| UTHEALTH HOME ABOUT SBMI A-Z WEBMAIL INSIDE THE UNIVERSITY |

|
|||||||
|
Fusion Protein:GNAI1-BRAF |
Fusion Protein Summary |
 Fusion gene summary Fusion gene summary |
| Fusion partner gene information | Fusion gene name: GNAI1-BRAF | FusionPDB ID: 33555 | FusionGDB2.0 ID: 33555 | Hgene | Tgene | Gene symbol | GNAI1 | BRAF | Gene ID | 2770 | 673 |
| Gene name | G protein subunit alpha i1 | B-Raf proto-oncogene, serine/threonine kinase | |
| Synonyms | Gi | B-RAF1|B-raf|BRAF1|NS7|RAFB1 | |
| Cytomap | 7q21.11 | 7q34 | |
| Type of gene | protein-coding | protein-coding | |
| Description | guanine nucleotide-binding protein G(i) subunit alpha-1Gi1 protein alpha subunitadenylate cyclase-inhibiting G alpha proteinguanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 1 | serine/threonine-protein kinase B-raf94 kDa B-raf proteinB-Raf proto-oncogene serine/threonine-protein kinase (p94)B-Raf serine/threonine-proteinmurine sarcoma viral (v-raf) oncogene homolog B1proto-oncogene B-Rafv-raf murine sarcoma viral oncogene | |
| Modification date | 20200329 | 20200329 | |
| UniProtAcc | P63096 | P15056 | |
| Ensembl transtripts involved in fusion gene | ENST ids | ENST00000490206, ENST00000351004, ENST00000457358, | ENST00000288602, |
| Fusion gene scores for assessment (based on all fusion genes of FusionGDB 2.0) | * DoF score | 2 X 2 X 2=8 | 48 X 58 X 16=44544 |
| # samples | 2 | 69 | |
| ** MAII score | log2(2/8*10)=1.32192809488736 | log2(69/44544*10)=-6.0124909441832 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context (manual curation of fusion genes in FusionPDB) | PubMed: GNAI1 [Title/Abstract] AND BRAF [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint (based on all fusion genes of FusionGDB 2.0) | |||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | BRAF | GO:0000186 | activation of MAPKK activity | 29433126 |
| Tgene | BRAF | GO:0006468 | protein phosphorylation | 17563371 |
| Tgene | BRAF | GO:0010828 | positive regulation of glucose transmembrane transport | 23010278 |
| Tgene | BRAF | GO:0033138 | positive regulation of peptidyl-serine phosphorylation | 19667065 |
| Tgene | BRAF | GO:0043066 | negative regulation of apoptotic process | 19667065 |
| Tgene | BRAF | GO:0070374 | positive regulation of ERK1 and ERK2 cascade | 22065586 |
| Tgene | BRAF | GO:0071277 | cellular response to calcium ion | 18567582 |
| Tgene | BRAF | GO:0090150 | establishment of protein localization to membrane | 23010278 |
 Fusion gene breakpoints across GNAI1 (5'-gene) Fusion gene breakpoints across GNAI1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 Fusion gene breakpoints across BRAF (3'-gene) Fusion gene breakpoints across BRAF (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
Top |
Fusion Gene Sample Information |
 Fusion gene information from FusionGDB2.0. Fusion gene information from FusionGDB2.0. |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerKB3 | . | . | GNAI1 | chr7 | 79764594 | + | BRAF | chr7 | 140482957 | - |
Top |
Fusion ORF Analysis |
 Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000351004 | GNAI1 | chr7 | 79764594 | + | ENST00000288602 | BRAF | chr7 | 140482957 | - | 1733 | 491 | 136 | 1614 | 492 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
Top |
Fusion Amino Acid Sequences |
 For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >FusionGDB ID_FusionGDB isoform ID_FGname_Hgene_Hchr_Hbp_Henst_Tgene_Tchr_Tbp_Tenst_length(fusion AA) seq_BP >33555_33555_1_GNAI1-BRAF_GNAI1_chr7_79764594_ENST00000351004_BRAF_chr7_140482957_ENST00000288602_length(amino acids)=492AA_BP=117 MGALSRAGKQSLVVRNSRPLLSAPLRTASPSTPLRKWWGRRGPRREAFERPLGERKDSPVLGARTRARRERRQALAFGTMGCTLSAEDKA AVERSKMIDRNLREDGEKAAREVKLLLLGSTTGLSATPPASLPGSLTNVKALQKSPGPQRERKSSSSSEDRNRMKTLGRRDSSDDWEIPD GQITVGQRIGSGSFGTVYKGKWHGDVAVKMLNVTAPTPQQLQAFKNEVGVLRKTRHVNILLFMGYSTKPQLAIVTQWCEGSSLYHHLHII ETKFEMIKLIDIARQTAQGMDYLHAKSIIHRDLKSNNIFLHEDLTVKIGDFGLATVKSRWSGSHQFEQLSGSILWMAPEVIRMQDKNPYS FQSDVYAFGIVLYELMTGQLPYSNINNRDQIIFMVGRGYLSPDLSKVRSNCPKAMKRLMAECLKKKRDERPLFPQILASIELLARSLPKI -------------------------------------------------------------- |
Top |
Fusion Protein Functional Features |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr7:/chr7:) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
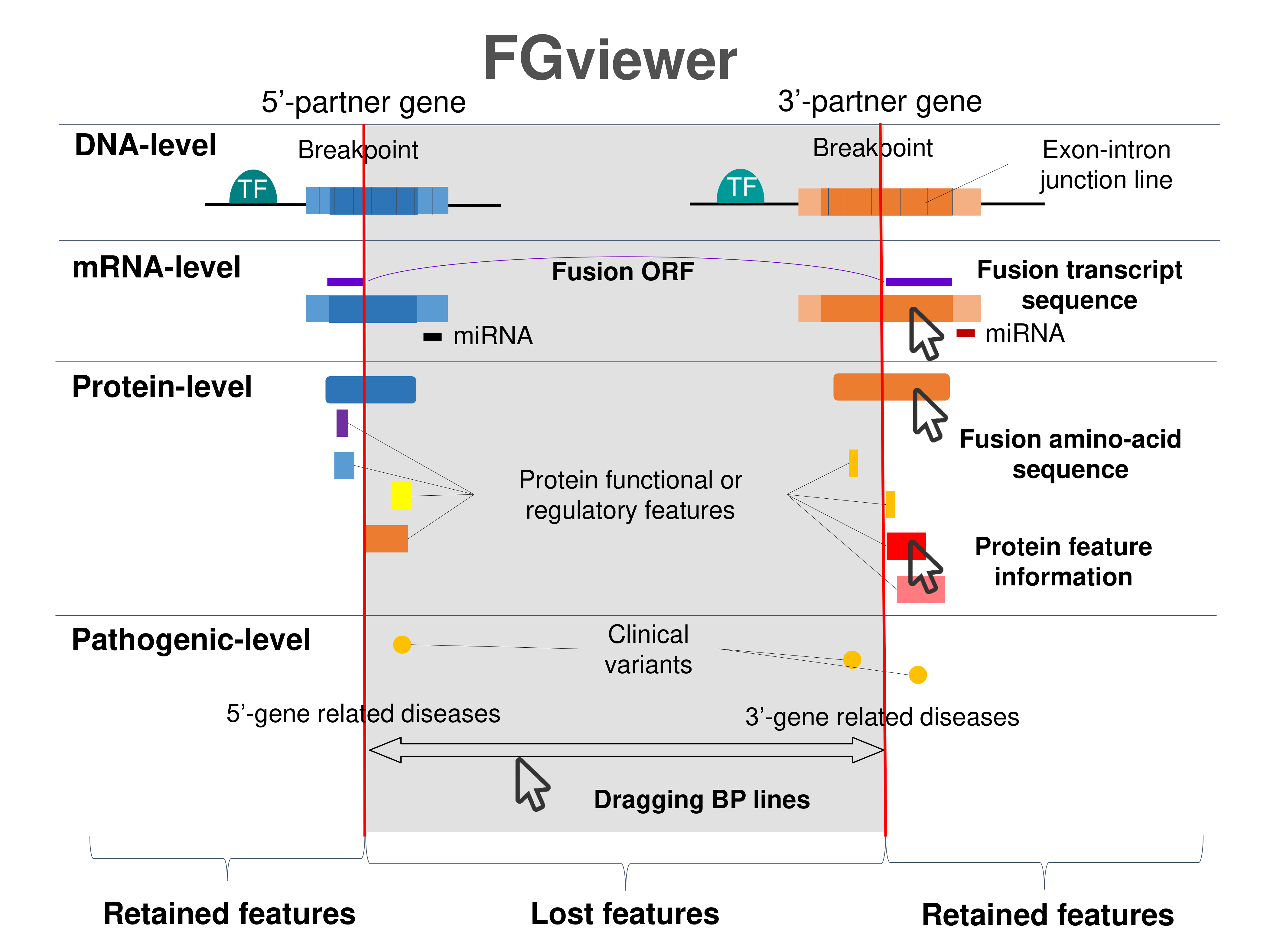 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| GNAI1 | BRAF |
| FUNCTION: Guanine nucleotide-binding proteins (G proteins) function as transducers downstream of G protein-coupled receptors (GPCRs) in numerous signaling cascades. The alpha chain contains the guanine nucleotide binding site and alternates between an active, GTP-bound state and an inactive, GDP-bound state. Signaling by an activated GPCR promotes GDP release and GTP binding. The alpha subunit has a low GTPase activity that converts bound GTP to GDP, thereby terminating the signal. Both GDP release and GTP hydrolysis are modulated by numerous regulatory proteins (PubMed:8774883, PubMed:18434541). Signaling is mediated via effector proteins, such as adenylate cyclase. Inhibits adenylate cyclase activity, leading to decreased intracellular cAMP levels (By similarity). The inactive GDP-bound form prevents the association of RGS14 with centrosomes and is required for the translocation of RGS14 from the cytoplasm to the plasma membrane. Required for normal cytokinesis during mitosis (PubMed:17635935). Required for cortical dynein-dynactin complex recruitment during metaphase (PubMed:22327364). {ECO:0000250|UniProtKB:P10824, ECO:0000269|PubMed:17635935, ECO:0000269|PubMed:18434541, ECO:0000269|PubMed:22327364, ECO:0000269|PubMed:8774883}. | FUNCTION: Protein kinase involved in the transduction of mitogenic signals from the cell membrane to the nucleus (Probable). Phosphorylates MAP2K1, and thereby activates the MAP kinase signal transduction pathway (PubMed:21441910, PubMed:29433126). May play a role in the postsynaptic responses of hippocampal neurons (PubMed:1508179). {ECO:0000269|PubMed:1508179, ECO:0000269|PubMed:21441910, ECO:0000269|PubMed:29433126, ECO:0000305}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - Retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| - Not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
Top |
Fusion Protein Structures |
 PDB and CIF files of the predicted fusion proteins PDB and CIF files of the predicted fusion proteins * Here we show the 3D structure of the fusion proteins using Mol*. AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. Model confidence is shown from the pLDDT values per residue. pLDDT corresponds to the model’s prediction of its score on the local Distance Difference Test. It is a measure of local accuracy (from AlphfaFold website). To color code individual residues, we transformed individual PDB files into CIF format. |
| Fusion protein PDB link (fusion AA seq ID in FusionPDB) | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | AA seq | Len(AA seq) |
