| UTHEALTH HOME ABOUT SBMI A-Z WEBMAIL INSIDE THE UNIVERSITY |

|
|||||||
|
Fusion Protein:EIF2AK4-PKM |
Fusion Protein Summary |
 Fusion gene summary Fusion gene summary |
| Fusion partner gene information | Fusion gene name: EIF2AK4-PKM | FusionPDB ID: 25682 | FusionGDB2.0 ID: 25682 | Hgene | Tgene | Gene symbol | EIF2AK4 | PKM | Gene ID | 440275 | 5315 |
| Gene name | eukaryotic translation initiation factor 2 alpha kinase 4 | pyruvate kinase M1/2 | |
| Synonyms | GCN2|PVOD2 | CTHBP|HEL-S-30|OIP3|PK3|PKM2|TCB|THBP1|p58 | |
| Cytomap | 15q15.1 | 15q23 | |
| Type of gene | protein-coding | protein-coding | |
| Description | eIF-2-alpha kinase GCN2GCN2 eIF2alpha kinaseGCN2-like proteingeneral control nonderepressible 2 | pyruvate kinase PKMOPA-interacting protein 3PK, muscle typecytosolic thyroid hormone-binding proteinepididymis secretory protein Li 30pyruvate kinase 2/3pyruvate kinase isozymes M1/M2pyruvate kinase muscle isozymepyruvate kinase, musclethyroid ho | |
| Modification date | 20200313 | 20200329 | |
| UniProtAcc | Q9P2K8 | Q99640 | |
| Ensembl transtripts involved in fusion gene | ENST ids | ENST00000263791, ENST00000382727, ENST00000559311, ENST00000559624, ENST00000560648, | ENST00000449901, ENST00000565154, ENST00000568459, ENST00000568883, ENST00000319622, ENST00000335181, ENST00000389093, ENST00000565184, |
| Fusion gene scores for assessment (based on all fusion genes of FusionGDB 2.0) | * DoF score | 15 X 17 X 10=2550 | 34 X 37 X 9=11322 |
| # samples | 20 | 41 | |
| ** MAII score | log2(20/2550*10)=-3.6724253419715 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(41/11322*10)=-4.78736110881616 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context (manual curation of fusion genes in FusionPDB) | PubMed: EIF2AK4 [Title/Abstract] AND PKM [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint (based on all fusion genes of FusionGDB 2.0) | EIF2AK4(40301931)-PKM(72502819), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | EIF2AK4-PKM seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. EIF2AK4-PKM seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. EIF2AK4-PKM seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. EIF2AK4-PKM seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | PKM | GO:0012501 | programmed cell death | 17308100 |
 Fusion gene breakpoints across EIF2AK4 (5'-gene) Fusion gene breakpoints across EIF2AK4 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
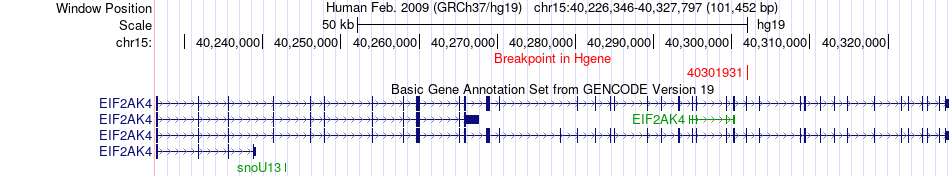 |
 Fusion gene breakpoints across PKM (3'-gene) Fusion gene breakpoints across PKM (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
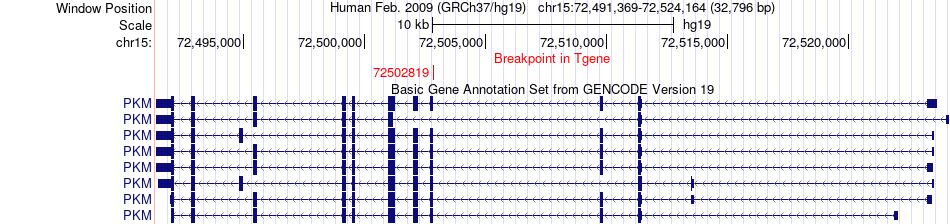 |
Top |
Fusion Gene Sample Information |
 Fusion gene information from FusionGDB2.0. Fusion gene information from FusionGDB2.0. |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | LGG | TCGA-DU-A7TC-01A | EIF2AK4 | chr15 | 40301931 | + | PKM | chr15 | 72502819 | - |
| ChimerDB4 | LGG | TCGA-DU-A7TC | EIF2AK4 | chr15 | 40301931 | + | PKM | chr15 | 72502819 | - |
Top |
Fusion ORF Analysis |
 Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000382727 | EIF2AK4 | chr15 | 40301931 | + | ENST00000319622 | PKM | chr15 | 72502819 | - | 5630 | 3659 | 50 | 5008 | 1652 |
| ENST00000382727 | EIF2AK4 | chr15 | 40301931 | + | ENST00000565184 | PKM | chr15 | 72502819 | - | 5627 | 3659 | 50 | 5008 | 1652 |
| ENST00000382727 | EIF2AK4 | chr15 | 40301931 | + | ENST00000389093 | PKM | chr15 | 72502819 | - | 5627 | 3659 | 50 | 5008 | 1652 |
| ENST00000382727 | EIF2AK4 | chr15 | 40301931 | + | ENST00000335181 | PKM | chr15 | 72502819 | - | 5627 | 3659 | 50 | 5008 | 1652 |
| ENST00000263791 | EIF2AK4 | chr15 | 40301931 | + | ENST00000319622 | PKM | chr15 | 72502819 | - | 5707 | 3736 | 43 | 5085 | 1680 |
| ENST00000263791 | EIF2AK4 | chr15 | 40301931 | + | ENST00000565184 | PKM | chr15 | 72502819 | - | 5704 | 3736 | 43 | 5085 | 1680 |
| ENST00000263791 | EIF2AK4 | chr15 | 40301931 | + | ENST00000389093 | PKM | chr15 | 72502819 | - | 5704 | 3736 | 43 | 5085 | 1680 |
| ENST00000263791 | EIF2AK4 | chr15 | 40301931 | + | ENST00000335181 | PKM | chr15 | 72502819 | - | 5704 | 3736 | 43 | 5085 | 1680 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000382727 | ENST00000319622 | EIF2AK4 | chr15 | 40301931 | + | PKM | chr15 | 72502819 | - | 0.001024933 | 0.99897504 |
| ENST00000382727 | ENST00000565184 | EIF2AK4 | chr15 | 40301931 | + | PKM | chr15 | 72502819 | - | 0.001014737 | 0.99898523 |
| ENST00000382727 | ENST00000389093 | EIF2AK4 | chr15 | 40301931 | + | PKM | chr15 | 72502819 | - | 0.001014737 | 0.99898523 |
| ENST00000382727 | ENST00000335181 | EIF2AK4 | chr15 | 40301931 | + | PKM | chr15 | 72502819 | - | 0.000975196 | 0.99902475 |
| ENST00000263791 | ENST00000319622 | EIF2AK4 | chr15 | 40301931 | + | PKM | chr15 | 72502819 | - | 0.00098113 | 0.9990189 |
| ENST00000263791 | ENST00000565184 | EIF2AK4 | chr15 | 40301931 | + | PKM | chr15 | 72502819 | - | 0.000970929 | 0.99902904 |
| ENST00000263791 | ENST00000389093 | EIF2AK4 | chr15 | 40301931 | + | PKM | chr15 | 72502819 | - | 0.000970929 | 0.99902904 |
| ENST00000263791 | ENST00000335181 | EIF2AK4 | chr15 | 40301931 | + | PKM | chr15 | 72502819 | - | 0.000927542 | 0.99907243 |
Top |
Fusion Amino Acid Sequences |
 For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >FusionGDB ID_FusionGDB isoform ID_FGname_Hgene_Hchr_Hbp_Henst_Tgene_Tchr_Tbp_Tenst_length(fusion AA) seq_BP >25682_25682_1_EIF2AK4-PKM_EIF2AK4_chr15_40301931_ENST00000263791_PKM_chr15_72502819_ENST00000319622_length(amino acids)=1680AA_BP=1231 MAGGRGAPGRGRDEPPESYPQRQDHELQALEAIYGADFQDLRPDACGPVKEPPEINLVLYPQGLTGEEVYVKVDLRVKCPPTYPDVVPEI ELKNAKGLSNESVNLLKSRLEELAKKHCGEVMIFELAYHVQSFLSEHNKPPPKSFHEEMLERRAQEEQQRLLEAKRKEEQEQREILHEIQ RRKEEIKEEKKRKEMAKQERLEIASLSNQDHTSKKDPGGHRTAAILHGGSPDFVGNGKHRANSSGRSRRERQYSVCNSEDSPGSCEILYF NMGSPDQLMVHKGKCIGSDEQLGKLVYNALETATGGFVLLYEWVLQWQKKMGPFLTSQEKEKIDKCKKQIQGTETEFNSLVKLSHPNVVR YLAMNLKEQDDSIVVDILVEHISGVSLAAHLSHSGPIPVHQLRRYTAQLLSGLDYLHSNSVVHKVLSASNVLVDAEGTVKITDYSISKRL ADICKEDVFEQTRVRFSDNALPYKTGKKGDVWRLGLLLLSLSQGQECGEYPVTIPSDLPADFQDFLKKCVCLDDKERWSPQQLLKHSFIN PQPKMPLVEQSPEDSEGQDYVETVIPSNRLPSAAFFSETQRQFSRYFIEFEELQLLGKGAFGAVIKVQNKLDGCCYAVKRIPINPASRQF RRIKGEVTLLSRLHHENIVRYYNAWIERHERPAGPGTPPPDSGPLAKDDRAARGQPASDTDGLDSVEAAAPPPILSSSVEWSTSGERSAS ARFPATGPGSSDDEDDDEDEHGGVFSQSFLPASDSESDIIFDNEDENSKSQNQDEDCNEKNGCHESEPSVTTEAVHYLYIQMEYCEKSTL RDTIDQGLYRDTVRLWRLFREILDGLAYIHEKGMIHRDLKPVNIFLDSDDHVKIGDFGLATDHLAFSADSKQDDQTGDLIKSDPSGHLTG MVGTALYVSPEVQGSTKSAYNQKVDLFSLGIIFFEMSYHPMVTASERIFVLNQLRDPTSPKFPEDFDDGEHAKQKSVISWLLNHDPAKRP TATELLKSELLPPPQMEESELHEVLHHTLTNVDGKAYRTMMAQIFSQRISPAIDYTYDSDILKGNFSIRTAKMQQHVCETIIRIFKRHGA VQLCTPLLLPRNRQIYEHNEAALFMDHSGMLVMLPFDLRIPFARYVARNNILNLKRYCIERVFRPRKLDRFHPKELLECAFDIVTSTTNS FLPTAEIIYTIYEIIQEFPALQERNYSIYLNHTMLLKAILLHCGIPEDKLSQVYIILYDAVYHAETIKNVRTATESFASDPILYRPVAVA LDTKGPEIRTGLIKGSGTAEVELKKGATLKITLDNAYMEKCDENILWLDYKNICKVVEVGSKIYVDDGLISLQVKQKGADFLVTEVENGG SLGSKKGVNLPGAAVDLPAVSEKDIQDLKFGVEQDVDMVFASFIRKASDVHEVRKVLGEKGKNIKIISKIENHEGVRRFDEILEASDGIM VARGDLGIEIPAEKVFLAQKMMIGRCNRAGKPVICATQMLESMIKKPRPTRAEGSDVANAVLDGADCIMLSGETAKGDYPLEAVRMQHLI AREAEAAMFHRKLFEELVRASSHSTDLMEAMAMGSVEASYKCLAAALIVLTESGRSAHQVARYRPRAPIIAVTRNPQTARQAHLYRGIFP -------------------------------------------------------------- >25682_25682_2_EIF2AK4-PKM_EIF2AK4_chr15_40301931_ENST00000263791_PKM_chr15_72502819_ENST00000335181_length(amino acids)=1680AA_BP=1231 MAGGRGAPGRGRDEPPESYPQRQDHELQALEAIYGADFQDLRPDACGPVKEPPEINLVLYPQGLTGEEVYVKVDLRVKCPPTYPDVVPEI ELKNAKGLSNESVNLLKSRLEELAKKHCGEVMIFELAYHVQSFLSEHNKPPPKSFHEEMLERRAQEEQQRLLEAKRKEEQEQREILHEIQ RRKEEIKEEKKRKEMAKQERLEIASLSNQDHTSKKDPGGHRTAAILHGGSPDFVGNGKHRANSSGRSRRERQYSVCNSEDSPGSCEILYF NMGSPDQLMVHKGKCIGSDEQLGKLVYNALETATGGFVLLYEWVLQWQKKMGPFLTSQEKEKIDKCKKQIQGTETEFNSLVKLSHPNVVR YLAMNLKEQDDSIVVDILVEHISGVSLAAHLSHSGPIPVHQLRRYTAQLLSGLDYLHSNSVVHKVLSASNVLVDAEGTVKITDYSISKRL ADICKEDVFEQTRVRFSDNALPYKTGKKGDVWRLGLLLLSLSQGQECGEYPVTIPSDLPADFQDFLKKCVCLDDKERWSPQQLLKHSFIN PQPKMPLVEQSPEDSEGQDYVETVIPSNRLPSAAFFSETQRQFSRYFIEFEELQLLGKGAFGAVIKVQNKLDGCCYAVKRIPINPASRQF RRIKGEVTLLSRLHHENIVRYYNAWIERHERPAGPGTPPPDSGPLAKDDRAARGQPASDTDGLDSVEAAAPPPILSSSVEWSTSGERSAS ARFPATGPGSSDDEDDDEDEHGGVFSQSFLPASDSESDIIFDNEDENSKSQNQDEDCNEKNGCHESEPSVTTEAVHYLYIQMEYCEKSTL RDTIDQGLYRDTVRLWRLFREILDGLAYIHEKGMIHRDLKPVNIFLDSDDHVKIGDFGLATDHLAFSADSKQDDQTGDLIKSDPSGHLTG MVGTALYVSPEVQGSTKSAYNQKVDLFSLGIIFFEMSYHPMVTASERIFVLNQLRDPTSPKFPEDFDDGEHAKQKSVISWLLNHDPAKRP TATELLKSELLPPPQMEESELHEVLHHTLTNVDGKAYRTMMAQIFSQRISPAIDYTYDSDILKGNFSIRTAKMQQHVCETIIRIFKRHGA VQLCTPLLLPRNRQIYEHNEAALFMDHSGMLVMLPFDLRIPFARYVARNNILNLKRYCIERVFRPRKLDRFHPKELLECAFDIVTSTTNS FLPTAEIIYTIYEIIQEFPALQERNYSIYLNHTMLLKAILLHCGIPEDKLSQVYIILYDAVYHAETIKNVRTATESFASDPILYRPVAVA LDTKGPEIRTGLIKGSGTAEVELKKGATLKITLDNAYMEKCDENILWLDYKNICKVVEVGSKIYVDDGLISLQVKQKGADFLVTEVENGG SLGSKKGVNLPGAAVDLPAVSEKDIQDLKFGVEQDVDMVFASFIRKASDVHEVRKVLGEKGKNIKIISKIENHEGVRRFDEILEASDGIM VARGDLGIEIPAEKVFLAQKMMIGRCNRAGKPVICATQMLESMIKKPRPTRAEGSDVANAVLDGADCIMLSGETAKGDYPLEAVRMQHLI AREAEAAIYHLQLFEELRRLAPITSDPTEATAVGAVEASFKCCSGAIIVLTKSGRSAHQVARYRPRAPIIAVTRNPQTARQAHLYRGIFP -------------------------------------------------------------- >25682_25682_3_EIF2AK4-PKM_EIF2AK4_chr15_40301931_ENST00000263791_PKM_chr15_72502819_ENST00000389093_length(amino acids)=1680AA_BP=1231 MAGGRGAPGRGRDEPPESYPQRQDHELQALEAIYGADFQDLRPDACGPVKEPPEINLVLYPQGLTGEEVYVKVDLRVKCPPTYPDVVPEI ELKNAKGLSNESVNLLKSRLEELAKKHCGEVMIFELAYHVQSFLSEHNKPPPKSFHEEMLERRAQEEQQRLLEAKRKEEQEQREILHEIQ RRKEEIKEEKKRKEMAKQERLEIASLSNQDHTSKKDPGGHRTAAILHGGSPDFVGNGKHRANSSGRSRRERQYSVCNSEDSPGSCEILYF NMGSPDQLMVHKGKCIGSDEQLGKLVYNALETATGGFVLLYEWVLQWQKKMGPFLTSQEKEKIDKCKKQIQGTETEFNSLVKLSHPNVVR YLAMNLKEQDDSIVVDILVEHISGVSLAAHLSHSGPIPVHQLRRYTAQLLSGLDYLHSNSVVHKVLSASNVLVDAEGTVKITDYSISKRL ADICKEDVFEQTRVRFSDNALPYKTGKKGDVWRLGLLLLSLSQGQECGEYPVTIPSDLPADFQDFLKKCVCLDDKERWSPQQLLKHSFIN PQPKMPLVEQSPEDSEGQDYVETVIPSNRLPSAAFFSETQRQFSRYFIEFEELQLLGKGAFGAVIKVQNKLDGCCYAVKRIPINPASRQF RRIKGEVTLLSRLHHENIVRYYNAWIERHERPAGPGTPPPDSGPLAKDDRAARGQPASDTDGLDSVEAAAPPPILSSSVEWSTSGERSAS ARFPATGPGSSDDEDDDEDEHGGVFSQSFLPASDSESDIIFDNEDENSKSQNQDEDCNEKNGCHESEPSVTTEAVHYLYIQMEYCEKSTL RDTIDQGLYRDTVRLWRLFREILDGLAYIHEKGMIHRDLKPVNIFLDSDDHVKIGDFGLATDHLAFSADSKQDDQTGDLIKSDPSGHLTG MVGTALYVSPEVQGSTKSAYNQKVDLFSLGIIFFEMSYHPMVTASERIFVLNQLRDPTSPKFPEDFDDGEHAKQKSVISWLLNHDPAKRP TATELLKSELLPPPQMEESELHEVLHHTLTNVDGKAYRTMMAQIFSQRISPAIDYTYDSDILKGNFSIRTAKMQQHVCETIIRIFKRHGA VQLCTPLLLPRNRQIYEHNEAALFMDHSGMLVMLPFDLRIPFARYVARNNILNLKRYCIERVFRPRKLDRFHPKELLECAFDIVTSTTNS FLPTAEIIYTIYEIIQEFPALQERNYSIYLNHTMLLKAILLHCGIPEDKLSQVYIILYDAVYHAETIKNVRTATESFASDPILYRPVAVA LDTKGPEIRTGLIKGSGTAEVELKKGATLKITLDNAYMEKCDENILWLDYKNICKVVEVGSKIYVDDGLISLQVKQKGADFLVTEVENGG SLGSKKGVNLPGAAVDLPAVSEKDIQDLKFGVEQDVDMVFASFIRKASDVHEVRKVLGEKGKNIKIISKIENHEGVRRFDEILEASDGIM VARGDLGIEIPAEKVFLAQKMMIGRCNRAGKPVICATQMLESMIKKPRPTRAEGSDVANAVLDGADCIMLSGETAKGDYPLEAVRMQHLI AREAEAAMFHRKLFEELVRASSHSTDLMEAMAMGSVEASYKCLAAALIVLTESGRSAHQVARYRPRAPIIAVTRNPQTARQAHLYRGIFP -------------------------------------------------------------- >25682_25682_4_EIF2AK4-PKM_EIF2AK4_chr15_40301931_ENST00000263791_PKM_chr15_72502819_ENST00000565184_length(amino acids)=1680AA_BP=1231 MAGGRGAPGRGRDEPPESYPQRQDHELQALEAIYGADFQDLRPDACGPVKEPPEINLVLYPQGLTGEEVYVKVDLRVKCPPTYPDVVPEI ELKNAKGLSNESVNLLKSRLEELAKKHCGEVMIFELAYHVQSFLSEHNKPPPKSFHEEMLERRAQEEQQRLLEAKRKEEQEQREILHEIQ RRKEEIKEEKKRKEMAKQERLEIASLSNQDHTSKKDPGGHRTAAILHGGSPDFVGNGKHRANSSGRSRRERQYSVCNSEDSPGSCEILYF NMGSPDQLMVHKGKCIGSDEQLGKLVYNALETATGGFVLLYEWVLQWQKKMGPFLTSQEKEKIDKCKKQIQGTETEFNSLVKLSHPNVVR YLAMNLKEQDDSIVVDILVEHISGVSLAAHLSHSGPIPVHQLRRYTAQLLSGLDYLHSNSVVHKVLSASNVLVDAEGTVKITDYSISKRL ADICKEDVFEQTRVRFSDNALPYKTGKKGDVWRLGLLLLSLSQGQECGEYPVTIPSDLPADFQDFLKKCVCLDDKERWSPQQLLKHSFIN PQPKMPLVEQSPEDSEGQDYVETVIPSNRLPSAAFFSETQRQFSRYFIEFEELQLLGKGAFGAVIKVQNKLDGCCYAVKRIPINPASRQF RRIKGEVTLLSRLHHENIVRYYNAWIERHERPAGPGTPPPDSGPLAKDDRAARGQPASDTDGLDSVEAAAPPPILSSSVEWSTSGERSAS ARFPATGPGSSDDEDDDEDEHGGVFSQSFLPASDSESDIIFDNEDENSKSQNQDEDCNEKNGCHESEPSVTTEAVHYLYIQMEYCEKSTL RDTIDQGLYRDTVRLWRLFREILDGLAYIHEKGMIHRDLKPVNIFLDSDDHVKIGDFGLATDHLAFSADSKQDDQTGDLIKSDPSGHLTG MVGTALYVSPEVQGSTKSAYNQKVDLFSLGIIFFEMSYHPMVTASERIFVLNQLRDPTSPKFPEDFDDGEHAKQKSVISWLLNHDPAKRP TATELLKSELLPPPQMEESELHEVLHHTLTNVDGKAYRTMMAQIFSQRISPAIDYTYDSDILKGNFSIRTAKMQQHVCETIIRIFKRHGA VQLCTPLLLPRNRQIYEHNEAALFMDHSGMLVMLPFDLRIPFARYVARNNILNLKRYCIERVFRPRKLDRFHPKELLECAFDIVTSTTNS FLPTAEIIYTIYEIIQEFPALQERNYSIYLNHTMLLKAILLHCGIPEDKLSQVYIILYDAVYHAETIKNVRTATESFASDPILYRPVAVA LDTKGPEIRTGLIKGSGTAEVELKKGATLKITLDNAYMEKCDENILWLDYKNICKVVEVGSKIYVDDGLISLQVKQKGADFLVTEVENGG SLGSKKGVNLPGAAVDLPAVSEKDIQDLKFGVEQDVDMVFASFIRKASDVHEVRKVLGEKGKNIKIISKIENHEGVRRFDEILEASDGIM VARGDLGIEIPAEKVFLAQKMMIGRCNRAGKPVICATQMLESMIKKPRPTRAEGSDVANAVLDGADCIMLSGETAKGDYPLEAVRMQHLI AREAEAAMFHRKLFEELVRASSHSTDLMEAMAMGSVEASYKCLAAALIVLTESGRSAHQVARYRPRAPIIAVTRNPQTARQAHLYRGIFP -------------------------------------------------------------- >25682_25682_5_EIF2AK4-PKM_EIF2AK4_chr15_40301931_ENST00000382727_PKM_chr15_72502819_ENST00000319622_length(amino acids)=1652AA_BP=1203 MAGGRGAPGRGRDEPPESYPQRQDHELQALEAIYGADFQDLRPDACGPVKEPPEINLVLYPQGLTGEEVYVKVDLRVKCPPTYPDVVPEI ELKNAKGLSNESVNLLKSRLEELAKKHCGEVMIFELAYHVQSFLSEHNKPPPKSFHEEMLERRAQEEQQRLLEAKRKEEQEQREILHEIQ RRKEEIKEEKKRKEMAKQERLEIASLSNQDHTSKKDPGGHRTAAILHGGSPDFVGNGKHRANSSGRSRRERQYSVCNSEDSPGSCEILYF NMGSPDQLMVHKGKCIGSDEQLGKLVYNALETATGGFVLLYEWVLQWQKKMGPFLTSQEKEKIDKCKKQIQGTETEFNSLVKLSHPNVVR YLAMNLKEQDDSIVVDILVEHISGVSLAAHLSHSGPIPVHQLRRYTAQLLSGLDYLHSNSVVHKVLSASNVLVDAEGTVKITDYSISKRL ADICKEDVFEQTRVRFSDNALPYKTGKKGDVWRLGLLLLSLSQGQECGEYPVTIPSDLPADFQDFLKKCVCLDDKERWSPQQLLKHSFIN PQPKMPLVEQSPEDSEGQDYVETVIPSNRLPSAAFFSETQRQFSRYFIEFEELQLLGKGAFGAVIKVQNKLDGCCYAVKRIPINPASRQF RRIKGEVTLLSRLHHENIVRYYNAWIERHERPAGPGTPPPDSGPLAKDDRAARGQPASDTDGLDSVEAAAPPPILSSSVEWSTSGERSAS ARFPATGPGSSDDEDDDEDEHGGVFSQSFLPASDSESDIIFDNEDENSKSQNQMEYCEKSTLRDTIDQGLYRDTVRLWRLFREILDGLAY IHEKGMIHRDLKPVNIFLDSDDHVKIGDFGLATDHLAFSADSKQDDQTGDLIKSDPSGHLTGMVGTALYVSPEVQGSTKSAYNQKVDLFS LGIIFFEMSYHPMVTASERIFVLNQLRDPTSPKFPEDFDDGEHAKQKSVISWLLNHDPAKRPTATELLKSELLPPPQMEESELHEVLHHT LTNVDGKAYRTMMAQIFSQRISPAIDYTYDSDILKGNFSIRTAKMQQHVCETIIRIFKRHGAVQLCTPLLLPRNRQIYEHNEAALFMDHS GMLVMLPFDLRIPFARYVARNNILNLKRYCIERVFRPRKLDRFHPKELLECAFDIVTSTTNSFLPTAEIIYTIYEIIQEFPALQERNYSI YLNHTMLLKAILLHCGIPEDKLSQVYIILYDAVYHAETIKNVRTATESFASDPILYRPVAVALDTKGPEIRTGLIKGSGTAEVELKKGAT LKITLDNAYMEKCDENILWLDYKNICKVVEVGSKIYVDDGLISLQVKQKGADFLVTEVENGGSLGSKKGVNLPGAAVDLPAVSEKDIQDL KFGVEQDVDMVFASFIRKASDVHEVRKVLGEKGKNIKIISKIENHEGVRRFDEILEASDGIMVARGDLGIEIPAEKVFLAQKMMIGRCNR AGKPVICATQMLESMIKKPRPTRAEGSDVANAVLDGADCIMLSGETAKGDYPLEAVRMQHLIAREAEAAMFHRKLFEELVRASSHSTDLM EAMAMGSVEASYKCLAAALIVLTESGRSAHQVARYRPRAPIIAVTRNPQTARQAHLYRGIFPVLCKDPVQEAWAEDVDLRVNFAMNVGKA -------------------------------------------------------------- >25682_25682_6_EIF2AK4-PKM_EIF2AK4_chr15_40301931_ENST00000382727_PKM_chr15_72502819_ENST00000335181_length(amino acids)=1652AA_BP=1203 MAGGRGAPGRGRDEPPESYPQRQDHELQALEAIYGADFQDLRPDACGPVKEPPEINLVLYPQGLTGEEVYVKVDLRVKCPPTYPDVVPEI ELKNAKGLSNESVNLLKSRLEELAKKHCGEVMIFELAYHVQSFLSEHNKPPPKSFHEEMLERRAQEEQQRLLEAKRKEEQEQREILHEIQ RRKEEIKEEKKRKEMAKQERLEIASLSNQDHTSKKDPGGHRTAAILHGGSPDFVGNGKHRANSSGRSRRERQYSVCNSEDSPGSCEILYF NMGSPDQLMVHKGKCIGSDEQLGKLVYNALETATGGFVLLYEWVLQWQKKMGPFLTSQEKEKIDKCKKQIQGTETEFNSLVKLSHPNVVR YLAMNLKEQDDSIVVDILVEHISGVSLAAHLSHSGPIPVHQLRRYTAQLLSGLDYLHSNSVVHKVLSASNVLVDAEGTVKITDYSISKRL ADICKEDVFEQTRVRFSDNALPYKTGKKGDVWRLGLLLLSLSQGQECGEYPVTIPSDLPADFQDFLKKCVCLDDKERWSPQQLLKHSFIN PQPKMPLVEQSPEDSEGQDYVETVIPSNRLPSAAFFSETQRQFSRYFIEFEELQLLGKGAFGAVIKVQNKLDGCCYAVKRIPINPASRQF RRIKGEVTLLSRLHHENIVRYYNAWIERHERPAGPGTPPPDSGPLAKDDRAARGQPASDTDGLDSVEAAAPPPILSSSVEWSTSGERSAS ARFPATGPGSSDDEDDDEDEHGGVFSQSFLPASDSESDIIFDNEDENSKSQNQMEYCEKSTLRDTIDQGLYRDTVRLWRLFREILDGLAY IHEKGMIHRDLKPVNIFLDSDDHVKIGDFGLATDHLAFSADSKQDDQTGDLIKSDPSGHLTGMVGTALYVSPEVQGSTKSAYNQKVDLFS LGIIFFEMSYHPMVTASERIFVLNQLRDPTSPKFPEDFDDGEHAKQKSVISWLLNHDPAKRPTATELLKSELLPPPQMEESELHEVLHHT LTNVDGKAYRTMMAQIFSQRISPAIDYTYDSDILKGNFSIRTAKMQQHVCETIIRIFKRHGAVQLCTPLLLPRNRQIYEHNEAALFMDHS GMLVMLPFDLRIPFARYVARNNILNLKRYCIERVFRPRKLDRFHPKELLECAFDIVTSTTNSFLPTAEIIYTIYEIIQEFPALQERNYSI YLNHTMLLKAILLHCGIPEDKLSQVYIILYDAVYHAETIKNVRTATESFASDPILYRPVAVALDTKGPEIRTGLIKGSGTAEVELKKGAT