| UTHEALTH HOME ABOUT SBMI A-Z WEBMAIL INSIDE THE UNIVERSITY |

|
|||||||
|
Fusion Protein:TMPRSS2-ETV5 |
Fusion Protein Summary |
 Fusion gene summary Fusion gene summary |
| Fusion partner gene information | Fusion gene name: TMPRSS2-ETV5 | FusionPDB ID: 92380 | FusionGDB2.0 ID: 92380 | Hgene | Tgene | Gene symbol | TMPRSS2 | ETV5 | Gene ID | 7113 | 2119 |
| Gene name | transmembrane serine protease 2 | ETS variant transcription factor 5 | |
| Synonyms | PP9284|PRSS10 | ERM | |
| Cytomap | 21q22.3 | 3q27.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | transmembrane protease serine 2epitheliasinserine protease 10transmembrane protease, serine 2 | ETS translocation variant 5ETS variant 5ets-related moleculeets-related protein ERM | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | O15393 | P41161 | |
| Ensembl transtripts involved in fusion gene | ENST ids | ENST00000332149, ENST00000398585, ENST00000458356, ENST00000497881, | ENST00000306376, ENST00000434744, ENST00000537818, ENST00000480706, |
| Fusion gene scores for assessment (based on all fusion genes of FusionGDB 2.0) | * DoF score | 40 X 73 X 10=29200 | 17 X 12 X 11=2244 |
| # samples | 130 | 22 | |
| ** MAII score | log2(130/29200*10)=-4.48938484073892 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(22/2244*10)=-3.35049724708413 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context (manual curation of fusion genes in FusionPDB) | PubMed: TMPRSS2 [Title/Abstract] AND ETV5 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint (based on all fusion genes of FusionGDB 2.0) | TMPRSS2(42866276)-ETV5(185823731), # samples:12 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | TMPRSS2 | GO:0006508 | proteolysis | 21068237|24227843 |
| Hgene | TMPRSS2 | GO:0046598 | positive regulation of viral entry into host cell | 21068237|24227843 |
| Tgene | ETV5 | GO:0034599 | cellular response to oxidative stress | 19443906 |
 Fusion gene breakpoints across TMPRSS2 (5'-gene) Fusion gene breakpoints across TMPRSS2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 Fusion gene breakpoints across ETV5 (3'-gene) Fusion gene breakpoints across ETV5 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
Top |
Fusion Gene Sample Information |
 Fusion gene information from FusionGDB2.0. Fusion gene information from FusionGDB2.0. |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | PRAD | TCGA-KK-A7AZ-01A | TMPRSS2 | chr21 | 42861433 | - | ETV5 | chr3 | 185823730 | - |
| ChimerDB4 | PRAD | TCGA-KK-A7AZ-01A | TMPRSS2 | chr21 | 42861434 | - | ETV5 | chr3 | 185823731 | - |
| ChimerDB4 | PRAD | TCGA-KK-A7AZ-01A | TMPRSS2 | chr21 | 42880007 | - | ETV5 | chr3 | 185823730 | - |
| ChimerDB4 | PRAD | TCGA-KK-A7AZ | TMPRSS2 | chr21 | 42861433 | - | ETV5 | chr3 | 185823731 | - |
| ChimerDB4 | PRAD | TCGA-KK-A7AZ | TMPRSS2 | chr21 | 42861434 | - | ETV5 | chr3 | 185823731 | - |
| ChimerDB4 | PRAD | TCGA-ZG-A9L5-01A | TMPRSS2 | chr21 | 42866282 | - | ETV5 | chr3 | 185823730 | - |
| ChimerDB4 | PRAD | TCGA-ZG-A9L5-01A | TMPRSS2 | chr21 | 42866283 | - | ETV5 | chr3 | 185823731 | - |
| ChimerDB4 | PRAD | TCGA-ZG-A9L5-01A | TMPRSS2 | chr21 | 42880007 | - | ETV5 | chr3 | 185823730 | - |
| ChimerDB4 | PRAD | TCGA-ZG-A9L5 | TMPRSS2 | chr21 | 42866282 | - | ETV5 | chr3 | 185823731 | - |
| ChimerDB4 | PRAD | TCGA-ZG-A9L5 | TMPRSS2 | chr21 | 42866283 | - | ETV5 | chr3 | 185823731 | - |
| ChimerDB4 | PRAD | TCGA-ZG-A9L5 | TMPRSS2 | chr21 | 42870045 | - | ETV5 | chr3 | 185823731 | - |
| ChimerDB4 | PRAD | TCGA-ZG-A9L5 | TMPRSS2 | chr21 | 42880007 | - | ETV5 | chr3 | 185823731 | - |
| ChimerDB4 | prostate cancer | EU314929 | TMPRSS2 | chr21 | 42880083 | ETV5 | chr3 | 185798922 | ||
| ChimerKB3 | . | . | TMPRSS2 | chr21 | 42866282 | - | ETV5 | chr3 | 185823731 | - |
| ChimerKB3 | . | . | TMPRSS2 | chr21 | 42879876 | - | ETV5 | chr3 | 185826901 | - |
| ChiTaRS5.0 | N/A | EU314929 | TMPRSS2 | chr21 | 42880007 | - | ETV5 | chr3 | 185823734 | - |
| ChiTaRS5.0 | N/A | EU314930 | TMPRSS2 | chr21 | 42866276 | - | ETV5 | chr3 | 185823731 | - |
| ChiTaRS5.0 | N/A | EU314931 | TMPRSS2 | chr21 | 42866276 | - | ETV5 | chr3 | 185823731 | - |
| ChiTaRS5.0 | N/A | FW581626 | TMPRSS2 | chr21 | 42880007 | - | ETV5 | chr3 | 185823734 | - |
| ChiTaRS5.0 | N/A | FW581627 | TMPRSS2 | chr21 | 42866276 | - | ETV5 | chr3 | 185823731 | - |
| ChiTaRS5.0 | N/A | FW581628 | TMPRSS2 | chr21 | 42866276 | - | ETV5 | chr3 | 185823731 | - |
| ChiTaRS5.0 | N/A | HZ416120 | TMPRSS2 | chr21 | 42880007 | - | ETV5 | chr3 | 185823734 | - |
| ChiTaRS5.0 | N/A | HZ416121 | TMPRSS2 | chr21 | 42866276 | - | ETV5 | chr3 | 185823731 | - |
| ChiTaRS5.0 | N/A | HZ416122 | TMPRSS2 | chr21 | 42866276 | - | ETV5 | chr3 | 185823731 | - |
| ChiTaRS5.0 | N/A | JB772367 | TMPRSS2 | chr21 | 42880007 | - | ETV5 | chr3 | 185823734 | - |
| ChiTaRS5.0 | N/A | JB772368 | TMPRSS2 | chr21 | 42866276 | - | ETV5 | chr3 | 185823731 | - |
| ChiTaRS5.0 | N/A | JB772369 | TMPRSS2 | chr21 | 42866276 | - | ETV5 | chr3 | 185823731 | - |
| ChiTaRS5.0 | N/A | LV708924 | TMPRSS2 | chr21 | 42880007 | - | ETV5 | chr3 | 185823734 | - |
| ChiTaRS5.0 | N/A | LV708925 | TMPRSS2 | chr21 | 42866276 | - | ETV5 | chr3 | 185823731 | - |
| ChiTaRS5.0 | N/A | LV708926 | TMPRSS2 | chr21 | 42866276 | - | ETV5 | chr3 | 185823731 | - |
| ChiTaRS5.0 | N/A | MA648838 | TMPRSS2 | chr21 | 42880007 | - | ETV5 | chr3 | 185823734 | - |
| ChiTaRS5.0 | N/A | MA648839 | TMPRSS2 | chr21 | 42866276 | - | ETV5 | chr3 | 185823731 | - |
| ChiTaRS5.0 | N/A | MA648840 | TMPRSS2 | chr21 | 42866276 | - | ETV5 | chr3 | 185823731 | - |
Top |
Fusion ORF Analysis |
 Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000398585 | TMPRSS2 | chr21 | 42879876 | - | ENST00000306376 | ETV5 | chr3 | 185826901 | - | 4227 | 116 | 243 | 1895 | 550 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
Top |
Fusion Amino Acid Sequences |
 For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >FusionGDB ID_FusionGDB isoform ID_FGname_Hgene_Hchr_Hbp_Henst_Tgene_Tchr_Tbp_Tenst_length(fusion AA) seq_BP >92380_92380_1_TMPRSS2-ETV5_TMPRSS2_chr21_42879876_ENST00000398585_ETV5_chr3_185826901_ENST00000306376_length(amino acids)=550AA_BP= MRRRGRCKGRREAASGGSLFPQKLLNAETSQSGIRDAESTMDGFYDQQVPFMVPGKSRSEECRGRPVIDRKRKFLDTDLAHDSEELFQDL SQLQEAWLAEAQVPDDEQFVPDFQSDNLVLHAPPPTKIKRELHSPSSELSSCSHEQALGANYGEKCLYNYCAYDRKPPSGFKPLTPPTTP LSPTHQNPLFPPPQATLPTSGHAPAAGPVQGVGPAPAPHSLPEPGPQQQTFAVPRPPHQPLQMPKMMPENQYPSEQRFQRQLSEPCHPFP PQPGVPGDNRPSYHRQMSEPIVPAAPPPPQGFKQEYHDPLYEHGVPGMPGPPAHGFQSPMGIKQEPRDYCVDSEVPNCQSSYMRGGYFSS SHEGFSYEKDPRLYFDDTCVVPERLEGKVKQEPTMYREGPPYQRRGSLQLWQFLVTLLDDPANAHFIAWTGRGMEFKLIEPEEVARRWGI QKNRPAMNYDKLSRSLRYYYEKGIMQKVAGERYVYKFVCDPDALFSMAFPDNQRPFLKAESECHLSEEDTLPLTHFEDSPAYLLDMDRCS -------------------------------------------------------------- |
Top |
Fusion Protein Functional Features |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr21:42866276/chr3:185823731) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
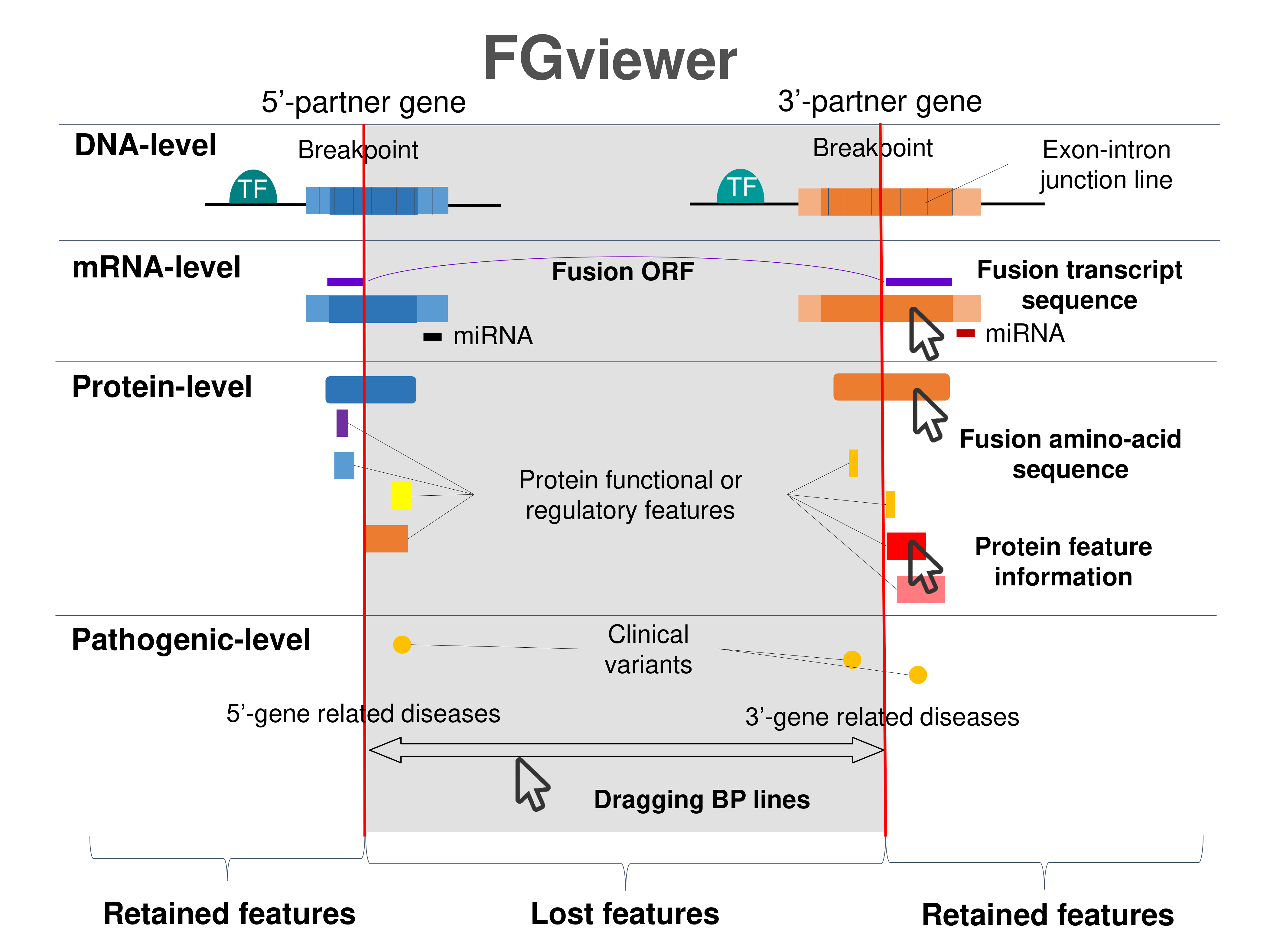 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| TMPRSS2 | ETV5 |
| FUNCTION: Plasma membrane-anchored serine protease that participates in proteolytic cascades of relevance for the normal physiologic function of the prostate (PubMed:25122198). Androgen-induced TMPRSS2 activates several substrates that include pro-hepatocyte growth factor/HGF, the protease activated receptor-2/F2RL1 or matriptase/ST14 leading to extracellular matrix disruption and metastasis of prostate cancer cells (PubMed:15537383, PubMed:26018085, PubMed:25122198). In addition, activates trigeminal neurons and contribute to both spontaneous pain and mechanical allodynia (By similarity). {ECO:0000250|UniProtKB:Q9JIQ8, ECO:0000269|PubMed:15537383, ECO:0000269|PubMed:25122198, ECO:0000269|PubMed:26018085}.; FUNCTION: (Microbial infection) Facilitates human coronaviruses SARS-CoV and SARS-CoV-2 infections via two independent mechanisms, proteolytic cleavage of ACE2 receptor which promotes viral uptake, and cleavage of coronavirus spike glycoproteins which activates the glycoprotein for host cell entry (PubMed:24227843, PubMed:32142651, PubMed:32404436, PubMed:34159616, PubMed:33051876). Upon SARS-CoV-2 infection, increases syncytia formation by accelerating the fusion process (PubMed:34159616, PubMed:33051876). Proteolytically cleaves and activates the spike glycoproteins of human coronavirus 229E (HCoV-229E) and human coronavirus EMC (HCoV-EMC) and the fusion glycoproteins F0 of Sendai virus (SeV), human metapneumovirus (HMPV), human parainfluenza 1, 2, 3, 4a and 4b viruses (HPIV). Essential for spread and pathogenesis of influenza A virus (strains H1N1, H3N2 and H7N9); involved in proteolytic cleavage and activation of hemagglutinin (HA) protein which is essential for viral infectivity. {ECO:0000269|PubMed:21068237, ECO:0000269|PubMed:21325420, ECO:0000269|PubMed:23536651, ECO:0000269|PubMed:23966399, ECO:0000269|PubMed:24027332, ECO:0000269|PubMed:24227843, ECO:0000269|PubMed:32142651, ECO:0000269|PubMed:32404436, ECO:0000269|PubMed:33051876, ECO:0000269|PubMed:34159616}. | FUNCTION: Binds to DNA sequences containing the consensus nucleotide core sequence 5'-GGAA.-3'. {ECO:0000269|PubMed:8152800}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - Retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| - Not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
Top |
Fusion Protein Structures |
 PDB and CIF files of the predicted fusion proteins PDB and CIF files of the predicted fusion proteins * Here we show the 3D structure of the fusion proteins using Mol*. AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. Model confidence is shown from the pLDDT values per residue. pLDDT corresponds to the model’s prediction of its score on the local Distance Difference Test. It is a measure of local accuracy (from AlphfaFold website). To color code individual residues, we transformed individual PDB files into CIF format. |
| Fusion protein PDB link (fusion AA seq ID in FusionPDB) | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | AA seq | Len(AA seq) |
| PDB file (390) >>>390.pdbFusion protein BP residue: CIF file (390) >>>390.cif | TMPRSS2 | chr21 | 42879876 | - | ETV5 | chr3 | 185826901 | - | MRRRGRCKGRREAASGGSLFPQKLLNAETSQSGIRDAESTMDGFYDQQVP FMVPGKSRSEECRGRPVIDRKRKFLDTDLAHDSEELFQDLSQLQEAWLAE AQVPDDEQFVPDFQSDNLVLHAPPPTKIKRELHSPSSELSSCSHEQALGA NYGEKCLYNYCAYDRKPPSGFKPLTPPTTPLSPTHQNPLFPPPQATLPTS GHAPAAGPVQGVGPAPAPHSLPEPGPQQQTFAVPRPPHQPLQMPKMMPEN QYPSEQRFQRQLSEPCHPFPPQPGVPGDNRPSYHRQMSEPIVPAAPPPPQ GFKQEYHDPLYEHGVPGMPGPPAHGFQSPMGIKQEPRDYCVDSEVPNCQS SYMRGGYFSSSHEGFSYEKDPRLYFDDTCVVPERLEGKVKQEPTMYREGP PYQRRGSLQLWQFLVTLLDDPANAHFIAWTGRGMEFKLIEPEEVARRWGI QKNRPAMNYDKLSRSLRYYYEKGIMQKVAGERYVYKFVCDPDALFSMAFP DNQRPFLKAESECHLSEEDTLPLTHFEDSPAYLLDMDRCSSLPYAEGFAY | 550 |
| 3D view using mol* of 390 (AA BP:) | ||||||||||
 | ||||||||||
Top |
pLDDT score distribution |
 pLDDT score distribution of the predicted wild-type structures of two partner proteins from AlphaFold2 pLDDT score distribution of the predicted wild-type structures of two partner proteins from AlphaFold2* AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. |
TMPRSS2_pLDDT.png |
ETV5_pLDDT.png |
 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2* AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. |
TMPRSS2_ETV5_390_pLDDT.png (AA BP:) |
TMPRSS2_ETV5_390_pLDDT_and_active_sites.png (AA BP:) |
TMPRSS2_ETV5_390_violinplot.png (AA BP:)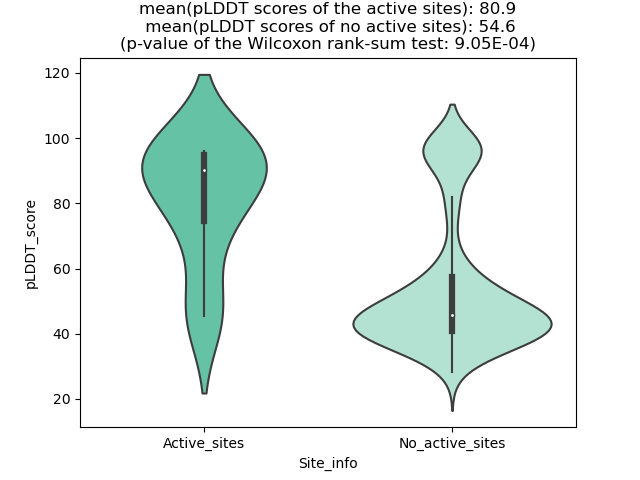 |
Top |
Ramachandran Plot of Fusion Protein Structure |
 Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. |
| Fusion AA seq ID in FusionPDB and their Ramachandran plots |
| TMPRSS2_ETV5_390.png |
 |
Top |
Potential Active Site Information |
 The potential binding sites of these fusion proteins were identified using SiteMap, a module of the Schrodinger suite. The potential binding sites of these fusion proteins were identified using SiteMap, a module of the Schrodinger suite. |
| Fusion AA seq ID in FusionPDB | Site score | Size | D score | Volume | Exposure | Enclosure | Contact | Phobic | Philic | Balance | Don/Acc | Residues |
| 390 | 0.747 | 37 | 0.745 | 131.369 | 0.713 | 0.631 | 0.851 | 0.74 | 0.632 | 1.171 | 1.068 | Chain A: 402,403,407,408,409,411,412,415,416,497,5 00 |
Top |
Potentially Interacting Small Molecules through Virtual Screening |
 The FDA-approved small molecule library molecules were subjected to virtual screening using the Glide. The FDA-approved small molecule library molecules were subjected to virtual screening using the Glide. |
| Fusion AA seq ID in FusionPDB | ZINC ID | DrugBank ID | Drug name | Docking score | Glide gscore |
Top |
 Drug information from DrugBank of the top 20 interacting small molecules. Drug information from DrugBank of the top 20 interacting small molecules. |
| ZINC ID | DrugBank ID | Drug name | Drug type | SMILES | Drug group |
Top |
Biochemical Features of Small Molecules |
 ADME (Absorption, Distribution, Metabolism, and Excretion) of drugs using QikProp(v3.9) ADME (Absorption, Distribution, Metabolism, and Excretion) of drugs using QikProp(v3.9) |
| ZINC ID | mol_MW | dipole | SASA | FOSA | FISA | PISA | WPSA | volume | donorHB | accptHB | IP | Human Oral Absorption | Percent Human Oral Absorption | Rule Of Five | Rule Of Three |
Top |
Drug Toxicity Information |
 Toxicity information of individual drugs using eToxPred Toxicity information of individual drugs using eToxPred |
| ZINC ID | Smile | Surface Accessibility | Toxicity |
Top |
Fusion Protein-Protein Interaction |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type from validated records (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type from validated records (BIOGRID-3.4.160) |
| Gene | PPI interactors |
| TMPRSS2 | CLK1, HNRNPL, DCAF4, S, ACE2, STX7, SCD, PLP2, PLLP, MOSPD3, UPK2, SELK, C3orf52, IGFBP5, SEC22A, PLP1, CLN6, TMEM128, C14orf1, CFHR5, TMEM243, TMEM60, CXCL9, BNIP2, CNIH3, PGAP2, TMEM218, TMEM86A, CMTM7, ZFPL1, BMP10, TMEM79, SLC35A1, SFTPC, VAMP5, FAXDC2, STX8, NINJ2, CLEC7A, C2CD2L, CNIH2, BNIP3, DEFB103A, DEFB103B, TMEM11, C17orf62, TMEM86B, ADIPOQ, EDDM3B, BRICD5, PTTG1IP, PTCH1, ANKRD46, TMEM222, C1QL4, TMEM120B, FAM3C, TMEM229B, PLN, CTXN3, TNF, SMIM1, AQP1, TMPRSS2, DNASE2, MYO1C, PGRMC1, NDUFA4, ADAM10, CLGN, SEC16A, SPINT2, COCH, B4GAT1, XPOT, MYO1B, LANCL1, CALU, PLXNA2, GPC4, BANF1, CPD, CLPX, STC2, SLIT2, MYO1D, ERLIN2, BAG2, TFRC, HLA-A, TP53, RPN1, RPN2, GNAI2, APP, ITGB1, INSR, GLA, GPI, ITGAV, CTSD, APRT, IGF1R, MET, GNAI3, CLTA, CALM2, HLA-C, DLAT, HSPA5, LAMC1, LAMP1, IGF2R, PDIA4, P4HA1, HSP90B1, PVR, RPA2, ATP2A2, PFKL, CDH2, MSH3, RPA1, CALR, RPA3, LONP1, TGFBR1, DDOST, HADHA, ECE1, DNM2, FXR2, PLXNA3, ATP1B3, HADHB, SEC61A1, GNB1, NOMO3, PRKDC, TFAM, SLC7A5, PFKP, KIF23, FMR1, PTK7, ADAM15, CUL2, SCARB2, WRN, DSC3, ITIH4, GANAB, KARS, RCN1, SAFB, IMMT, HSD17B12, UBR4, QSOX2, RRM2B, MON2, YTHDF3, PABPN1, PLD3, PELP1, TTC13, CNNM3, IPO4, FAR1, GGH, ERLEC1, RBM33, DNAJA3, RBM14, PPP1R9B, SDF2, NEU1, GRWD1, AIF1L, NTPCR, FUCA2, EDEM3, RACGAP1, RAB18, TMEM106B, CHPF2, GNG12, DNAJB11, EGFL7, DDX20, ADAMTS1, LNPEP, ITM2B, MRPL11, HYOU1, AFG3L2, YTHDF2, SUPT16H, PDHX, IPO5, SAP18, HGS, SPTBN2, PLXNB2, ARPC3, ACTN4, TRIM13, PDCD6, MPDU1, SEC22B, ATP6AP2, STAM2, ARL6IP5, UFL1, SEC31A, STBD1, SMC2, SEC24A, SEC24B, ACSL3, CDK1, SRPR, SPTB, MTHFD1, ACTN1, ETFA, RAB6A, VDAC1, HMOX2, MYH10, DEK, ATP5C1, NAMPT, VDAC2, QARS, TUFM, SERPINH1, BCAP31, PSMD7, CLCN7, SLC25A1, YARS, EPPK1, RAB10, ABCE1, RHOA, CNBP, TPM4, SLC25A11, LMNB2, CKAP4, GOLGA3, TWF1, TRAP1, STX5, DDX39B, RAB3GAP1, PON2, PPA1, SEC23A, SEC23B, NDUFA9, ATP6V1F, INF2, ANO6, ACTBL2, MIA3, CDKAL1, KIAA0368, C19orf70, SLC25A24, HUWE1, MEGF8, KTN1, COMTD1, ANKLE2, VRK2, ZDHHC17, DNAH10, NUP93, MMGT1, TMEM199, ATPAF2, EMC1, PDZD8, NBEA, MOSPD2, PDCD6IP, SYNE2, UBXN4, CCDC47, LRRC59, C7orf55, DDRGK1, NGLY1, CDK5RAP3, CCDC115, CLCC1, OSBPL9, RBM15, CGRRF1, ESYT1, NDC1, EFHD1, OSBPL8, KIAA1715, LSG1, RAB1B, TMX1, GORASP2, SFXN1, XPO5, VEZT, JPH1, BIRC6, AAAS, TOMM22, DDX19A, SMPD4, TMOD3, EMC3, VAPA, CEP72, LARS, DPM3, DHCR7, UBA2, AASS, ATP6V1H, SLC25A13, STOML2, RAB21, MYO6, VDAC3, AUP1, SEC23IP, |
 Protein-protein interactors based on sequence similarity (STRING) Protein-protein interactors based on sequence similarity (STRING) |
| Gene | STRING network |
| TMPRSS2 |  |
| ETV5 |  |
 - Retained interactions in fusion protein (protein functional feature from UniProt). - Retained interactions in fusion protein (protein functional feature from UniProt). |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost interactions due to fusion (protein functional feature from UniProt). - Lost interactions due to fusion (protein functional feature from UniProt). |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs to TMPRSS2-ETV5 |
 Drugs used for this fusion-positive patient. Drugs used for this fusion-positive patient. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Drug | Source | PMID |
Top |
Related Diseases to TMPRSS2-ETV5 |
 Diseases that have this fusion gene. Diseases that have this fusion gene. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Disease | Source | PMID |
| TMPRSS2 | ETV5 | Prostate Adenocarcinoma | MyCancerGenome | |
| TMPRSS2 | ETV5 | Lung Adenocarcinoma | MyCancerGenome | |
| TMPRSS2 | ETV5 | Cancer Of Unknown Primary | MyCancerGenome | |
| TMPRSS2 | ETV5 | Adenocarcinoma Of Unknown Primary | MyCancerGenome | |
| TMPRSS2 | ETV5 | Breast Invasive Ductal Carcinoma | MyCancerGenome |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
| Hgene | TMPRSS2 | C0033578 | Prostatic Neoplasms | 4 | CTD_human |
| Hgene | TMPRSS2 | C0376358 | Malignant neoplasm of prostate | 4 | CTD_human |
| Tgene | ETV5 | C0033578 | Prostatic Neoplasms | 1 | CTD_human |
| Tgene | ETV5 | C0376358 | Malignant neoplasm of prostate | 1 | CTD_human |