| PDB file (286) >>>286.pdbFusion protein BP residue: 117 CIF file (286) >>>286.cif | GNAI1 | chr7 | 79764594 | + | BRAF | chr7 | 140482957 | - | MGALSRAGKQSLVVRNSRPLLSAPLRTASPSTPLRKWWGRRGPRREAFER PLGERKDSPVLGARTRARRERRQALAFGTMGCTLSAEDKAAVERSKMIDR NLREDGEKAAREVKLLLLGSTTGLSATPPASLPGSLTNVKALQKSPGPQR ERKSSSSSEDRNRMKTLGRRDSSDDWEIPDGQITVGQRIGSGSFGTVYKG KWHGDVAVKMLNVTAPTPQQLQAFKNEVGVLRKTRHVNILLFMGYSTKPQ LAIVTQWCEGSSLYHHLHIIETKFEMIKLIDIARQTAQGMDYLHAKSIIH RDLKSNNIFLHEDLTVKIGDFGLATVKSRWSGSHQFEQLSGSILWMAPEV IRMQDKNPYSFQSDVYAFGIVLYELMTGQLPYSNINNRDQIIFMVGRGYL SPDLSKVRSNCPKAMKRLMAECLKKKRDERPLFPQILASIELLARSLPKI | 492 |
| 3D view using mol* of 286 (AA BP:117) | ||||||||||
 | ||||||||||
Top |
pLDDT score distribution |
 pLDDT score distribution of the predicted wild-type structures of two partner proteins from AlphaFold2 pLDDT score distribution of the predicted wild-type structures of two partner proteins from AlphaFold2* AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. |
BRAF_pLDDT.png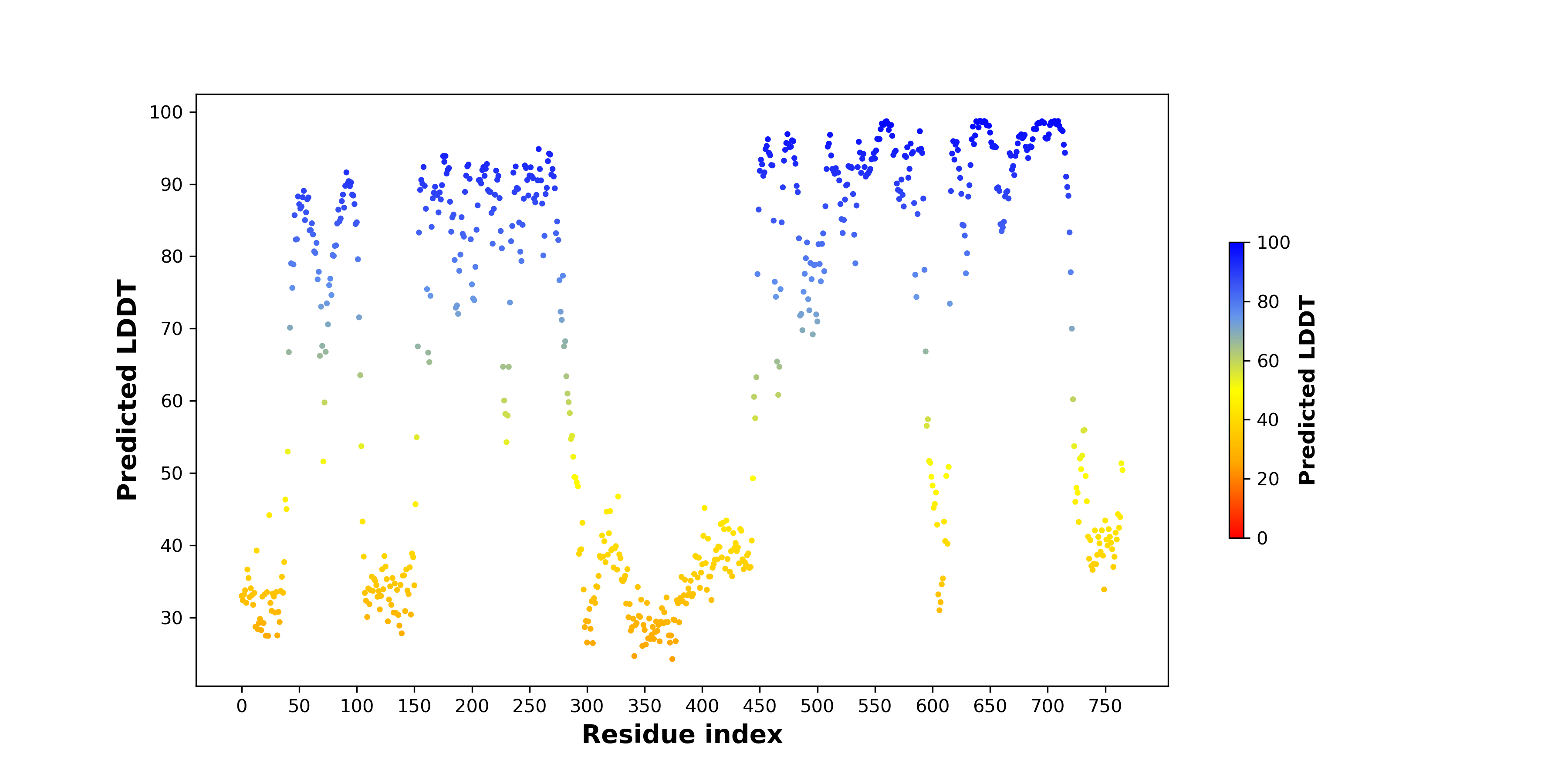 |
 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2 pLDDT score distribution of the predicted fusion protein structures from AlphaFold2* AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. |
GNAI1_BRAF_286_pLDDT.png (AA BP:117)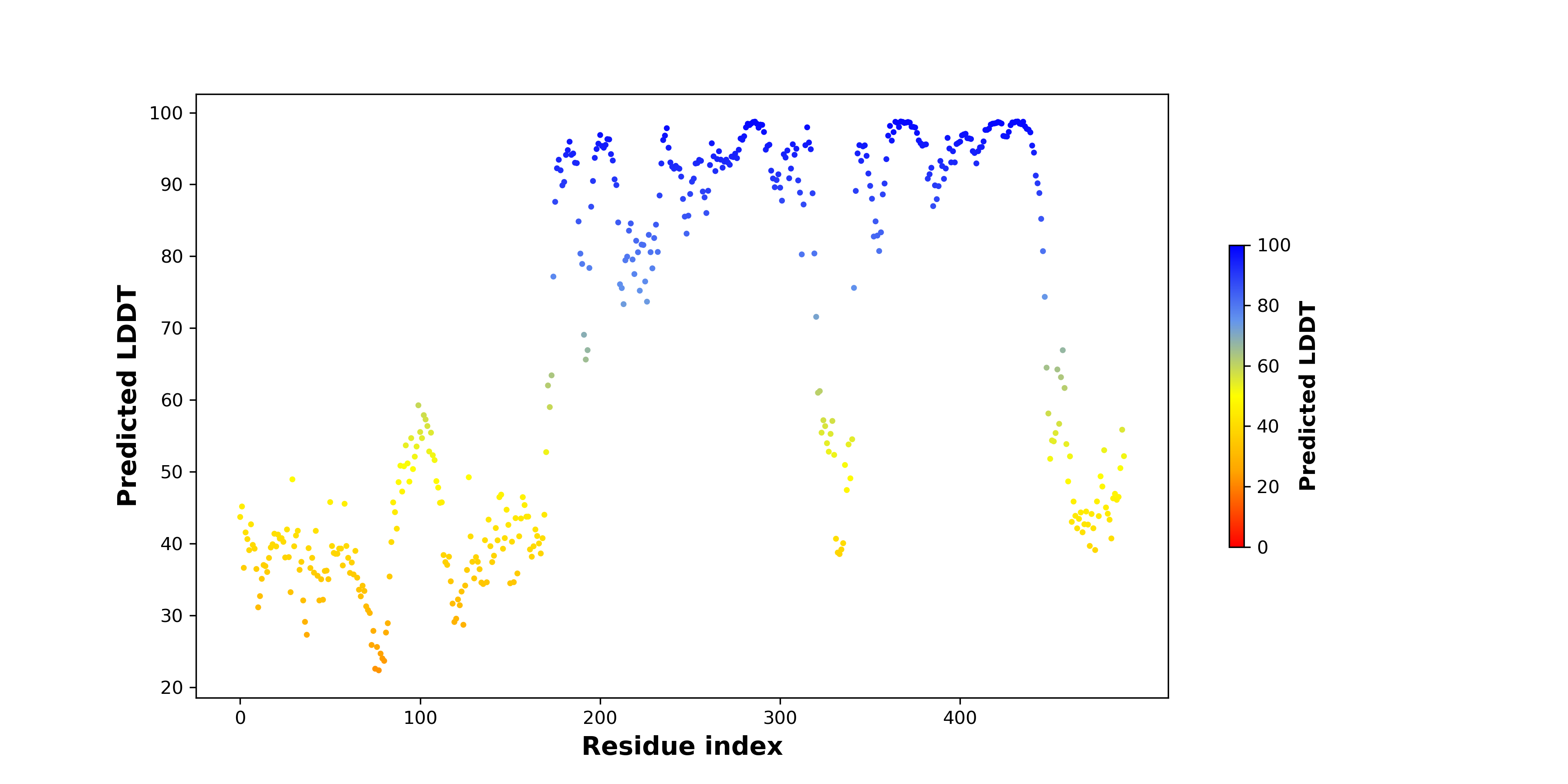 |
GNAI1_BRAF_286_pLDDT_and_active_sites.png (AA BP:117)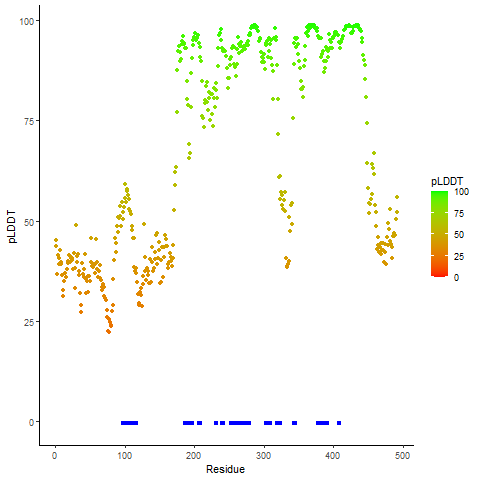 |
GNAI1_BRAF_286_violinplot.png (AA BP:117)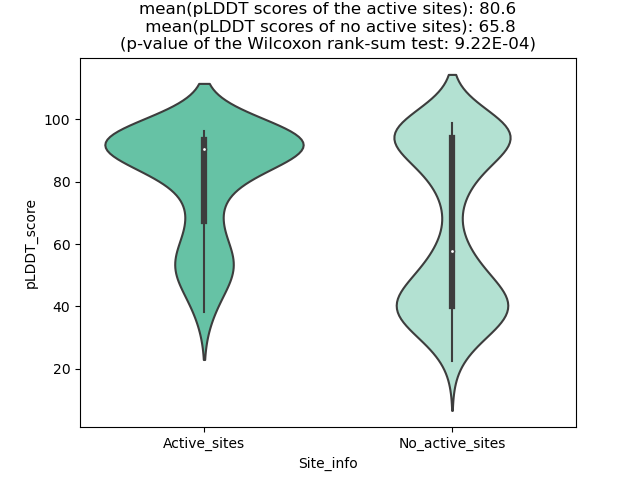 |
Top |
Ramachandran Plot of Fusion Protein Structure |
 Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. Ramachandran plot of the torsional angles - phi (φ)and psi (ψ) - of the residues (amino acids) contained in this fusion protein peptide. |
| Fusion AA seq ID in FusionPDB and their Ramachandran plots |
| GNAI1_BRAF_286.png |
