| UTHEALTH HOME ABOUT SBMI A-Z WEBMAIL INSIDE THE UNIVERSITY |

|
|||||||
|
Fusion Protein:CLIP1-ROS1 |
Fusion Protein Summary |
 Fusion gene summary Fusion gene summary |
| Fusion partner gene information | Fusion gene name: CLIP1-ROS1 | FusionPDB ID: 17164 | FusionGDB2.0 ID: 17164 | Hgene | Tgene | Gene symbol | CLIP1 | ROS1 | Gene ID | 6249 | 6098 |
| Gene name | CAP-Gly domain containing linker protein 1 | ROS proto-oncogene 1, receptor tyrosine kinase | |
| Synonyms | CLIP|CLIP-170|CLIP170|CYLN1|RSN | MCF3|ROS|c-ros-1 | |
| Cytomap | 12q24.31 | 6q22.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | CAP-Gly domain-containing linker protein 1cytoplasmic linker protein 1cytoplasmic linker protein 170 alpha-2cytoplasmic linker protein CLIP-170restin (Reed-Steinberg cell-expressed intermediate filament-associated protein) | proto-oncogene tyrosine-protein kinase ROSROS proto-oncogene 1 , receptor tyrosine kinasec-ros oncogene 1 , receptor tyrosine kinaseproto-oncogene c-Ros-1transmembrane tyrosine-specific protein kinasev-ros avian UR2 sarcoma virus oncogene homolog 1 | |
| Modification date | 20200313 | 20200314 | |
| UniProtAcc | P30622 | P08922 | |
| Ensembl transtripts involved in fusion gene | ENST ids | ENST00000302528, ENST00000540338, ENST00000545889, ENST00000358808, ENST00000361654, ENST00000537178, ENST00000536634, ENST00000540539, | ENST00000368507, ENST00000368508, |
| Fusion gene scores for assessment (based on all fusion genes of FusionGDB 2.0) | * DoF score | 22 X 22 X 13=6292 | 18 X 20 X 5=1800 |
| # samples | 31 | 20 | |
| ** MAII score | log2(31/6292*10)=-4.34317855014145 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(20/1800*10)=-3.16992500144231 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context (manual curation of fusion genes in FusionPDB) | PubMed: CLIP1 [Title/Abstract] AND ROS1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint (based on all fusion genes of FusionGDB 2.0) | |||
| Anticipated loss of major functional domain due to fusion event. | CLIP1-ROS1 seems lost the major protein functional domain in Hgene partner, which is a CGC due to the frame-shifted ORF. CLIP1-ROS1 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. CLIP1-ROS1 seems lost the major protein functional domain in Tgene partner, which is a CGC due to the frame-shifted ORF. CLIP1-ROS1 seems lost the major protein functional domain in Tgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. CLIP1-ROS1 seems lost the major protein functional domain in Tgene partner, which is a kinase due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | ROS1 | GO:0001558 | regulation of cell growth | 16885344 |
| Tgene | ROS1 | GO:0006468 | protein phosphorylation | 16885344 |
| Tgene | ROS1 | GO:0032006 | regulation of TOR signaling | 16885344 |
 Fusion gene breakpoints across CLIP1 (5'-gene) Fusion gene breakpoints across CLIP1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 Fusion gene breakpoints across ROS1 (3'-gene) Fusion gene breakpoints across ROS1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
Top |
Fusion Gene Sample Information |
 Fusion gene information from FusionGDB2.0. Fusion gene information from FusionGDB2.0. |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerKB3 | . | . | CLIP1 | chr12 | 122794308 | - | ROS1 | chr6 | 117641193 | - |
Top |
Fusion ORF Analysis |
 Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000361654 | CLIP1 | chr12 | 122794308 | - | ENST00000368508 | ROS1 | chr6 | 117641193 | - | 4861 | 3402 | 174 | 3428 | 1084 |
| ENST00000361654 | CLIP1 | chr12 | 122794308 | - | ENST00000368507 | ROS1 | chr6 | 117641193 | - | 4861 | 3402 | 174 | 3428 | 1084 |
| ENST00000358808 | CLIP1 | chr12 | 122794308 | - | ENST00000368508 | ROS1 | chr6 | 117641193 | - | 5175 | 3716 | 155 | 3742 | 1195 |
| ENST00000358808 | CLIP1 | chr12 | 122794308 | - | ENST00000368507 | ROS1 | chr6 | 117641193 | - | 5175 | 3716 | 155 | 3742 | 1195 |
| ENST00000537178 | CLIP1 | chr12 | 122794308 | - | ENST00000368508 | ROS1 | chr6 | 117641193 | - | 5133 | 3674 | 218 | 3700 | 1160 |
| ENST00000537178 | CLIP1 | chr12 | 122794308 | - | ENST00000368507 | ROS1 | chr6 | 117641193 | - | 5133 | 3674 | 218 | 3700 | 1160 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
Top |
Fusion Amino Acid Sequences |
 For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >FusionGDB ID_FusionGDB isoform ID_FGname_Hgene_Hchr_Hbp_Henst_Tgene_Tchr_Tbp_Tenst_length(fusion AA) seq_BP >17164_17164_1_CLIP1-ROS1_CLIP1_chr12_122794308_ENST00000358808_ROS1_chr6_117641193_ENST00000368507_length(amino acids)=1195AA_BP= MSMLKPSGLKAPTKILKPGSTALKTPTAVVAPVEKTISSEKASSTPSSETQEEFVDDFRVGERVWVNGNKPGFIQFLGETQFAPGQWAGI VLDEPIGKNDGSVAGVRYFQCEPLKGIFTRPSKLTRKVQAEDEANGLQTTPASRATSPLCTSTASMVSSSPSTPSNIPQKPSQPAAKEPS ATPPISNLTKTASESISNLSEAGSIKKGERELKIGDRVLVGGTKAGVVRFLGETDFAKGEWCGVELDEPLGKNDGAVAGTRYFQCQPKYG LFAPVHKVTKIGFPSTTPAKAKANAVRRVMATTSASLKRSPSASSLSSMSSVASSVSSRPSRTGLLTETSSRYARKISGTTALQEALKEK QQHIEQLLAERDLERAEVAKATSHVGEIEQELALARDGHDQHVLELEAKMDQLRTMVEAADREKVELLNQLEEEKRKVEDLQFRVEEESI TKGDLETQTKLEHARIKELEQSLLFEKTKADKLQRELEDTRVATVSEKSRIMELEKDLALRVQEVAELRRRLESNKPAGDVDMSLSLLQE ISSLQEKLEVTRTDHQREITSLKEHFGAREETHQKEIKALYTATEKLSKENESLKSKLEHANKENSDVIALWKSKLETAIASHQQAMEEL KVSFSKGLGTETAEFAELKTQIEKMRLDYQHEIENLQNQQDSERAAHAKEMEALRAKLMKVIKEKENSLEAIRSKLDKAEDQHLVEMEDT LNKLQEAEIKVKELEVLQAKCNEQTKVIDNFTSQLKATEEKLLDLDALRKASSEGKSEMKKLRQQLEAAEKQIKHLEIEKNAESSKASSI TRELQGRELKLTNLQENLSEVSQVKETLEKELQILKEKFAEASEEAVSVQRSMQETVNKLHQKEEQFNMLSSDLEKLRENLADMEAKFRE KDEREEQLIKAKEKLENDIAEIMKMSGDNSSQLTKMNDELRLKERDVEELQLKLTKANENASFLQKSIEDMTVKAEQSQQEAAKKHEEEK KELERKLSDLEKKMETSHNQCQELKARYERATSETKTKHEEILQNLQKTLLDTEDKLKGAREENSGLLQELEELRKQADKAKAAQTAEDA MQIMEQMTKEKTETLASLEDTKQTNAKLQNELDTLKENNLKNVEELNKSKELLTVENQKMEEFRKEIETLKQAAAQKSQQLSALQEENVK -------------------------------------------------------------- >17164_17164_2_CLIP1-ROS1_CLIP1_chr12_122794308_ENST00000358808_ROS1_chr6_117641193_ENST00000368508_length(amino acids)=1195AA_BP= MSMLKPSGLKAPTKILKPGSTALKTPTAVVAPVEKTISSEKASSTPSSETQEEFVDDFRVGERVWVNGNKPGFIQFLGETQFAPGQWAGI VLDEPIGKNDGSVAGVRYFQCEPLKGIFTRPSKLTRKVQAEDEANGLQTTPASRATSPLCTSTASMVSSSPSTPSNIPQKPSQPAAKEPS ATPPISNLTKTASESISNLSEAGSIKKGERELKIGDRVLVGGTKAGVVRFLGETDFAKGEWCGVELDEPLGKNDGAVAGTRYFQCQPKYG LFAPVHKVTKIGFPSTTPAKAKANAVRRVMATTSASLKRSPSASSLSSMSSVASSVSSRPSRTGLLTETSSRYARKISGTTALQEALKEK QQHIEQLLAERDLERAEVAKATSHVGEIEQELALARDGHDQHVLELEAKMDQLRTMVEAADREKVELLNQLEEEKRKVEDLQFRVEEESI TKGDLETQTKLEHARIKELEQSLLFEKTKADKLQRELEDTRVATVSEKSRIMELEKDLALRVQEVAELRRRLESNKPAGDVDMSLSLLQE ISSLQEKLEVTRTDHQREITSLKEHFGAREETHQKEIKALYTATEKLSKENESLKSKLEHANKENSDVIALWKSKLETAIASHQQAMEEL KVSFSKGLGTETAEFAELKTQIEKMRLDYQHEIENLQNQQDSERAAHAKEMEALRAKLMKVIKEKENSLEAIRSKLDKAEDQHLVEMEDT LNKLQEAEIKVKELEVLQAKCNEQTKVIDNFTSQLKATEEKLLDLDALRKASSEGKSEMKKLRQQLEAAEKQIKHLEIEKNAESSKASSI TRELQGRELKLTNLQENLSEVSQVKETLEKELQILKEKFAEASEEAVSVQRSMQETVNKLHQKEEQFNMLSSDLEKLRENLADMEAKFRE KDEREEQLIKAKEKLENDIAEIMKMSGDNSSQLTKMNDELRLKERDVEELQLKLTKANENASFLQKSIEDMTVKAEQSQQEAAKKHEEEK KELERKLSDLEKKMETSHNQCQELKARYERATSETKTKHEEILQNLQKTLLDTEDKLKGAREENSGLLQELEELRKQADKAKAAQTAEDA MQIMEQMTKEKTETLASLEDTKQTNAKLQNELDTLKENNLKNVEELNKSKELLTVENQKMEEFRKEIETLKQAAAQKSQQLSALQEENVK -------------------------------------------------------------- >17164_17164_3_CLIP1-ROS1_CLIP1_chr12_122794308_ENST00000361654_ROS1_chr6_117641193_ENST00000368507_length(amino acids)=1084AA_BP= MSMLKPSGLKAPTKILKPGSTALKTPTAVVAPVEKTISSEKASSTPSSETQEEFVDDFRVGERVWVNGNKPGFIQFLGETQFAPGQWAGI VLDEPIGKNDGSVAGVRYFQCEPLKGIFTRPSKLTRKVQAEDEANGLQTTPASRATSPLCTSTASMVSSSPSTPSNIPQKPSQPAAKEPS ATPPISNLTKTASESISNLSEAGSIKKGERELKIGDRVLVGGTKAGVVRFLGETDFAKGEWCGVELDEPLGKNDGAVAGTRYFQCQPKYG LFAPVHKVTKIGFPSTTPAKAKANAVRRVMATTSASLKRSPSASSLSSMSSVASSVSSRPSRTGLLTETSSRYARKISGTTALQEALKEK QQHIEQLLAERDLERAEVAKATSHVGEIEQELALARDGHDQHVLELEAKMDQLRTMVEAADREKVELLNQLEEEKRKVEDLQFRVEEESI TKGDLEVATVSEKSRIMELEKDLALRVQEVAELRRRLESNKPAGDVDMSLSLLQEISSLQEKLEVTRTDHQREITSLKEHFGAREETHQK EIKALYTATEKLSKENESLKSKLEHANKENSDVIALWKSKLETAIASHQQAMEELKVSFSKGLGTETAEFAELKTQIEKMRLDYQHEIEN LQNQQDSERAAHAKEMEALRAKLMKVIKEKENSLEAIRSKLDKAEDQHLVEMEDTLNKLQEAEIKASSITRELQGRELKLTNLQENLSEV SQVKETLEKELQILKEKFAEASEEAVSVQRSMQETVNKLHQKEEQFNMLSSDLEKLRENLADMEAKFREKDEREEQLIKAKEKLENDIAE IMKMSGDNSSQLTKMNDELRLKERDVEELQLKLTKANENASFLQKSIEDMTVKAEQSQQEAAKKHEEEKKELERKLSDLEKKMETSHNQC QELKARYERATSETKTKHEEILQNLQKTLLDTEDKLKGAREENSGLLQELEELRKQADKAKAAQTAEDAMQIMEQMTKEKTETLASLEDT KQTNAKLQNELDTLKENNLKNVEELNKSKELLTVENQKMEEFRKEIETLKQAAAQKSQQLSALQEENVKLAEELGRSRDEVTSHQKYSSN -------------------------------------------------------------- >17164_17164_4_CLIP1-ROS1_CLIP1_chr12_122794308_ENST00000361654_ROS1_chr6_117641193_ENST00000368508_length(amino acids)=1084AA_BP= MSMLKPSGLKAPTKILKPGSTALKTPTAVVAPVEKTISSEKASSTPSSETQEEFVDDFRVGERVWVNGNKPGFIQFLGETQFAPGQWAGI VLDEPIGKNDGSVAGVRYFQCEPLKGIFTRPSKLTRKVQAEDEANGLQTTPASRATSPLCTSTASMVSSSPSTPSNIPQKPSQPAAKEPS ATPPISNLTKTASESISNLSEAGSIKKGERELKIGDRVLVGGTKAGVVRFLGETDFAKGEWCGVELDEPLGKNDGAVAGTRYFQCQPKYG LFAPVHKVTKIGFPSTTPAKAKANAVRRVMATTSASLKRSPSASSLSSMSSVASSVSSRPSRTGLLTETSSRYARKISGTTALQEALKEK QQHIEQLLAERDLERAEVAKATSHVGEIEQELALARDGHDQHVLELEAKMDQLRTMVEAADREKVELLNQLEEEKRKVEDLQFRVEEESI TKGDLEVATVSEKSRIMELEKDLALRVQEVAELRRRLESNKPAGDVDMSLSLLQEISSLQEKLEVTRTDHQREITSLKEHFGAREETHQK EIKALYTATEKLSKENESLKSKLEHANKENSDVIALWKSKLETAIASHQQAMEELKVSFSKGLGTETAEFAELKTQIEKMRLDYQHEIEN LQNQQDSERAAHAKEMEALRAKLMKVIKEKENSLEAIRSKLDKAEDQHLVEMEDTLNKLQEAEIKASSITRELQGRELKLTNLQENLSEV SQVKETLEKELQILKEKFAEASEEAVSVQRSMQETVNKLHQKEEQFNMLSSDLEKLRENLADMEAKFREKDEREEQLIKAKEKLENDIAE IMKMSGDNSSQLTKMNDELRLKERDVEELQLKLTKANENASFLQKSIEDMTVKAEQSQQEAAKKHEEEKKELERKLSDLEKKMETSHNQC QELKARYERATSETKTKHEEILQNLQKTLLDTEDKLKGAREENSGLLQELEELRKQADKAKAAQTAEDAMQIMEQMTKEKTETLASLEDT KQTNAKLQNELDTLKENNLKNVEELNKSKELLTVENQKMEEFRKEIETLKQAAAQKSQQLSALQEENVKLAEELGRSRDEVTSHQKYSSN -------------------------------------------------------------- >17164_17164_5_CLIP1-ROS1_CLIP1_chr12_122794308_ENST00000537178_ROS1_chr6_117641193_ENST00000368507_length(amino acids)=1160AA_BP= MSMLKPSGLKAPTKILKPGSTALKTPTAVVAPVEKTISSEKASSTPSSETQEEFVDDFRVGERVWVNGNKPGFIQFLGETQFAPGQWAGI VLDEPIGKNDGSVAGVRYFQCEPLKGIFTRPSKLTRKVQAEDEANGLQTTPASRATSPLCTSTASMVSSSPSTPSNIPQKPSQPAAKEPS ATPPISNLTKTASESISNLSEAGSIKKGERELKIGDRVLVGGTKAGVVRFLGETDFAKGEWCGVELDEPLGKNDGAVAGTRYFQCQPKYG LFAPVHKVTKIGFPSTTPAKAKANAVRRVMATTSASLKRSPSASSLSSMSSVASSVSSRPSRTGLLTETSSRYARKISGTTALQEALKEK QQHIEQLLAERDLERAEVAKATSHVGEIEQELALARDGHDQHVLELEAKMDQLRTMVEAADREKVELLNQLEEEKRKVEDLQFRVEEESI TKGDLEVATVSEKSRIMELEKDLALRVQEVAELRRRLESNKPAGDVDMSLSLLQEISSLQEKLEVTRTDHQREITSLKEHFGAREETHQK EIKALYTATEKLSKENESLKSKLEHANKENSDVIALWKSKLETAIASHQQAMEELKVSFSKGLGTETAEFAELKTQIEKMRLDYQHEIEN LQNQQDSERAAHAKEMEALRAKLMKVIKEKENSLEAIRSKLDKAEDQHLVEMEDTLNKLQEAEIKVKELEVLQAKCNEQTKVIDNFTSQL KATEEKLLDLDALRKASSEGKSEMKKLRQQLEAAEKQIKHLEIEKNAESSKASSITRELQGRELKLTNLQENLSEVSQVKETLEKELQIL KEKFAEASEEAVSVQRSMQETVNKLHQKEEQFNMLSSDLEKLRENLADMEAKFREKDEREEQLIKAKEKLENDIAEIMKMSGDNSSQLTK MNDELRLKERDVEELQLKLTKANENASFLQKSIEDMTVKAEQSQQEAAKKHEEEKKELERKLSDLEKKMETSHNQCQELKARYERATSET KTKHEEILQNLQKTLLDTEDKLKGAREENSGLLQELEELRKQADKAKAAQTAEDAMQIMEQMTKEKTETLASLEDTKQTNAKLQNELDTL -------------------------------------------------------------- >17164_17164_6_CLIP1-ROS1_CLIP1_chr12_122794308_ENST00000537178_ROS1_chr6_117641193_ENST00000368508_length(amino acids)=1160AA_BP= MSMLKPSGLKAPTKILKPGSTALKTPTAVVAPVEKTISSEKASSTPSSETQEEFVDDFRVGERVWVNGNKPGFIQFLGETQFAPGQWAGI VLDEPIGKNDGSVAGVRYFQCEPLKGIFTRPSKLTRKVQAEDEANGLQTTPASRATSPLCTSTASMVSSSPSTPSNIPQKPSQPAAKEPS ATPPISNLTKTASESISNLSEAGSIKKGERELKIGDRVLVGGTKAGVVRFLGETDFAKGEWCGVELDEPLGKNDGAVAGTRYFQCQPKYG LFAPVHKVTKIGFPSTTPAKAKANAVRRVMATTSASLKRSPSASSLSSMSSVASSVSSRPSRTGLLTETSSRYARKISGTTALQEALKEK QQHIEQLLAERDLERAEVAKATSHVGEIEQELALARDGHDQHVLELEAKMDQLRTMVEAADREKVELLNQLEEEKRKVEDLQFRVEEESI TKGDLEVATVSEKSRIMELEKDLALRVQEVAELRRRLESNKPAGDVDMSLSLLQEISSLQEKLEVTRTDHQREITSLKEHFGAREETHQK EIKALYTATEKLSKENESLKSKLEHANKENSDVIALWKSKLETAIASHQQAMEELKVSFSKGLGTETAEFAELKTQIEKMRLDYQHEIEN LQNQQDSERAAHAKEMEALRAKLMKVIKEKENSLEAIRSKLDKAEDQHLVEMEDTLNKLQEAEIKVKELEVLQAKCNEQTKVIDNFTSQL KATEEKLLDLDALRKASSEGKSEMKKLRQQLEAAEKQIKHLEIEKNAESSKASSITRELQGRELKLTNLQENLSEVSQVKETLEKELQIL KEKFAEASEEAVSVQRSMQETVNKLHQKEEQFNMLSSDLEKLRENLADMEAKFREKDEREEQLIKAKEKLENDIAEIMKMSGDNSSQLTK MNDELRLKERDVEELQLKLTKANENASFLQKSIEDMTVKAEQSQQEAAKKHEEEKKELERKLSDLEKKMETSHNQCQELKARYERATSET KTKHEEILQNLQKTLLDTEDKLKGAREENSGLLQELEELRKQADKAKAAQTAEDAMQIMEQMTKEKTETLASLEDTKQTNAKLQNELDTL -------------------------------------------------------------- |
Top |
Fusion Protein Functional Features |
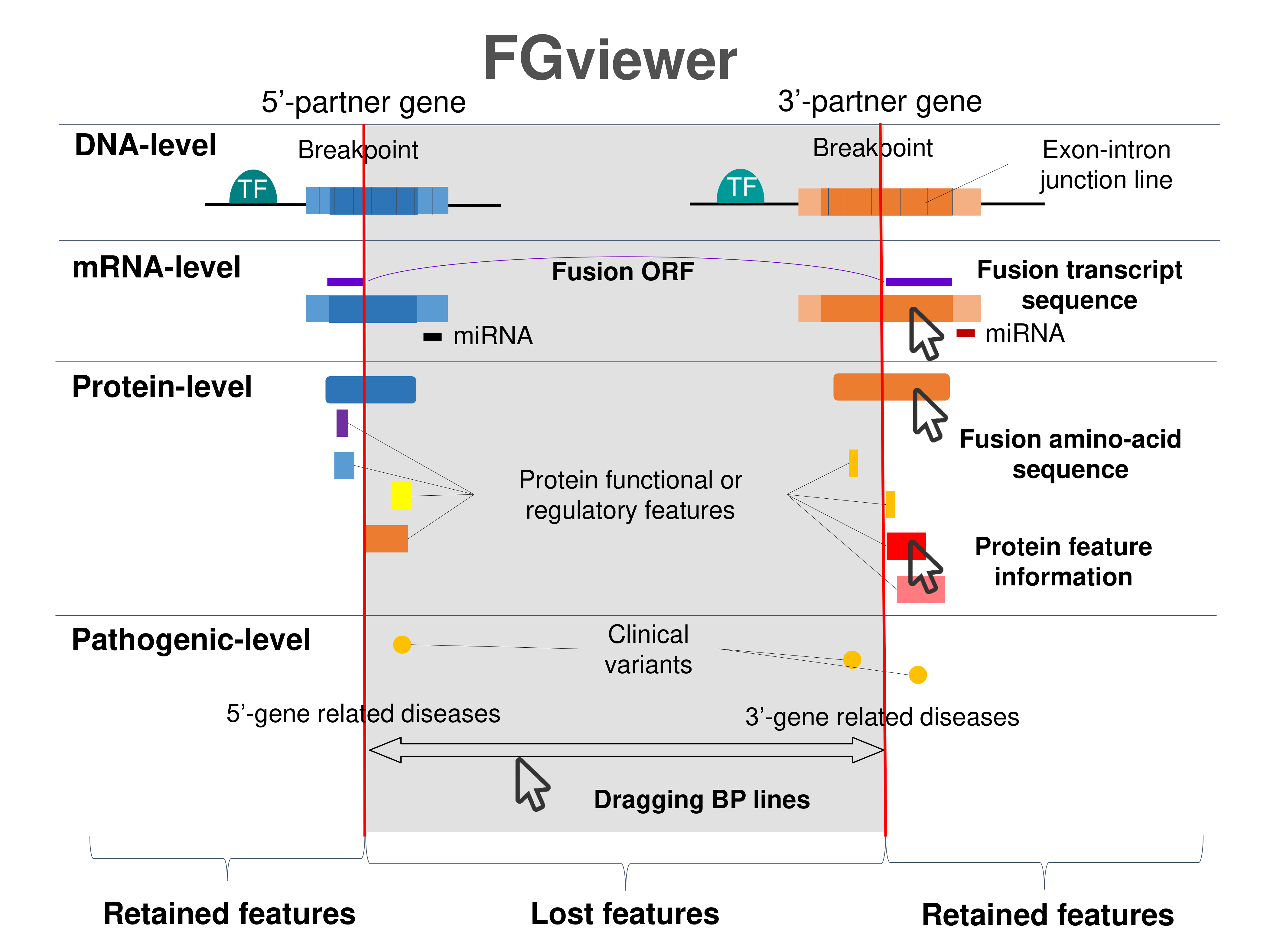
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr12:/chr6:) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| CLIP1 | ROS1 |
| FUNCTION: Binds to the plus end of microtubules and regulates the dynamics of the microtubule cytoskeleton. Promotes microtubule growth and microtubule bundling. Links cytoplasmic vesicles to microtubules and thereby plays an important role in intracellular vesicle trafficking. Plays a role macropinocytosis and endosome trafficking. {ECO:0000269|PubMed:12433698, ECO:0000269|PubMed:17563362, ECO:0000269|PubMed:17889670}. | FUNCTION: Orphan receptor tyrosine kinase (RTK) that plays a role in epithelial cell differentiation and regionalization of the proximal epididymal epithelium. May activate several downstream signaling pathways related to cell differentiation, proliferation, growth and survival including the PI3 kinase-mTOR signaling pathway. Mediates the phosphorylation of PTPN11, an activator of this pathway. May also phosphorylate and activate the transcription factor STAT3 to control anchorage-independent cell growth. Mediates the phosphorylation and the activation of VAV3, a guanine nucleotide exchange factor regulating cell morphology. May activate other downstream signaling proteins including AKT1, MAPK1, MAPK3, IRS1 and PLCG2. {ECO:0000269|PubMed:11094073, ECO:0000269|PubMed:16885344}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - Retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| - Not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
Top |
Fusion Protein Structures |
 PDB and CIF files of the predicted fusion proteins PDB and CIF files of the predicted fusion proteins * Here we show the 3D structure of the fusion proteins using Mol*. AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. Model confidence is shown from the pLDDT values per residue. pLDDT corresponds to the model’s prediction of its score on the local Distance Difference Test. It is a measure of local accuracy (from AlphfaFold website). To color code individual residues, we transformed individual PDB files into CIF format. |
| Fusion protein PDB link (fusion AA seq ID in FusionPDB) | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | AA seq | Len(AA seq) |
| PDB file (795) >>>795.pdbFusion protein BP residue: CIF file (795) >>>795.cif | CLIP1 | chr12 | 122794308 | - | ROS1 | chr6 | 117641193 | - | MSMLKPSGLKAPTKILKPGSTALKTPTAVVAPVEKTISSEKASSTPSSET QEEFVDDFRVGERVWVNGNKPGFIQFLGETQFAPGQWAGIVLDEPIGKND GSVAGVRYFQCEPLKGIFTRPSKLTRKVQAEDEANGLQTTPASRATSPLC TSTASMVSSSPSTPSNIPQKPSQPAAKEPSATPPISNLTKTASESISNLS EAGSIKKGERELKIGDRVLVGGTKAGVVRFLGETDFAKGEWCGVELDEPL GKNDGAVAGTRYFQCQPKYGLFAPVHKVTKIGFPSTTPAKAKANAVRRVM ATTSASLKRSPSASSLSSMSSVASSVSSRPSRTGLLTETSSRYARKISGT TALQEALKEKQQHIEQLLAERDLERAEVAKATSHVGEIEQELALARDGHD QHVLELEAKMDQLRTMVEAADREKVELLNQLEEEKRKVEDLQFRVEEESI TKGDLEVATVSEKSRIMELEKDLALRVQEVAELRRRLESNKPAGDVDMSL SLLQEISSLQEKLEVTRTDHQREITSLKEHFGAREETHQKEIKALYTATE KLSKENESLKSKLEHANKENSDVIALWKSKLETAIASHQQAMEELKVSFS KGLGTETAEFAELKTQIEKMRLDYQHEIENLQNQQDSERAAHAKEMEALR AKLMKVIKEKENSLEAIRSKLDKAEDQHLVEMEDTLNKLQEAEIKASSIT RELQGRELKLTNLQENLSEVSQVKETLEKELQILKEKFAEASEEAVSVQR SMQETVNKLHQKEEQFNMLSSDLEKLRENLADMEAKFREKDEREEQLIKA KEKLENDIAEIMKMSGDNSSQLTKMNDELRLKERDVEELQLKLTKANENA SFLQKSIEDMTVKAEQSQQEAAKKHEEEKKELERKLSDLEKKMETSHNQC QELKARYERATSETKTKHEEILQNLQKTLLDTEDKLKGAREENSGLLQEL EELRKQADKAKAAQTAEDAMQIMEQMTKEKTETLASLEDTKQTNAKLQNE LDTLKENNLKNVEELNKSKELLTVENQKMEEFRKEIETLKQAAAQKSQQL | 1084 |
| 3D view using mol* of 795 (AA BP:) | ||||||||||
 | ||||||||||
| PDB file (836) >>>836.pdbFusion protein BP residue: CIF file (836) >>>836.cif | CLIP1 | chr12 | 122794308 | - | ROS1 | chr6 | 117641193 | - | MSMLKPSGLKAPTKILKPGSTALKTPTAVVAPVEKTISSEKASSTPSSET QEEFVDDFRVGERVWVNGNKPGFIQFLGETQFAPGQWAGIVLDEPIGKND GSVAGVRYFQCEPLKGIFTRPSKLTRKVQAEDEANGLQTTPASRATSPLC TSTASMVSSSPSTPSNIPQKPSQPAAKEPSATPPISNLTKTASESISNLS EAGSIKKGERELKIGDRVLVGGTKAGVVRFLGETDFAKGEWCGVELDEPL GKNDGAVAGTRYFQCQPKYGLFAPVHKVTKIGFPSTTPAKAKANAVRRVM ATTSASLKRSPSASSLSSMSSVASSVSSRPSRTGLLTETSSRYARKISGT TALQEALKEKQQHIEQLLAERDLERAEVAKATSHVGEIEQELALARDGHD QHVLELEAKMDQLRTMVEAADREKVELLNQLEEEKRKVEDLQFRVEEESI TKGDLEVATVSEKSRIMELEKDLALRVQEVAELRRRLESNKPAGDVDMSL SLLQEISSLQEKLEVTRTDHQREITSLKEHFGAREETHQKEIKALYTATE KLSKENESLKSKLEHANKENSDVIALWKSKLETAIASHQQAMEELKVSFS KGLGTETAEFAELKTQIEKMRLDYQHEIENLQNQQDSERAAHAKEMEALR AKLMKVIKEKENSLEAIRSKLDKAEDQHLVEMEDTLNKLQEAEIKVKELE VLQAKCNEQTKVIDNFTSQLKATEEKLLDLDALRKASSEGKSEMKKLRQQ LEAAEKQIKHLEIEKNAESSKASSITRELQGRELKLTNLQENLSEVSQVK ETLEKELQILKEKFAEASEEAVSVQRSMQETVNKLHQKEEQFNMLSSDLE KLRENLADMEAKFREKDEREEQLIKAKEKLENDIAEIMKMSGDNSSQLTK MNDELRLKERDVEELQLKLTKANENASFLQKSIEDMTVKAEQSQQEAAKK HEEEKKELERKLSDLEKKMETSHNQCQELKARYERATSETKTKHEEILQN LQKTLLDTEDKLKGAREENSGLLQELEELRKQADKAKAAQTAEDAMQIME QMTKEKTETLASLEDTKQTNAKLQNELDTLKENNLKNVEELNKSKELLTV ENQKMEEFRKEIETLKQAAAQKSQQLSALQEENVKLAEELGRSRDEVTSH | 1160 |
| 3D view using mol* of 836 (AA BP:) | ||||||||||
 | ||||||||||
| PDB file (845) >>>845.pdbFusion protein BP residue: CIF file (845) >>>845.cif | CLIP1 | chr12 | 122794308 | - | ROS1 | chr6 | 117641193 | - | MSMLKPSGLKAPTKILKPGSTALKTPTAVVAPVEKTISSEKASSTPSSET QEEFVDDFRVGERVWVNGNKPGFIQFLGETQFAPGQWAGIVLDEPIGKND GSVAGVRYFQCEPLKGIFTRPSKLTRKVQAEDEANGLQTTPASRATSPLC TSTASMVSSSPSTPSNIPQKPSQPAAKEPSATPPISNLTKTASESISNLS EAGSIKKGERELKIGDRVLVGGTKAGVVRFLGETDFAKGEWCGVELDEPL GKNDGAVAGTRYFQCQPKYGLFAPVHKVTKIGFPSTTPAKAKANAVRRVM ATTSASLKRSPSASSLSSMSSVASSVSSRPSRTGLLTETSSRYARKISGT TALQEALKEKQQHIEQLLAERDLERAEVAKATSHVGEIEQELALARDGHD QHVLELEAKMDQLRTMVEAADREKVELLNQLEEEKRKVEDLQFRVEEESI TKGDLETQTKLEHARIKELEQSLLFEKTKADKLQRELEDTRVATVSEKSR IMELEKDLALRVQEVAELRRRLESNKPAGDVDMSLSLLQEISSLQEKLEV TRTDHQREITSLKEHFGAREETHQKEIKALYTATEKLSKENESLKSKLEH ANKENSDVIALWKSKLETAIASHQQAMEELKVSFSKGLGTETAEFAELKT QIEKMRLDYQHEIENLQNQQDSERAAHAKEMEALRAKLMKVIKEKENSLE AIRSKLDKAEDQHLVEMEDTLNKLQEAEIKVKELEVLQAKCNEQTKVIDN FTSQLKATEEKLLDLDALRKASSEGKSEMKKLRQQLEAAEKQIKHLEIEK NAESSKASSITRELQGRELKLTNLQENLSEVSQVKETLEKELQILKEKFA EASEEAVSVQRSMQETVNKLHQKEEQFNMLSSDLEKLRENLADMEAKFRE KDEREEQLIKAKEKLENDIAEIMKMSGDNSSQLTKMNDELRLKERDVEEL QLKLTKANENASFLQKSIEDMTVKAEQSQQEAAKKHEEEKKELERKLSDL EKKMETSHNQCQELKARYERATSETKTKHEEILQNLQKTLLDTEDKLKGA REENSGLLQELEELRKQADKAKAAQTAEDAMQIMEQMTKEKTETLASLED TKQTNAKLQNELDTLKENNLKNVEELNKSKELLTVENQKMEEFRKEIETL | 1195 |
| 3D view using mol* of 845 (AA BP:) | ||||||||||
 | ||||||||||
Top |
pLDDT score distribution |
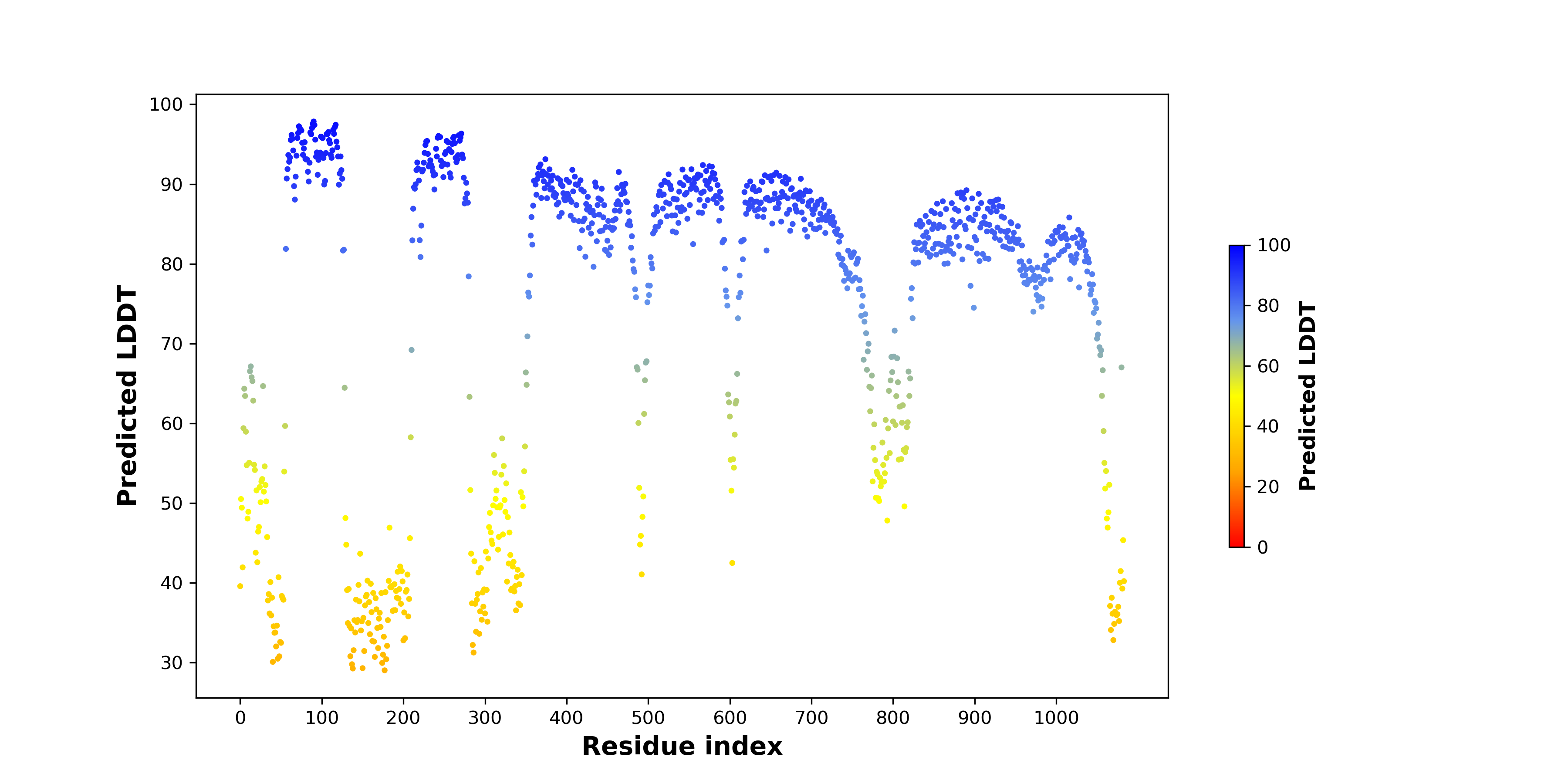
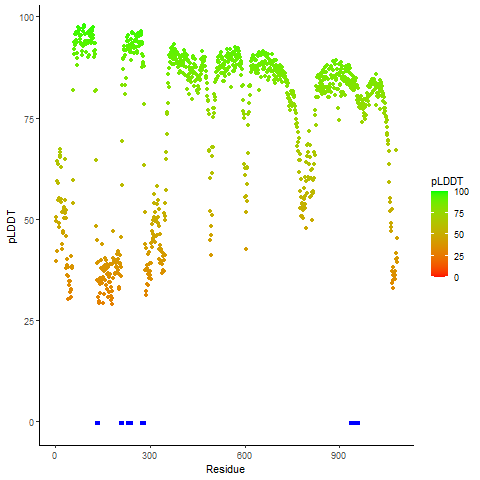
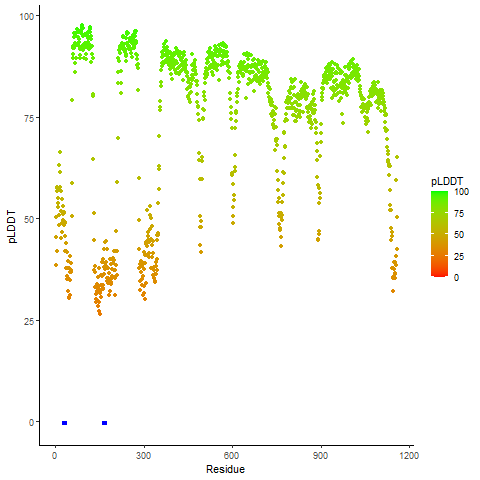
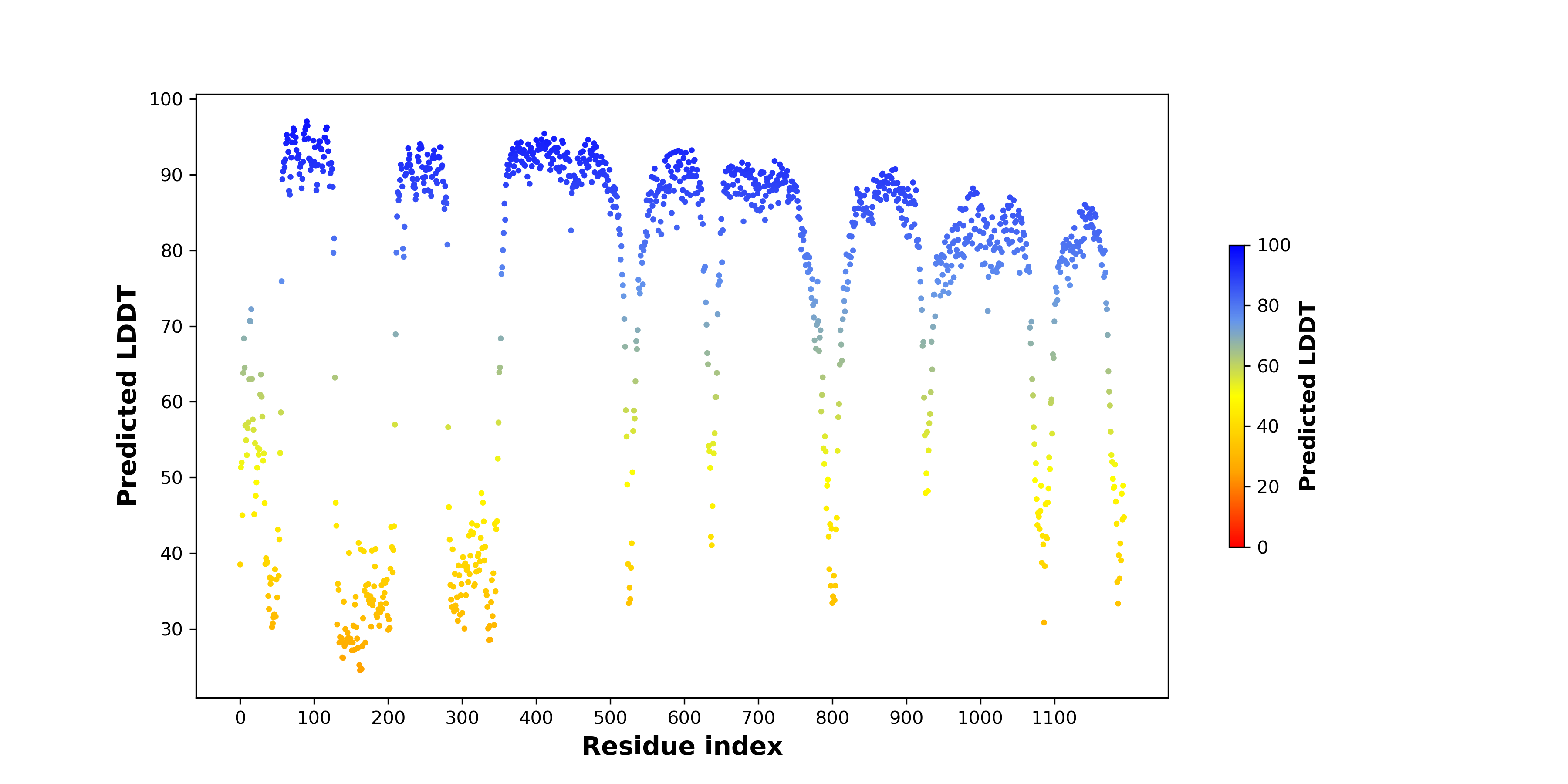
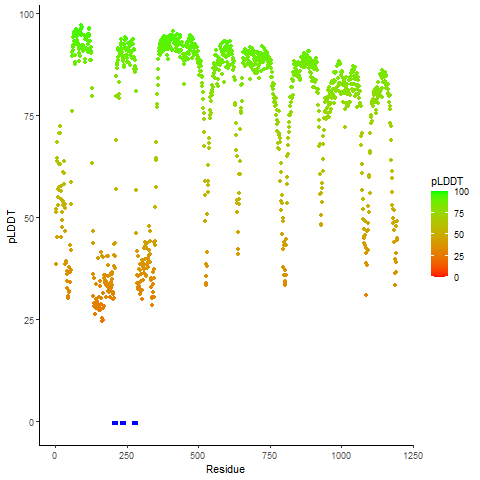
 pLDDT score distribution of the predicted wild-type structures of two partner proteins from AlphaFold2 pLDDT score distribution of the predicted wild-type structures of two partner proteins from AlphaFold2* AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. |
CLIP1_pLDDT.png |
ROS1_pLDDT.png |
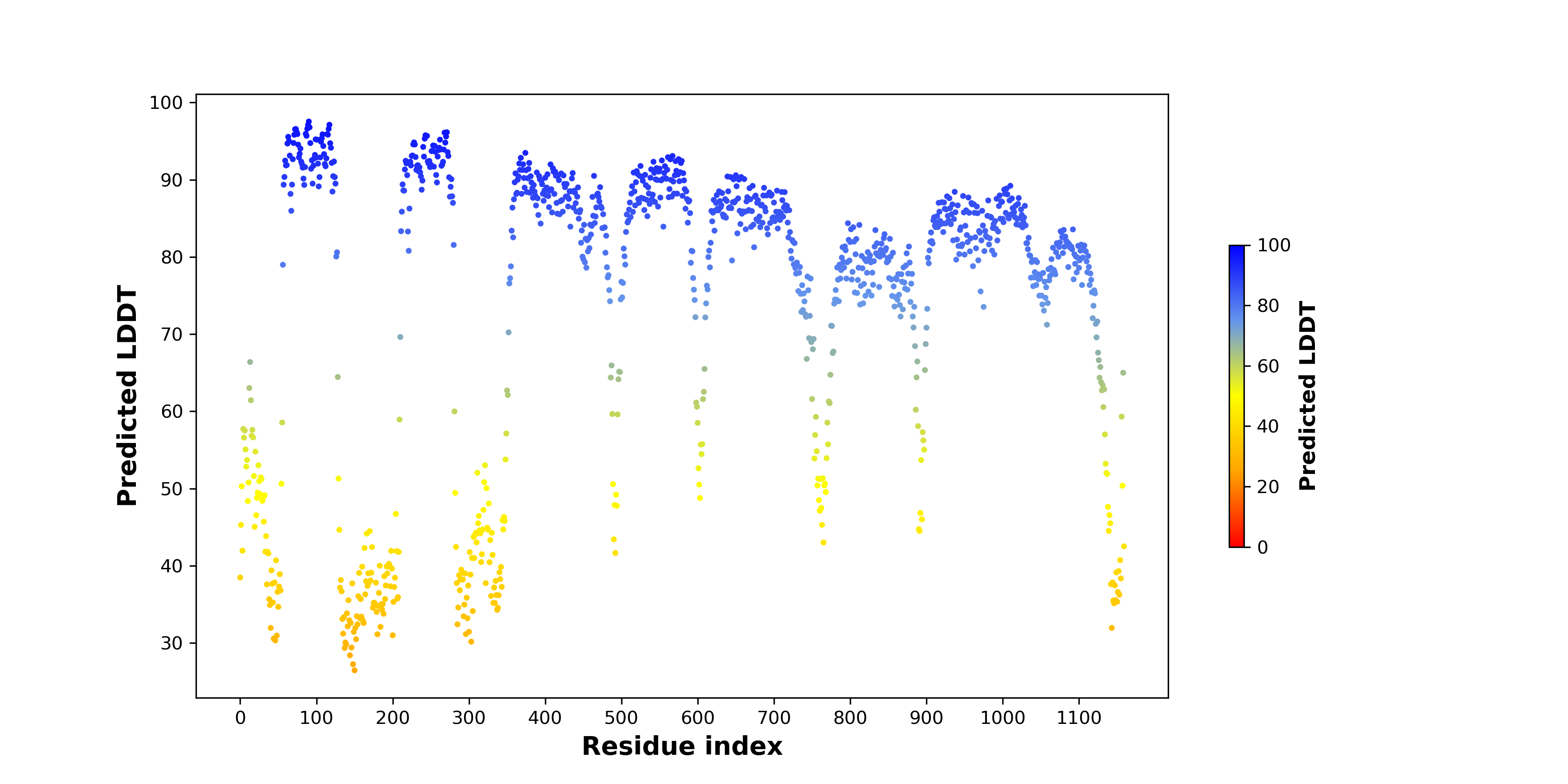
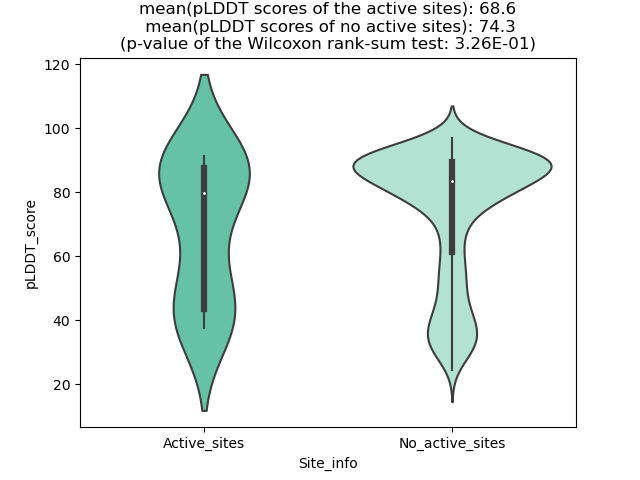
 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2* AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. |
Top |
Ramachandran Plot of Fusion Protein Structure |
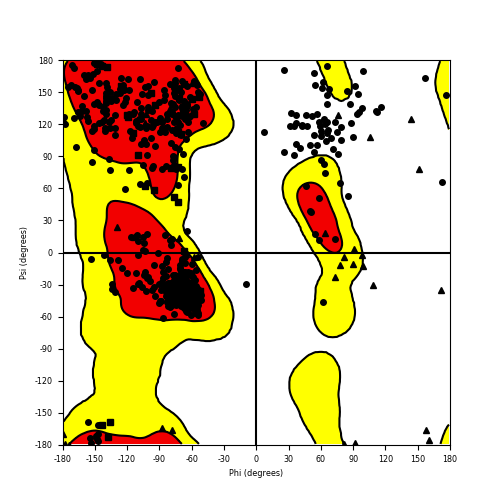
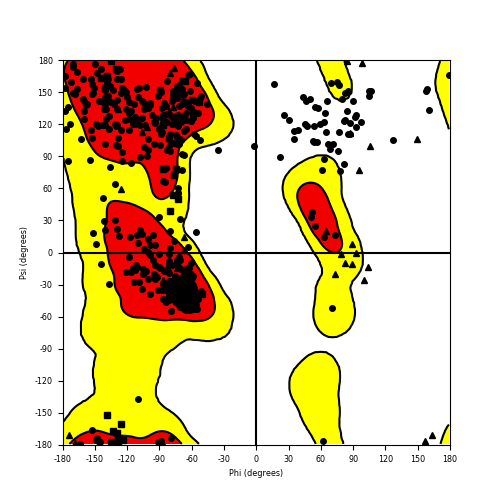
 Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. |
| Fusion AA seq ID in FusionPDB and their Ramachandran plots |
| CLIP1_ROS1_836.png |
 |
| CLIP1_ROS1_845.png |
 |
Top |
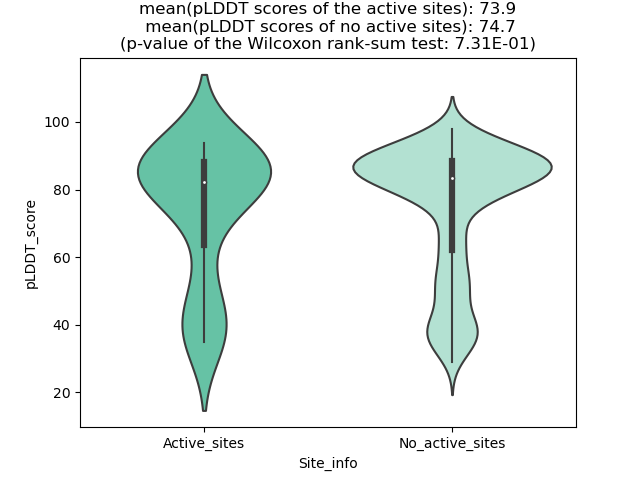
Potential Active Site Information |
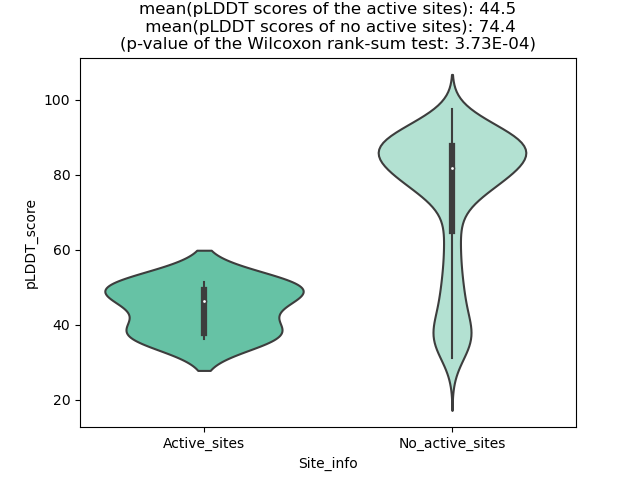
 The potential binding sites of these fusion proteins were identified using SiteMap, a module of the Schrodinger suite. The potential binding sites of these fusion proteins were identified using SiteMap, a module of the Schrodinger suite. |
| Fusion AA seq ID in FusionPDB | Site score | Size | D score | Volume | Exposure | Enclosure | Contact | Phobic | Philic | Balance | Don/Acc | Residues |
| 795 | 0.962 | 183 | 0.966 | 474.026 | 0.565 | 0.641 | 0.886 | 0.231 | 1.099 | 0.21 | 0.833 | Chain A: 131,132,133,134,207,208,209,210,211,212,2 31,233,238,239,240,275,276,278,279,280,936,939,940 ,943,946,947,950,953,956,957,961 |
| 836 | 0.6524 | 47 | 0.6018 | 11.662 | 0.5766 | 0.523 | 0.7731 | 0.146 | 1.0934 | 0.1335 | 0.4436 | Chain A: 28,29,30,31,165,166,167,168 |
| 845 | 0.8098 | 68 | 0.8025 | 171.5 | 0.6566 | 0.5994 | 0.8713 | 0.2578 | 1.0191 | 0.253 | 1.5495 | Chain A: 205,206,207,208,209,210,211,212,231,240,2 75,276,278,279,280 |
Top |
Potentially Interacting Small Molecules through Virtual Screening |
 The FDA-approved small molecule library molecules were subjected to virtual screening using the Glide. The FDA-approved small molecule library molecules were subjected to virtual screening using the Glide. |
| Fusion AA seq ID in FusionPDB | ZINC ID | DrugBank ID | Drug name | Docking score | Glide gscore |
Top |
 Drug information from DrugBank of the top 20 interacting small molecules. Drug information from DrugBank of the top 20 interacting small molecules. |
| ZINC ID | DrugBank ID | Drug name | Drug type | SMILES | Drug group |
Top |
Biochemical Features of Small Molecules |
 ADME (Absorption, Distribution, Metabolism, and Excretion) of drugs using QikProp(v3.9) ADME (Absorption, Distribution, Metabolism, and Excretion) of drugs using QikProp(v3.9) |
| ZINC ID | mol_MW | dipole | SASA | FOSA | FISA | PISA | WPSA | volume | donorHB | accptHB | IP | Human Oral Absorption | Percent Human Oral Absorption | Rule Of Five | Rule Of Three |
Top |
Drug Toxicity Information |
 Toxicity information of individual drugs using eToxPred Toxicity information of individual drugs using eToxPred |
| ZINC ID | Smile | Surface Accessibility | Toxicity |
Top |
Fusion Protein-Protein Interaction |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type from validated records (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type from validated records (BIOGRID-3.4.160) |
| Gene | PPI interactors |
| CLIP1 | MTOR, IQGAP1, CDC42, RAC1, PAFAH1B1, Gorasp1, CALM1, BIN1, MAPRE1, TUBA1A, TUBB, AMOT, BUB3, MAEA, PSMC2, HDAC6, CADPS, CHD4, GPS1, DCTN1, DYNC1H1, Mapre1, Smurf2, PCGF1, DPPA4, NANOG, POU5F1, TSC22D2, GORASP1, BMP1, DAB2IP, RUNDC3A, BRCA1, ZNF598, UBE2M, KIAA1429, DCAF15, BICD1, REL, ORF3a, SERBP1, nsp2, CLIP4, NINL, ORF6, DDX58, AGAP3, CKAP2, CLASP1, CLASP2, DNAJA2, IKBKAP, ELP4, EXOC4, HINT1, HMMR, HNRNPA1L2, IARS2, KIF15, MAP7, MAST4, PLEKHA5, RPS6KA3, SIRT2, SLAIN1, TCHP, TEX9, BIRC6, CAMSAP1, DRG1, KIAA0368, MAP7D1, MAP7D2, MAP7D3, SF3B3, SLAIN2, MAPRE3, TRIM36, PPP2R2B, RABEP1, SRRT, OR2A4, GOT1, RABEP2, SYCE1, MBNL1, SSUH2, C18orf21, AGPAT1, CCR1, FBXO32, CDK1, |
 Protein-protein interactors based on sequence similarity (STRING) Protein-protein interactors based on sequence similarity (STRING) |
| Gene | STRING network |
| CLIP1 |  |
| ROS1 |  |
 - Retained interactions in fusion protein (protein functional feature from UniProt). - Retained interactions in fusion protein (protein functional feature from UniProt). |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost interactions due to fusion (protein functional feature from UniProt). - Lost interactions due to fusion (protein functional feature from UniProt). |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs to CLIP1-ROS1 |
 Drugs used for this fusion-positive patient. Drugs used for this fusion-positive patient. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Drug | Source | PMID |
Top |
Related Diseases to CLIP1-ROS1 |
 Diseases that have this fusion gene. Diseases that have this fusion gene. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Disease | Source | PMID |
| CLIP1 | ROS1 | Lung Adenocarcinoma | MyCancerGenome | |
| CLIP1 | ROS1 | Dedifferentiated Liposarcoma | MyCancerGenome | |
| CLIP1 | ROS1 | Breast Invasive Ductal Carcinoma | MyCancerGenome | |
| CLIP1 | ROS1 | Anaplastic Ganglioglioma | MyCancerGenome | |
| CLIP1 | ROS1 | Cancer Of Unknown Primary | MyCancerGenome |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
| Tgene | ROS1 | C0007131 | Non-Small Cell Lung Carcinoma | 4 | CTD_human |
| Tgene | ROS1 | C0017638 | Glioma | 1 | CTD_human |
| Tgene | ROS1 | C0025202 | melanoma | 1 | CTD_human |
| Tgene | ROS1 | C0152013 | Adenocarcinoma of lung (disorder) | 1 | CGI;CTD_human |
| Tgene | ROS1 | C0259783 | mixed gliomas | 1 | CTD_human |
| Tgene | ROS1 | C0555198 | Malignant Glioma | 1 | CTD_human |