LKITLDNAYMEKCDENILWLDYKNICKVVEVGSKIYVDDGLISLQVKQKGADFLVTEVENGGSLGSKKGVNLPGAAVDLPAVSEKDIQDL KFGVEQDVDMVFASFIRKASDVHEVRKVLGEKGKNIKIISKIENHEGVRRFDEILEASDGIMVARGDLGIEIPAEKVFLAQKMMIGRCNR AGKPVICATQMLESMIKKPRPTRAEGSDVANAVLDGADCIMLSGETAKGDYPLEAVRMQHLIAREAEAAIYHLQLFEELRRLAPITSDPT EATAVGAVEASFKCCSGAIIVLTKSGRSAHQVARYRPRAPIIAVTRNPQTARQAHLYRGIFPVLCKDPVQEAWAEDVDLRVNFAMNVGKA -------------------------------------------------------------- >25682_25682_7_EIF2AK4-PKM_EIF2AK4_chr15_40301931_ENST00000382727_PKM_chr15_72502819_ENST00000389093_length(amino acids)=1652AA_BP=1203 MAGGRGAPGRGRDEPPESYPQRQDHELQALEAIYGADFQDLRPDACGPVKEPPEINLVLYPQGLTGEEVYVKVDLRVKCPPTYPDVVPEI ELKNAKGLSNESVNLLKSRLEELAKKHCGEVMIFELAYHVQSFLSEHNKPPPKSFHEEMLERRAQEEQQRLLEAKRKEEQEQREILHEIQ RRKEEIKEEKKRKEMAKQERLEIASLSNQDHTSKKDPGGHRTAAILHGGSPDFVGNGKHRANSSGRSRRERQYSVCNSEDSPGSCEILYF NMGSPDQLMVHKGKCIGSDEQLGKLVYNALETATGGFVLLYEWVLQWQKKMGPFLTSQEKEKIDKCKKQIQGTETEFNSLVKLSHPNVVR YLAMNLKEQDDSIVVDILVEHISGVSLAAHLSHSGPIPVHQLRRYTAQLLSGLDYLHSNSVVHKVLSASNVLVDAEGTVKITDYSISKRL ADICKEDVFEQTRVRFSDNALPYKTGKKGDVWRLGLLLLSLSQGQECGEYPVTIPSDLPADFQDFLKKCVCLDDKERWSPQQLLKHSFIN PQPKMPLVEQSPEDSEGQDYVETVIPSNRLPSAAFFSETQRQFSRYFIEFEELQLLGKGAFGAVIKVQNKLDGCCYAVKRIPINPASRQF RRIKGEVTLLSRLHHENIVRYYNAWIERHERPAGPGTPPPDSGPLAKDDRAARGQPASDTDGLDSVEAAAPPPILSSSVEWSTSGERSAS ARFPATGPGSSDDEDDDEDEHGGVFSQSFLPASDSESDIIFDNEDENSKSQNQMEYCEKSTLRDTIDQGLYRDTVRLWRLFREILDGLAY IHEKGMIHRDLKPVNIFLDSDDHVKIGDFGLATDHLAFSADSKQDDQTGDLIKSDPSGHLTGMVGTALYVSPEVQGSTKSAYNQKVDLFS LGIIFFEMSYHPMVTASERIFVLNQLRDPTSPKFPEDFDDGEHAKQKSVISWLLNHDPAKRPTATELLKSELLPPPQMEESELHEVLHHT LTNVDGKAYRTMMAQIFSQRISPAIDYTYDSDILKGNFSIRTAKMQQHVCETIIRIFKRHGAVQLCTPLLLPRNRQIYEHNEAALFMDHS GMLVMLPFDLRIPFARYVARNNILNLKRYCIERVFRPRKLDRFHPKELLECAFDIVTSTTNSFLPTAEIIYTIYEIIQEFPALQERNYSI YLNHTMLLKAILLHCGIPEDKLSQVYIILYDAVYHAETIKNVRTATESFASDPILYRPVAVALDTKGPEIRTGLIKGSGTAEVELKKGAT LKITLDNAYMEKCDENILWLDYKNICKVVEVGSKIYVDDGLISLQVKQKGADFLVTEVENGGSLGSKKGVNLPGAAVDLPAVSEKDIQDL KFGVEQDVDMVFASFIRKASDVHEVRKVLGEKGKNIKIISKIENHEGVRRFDEILEASDGIMVARGDLGIEIPAEKVFLAQKMMIGRCNR AGKPVICATQMLESMIKKPRPTRAEGSDVANAVLDGADCIMLSGETAKGDYPLEAVRMQHLIAREAEAAMFHRKLFEELVRASSHSTDLM EAMAMGSVEASYKCLAAALIVLTESGRSAHQVARYRPRAPIIAVTRNPQTARQAHLYRGIFPVLCKDPVQEAWAEDVDLRVNFAMNVGKA -------------------------------------------------------------- >25682_25682_8_EIF2AK4-PKM_EIF2AK4_chr15_40301931_ENST00000382727_PKM_chr15_72502819_ENST00000565184_length(amino acids)=1652AA_BP=1203 MAGGRGAPGRGRDEPPESYPQRQDHELQALEAIYGADFQDLRPDACGPVKEPPEINLVLYPQGLTGEEVYVKVDLRVKCPPTYPDVVPEI ELKNAKGLSNESVNLLKSRLEELAKKHCGEVMIFELAYHVQSFLSEHNKPPPKSFHEEMLERRAQEEQQRLLEAKRKEEQEQREILHEIQ RRKEEIKEEKKRKEMAKQERLEIASLSNQDHTSKKDPGGHRTAAILHGGSPDFVGNGKHRANSSGRSRRERQYSVCNSEDSPGSCEILYF NMGSPDQLMVHKGKCIGSDEQLGKLVYNALETATGGFVLLYEWVLQWQKKMGPFLTSQEKEKIDKCKKQIQGTETEFNSLVKLSHPNVVR YLAMNLKEQDDSIVVDILVEHISGVSLAAHLSHSGPIPVHQLRRYTAQLLSGLDYLHSNSVVHKVLSASNVLVDAEGTVKITDYSISKRL ADICKEDVFEQTRVRFSDNALPYKTGKKGDVWRLGLLLLSLSQGQECGEYPVTIPSDLPADFQDFLKKCVCLDDKERWSPQQLLKHSFIN PQPKMPLVEQSPEDSEGQDYVETVIPSNRLPSAAFFSETQRQFSRYFIEFEELQLLGKGAFGAVIKVQNKLDGCCYAVKRIPINPASRQF RRIKGEVTLLSRLHHENIVRYYNAWIERHERPAGPGTPPPDSGPLAKDDRAARGQPASDTDGLDSVEAAAPPPILSSSVEWSTSGERSAS ARFPATGPGSSDDEDDDEDEHGGVFSQSFLPASDSESDIIFDNEDENSKSQNQMEYCEKSTLRDTIDQGLYRDTVRLWRLFREILDGLAY IHEKGMIHRDLKPVNIFLDSDDHVKIGDFGLATDHLAFSADSKQDDQTGDLIKSDPSGHLTGMVGTALYVSPEVQGSTKSAYNQKVDLFS LGIIFFEMSYHPMVTASERIFVLNQLRDPTSPKFPEDFDDGEHAKQKSVISWLLNHDPAKRPTATELLKSELLPPPQMEESELHEVLHHT LTNVDGKAYRTMMAQIFSQRISPAIDYTYDSDILKGNFSIRTAKMQQHVCETIIRIFKRHGAVQLCTPLLLPRNRQIYEHNEAALFMDHS GMLVMLPFDLRIPFARYVARNNILNLKRYCIERVFRPRKLDRFHPKELLECAFDIVTSTTNSFLPTAEIIYTIYEIIQEFPALQERNYSI YLNHTMLLKAILLHCGIPEDKLSQVYIILYDAVYHAETIKNVRTATESFASDPILYRPVAVALDTKGPEIRTGLIKGSGTAEVELKKGAT LKITLDNAYMEKCDENILWLDYKNICKVVEVGSKIYVDDGLISLQVKQKGADFLVTEVENGGSLGSKKGVNLPGAAVDLPAVSEKDIQDL KFGVEQDVDMVFASFIRKASDVHEVRKVLGEKGKNIKIISKIENHEGVRRFDEILEASDGIMVARGDLGIEIPAEKVFLAQKMMIGRCNR AGKPVICATQMLESMIKKPRPTRAEGSDVANAVLDGADCIMLSGETAKGDYPLEAVRMQHLIAREAEAAMFHRKLFEELVRASSHSTDLM EAMAMGSVEASYKCLAAALIVLTESGRSAHQVARYRPRAPIIAVTRNPQTARQAHLYRGIFPVLCKDPVQEAWAEDVDLRVNFAMNVGKA -------------------------------------------------------------- |
Top |
Fusion Protein Functional Features |
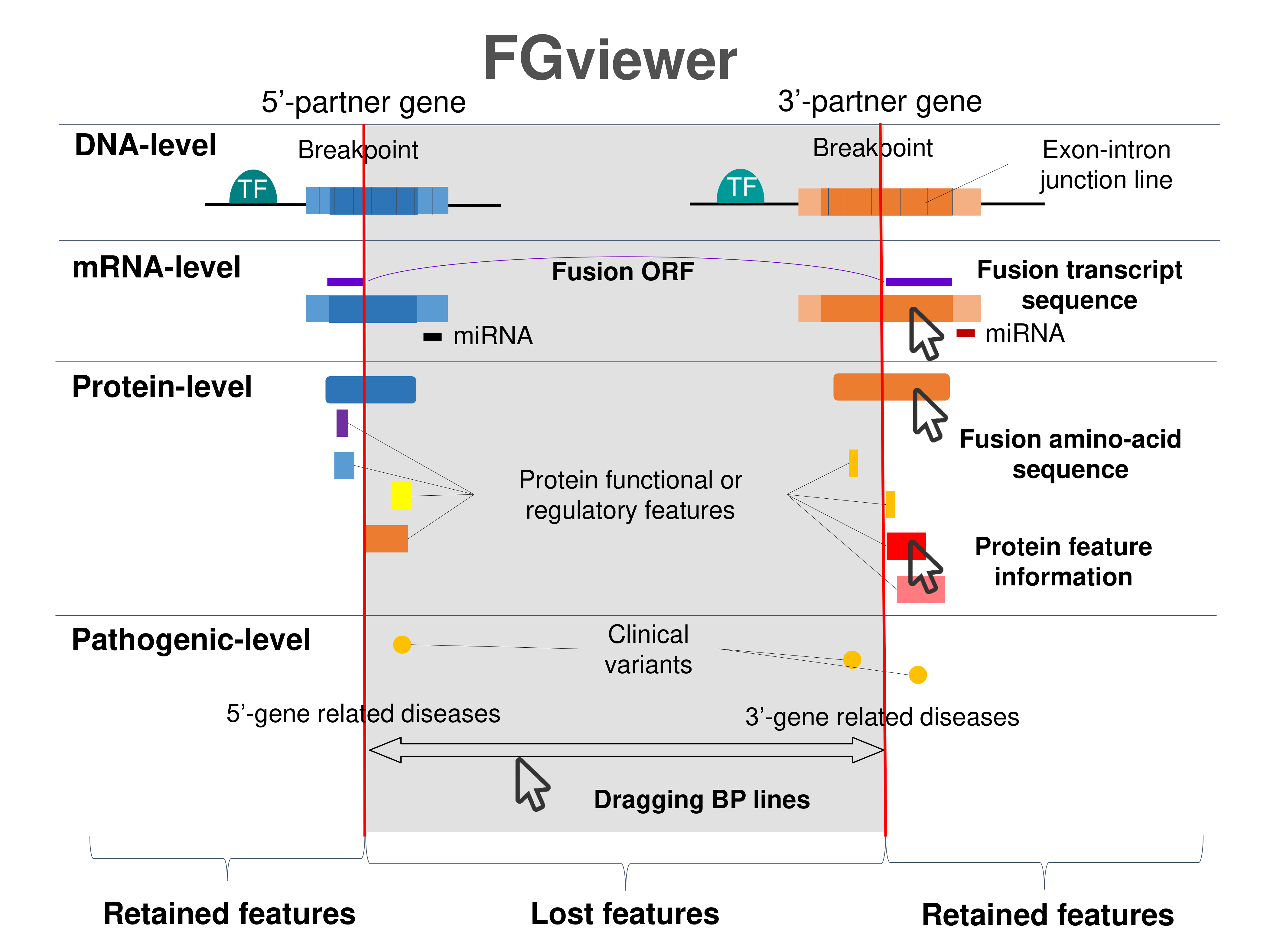
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr15:40301931/chr15:72502819) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| EIF2AK4 | PKM |
| FUNCTION: Metabolic-stress sensing protein kinase that phosphorylates the alpha subunit of eukaryotic translation initiation factor 2 (EIF2S1/eIF-2-alpha) in response to low amino acid availability (PubMed:25329545). Plays a role as an activator of the integrated stress response (ISR) required for adaptation to amino acid starvation (By similarity). EIF2S1/eIF-2-alpha phosphorylation in response to stress converts EIF2S1/eIF-2-alpha in a global protein synthesis inhibitor, leading to a global attenuation of cap-dependent translation, and thus to a reduced overall utilization of amino acids, while concomitantly initiating the preferential translation of ISR-specific mRNAs, such as the transcriptional activator ATF4, and hence allowing ATF4-mediated reprogramming of amino acid biosynthetic gene expression to alleviate nutrient depletion (By similarity). Binds uncharged tRNAs (By similarity). Involved in cell cycle arrest by promoting cyclin D1 mRNA translation repression after the unfolded protein response pathway (UPR) activation or cell cycle inhibitor CDKN1A/p21 mRNA translation activation in response to amino acid deprivation (PubMed:26102367). Plays a role in the consolidation of synaptic plasticity, learning as well as formation of long-term memory (By similarity). Plays a role in neurite outgrowth inhibition (By similarity). Plays a proapoptotic role in response to glucose deprivation (By similarity). Promotes global cellular protein synthesis repression in response to UV irradiation independently of the stress-activated protein kinase/c-Jun N-terminal kinase (SAPK/JNK) and p38 MAPK signaling pathways (By similarity). Plays a role in the antiviral response against alphavirus infection; impairs early viral mRNA translation of the incoming genomic virus RNA, thus preventing alphavirus replication (By similarity). {ECO:0000250|UniProtKB:P15442, ECO:0000250|UniProtKB:Q9QZ05, ECO:0000269|PubMed:25329545, ECO:0000269|PubMed:26102367}.; FUNCTION: (Microbial infection) Plays a role in modulating the adaptive immune response to yellow fever virus infection; promotes dendritic cells to initiate autophagy and antigene presentation to both CD4(+) and CD8(+) T-cells under amino acid starvation (PubMed:24310610). {ECO:0000269|PubMed:24310610}. | FUNCTION: Acts as a negative regulator of entry into mitosis (G2 to M transition) by phosphorylation of the CDK1 kinase specifically when CDK1 is complexed to cyclins. Mediates phosphorylation of CDK1 predominantly on 'Thr-14'. Also involved in Golgi fragmentation. May be involved in phosphorylation of CDK1 on 'Tyr-15' to a lesser degree, however tyrosine kinase activity is unclear and may be indirect. May be a downstream target of Notch signaling pathway during eye development. {ECO:0000269|PubMed:10373560, ECO:0000269|PubMed:9001210}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - Retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000263791 | + | 26 | 39 | 146_205 | 1231.0 | 1650.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000382727 | + | 25 | 38 | 146_205 | 1203.0 | 1622.