| UTHEALTH HOME ABOUT SBMI A-Z WEBMAIL INSIDE THE UNIVERSITY |

|
|||||||
|
Fusion Protein:EPHA2-LSM14B |
Fusion Protein Summary |
 Fusion gene summary Fusion gene summary |
| Fusion partner gene information | Fusion gene name: EPHA2-LSM14B | FusionPDB ID: 26954 | FusionGDB2.0 ID: 26954 | Hgene | Tgene | Gene symbol | EPHA2 | LSM14B | Gene ID | 1969 | 149986 |
| Gene name | EPH receptor A2 | LSM family member 14B | |
| Synonyms | ARCC2|CTPA|CTPP1|CTRCT6|ECK | C20orf40|FAM61B|FT005|LSM13|RAP55B|bA11M20.3 | |
| Cytomap | 1p36.13 | 20q13.33 | |
| Type of gene | protein-coding | protein-coding | |
| Description | ephrin type-A receptor 2epithelial cell receptor protein tyrosine kinasesoluble EPHA2 variant 1tyrosine-protein kinase receptor ECK | protein LSM14 homolog BLSM14 homolog BLSM14B, SCD6 homolog BRNA-associated protein 55Bfamily with sequence similarity 61, member BhRAP55B | |
| Modification date | 20200313 | 20200327 | |
| UniProtAcc | P29317 | Q9BX40 | |
| Ensembl transtripts involved in fusion gene | ENST ids | ENST00000358432, ENST00000461614, | ENST00000253001, ENST00000279068, ENST00000370915, |
| Fusion gene scores for assessment (based on all fusion genes of FusionGDB 2.0) | * DoF score | 6 X 5 X 4=120 | 3 X 4 X 2=24 |
| # samples | 8 | 4 | |
| ** MAII score | log2(8/120*10)=-0.584962500721156 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(4/24*10)=0.736965594166206 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | |
| Context (manual curation of fusion genes in FusionPDB) | PubMed: EPHA2 [Title/Abstract] AND LSM14B [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint (based on all fusion genes of FusionGDB 2.0) | EPHA2(16474873)-LSM14B(60699673), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | EPHA2-LSM14B seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. EPHA2-LSM14B seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. EPHA2-LSM14B seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. EPHA2-LSM14B seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | EPHA2 | GO:0008630 | intrinsic apoptotic signaling pathway in response to DNA damage | 18339848 |
| Hgene | EPHA2 | GO:0033628 | regulation of cell adhesion mediated by integrin | 10655584 |
| Hgene | EPHA2 | GO:0043491 | protein kinase B signaling | 19573808 |
| Hgene | EPHA2 | GO:0048013 | ephrin receptor signaling pathway | 10655584|20861311 |
| Hgene | EPHA2 | GO:0051898 | negative regulation of protein kinase B signaling | 19573808 |
 Fusion gene breakpoints across EPHA2 (5'-gene) Fusion gene breakpoints across EPHA2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
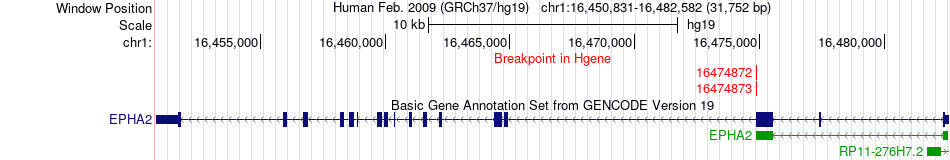 |
 Fusion gene breakpoints across LSM14B (3'-gene) Fusion gene breakpoints across LSM14B (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
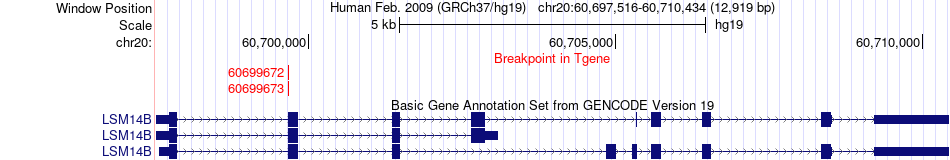 |
Top |
Fusion Gene Sample Information |
 Fusion gene information from FusionGDB2.0. Fusion gene information from FusionGDB2.0. |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | UCS | TCGA-N9-A4PZ-01A | EPHA2 | chr1 | 16474873 | - | LSM14B | chr20 | 60699673 | + |
| ChimerDB4 | UCS | TCGA-N9-A4PZ | EPHA2 | chr1 | 16474872 | - | LSM14B | chr20 | 60699672 | + |
| ChimerDB4 | UCS | TCGA-N9-A4PZ | EPHA2 | chr1 | 16474873 | - | LSM14B | chr20 | 60699673 | + |
Top |
Fusion ORF Analysis |
 Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000358432 | EPHA2 | chr1 | 16474873 | - | ENST00000370915 | LSM14B | chr20 | 60699673 | + | 1724 | 978 | 155 | 1516 | 453 |
| ENST00000358432 | EPHA2 | chr1 | 16474873 | - | ENST00000253001 | LSM14B | chr20 | 60699673 | + | 3239 | 978 | 155 | 2008 | 617 |
| ENST00000358432 | EPHA2 | chr1 | 16474873 | - | ENST00000279068 | LSM14B | chr20 | 60699673 | + | 3239 | 978 | 155 | 2008 | 617 |
| ENST00000461614 | EPHA2 | chr1 | 16474873 | - | ENST00000370915 | LSM14B | chr20 | 60699673 | + | 1621 | 875 | 13 | 1413 | 466 |
| ENST00000461614 | EPHA2 | chr1 | 16474873 | - | ENST00000253001 | LSM14B | chr20 | 60699673 | + | 3136 | 875 | 13 | 1905 | 630 |
| ENST00000461614 | EPHA2 | chr1 | 16474873 | - | ENST00000279068 | LSM14B | chr20 | 60699673 | + | 3136 | 875 | 13 | 1905 | 630 |
| ENST00000358432 | EPHA2 | chr1 | 16474872 | - | ENST00000370915 | LSM14B | chr20 | 60699672 | + | 1724 | 978 | 155 | 1516 | 453 |
| ENST00000358432 | EPHA2 | chr1 | 16474872 | - | ENST00000253001 | LSM14B | chr20 | 60699672 | + | 3239 | 978 | 155 | 2008 | 617 |
| ENST00000358432 | EPHA2 | chr1 | 16474872 | - | ENST00000279068 | LSM14B | chr20 | 60699672 | + | 3239 | 978 | 155 | 2008 | 617 |
| ENST00000461614 | EPHA2 | chr1 | 16474872 | - | ENST00000370915 | LSM14B | chr20 | 60699672 | + | 1621 | 875 | 13 | 1413 | 466 |
| ENST00000461614 | EPHA2 | chr1 | 16474872 | - | ENST00000253001 | LSM14B | chr20 | 60699672 | + | 3136 | 875 | 13 | 1905 | 630 |
| ENST00000461614 | EPHA2 | chr1 | 16474872 | - | ENST00000279068 | LSM14B | chr20 | 60699672 | + | 3136 | 875 | 13 | 1905 | 630 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000358432 | ENST00000370915 | EPHA2 | chr1 | 16474873 | - | LSM14B | chr20 | 60699673 | + | 0.009616244 | 0.99038374 |
| ENST00000358432 | ENST00000253001 | EPHA2 | chr1 | 16474873 | - | LSM14B | chr20 | 60699673 | + | 0.001953909 | 0.99804604 |
| ENST00000358432 | ENST00000279068 | EPHA2 | chr1 | 16474873 | - | LSM14B | chr20 | 60699673 | + | 0.001545863 | 0.9984541 |
| ENST00000461614 | ENST00000370915 | EPHA2 | chr1 | 16474873 | - | LSM14B | chr20 | 60699673 | + | 0.012815283 | 0.9871847 |
| ENST00000461614 | ENST00000253001 | EPHA2 | chr1 | 16474873 | - | LSM14B | chr20 | 60699673 | + | 0.002736484 | 0.99726355 |
| ENST00000461614 | ENST00000279068 | EPHA2 | chr1 | 16474873 | - | LSM14B | chr20 | 60699673 | + | 0.002154937 | 0.99784505 |
| ENST00000358432 | ENST00000370915 | EPHA2 | chr1 | 16474872 | - | LSM14B | chr20 | 60699672 | + | 0.009616244 | 0.99038374 |
| ENST00000358432 | ENST00000253001 | EPHA2 | chr1 | 16474872 | - | LSM14B | chr20 | 60699672 | + | 0.001953909 | 0.99804604 |
| ENST00000358432 | ENST00000279068 | EPHA2 | chr1 | 16474872 | - | LSM14B | chr20 | 60699672 | + | 0.001545863 | 0.9984541 |
| ENST00000461614 | ENST00000370915 | EPHA2 | chr1 | 16474872 | - | LSM14B | chr20 | 60699672 | + | 0.012815283 | 0.9871847 |
| ENST00000461614 | ENST00000253001 | EPHA2 | chr1 | 16474872 | - | LSM14B | chr20 | 60699672 | + | 0.002736484 | 0.99726355 |
| ENST00000461614 | ENST00000279068 | EPHA2 | chr1 | 16474872 | - | LSM14B | chr20 | 60699672 | + | 0.002154937 | 0.99784505 |
Top |
Fusion Amino Acid Sequences |
 For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >FusionGDB ID_FusionGDB isoform ID_FGname_Hgene_Hchr_Hbp_Henst_Tgene_Tchr_Tbp_Tenst_length(fusion AA) seq_BP >26954_26954_1_EPHA2-LSM14B_EPHA2_chr1_16474872_ENST00000358432_LSM14B_chr20_60699672_ENST00000253001_length(amino acids)=617AA_BP=274 MELQAARACFALLWGCALAAAAAAQGKEVVLLDFAAAGGELGWLTHPYGKGWDLMQNIMNDMPIYMYSVCNVMSGDQDNWLRTNWVYRGE AERIFIELKFTVRDCNSFPGGASSCKETFNLYYAESDLDYGTNFQKRLFTKIDTIAPDEITVSSDFEARHVKLNVEERSVGPLTRKGFYL AFQDIGACVALLSVRVYYKKCPELLQGLAHFPETIAGSDAPSLATVAGTCVDHAVVPPGGEEPRMHCAVDGEWLVPIGQCLCQAGYEKVE DACQVRSFGTEDRPTDRPAPPREEIYEYIIFRGSDIKDITVCEPPKAQHTLPQDPAIVQSSLGSASASPFQPHVPYSPFRGMAPYGPLAA SSLLSQQYAASLGLEKLVSPPASAAASSPSSSPSPQPVSELDLSSEPQQLTAKGNSSLGELHAVLQTILRARGKAADRMTVAVADHLPSP CSRRRSGNRRTRNRSRGQNRPTNVKENTIKFEGDFDFESANAQFNREELDKEFKKKLNFKDDKAEKGEEKDLAVVTQSAEAPAEEDLLGP -------------------------------------------------------------- >26954_26954_2_EPHA2-LSM14B_EPHA2_chr1_16474872_ENST00000358432_LSM14B_chr20_60699672_ENST00000279068_length(amino acids)=617AA_BP=274 MELQAARACFALLWGCALAAAAAAQGKEVVLLDFAAAGGELGWLTHPYGKGWDLMQNIMNDMPIYMYSVCNVMSGDQDNWLRTNWVYRGE AERIFIELKFTVRDCNSFPGGASSCKETFNLYYAESDLDYGTNFQKRLFTKIDTIAPDEITVSSDFEARHVKLNVEERSVGPLTRKGFYL AFQDIGACVALLSVRVYYKKCPELLQGLAHFPETIAGSDAPSLATVAGTCVDHAVVPPGGEEPRMHCAVDGEWLVPIGQCLCQAGYEKVE DACQVRSFGTEDRPTDRPAPPREEIYEYIIFRGSDIKDITVCEPPKAQHTLPQDPAIVQSSLGSASASPFQPHVPYSPFRGMAPYGPLAA SSLLSQQYAASLGLGAGFPSIPVGKSPMVEQAVQTGSADNLNAKKLLPGKGTTGTQLNGRQAQPSSKTASDVVQPAAVQAQGQVNDENRR PQRRRSGNRRTRNRSRGQNRPTNVKENTIKFEGDFDFESANAQFNREELDKEFKKKLNFKDDKAEKGEEKDLAVVTQSAEAPAEEDLLGP -------------------------------------------------------------- >26954_26954_3_EPHA2-LSM14B_EPHA2_chr1_16474872_ENST00000358432_LSM14B_chr20_60699672_ENST00000370915_length(amino acids)=453AA_BP=274 MELQAARACFALLWGCALAAAAAAQGKEVVLLDFAAAGGELGWLTHPYGKGWDLMQNIMNDMPIYMYSVCNVMSGDQDNWLRTNWVYRGE AERIFIELKFTVRDCNSFPGGASSCKETFNLYYAESDLDYGTNFQKRLFTKIDTIAPDEITVSSDFEARHVKLNVEERSVGPLTRKGFYL AFQDIGACVALLSVRVYYKKCPELLQGLAHFPETIAGSDAPSLATVAGTCVDHAVVPPGGEEPRMHCAVDGEWLVPIGQCLCQAGYEKVE DACQVRSFGTEDRPTDRPAPPREEIYEYIIFRGSDIKDITVCEPPKAQHTLPQDPAIVQSSLGSASASPFQPHVPYSPFRGMAPYGPLAA SSLLSQQYAASLGLEKLVSPPASAAASSPSSSPSPQPVSELDLSSEPQQLTAKGNSSLGELHAVLQTILRARGKAADRMTVAVADHLPSP -------------------------------------------------------------- >26954_26954_4_EPHA2-LSM14B_EPHA2_chr1_16474872_ENST00000461614_LSM14B_chr20_60699672_ENST00000253001_length(amino acids)=630AA_BP=287 MRAGRRAGADTEAGVQACGCAGAGLGGIGPRARSAAWSSRQPAPASPCCGAVRWPRPRRRRARKWDLMQNIMNDMPIYMYSVCNVMSGDQ DNWLRTNWVYRGEAERIFIELKFTVRDCNSFPGGASSCKETFNLYYAESDLDYGTNFQKRLFTKIDTIAPDEITVSSDFEARHVKLNVEE RSVGPLTRKGFYLAFQDIGACVALLSVRVYYKKCPELLQGLAHFPETIAGSDAPSLATVAGTCVDHAVVPPGGEEPRMHCAVDGEWLVPI GQCLCQAGYEKVEDACQVRSFGTEDRPTDRPAPPREEIYEYIIFRGSDIKDITVCEPPKAQHTLPQDPAIVQSSLGSASASPFQPHVPYS PFRGMAPYGPLAASSLLSQQYAASLGLEKLVSPPASAAASSPSSSPSPQPVSELDLSSEPQQLTAKGNSSLGELHAVLQTILRARGKAAD RMTVAVADHLPSPCSRRRSGNRRTRNRSRGQNRPTNVKENTIKFEGDFDFESANAQFNREELDKEFKKKLNFKDDKAEKGEEKDLAVVTQ SAEAPAEEDLLGPNCYYDKSKSFFDNISSELKTSSRRTTWAEERKLNTETFGVSGRFLRGRSSRGGFRGGRGNGTTRRNPTSHRAGTGRV -------------------------------------------------------------- >26954_26954_5_EPHA2-LSM14B_EPHA2_chr1_16474872_ENST00000461614_LSM14B_chr20_60699672_ENST00000279068_length(amino acids)=630AA_BP=287 MRAGRRAGADTEAGVQACGCAGAGLGGIGPRARSAAWSSRQPAPASPCCGAVRWPRPRRRRARKWDLMQNIMNDMPIYMYSVCNVMSGDQ DNWLRTNWVYRGEAERIFIELKFTVRDCNSFPGGASSCKETFNLYYAESDLDYGTNFQKRLFTKIDTIAPDEITVSSDFEARHVKLNVEE RSVGPLTRKGFYLAFQDIGACVALLSVRVYYKKCPELLQGLAHFPETIAGSDAPSLATVAGTCVDHAVVPPGGEEPRMHCAVDGEWLVPI GQCLCQAGYEKVEDACQVRSFGTEDRPTDRPAPPREEIYEYIIFRGSDIKDITVCEPPKAQHTLPQDPAIVQSSLGSASASPFQPHVPYS PFRGMAPYGPLAASSLLSQQYAASLGLGAGFPSIPVGKSPMVEQAVQTGSADNLNAKKLLPGKGTTGTQLNGRQAQPSSKTASDVVQPAA VQAQGQVNDENRRPQRRRSGNRRTRNRSRGQNRPTNVKENTIKFEGDFDFESANAQFNREELDKEFKKKLNFKDDKAEKGEEKDLAVVTQ SAEAPAEEDLLGPNCYYDKSKSFFDNISSELKTSSRRTTWAEERKLNTETFGVSGRFLRGRSSRGGFRGGRGNGTTRRNPTSHRAGTGRV -------------------------------------------------------------- >26954_26954_6_EPHA2-LSM14B_EPHA2_chr1_16474872_ENST00000461614_LSM14B_chr20_60699672_ENST00000370915_length(amino acids)=466AA_BP=287 MRAGRRAGADTEAGVQACGCAGAGLGGIGPRARSAAWSSRQPAPASPCCGAVRWPRPRRRRARKWDLMQNIMNDMPIYMYSVCNVMSGDQ DNWLRTNWVYRGEAERIFIELKFTVRDCNSFPGGASSCKETFNLYYAESDLDYGTNFQKRLFTKIDTIAPDEITVSSDFEARHVKLNVEE RSVGPLTRKGFYLAFQDIGACVALLSVRVYYKKCPELLQGLAHFPETIAGSDAPSLATVAGTCVDHAVVPPGGEEPRMHCAVDGEWLVPI GQCLCQAGYEKVEDACQVRSFGTEDRPTDRPAPPREEIYEYIIFRGSDIKDITVCEPPKAQHTLPQDPAIVQSSLGSASASPFQPHVPYS PFRGMAPYGPLAASSLLSQQYAASLGLEKLVSPPASAAASSPSSSPSPQPVSELDLSSEPQQLTAKGNSSLGELHAVLQTILRARGKAAD -------------------------------------------------------------- >26954_26954_7_EPHA2-LSM14B_EPHA2_chr1_16474873_ENST00000358432_LSM14B_chr20_60699673_ENST00000253001_length(amino acids)=617AA_BP=274 MELQAARACFALLWGCALAAAAAAQGKEVVLLDFAAAGGELGWLTHPYGKGWDLMQNIMNDMPIYMYSVCNVMSGDQDNWLRTNWVYRGE AERIFIELKFTVRDCNSFPGGASSCKETFNLYYAESDLDYGTNFQKRLFTKIDTIAPDEITVSSDFEARHVKLNVEERSVGPLTRKGFYL AFQDIGACVALLSVRVYYKKCPELLQGLAHFPETIAGSDAPSLATVAGTCVDHAVVPPGGEEPRMHCAVDGEWLVPIGQCLCQAGYEKVE DACQVRSFGTEDRPTDRPAPPREEIYEYIIFRGSDIKDITVCEPPKAQHTLPQDPAIVQSSLGSASASPFQPHVPYSPFRGMAPYGPLAA SSLLSQQYAASLGLEKLVSPPASAAASSPSSSPSPQPVSELDLSSEPQQLTAKGNSSLGELHAVLQTILRARGKAADRMTVAVADHLPSP CSRRRSGNRRTRNRSRGQNRPTNVKENTIKFEGDFDFESANAQFNREELDKEFKKKLNFKDDKAEKGEEKDLAVVTQSAEAPAEEDLLGP -------------------------------------------------------------- >26954_26954_8_EPHA2-LSM14B_EPHA2_chr1_16474873_ENST00000358432_LSM14B_chr20_60699673_ENST00000279068_length(amino acids)=617AA_BP=274 MELQAARACFALLWGCALAAAAAAQGKEVVLLDFAAAGGELGWLTHPYGKGWDLMQNIMNDMPIYMYSVCNVMSGDQDNWLRTNWVYRGE AERIFIELKFTVRDCNSFPGGASSCKETFNLYYAESDLDYGTNFQKRLFTKIDTIAPDEITVSSDFEARHVKLNVEERSVGPLTRKGFYL AFQDIGACVALLSVRVYYKKCPELLQGLAHFPETIAGSDAPSLATVAGTCVDHAVVPPGGEEPRMHCAVDGEWLVPIGQCLCQAGYEKVE DACQVRSFGTEDRPTDRPAPPREEIYEYIIFRGSDIKDITVCEPPKAQHTLPQDPAIVQSSLGSASASPFQPHVPYSPFRGMAPYGPLAA SSLLSQQYAASLGLGAGFPSIPVGKSPMVEQAVQTGSADNLNAKKLLPGKGTTGTQLNGRQAQPSSKTASDVVQPAAVQAQGQVNDENRR PQRRRSGNRRTRNRSRGQNRPTNVKENTIKFEGDFDFESANAQFNREELDKEFKKKLNFKDDKAEKGEEKDLAVVTQSAEAPAEEDLLGP -------------------------------------------------------------- >26954_26954_9_EPHA2-LSM14B_EPHA2_chr1_16474873_ENST00000358432_LSM14B_chr20_60699673_ENST00000370915_length(amino acids)=453AA_BP=274 MELQAARACFALLWGCALAAAAAAQGKEVVLLDFAAAGGELGWLTHPYGKGWDLMQNIMNDMPIYMYSVCNVMSGDQDNWLRTNWVYRGE AERIFIELKFTVRDCNSFPGGASSCKETFNLYYAESDLDYGTNFQKRLFTKIDTIAPDEITVSSDFEARHVKLNVEERSVGPLTRKGFYL AFQDIGACVALLSVRVYYKKCPELLQGLAHFPETIAGSDAPSLATVAGTCVDHAVVPPGGEEPRMHCAVDGEWLVPIGQCLCQAGYEKVE DACQVRSFGTEDRPTDRPAPPREEIYEYIIFRGSDIKDITVCEPPKAQHTLPQDPAIVQSSLGSASASPFQPHVPYSPFRGMAPYGPLAA SSLLSQQYAASLGLEKLVSPPASAAASSPSSSPSPQPVSELDLSSEPQQLTAKGNSSLGELHAVLQTILRARGKAADRMTVAVADHLPSP -------------------------------------------------------------- >26954_26954_10_EPHA2-LSM14B_EPHA2_chr1_16474873_ENST00000461614_LSM14B_chr20_60699673_ENST00000253001_length(amino acids)=630AA_BP=287 MRAGRRAGADTEAGVQACGCAGAGLGGIGPRARSAAWSSRQPAPASPCCGAVRWPRPRRRRARKWDLMQNIMNDMPIYMYSVCNVMSGDQ DNWLRTNWVYRGEAERIFIELKFTVRDCNSFPGGASSCKETFNLYYAESDLDYGTNFQKRLFTKIDTIAPDEITVSSDFEARHVKLNVEE RSVGPLTRKGFYLAFQDIGACVALLSVRVYYKKCPELLQGLAHFPETIAGSDAPSLATVAGTCVDHAVVPPGGEEPRMHCAVDGEWLVPI GQCLCQAGYEKVEDACQVRSFGTEDRPTDRPAPPREEIYEYIIFRGSDIKDITVCEPPKAQHTLPQDPAIVQSSLGSASASPFQPHVPYS PFRGMAPYGPLAASSLLSQQYAASLGLEKLVSPPASAAASSPSSSPSPQPVSELDLSSEPQQLTAKGNSSLGELHAVLQTILRARGKAAD RMTVAVADHLPSPCSRRRSGNRRTRNRSRGQNRPTNVKENTIKFEGDFDFESANAQFNREELDKEFKKKLNFKDDKAEKGEEKDLAVVTQ SAEAPAEEDLLGPNCYYDKSKSFFDNISSELKTSSRRTTWAEERKLNTETFGVSGRFLRGRSSRGGFRGGRGNGTTRRNPTSHRAGTGRV -------------------------------------------------------------- >26954_26954_11_EPHA2-LSM14B_EPHA2_chr1_16474873_ENST00000461614_LSM14B_chr20_60699673_ENST00000279068_length(amino acids)=630AA_BP=287 MRAGRRAGADTEAGVQACGCAGAGLGGIGPRARSAAWSSRQPAPASPCCGAVRWPRPRRRRARKWDLMQNIMNDMPIYMYSVCNVMSGDQ DNWLRTNWVYRGEAERIFIELKFTVRDCNSFPGGASSCKETFNLYYAESDLDYGTNFQKRLFTKIDTIAPDEITVSSDFEARHVKLNVEE RSVGPLTRKGFYLAFQDIGACVALLSVRVYYKKCPELLQGLAHFPETIAGSDAPSLATVAGTCVDHAVVPPGGEEPRMHCAVDGEWLVPI GQCLCQAGYEKVEDACQVRSFGTEDRPTDRPAPPREEIYEYIIFRGSDIKDITVCEPPKAQHTLPQDPAIVQSSLGSASASPFQPHVPYS PFRGMAPYGPLAASSLLSQQYAASLGLGAGFPSIPVGKSPMVEQAVQTGSADNLNAKKLLPGKGTTGTQLNGRQAQPSSKTASDVVQPAA VQAQGQVNDENRRPQRRRSGNRRTRNRSRGQNRPTNVKENTIKFEGDFDFESANAQFNREELDKEFKKKLNFKDDKAEKGEEKDLAVVTQ SAEAPAEEDLLGPNCYYDKSKSFFDNISSELKTSSRRTTWAEERKLNTETFGVSGRFLRGRSSRGGFRGGRGNGTTRRNPTSHRAGTGRV -------------------------------------------------------------- >26954_26954_12_EPHA2-LSM14B_EPHA2_chr1_16474873_ENST00000461614_LSM14B_chr20_60699673_ENST00000370915_length(amino acids)=466AA_BP=287 MRAGRRAGADTEAGVQACGCAGAGLGGIGPRARSAAWSSRQPAPASPCCGAVRWPRPRRRRARKWDLMQNIMNDMPIYMYSVCNVMSGDQ DNWLRTNWVYRGEAERIFIELKFTVRDCNSFPGGASSCKETFNLYYAESDLDYGTNFQKRLFTKIDTIAPDEITVSSDFEARHVKLNVEE RSVGPLTRKGFYLAFQDIGACVALLSVRVYYKKCPELLQGLAHFPETIAGSDAPSLATVAGTCVDHAVVPPGGEEPRMHCAVDGEWLVPI GQCLCQAGYEKVEDACQVRSFGTEDRPTDRPAPPREEIYEYIIFRGSDIKDITVCEPPKAQHTLPQDPAIVQSSLGSASASPFQPHVPYS PFRGMAPYGPLAASSLLSQQYAASLGLEKLVSPPASAAASSPSSSPSPQPVSELDLSSEPQQLTAKGNSSLGELHAVLQTILRARGKAAD -------------------------------------------------------------- |
Top |
Fusion Protein Functional Features |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr1:16474873/chr20:60699673) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
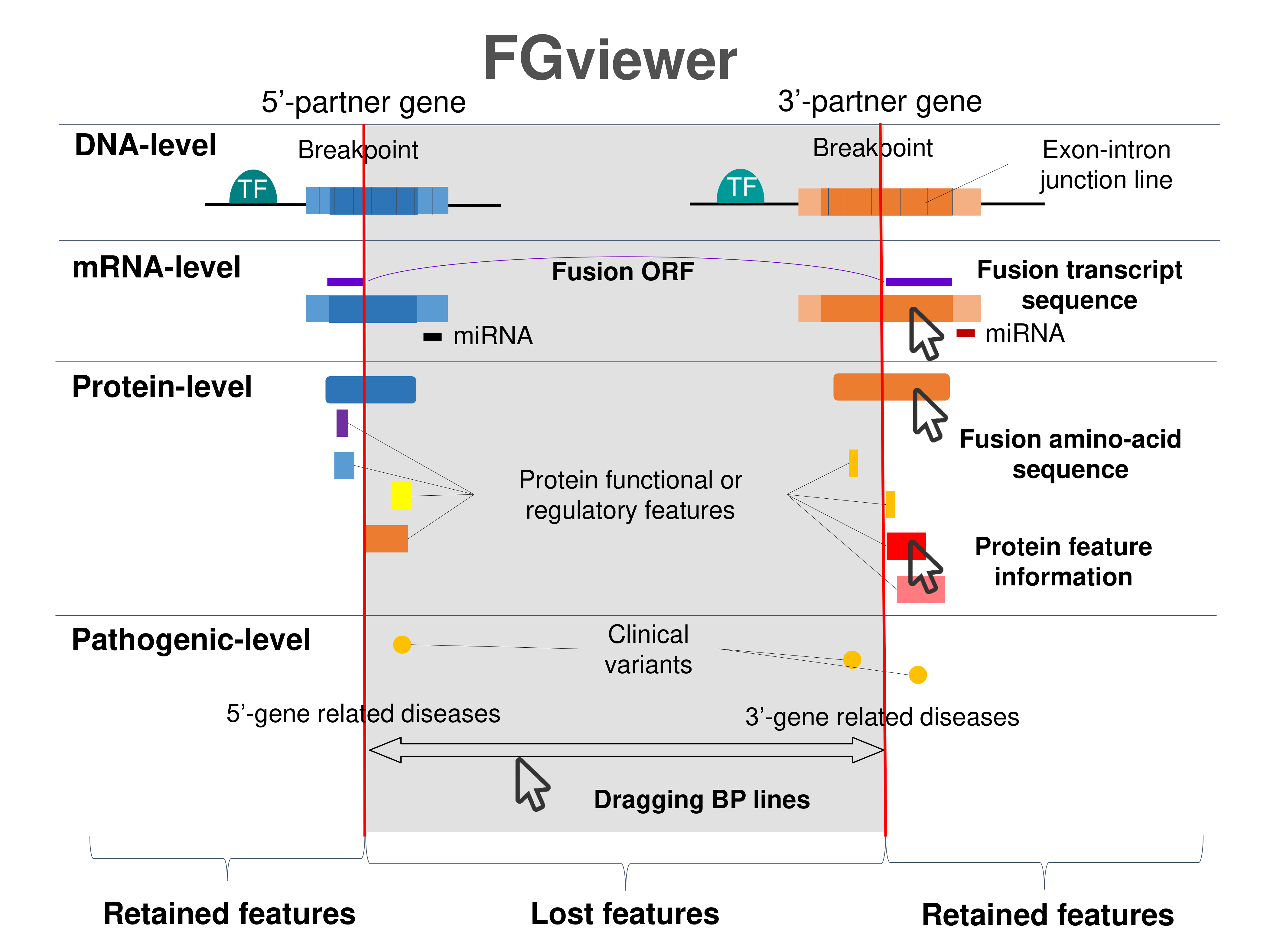 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| EPHA2 | LSM14B |
| FUNCTION: Receptor tyrosine kinase which binds promiscuously membrane-bound ephrin-A family ligands residing on adjacent cells, leading to contact-dependent bidirectional signaling into neighboring cells. The signaling pathway downstream of the receptor is referred to as forward signaling while the signaling pathway downstream of the ephrin ligand is referred to as reverse signaling. Activated by the ligand ephrin-A1/EFNA1 regulates migration, integrin-mediated adhesion, proliferation and differentiation of cells. Regulates cell adhesion and differentiation through DSG1/desmoglein-1 and inhibition of the ERK1/ERK2 (MAPK3/MAPK1, respectively) signaling pathway. May also participate in UV radiation-induced apoptosis and have a ligand-independent stimulatory effect on chemotactic cell migration. During development, may function in distinctive aspects of pattern formation and subsequently in development of several fetal tissues. Involved for instance in angiogenesis, in early hindbrain development and epithelial proliferation and branching morphogenesis during mammary gland development. Engaged by the ligand ephrin-A5/EFNA5 may regulate lens fiber cells shape and interactions and be important for lens transparency development and maintenance. With ephrin-A2/EFNA2 may play a role in bone remodeling through regulation of osteoclastogenesis and osteoblastogenesis. {ECO:0000269|PubMed:10655584, ECO:0000269|PubMed:16236711, ECO:0000269|PubMed:18339848, ECO:0000269|PubMed:19573808, ECO:0000269|PubMed:20679435, ECO:0000269|PubMed:20861311, ECO:0000269|PubMed:23358419, ECO:0000269|PubMed:26158630, ECO:0000269|PubMed:27385333}.; FUNCTION: (Microbial infection) Acts as a receptor for hepatitis C virus (HCV) in hepatocytes and facilitates its cell entry. Mediates HCV entry by promoting the formation of the CD81-CLDN1 receptor complexes that are essential for HCV entry and by enhancing membrane fusion of cells expressing HCV envelope glycoproteins. {ECO:0000269|PubMed:21516087}. | FUNCTION: Required for oocyte meiotic maturation. May be involved in the storage of translationally inactive mRNAs and protect them from degradation (By similarity). Plays a role in control of mRNA translation (By similarity). {ECO:0000250|UniProtKB:Q68FI1, ECO:0000250|UniProtKB:Q8CGC4}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - Retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | EPHA2 | chr1:16474872 | chr20:60699672 | ENST00000358432 | - | 3 | 17 | 28_206 | 274.3333333333333 | 977.0 | Domain | Eph LBD |
| Hgene | EPHA2 | chr1:16474873 | chr20:60699673 | ENST00000358432 | - | 3 | 17 | 28_206 | 274.3333333333333 | 977.0 | Domain | Eph LBD |
| Tgene | LSM14B | chr1:16474872 | chr20:60699672 | ENST00000253001 | 0 | 9 | 241_277 | 42.333333333333336 | 680.3333333333334 | Domain | DFDF | |
| Tgene | LSM14B | chr1:16474872 | chr20:60699672 | ENST00000279068 | 0 | 9 | 241_277 | 42.333333333333336 | 695.6666666666666 | Domain | DFDF | |
| Tgene | LSM14B | chr1:16474872 | chr20:60699672 | ENST00000370915 | 0 | 4 | 241_277 | 42.333333333333336 | 222.0 | Domain | DFDF | |
| Tgene | LSM14B | chr1:16474873 | chr20:60699673 | ENST00000253001 | 0 | 9 | 241_277 | 42.333333333333336 | 680.3333333333334 | Domain | DFDF | |
| Tgene | LSM14B | chr1:16474873 | chr20:60699673 | ENST00000279068 | 0 | 9 | 241_277 | 42.333333333333336 | 695.6666666666666 | Domain | DFDF | |
| Tgene | LSM14B | chr1:16474873 | chr20:60699673 | ENST00000370915 | 0 | 4 | 241_277 | 42.333333333333336 | 222.0 | Domain | DFDF | |
| Tgene | LSM14B | chr1:16474872 | chr20:60699672 | ENST00000253001 | 0 | 9 | 310_326 | 42.333333333333336 | 680.3333333333334 | Motif | Note=FFD box | |
| Tgene | LSM14B | chr1:16474872 | chr20:60699672 | ENST00000253001 | 0 | 9 | 330_350 | 42.333333333333336 | 680.3333333333334 | Motif | Note=TFG box | |
| Tgene | LSM14B | chr1:16474872 | chr20:60699672 | ENST00000279068 | 0 | 9 | 310_326 | 42.333333333333336 | 695.6666666666666 | Motif | Note=FFD box | |
| Tgene | LSM14B | chr1:16474872 | chr20:60699672 | ENST00000279068 | 0 | 9 | 330_350 | 42.333333333333336 | 695.6666666666666 | Motif | Note=TFG box | |
| Tgene | LSM14B | chr1:16474872 | chr20:60699672 | ENST00000370915 | 0 | 4 | 310_326 | 42.333333333333336 | 222.0 | Motif | Note=FFD box | |
| Tgene | LSM14B | chr1:16474872 | chr20:60699672 | ENST00000370915 | 0 | 4 | 330_350 | 42.333333333333336 | 222.0 | Motif | Note=TFG box | |
| Tgene | LSM14B | chr1:16474873 | chr20:60699673 | ENST00000253001 | 0 | 9 | 310_326 | 42.333333333333336 | 680.3333333333334 | Motif | Note=FFD box | |
| Tgene | LSM14B | chr1:16474873 | chr20:60699673 | ENST00000253001 | 0 | 9 | 330_350 | 42.333333333333336 | 680.3333333333334 | Motif | Note=TFG box | |
| Tgene | LSM14B | chr1:16474873 | chr20:60699673 | ENST00000279068 | 0 | 9 | 310_326 | 42.333333333333336 | 695.6666666666666 | Motif | Note=FFD box | |
| Tgene | LSM14B | chr1:16474873 | chr20:60699673 | ENST00000279068 | 0 | 9 | 330_350 | 42.333333333333336 | 695.6666666666666 | Motif | Note=TFG box | |
| Tgene | LSM14B | chr1:16474873 | chr20:60699673 | ENST00000370915 | 0 | 4 | 310_326 | 42.333333333333336 | 222.0 | Motif | Note=FFD box | |
| Tgene | LSM14B | chr1:16474873 | chr20:60699673 | ENST00000370915 | 0 | 4 | 330_350 | 42.333333333333336 | 222.0 | Motif | Note=TFG box |
| - Not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | EPHA2 | chr1:16474872 | chr20:60699672 | ENST00000358432 | - | 3 | 17 | 188_325 | 274.3333333333333 | 977.0 | Compositional bias | Note=Cys-rich |
| Hgene | EPHA2 | chr1:16474873 | chr20:60699673 | ENST00000358432 | - | 3 | 17 | 188_325 | 274.3333333333333 | 977.0 | Compositional bias | Note=Cys-rich |
| Hgene | EPHA2 | chr1:16474872 | chr20:60699672 | ENST00000358432 | - | 3 | 17 | 328_432 | 274.3333333333333 | 977.0 | Domain | Fibronectin type-III 1 |
| Hgene | EPHA2 | chr1:16474872 | chr20:60699672 | ENST00000358432 | - | 3 | 17 | 438_529 | 274.3333333333333 | 977.0 | Domain | Fibronectin type-III 2 |
| Hgene | EPHA2 | chr1:16474872 | chr20:60699672 | ENST00000358432 | - | 3 | 17 | 613_875 | 274.3333333333333 | 977.0 | Domain | Protein kinase |
| Hgene | EPHA2 | chr1:16474872 | chr20:60699672 | ENST00000358432 | - | 3 | 17 | 904_968 | 274.3333333333333 | 977.0 | Domain | SAM |
| Hgene | EPHA2 | chr1:16474873 | chr20:60699673 | ENST00000358432 | - | 3 | 17 | 328_432 | 274.3333333333333 | 977.0 | Domain | Fibronectin type-III 1 |
| Hgene | EPHA2 | chr1:16474873 | chr20:60699673 | ENST00000358432 | - | 3 | 17 | 438_529 | 274.3333333333333 | 977.0 | Domain | Fibronectin type-III 2 |
| Hgene | EPHA2 | chr1:16474873 | chr20:60699673 | ENST00000358432 | - | 3 | 17 | 613_875 | 274.3333333333333 | 977.0 | Domain | Protein kinase |
| Hgene | EPHA2 | chr1:16474873 | chr20:60699673 | ENST00000358432 | - | 3 | 17 | 904_968 | 274.3333333333333 | 977.0 | Domain | SAM |
| Hgene | EPHA2 | chr1:16474872 | chr20:60699672 | ENST00000358432 | - | 3 | 17 | 974_976 | 274.3333333333333 | 977.0 | Motif | PDZ-binding |
| Hgene | EPHA2 | chr1:16474873 | chr20:60699673 | ENST00000358432 | - | 3 | 17 | 974_976 | 274.3333333333333 | 977.0 | Motif | PDZ-binding |
| Hgene | EPHA2 | chr1:16474872 | chr20:60699672 | ENST00000358432 | - | 3 | 17 | 619_627 | 274.3333333333333 | 977.0 | Nucleotide binding | ATP |
| Hgene | EPHA2 | chr1:16474873 | chr20:60699673 | ENST00000358432 | - | 3 | 17 | 619_627 | 274.3333333333333 | 977.0 | Nucleotide binding | ATP |
| Hgene | EPHA2 | chr1:16474872 | chr20:60699672 | ENST00000358432 | - | 3 | 17 | 24_537 | 274.3333333333333 | 977.0 | Topological domain | Extracellular |
| Hgene | EPHA2 | chr1:16474872 | chr20:60699672 | ENST00000358432 | - | 3 | 17 | 559_976 | 274.3333333333333 | 977.0 | Topological domain | Cytoplasmic |
| Hgene | EPHA2 | chr1:16474873 | chr20:60699673 | ENST00000358432 | - | 3 | 17 | 24_537 | 274.3333333333333 | 977.0 | Topological domain | Extracellular |
| Hgene | EPHA2 | chr1:16474873 | chr20:60699673 | ENST00000358432 | - | 3 | 17 | 559_976 | 274.3333333333333 | 977.0 | Topological domain | Cytoplasmic |
| Hgene | EPHA2 | chr1:16474872 | chr20:60699672 | ENST00000358432 | - | 3 | 17 | 538_558 | 274.3333333333333 | 977.0 | Transmembrane | Helical |
| Hgene | EPHA2 | chr1:16474873 | chr20:60699673 | ENST00000358432 | - | 3 | 17 | 538_558 | 274.3333333333333 | 977.0 | Transmembrane | Helical |
Top |
Fusion Protein Structures |
 PDB and CIF files of the predicted fusion proteins PDB and CIF files of the predicted fusion proteins * Here we show the 3D structure of the fusion proteins using Mol*. AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. Model confidence is shown from the pLDDT values per residue. pLDDT corresponds to the model’s prediction of its score on the local Distance Difference Test. It is a measure of local accuracy (from AlphfaFold website). To color code individual residues, we transformed individual PDB files into CIF format. |
| Fusion protein PDB link (fusion AA seq ID in FusionPDB) | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | AA seq | Len(AA seq) |
| PDB file >>>1326_EPHA2_16474873_LSM14B_60699673_1326_EPHA2_16474873_LSM14B_60699673_ranked_0.pdb | EPHA2 | 16474872 | 16474873 | ENST00000279068 | LSM14B | chr20 | 60699673 | + | MRAGRRAGADTEAGVQACGCAGAGLGGIGPRARSAAWSSRQPAPASPCCGAVRWPRPRRRRARKWDLMQNIMNDMPIYMYSVCNVMSGDQ DNWLRTNWVYRGEAERIFIELKFTVRDCNSFPGGASSCKETFNLYYAESDLDYGTNFQKRLFTKIDTIAPDEITVSSDFEARHVKLNVEE RSVGPLTRKGFYLAFQDIGACVALLSVRVYYKKCPELLQGLAHFPETIAGSDAPSLATVAGTCVDHAVVPPGGEEPRMHCAVDGEWLVPI GQCLCQAGYEKVEDACQVRSFGTEDRPTDRPAPPREEIYEYIIFRGSDIKDITVCEPPKAQHTLPQDPAIVQSSLGSASASPFQPHVPYS PFRGMAPYGPLAASSLLSQQYAASLGLEKLVSPPASAAASSPSSSPSPQPVSELDLSSEPQQLTAKGNSSLGELHAVLQTILRARGKAAD RMTVAVADHLPSPCSRRRSGNRRTRNRSRGQNRPTNVKENTIKFEGDFDFESANAQFNREELDKEFKKKLNFKDDKAEKGEEKDLAVVTQ SAEAPAEEDLLGPNCYYDKSKSFFDNISSELKTSSRRTTWAEERKLNTETFGVSGRFLRGRSSRGGFRGGRGNGTTRRNPTSHRAGTGRV | 630 |
Top |
pLDDT score distribution |
 pLDDT score distribution of the predicted wild-type structures of two partner proteins from AlphaFold2 pLDDT score distribution of the predicted wild-type structures of two partner proteins from AlphaFold2* AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. |
EPHA2_pLDDT.png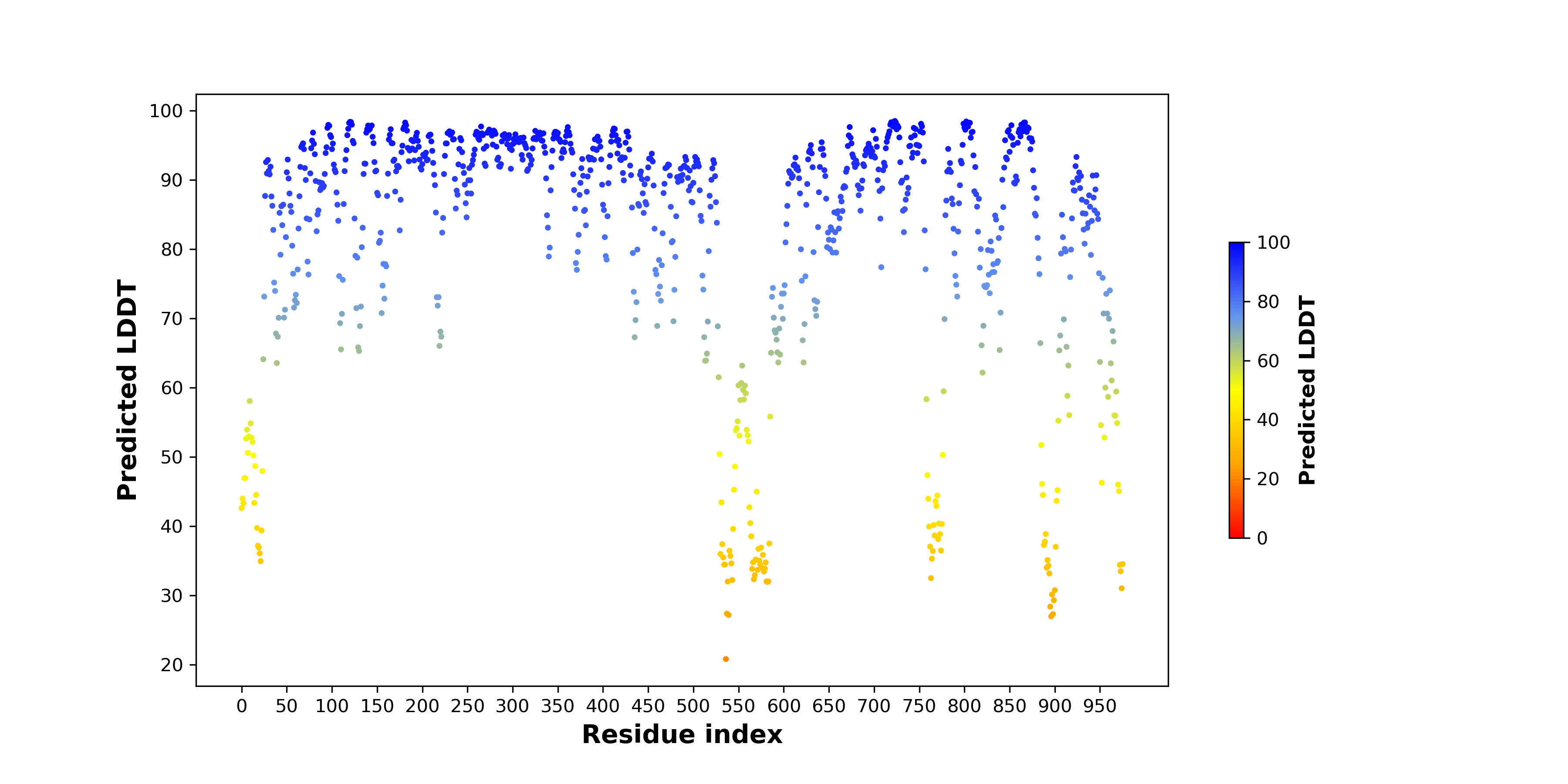 |
LSM14B_pLDDT.png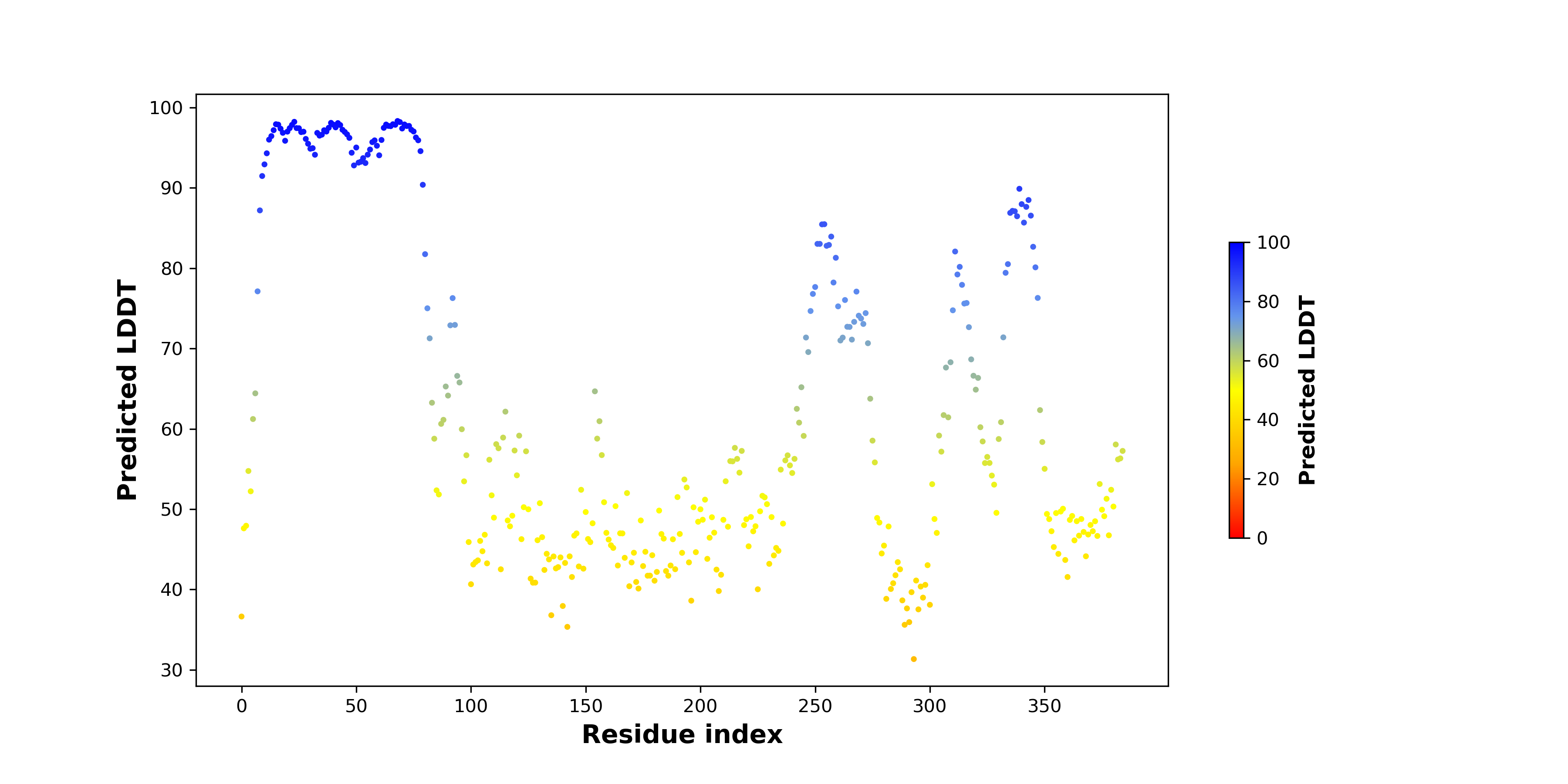 |
 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2* AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. |
 |
Top |
Ramachandran Plot of Fusion Protein Structure |
 Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. |
| Fusion AA seq ID in FusionPDB and their Ramachandran plots |
Top |
Fusion Protein-Protein Interaction |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type from validated records (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type from validated records (BIOGRID-3.4.160) |
| Gene | PPI interactors |
 Protein-protein interactors based on sequence similarity (STRING) Protein-protein interactors based on sequence similarity (STRING) |
| Gene | STRING network |
| EPHA2 | |
| LSM14B |
 - Retained interactions in fusion protein (protein functional feature from UniProt). - Retained interactions in fusion protein (protein functional feature from UniProt). |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost interactions due to fusion (protein functional feature from UniProt). - Lost interactions due to fusion (protein functional feature from UniProt). |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs to EPHA2-LSM14B |
 Drugs used for this fusion-positive patient. Drugs used for this fusion-positive patient. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Drug | Source | PMID |
Top |
Related Diseases to EPHA2-LSM14B |
 Diseases that have this fusion gene. Diseases that have this fusion gene. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Disease | Source | PMID |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |

