| UTHEALTH HOME ABOUT SBMI A-Z WEBMAIL INSIDE THE UNIVERSITY |

|
|||||||
|
Fusion Protein:DDX17-CDH10 |
Fusion Protein Summary |
 Fusion gene summary Fusion gene summary |
| Fusion partner gene information | Fusion gene name: DDX17-CDH10 | FusionPDB ID: 21897 | FusionGDB2.0 ID: 21897 | Hgene | Tgene | Gene symbol | DDX17 | CDH10 | Gene ID | 10521 | 1008 |
| Gene name | DEAD-box helicase 17 | cadherin 10 | |
| Synonyms | P72|RH70 | - | |
| Cytomap | 22q13.1 | 5p14.2-p14.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | probable ATP-dependent RNA helicase DDX17DEAD (Asp-Glu-Ala-Asp) box helicase 17DEAD (Asp-Glu-Ala-Asp) box polypeptide 17DEAD box protein p72DEAD box protein p82DEAD/H (Asp-Glu-Ala-Asp/His) box polypeptide 17 (72kD)RNA-dependent helicase p72 | cadherin-10T2-cadherincadherin 10 type 2 | |
| Modification date | 20200327 | 20200313 | |
| UniProtAcc | Q92841 | Q9Y6N8 | |
| Ensembl transtripts involved in fusion gene | ENST ids | ENST00000381633, ENST00000396821, ENST00000432525, ENST00000444597, | ENST00000502921, ENST00000264463, |
| Fusion gene scores for assessment (based on all fusion genes of FusionGDB 2.0) | * DoF score | 17 X 18 X 8=2448 | 6 X 6 X 3=108 |
| # samples | 20 | 5 | |
| ** MAII score | log2(20/2448*10)=-3.61353165291793 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(5/108*10)=-1.11103131238874 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context (manual curation of fusion genes in FusionPDB) | PubMed: DDX17 [Title/Abstract] AND CDH10 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint (based on all fusion genes of FusionGDB 2.0) | DDX17(38888061)-CDH10(24498628), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | DDX17-CDH10 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. DDX17-CDH10 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. DDX17-CDH10 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. DDX17-CDH10 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. DDX17-CDH10 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. DDX17-CDH10 seems lost the major protein functional domain in Tgene partner, which is a CGC due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | DDX17 | GO:0045944 | positive regulation of transcription by RNA polymerase II | 17226766 |
 Fusion gene breakpoints across DDX17 (5'-gene) Fusion gene breakpoints across DDX17 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
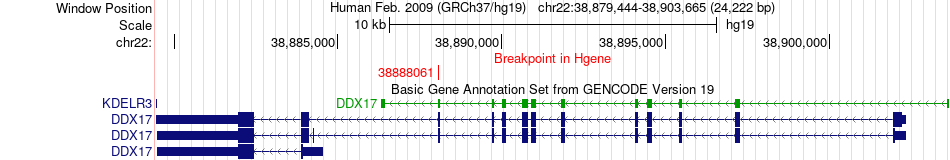 |
 Fusion gene breakpoints across CDH10 (3'-gene) Fusion gene breakpoints across CDH10 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
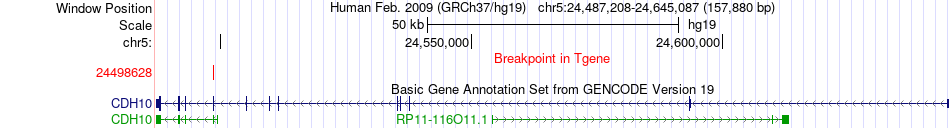 |
Top |
Fusion Gene Sample Information |
 Fusion gene information from FusionGDB2.0. Fusion gene information from FusionGDB2.0. |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | SKCM | TCGA-D3-A1Q3-06A | DDX17 | chr22 | 38888061 | - | CDH10 | chr5 | 24498628 | - |
Top |
Fusion ORF Analysis |
 Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000396821 | DDX17 | chr22 | 38888061 | - | ENST00000264463 | CDH10 | chr5 | 24498628 | - | 3084 | 1547 | 100 | 2520 | 806 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000396821 | ENST00000264463 | DDX17 | chr22 | 38888061 | - | CDH10 | chr5 | 24498628 | - | 0.000594043 | 0.999406 |
Top |
Fusion Amino Acid Sequences |
 For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >FusionGDB ID_FusionGDB isoform ID_FGname_Hgene_Hchr_Hbp_Henst_Tgene_Tchr_Tbp_Tenst_length(fusion AA) seq_BP >21897_21897_1_DDX17-CDH10_DDX17_chr22_38888061_ENST00000396821_CDH10_chr5_24498628_ENST00000264463_length(amino acids)=806AA_BP=482 MPTGFVAPILCVLLPSPTREAATVASATGDSASERESAAPAAAPTAEAPPPSVVTRPEPQALPSPAIRAPLPDLYPFGTMRGGGFGDRDR DRDRGGFGARGGGGLPPKKFGNPGERLRKKKWDLSELPKFEKNFYVEHPEVARLTPYEVDELRRKKEITVRGGDVCPKPVFAFHHANFPQ YVMDVLMDQHFTEPTPIQCQGFPLALSGRDMVGIAQTGSGKTLAYLLPAIVHINHQPYLERGDGPICLVLAPTRELAQQVQQVADDYGKC SRLKSTCIYGGAPKGPQIRDLERGVEICIATPGRLIDFLESGKTNLRRCTYLVLDEADRMLDMGFEPQIRKIVDQIRPDRQTLMWSATWP KEVRQLAEDFLRDYTQINVGNLELSANHNILQIVDVCMESEKDHKLIQLMEEIMAEKENKTIIFVETKRRCDDLTRRMRRDGWPAMCIHG DKSQPERDWVLNEFRSGKAPILIATDVASRGLDNPKETTRVAVFVRILDVNDNAPQFAVFYDTFVCENARPGQLIQTISAVDKDDPLGGQ KFFFSLAAVNPNFTVQDNEDNTARILTRKNGFNRHEISTYLLPVVISDNDYPIQSSTGTLTIRVCACDSQGNMQSCSAEALLLPAGLSTG ALIAILLCIIILLVIVVLFAALKRQRKKEPLILSKEDIRDNIVSYNDEGGGEEDTQAFDIGTLRNPAAIEEKKLRRDIIPETLFIPRRTP -------------------------------------------------------------- |
Top |
Fusion Protein Functional Features |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr22:38888061/chr5:24498628) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
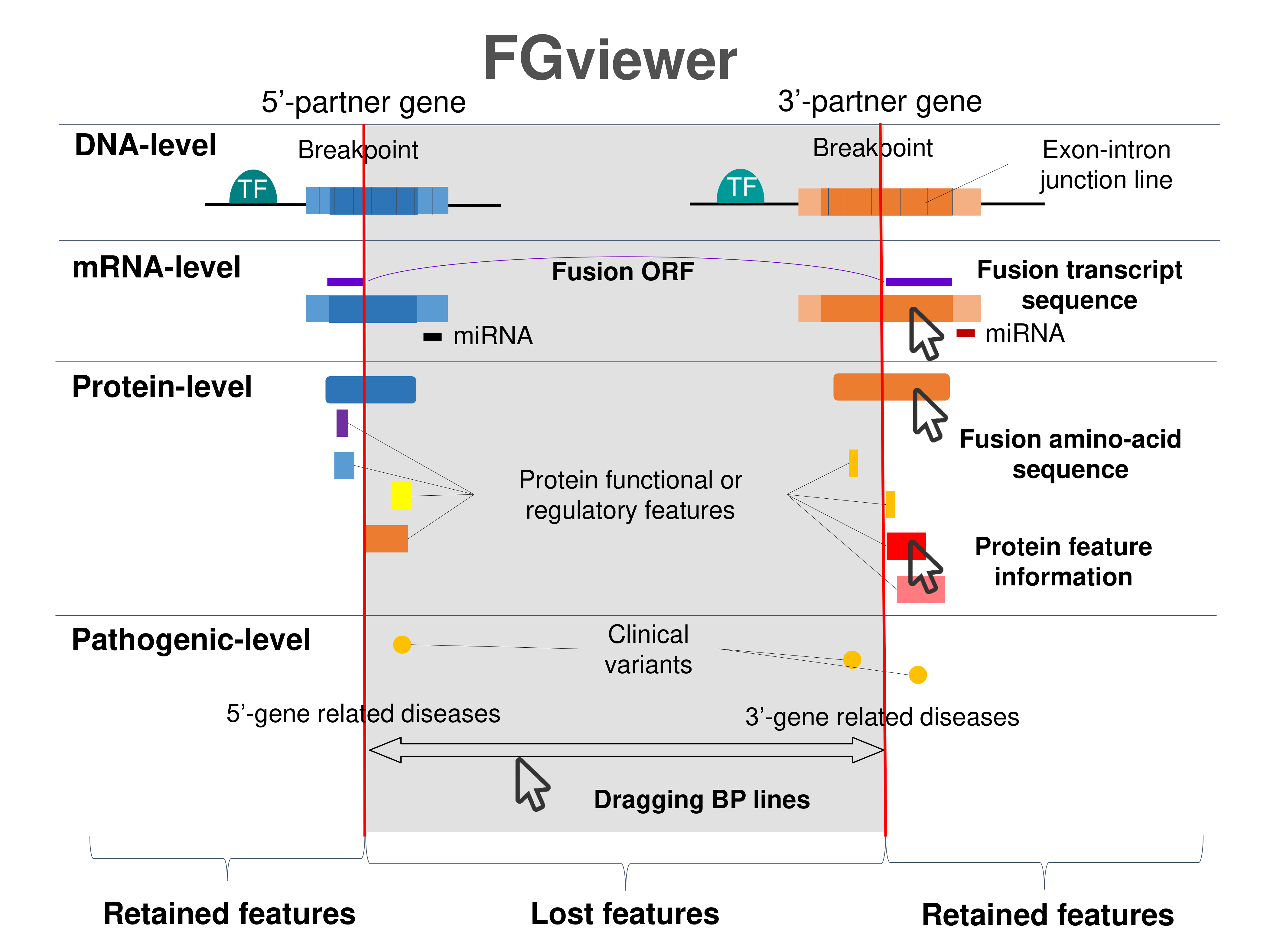 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| DDX17 | CDH10 |
| FUNCTION: As an RNA helicase, unwinds RNA and alters RNA structures through ATP binding and hydrolysis. Involved in multiple cellular processes, including pre-mRNA splicing, alternative splicing, ribosomal RNA processing and miRNA processing, as well as transcription regulation. Regulates the alternative splicing of exons exhibiting specific features (PubMed:12138182, PubMed:23022728, PubMed:24910439, PubMed:22266867). For instance, promotes the inclusion of AC-rich alternative exons in CD44 transcripts (PubMed:12138182). This function requires the RNA helicase activity (PubMed:12138182, PubMed:23022728, PubMed:24910439, PubMed:22266867). Affects NFAT5 and histone macro-H2A.1/MACROH2A1 alternative splicing in a CDK9-dependent manner (PubMed:26209609, PubMed:22266867). In NFAT5, promotes the introduction of alternative exon 4, which contains 2 stop codons and may target NFAT5 exon 4-containing transcripts to nonsense-mediated mRNA decay, leading to the down-regulation of NFAT5 protein (PubMed:22266867). Affects splicing of mediators of steroid hormone signaling pathway, including kinases that phosphorylates ESR1, such as CDK2, MAPK1 and GSK3B, and transcriptional regulators, such as CREBBP, MED1, NCOR1 and NCOR2. By affecting GSK3B splicing, participates in ESR1 and AR stabilization (PubMed:24275493). In myoblasts and epithelial cells, cooperates with HNRNPH1 to control the splicing of specific subsets of exons (PubMed:24910439). In addition to binding mature mRNAs, also interacts with certain pri-microRNAs, including MIR663/miR-663a, MIR99B/miR-99b, and MIR6087/miR-6087 (PubMed:25126784). Binds pri-microRNAs on the 3' segment flanking the stem loop via the 5'-[ACG]CAUC[ACU]-3' consensus sequence (PubMed:24581491). Required for the production of subsets of microRNAs, including MIR21 and MIR125B1 (PubMed:24581491, PubMed:27478153). May be involved not only in microRNA primary transcript processing, but also stabilization (By similarity). Participates in MYC down-regulation at high cell density through the production of MYC-targeting microRNAs (PubMed:24581491). Along with DDX5, may be involved in the processing of the 32S intermediate into the mature 28S ribosomal RNA (PubMed:17485482). Promoter-specific transcription regulator, functioning as a coactivator or corepressor depending on the context of the promoter and the transcriptional complex in which it exists (PubMed:15298701). Enhances NFAT5 transcriptional activity (PubMed:22266867). Synergizes with TP53 in the activation of the MDM2 promoter; this activity requires acetylation on lysine residues (PubMed:17226766, PubMed:20663877, PubMed:19995069). May also coactivate MDM2 transcription through a TP53-independent pathway (PubMed:17226766). Coactivates MMP7 transcription (PubMed:17226766). Along with CTNNB1, coactivates MYC, JUN, FOSL1 and cyclin D1/CCND1 transcription (PubMed:17699760). Alone or in combination with DDX5 and/or SRA1 non-coding RNA, plays a critical role in promoting the assembly of proteins required for the formation of the transcription initiation complex and chromatin remodeling leading to coactivation of MYOD1-dependent transcription. This helicase-independent activity is required for skeletal muscle cells to properly differentiate into myotubes (PubMed:17011493, PubMed:24910439). During epithelial-to-mesenchymal transition, coregulates SMAD-dependent transcriptional activity, directly controlling key effectors of differentiation, including miRNAs which in turn directly repress its expression (PubMed:24910439). Plays a role in estrogen and testosterone signaling pathway at several levels. Mediates the use of alternative promoters in estrogen-responsive genes and regulates transcription and splicing of a large number of steroid hormone target genes (PubMed:24275493, PubMed:20406972, PubMed:20663877, PubMed:19995069). Contrary to splicing regulation activity, transcriptional coregulation of the estrogen receptor ESR1 is helicase-independent (PubMed:19718048, PubMed:24275493). Plays a role in innate immunity. Specifically restricts bunyavirus infection, including Rift Valley fever virus (RVFV) or La Crosse virus (LACV), but not vesicular stomatitis virus (VSV), in an interferon- and DROSHA-independent manner (PubMed:25126784). Binds to RVFV RNA, likely via structured viral RNA elements (PubMed:25126784). Promotes mRNA degradation mediated by the antiviral zinc-finger protein ZC3HAV1, in an ATPase-dependent manner (PubMed:18334637). {ECO:0000250|UniProtKB:Q501J6, ECO:0000269|PubMed:12138182, ECO:0000269|PubMed:15298701, ECO:0000269|PubMed:17011493, ECO:0000269|PubMed:17226766, ECO:0000269|PubMed:17485482, ECO:0000269|PubMed:17699760, ECO:0000269|PubMed:18334637, ECO:0000269|PubMed:19718048, ECO:0000269|PubMed:19995069, ECO:0000269|PubMed:20406972, ECO:0000269|PubMed:20663877, ECO:0000269|PubMed:22266867, ECO:0000269|PubMed:23022728, ECO:0000269|PubMed:24275493, ECO:0000269|PubMed:24581491, ECO:0000269|PubMed:24910439, ECO:0000269|PubMed:25126784, ECO:0000269|PubMed:26209609, ECO:0000269|PubMed:27478153, ECO:0000305}. | FUNCTION: Cadherins are calcium-dependent cell adhesion proteins. They preferentially interact with themselves in a homophilic manner in connecting cells; cadherins may thus contribute to the sorting of heterogeneous cell types. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - Retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | CDH10 | chr22:38888061 | chr5:24498628 | ENST00000264463 | 7 | 12 | 488_606 | 464.3333333333333 | 789.0 | Domain | Cadherin 5 | |
| Tgene | CDH10 | chr22:38888061 | chr5:24498628 | ENST00000264463 | 7 | 12 | 635_788 | 464.3333333333333 | 789.0 | Topological domain | Cytoplasmic | |
| Tgene | CDH10 | chr22:38888061 | chr5:24498628 | ENST00000264463 | 7 | 12 | 614_634 | 464.3333333333333 | 789.0 | Transmembrane | Helical |
| - Not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | CDH10 | chr22:38888061 | chr5:24498628 | ENST00000264463 | 7 | 12 | 161_269 | 464.3333333333333 | 789.0 | Domain | Cadherin 2 | |
| Tgene | CDH10 | chr22:38888061 | chr5:24498628 | ENST00000264463 | 7 | 12 | 270_384 | 464.3333333333333 | 789.0 | Domain | Cadherin 3 | |
| Tgene | CDH10 | chr22:38888061 | chr5:24498628 | ENST00000264463 | 7 | 12 | 385_487 | 464.3333333333333 | 789.0 | Domain | Cadherin 4 | |
| Tgene | CDH10 | chr22:38888061 | chr5:24498628 | ENST00000264463 | 7 | 12 | 55_160 | 464.3333333333333 | 789.0 | Domain | Cadherin 1 | |
| Tgene | CDH10 | chr22:38888061 | chr5:24498628 | ENST00000264463 | 7 | 12 | 55_613 | 464.3333333333333 | 789.0 | Topological domain | Extracellular |
Top |
Fusion Protein-Protein Interaction |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type from validated records (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type from validated records (BIOGRID-3.4.160) |
| Gene | PPI interactors |
 Protein-protein interactors based on sequence similarity (STRING) Protein-protein interactors based on sequence similarity (STRING) |
| Gene | STRING network |
| DDX17 | |
| CDH10 |
 - Retained interactions in fusion protein (protein functional feature from UniProt). - Retained interactions in fusion protein (protein functional feature from UniProt). |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost interactions due to fusion (protein functional feature from UniProt). - Lost interactions due to fusion (protein functional feature from UniProt). |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs to DDX17-CDH10 |
 Drugs used for this fusion-positive patient. Drugs used for this fusion-positive patient. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Drug | Source | PMID |
Top |
Related Diseases to DDX17-CDH10 |
 Diseases that have this fusion gene. Diseases that have this fusion gene. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Disease | Source | PMID |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |