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000263791 | + | 26 | 39 | 484_491 | 1231.0 | 1650.0 | Compositional bias | Note=Poly-Leu |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000382727 | + | 25 | 38 | 484_491 | 1203.0 | 1622.0 | Compositional bias | Note=Poly-Leu |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000263791 | + | 26 | 39 | 25_137 | 1231.0 | 1650.0 | Domain | RWD |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000263791 | + | 26 | 39 | 296_539 | 1231.0 | 1650.0 | Domain | Protein kinase 1 |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000263791 | + | 26 | 39 | 590_1001 | 1231.0 | 1650.0 | Domain | Protein kinase 2 |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000382727 | + | 25 | 38 | 25_137 | 1203.0 | 1622.0 | Domain | RWD |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000382727 | + | 25 | 38 | 296_539 | 1203.0 | 1622.0 | Domain | Protein kinase 1 |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000382727 | + | 25 | 38 | 590_1001 | 1203.0 | 1622.0 | Domain | Protein kinase 2 |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000263791 | + | 26 | 39 | 596_604 | 1231.0 | 1650.0 | Nucleotide binding | ATP |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000382727 | + | 25 | 38 | 596_604 | 1203.0 | 1622.0 | Nucleotide binding | ATP |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000449901 | 2 | 11 | 75_78 | 67.0 | 517.0 | Nucleotide binding | ATP | |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000319622 | 2 | 11 | 432_437 | 82.0 | 532.0 | Region | D-fructose 1%2C6-bisphosphate binding%3B part of allosteric site | |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000319622 | 2 | 11 | 516_521 | 82.0 | 532.0 | Region | D-fructose 1%2C6-bisphosphate binding%3B part of allosteric site | |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000335181 | 2 | 11 | 432_437 | 82.0 | 532.0 | Region | D-fructose 1%2C6-bisphosphate binding%3B part of allosteric site | |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000335181 | 2 | 11 | 516_521 | 82.0 | 532.0 | Region | D-fructose 1%2C6-bisphosphate binding%3B part of allosteric site | |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000389093 | 2 | 11 | 432_437 | 82.0 | 532.0 | Region | D-fructose 1%2C6-bisphosphate binding%3B part of allosteric site | |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000389093 | 2 | 11 | 516_521 | 82.0 | 532.0 | Region | D-fructose 1%2C6-bisphosphate binding%3B part of allosteric site | |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000449901 | 2 | 11 | 432_437 | 67.0 | 517.0 | Region | D-fructose 1%2C6-bisphosphate binding%3B part of allosteric site | |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000449901 | 2 | 11 | 516_521 | 67.0 | 517.0 | Region | D-fructose 1%2C6-bisphosphate binding%3B part of allosteric site | |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000565154 | 3 | 12 | 432_437 | 82.0 | 532.0 | Region | D-fructose 1%2C6-bisphosphate binding%3B part of allosteric site | |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000565154 | 3 | 12 | 516_521 | 82.0 | 532.0 | Region | D-fructose 1%2C6-bisphosphate binding%3B part of allosteric site | |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000565184 | 2 | 11 | 432_437 | 82.0 | 532.0 | Region | D-fructose 1%2C6-bisphosphate binding%3B part of allosteric site | |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000565184 | 2 | 11 | 516_521 | 82.0 | 532.0 | Region | D-fructose 1%2C6-bisphosphate binding%3B part of allosteric site | |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000568459 | 2 | 11 | 432_437 | 82.0 | 532.0 | Region | D-fructose 1%2C6-bisphosphate binding%3B part of allosteric site | |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000568459 | 2 | 11 | 516_521 | 82.0 | 532.0 | Region | D-fructose 1%2C6-bisphosphate binding%3B part of allosteric site |
| - Not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000559624 | + | 1 | 11 | 146_205 | 0 | 617.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000559624 | + | 1 | 11 | 484_491 | 0 | 617.0 | Compositional bias | Note=Poly-Leu |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000559624 | + | 1 | 11 | 25_137 | 0 | 617.0 | Domain | RWD |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000559624 | + | 1 | 11 | 296_539 | 0 | 617.0 | Domain | Protein kinase 1 |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000559624 | + | 1 | 11 | 590_1001 | 0 | 617.0 | Domain | Protein kinase 2 |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000559624 | + | 1 | 11 | 596_604 | 0 | 617.0 | Nucleotide binding | ATP |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000263791 | + | 26 | 39 | 1022_1493 | 1231.0 | 1650.0 | Region | Note=Histidyl-tRNA synthetase-like |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000382727 | + | 25 | 38 | 1022_1493 | 1203.0 | 1622.0 | Region | Note=Histidyl-tRNA synthetase-like |
| Hgene | EIF2AK4 | chr15:40301931 | chr15:72502819 | ENST00000559624 | + | 1 | 11 | 1022_1493 | 0 | 617.0 | Region | Note=Histidyl-tRNA synthetase-like |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000319622 | 2 | 11 | 75_78 | 82.0 | 532.0 | Nucleotide binding | ATP | |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000335181 | 2 | 11 | 75_78 | 82.0 | 532.0 | Nucleotide binding | ATP | |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000389093 | 2 | 11 | 75_78 | 82.0 | 532.0 | Nucleotide binding | ATP | |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000565154 | 3 | 12 | 75_78 | 82.0 | 532.0 | Nucleotide binding | ATP | |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000565184 | 2 | 11 | 75_78 | 82.0 | 532.0 | Nucleotide binding | ATP | |
| Tgene | PKM | chr15:40301931 | chr15:72502819 | ENST00000568459 | 2 | 11 | 75_78 | 82.0 | 532.0 | Nucleotide binding | ATP |
Top |
Fusion Protein Structures |
 PDB and CIF files of the predicted fusion proteins PDB and CIF files of the predicted fusion proteins * Here we show the 3D structure of the fusion proteins using Mol*. AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. Model confidence is shown from the pLDDT values per residue. pLDDT corresponds to the model’s prediction of its score on the local Distance Difference Test. It is a measure of local accuracy (from AlphfaFold website). To color code individual residues, we transformed individual PDB files into CIF format. |
| Fusion protein PDB link (fusion AA seq ID in FusionPDB) | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | AA seq | Len(AA seq) |
Top |
pLDDT score distribution |
 pLDDT score distribution of the predicted wild-type structures of two partner proteins from AlphaFold2 pLDDT score distribution of the predicted wild-type structures of two partner proteins from AlphaFold2* AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. |
EIF2AK4_pLDDT.png |
PKM_pLDDT.png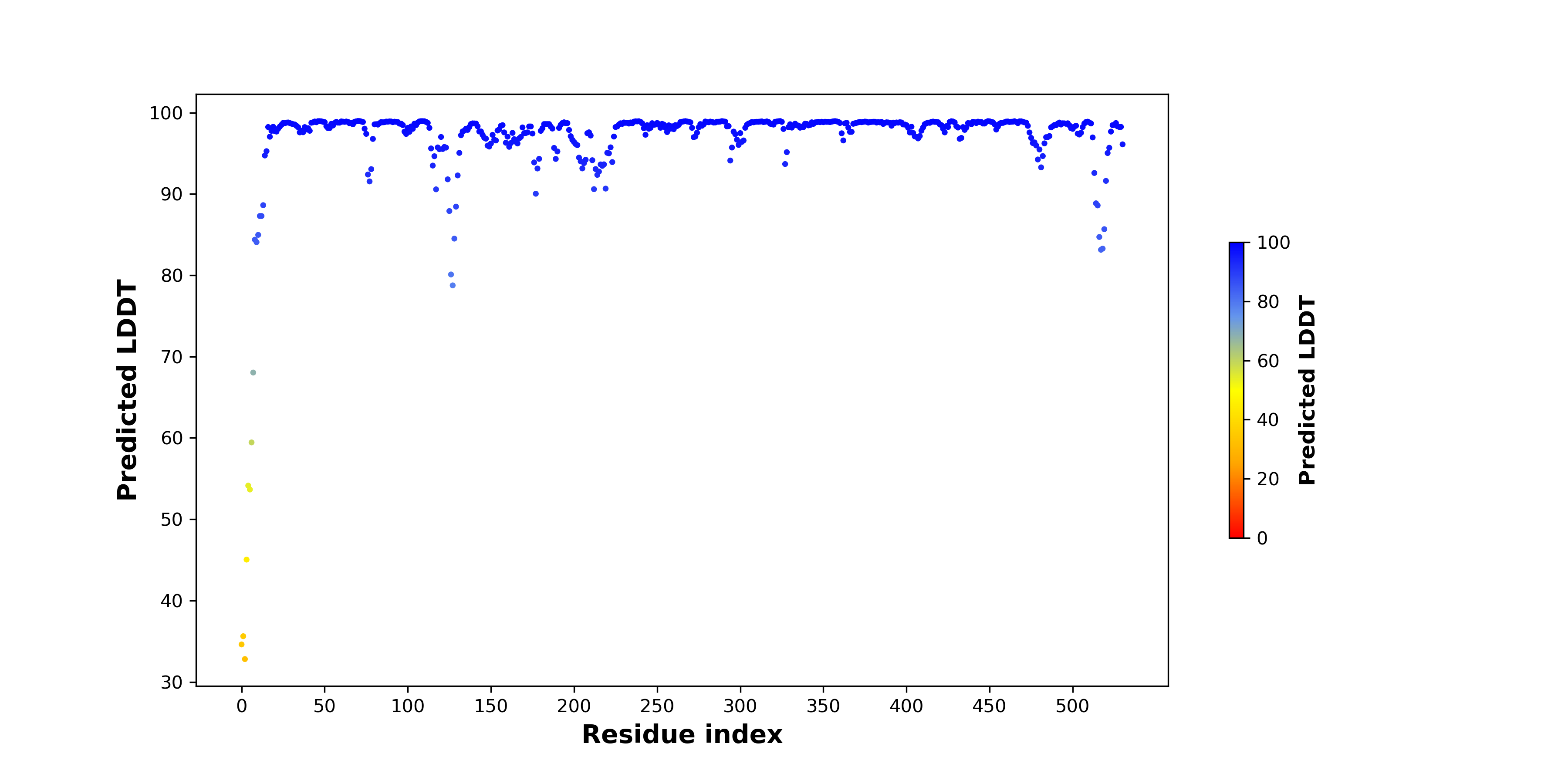 |
 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2* AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. |
Top |
Ramachandran Plot of Fusion Protein Structure |
 Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. |
| Fusion AA seq ID in FusionPDB and their Ramachandran plots |
Top |
Fusion Protein-Protein Interaction |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type from validated records (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type from validated records (BIOGRID-3.4.160) |
| Gene | PPI interactors |
 Protein-protein interactors based on sequence similarity (STRING) Protein-protein interactors based on sequence similarity (STRING) |
| Gene | STRING network |
| EIF2AK4 | |
| PKM |
 - Retained interactions in fusion protein (protein functional feature from UniProt). - Retained interactions in fusion protein (protein functional feature from UniProt). |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost interactions due to fusion (protein functional feature from UniProt). - Lost interactions due to fusion (protein functional feature from UniProt). |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs to EIF2AK4-PKM |
 Drugs used for this fusion-positive patient. Drugs used for this fusion-positive patient. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Drug | Source | PMID |
Top |
Related Diseases to EIF2AK4-PKM |
 Diseases that have this fusion gene. Diseases that have this fusion gene. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Disease | Source | PMID |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |

