcinoma | 6 | CGI;CTD_human;UNIPROT |
| Hgene | SETD2 | C4085873 | LUSCAN-LUMISH SYNDROME | 4 | CTD_human;GENOMICS_ENGLAND;UNIPROT |
| Hgene | SETD2 | C0007134 | Renal Cell Carcinoma | 2 | CTD_human |
| Hgene | SETD2 | C0023467 | Leukemia, Myelocytic, Acute | 2 | UNIPROT |
| Hgene | SETD2 | C1266042 | Chromophobe Renal Cell Carcinoma | 2 | CTD_human |
| Hgene | SETD2 | C1266043 | Sarcomatoid Renal Cell Carcinoma | 2 | CTD_human |
| Hgene | SETD2 | C1266044 | Collecting Duct Carcinoma of the Kidney | 2 | CTD_human |
| Hgene | SETD2 | C1306837 | Papillary Renal Cell Carcinoma | 2 | CTD_human |
| Hgene | SETD2 | C1961102 | Precursor Cell Lymphoblastic Leukemia Lymphoma | 2 | UNIPROT |
| Hgene | SETD2 | C0006142 | Malignant neoplasm of breast | 1 | CTD_human |
| Hgene | SETD2 | C0010606 | Adenoid Cystic Carcinoma | 1 | CTD_human |
| Hgene | SETD2 | C0010701 | Phyllodes Tumor | 1 | CTD_human |
| Hgene | SETD2 | C0023418 | leukemia | 1 | CTD_human |
| Hgene | SETD2 | C0033578 | Prostatic Neoplasms | 1 | CTD_human |
| Hgene | SETD2 | C0175695 | Sotos' syndrome | 1 | ORPHANET |
| Hgene | SETD2 | C0206656 | Embryonal Rhabdomyosarcoma | 1 | CTD_human |
| Hgene | SETD2 | C0345967 | Malignant mesothelioma | 1 | CTD_human |
| Hgene | SETD2 | C0376358 | Malignant neoplasm of prostate | 1 | CTD_human |
| Hgene | SETD2 | C0600066 | Malignant Cystosarcoma Phyllodes | 1 | CTD_human |
| Hgene | SETD2 | C0678222 | Breast Carcinoma | 1 | CTD_human |
| Hgene | SETD2 | C0920269 | Microsatellite Instability | 1 | CTD_human |
| Hgene | SETD2 | C1257931 | Mammary Neoplasms, Human | 1 | CTD_human |
| Hgene | SETD2 | C1458155 | Mammary Neoplasms | 1 | CTD_human |
| Hgene | SETD2 | C1535926 | Neurodevelopmental Disorders | 1 | CTD_human |
| Hgene | SETD2 | C1721098 | Replication Error Phenotype | 1 | CTD_human |
| Hgene | SETD2 | C3714756 | Intellectual Disability | 1 | GENOMICS_ENGLAND |
| Hgene | SETD2 | C4704874 | Mammary Carcinoma, Human | 1 | CTD_human |