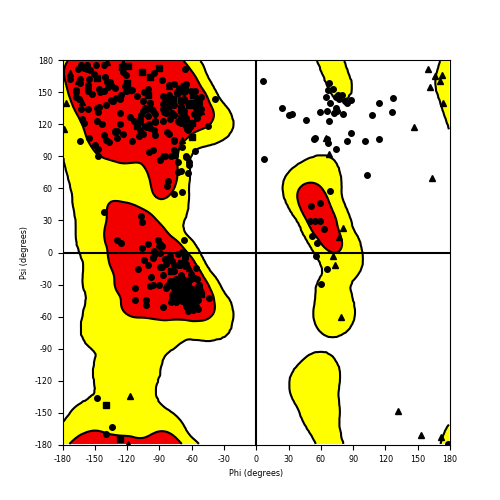 |
Top |
Potential Active Site Information |
 The potential binding sites of these fusion proteins were identified using SiteMap, a module of the Schrodinger suite. The potential binding sites of these fusion proteins were identified using SiteMap, a module of the Schrodinger suite. |
| Fusion AA seq ID in FusionPDB | Site score | Size | D score | Volume | Exposure | Enclosure | Contact | Phobic | Philic | Balance | Don/Acc | Residues |
| 286 | 1.08 | 333 | 1.13 | 1136.702 | 0.582 | 0.745 | 0.917 | 1.115 | 0.787 | 1.416 | 0.987 | Chain A: 98,99,101,102,103,105,106,107,109,110,111 ,112,113,114,117,187,189,190,191,192,193,194,195,1 97,207,209,231,240,242,253,255,256,257,258,259,260 ,261,262,264,265,267,268,269,271,272,273,274,279,3 04,306,307,309,319,320,321,323,343,344,345,378,379 ,380,381,382,383,385,386,388,391,408 |
Top |
Potentially Interacting Small Molecules through Virtual Screening |
 The FDA-approved small molecule library molecules were subjected to virtual screening using the Glide. The FDA-approved small molecule library molecules were subjected to virtual screening using the Glide. |
| Fusion AA seq ID in FusionPDB | ZINC ID | DrugBank ID | Drug name | Docking score | Glide gscore |
Top |
 Drug information from DrugBank of the top 20 interacting small molecules. Drug information from DrugBank of the top 20 interacting small molecules. |
| ZINC ID | DrugBank ID | Drug name | Drug type | SMILES | Drug group |
Top |
Biochemical Features of Small Molecules |
 ADME (Absorption, Distribution, Metabolism, and Excretion) of drugs using QikProp(v3.9) ADME (Absorption, Distribution, Metabolism, and Excretion) of drugs using QikProp(v3.9) |
| ZINC ID | mol_MW | dipole | SASA | FOSA | FISA | PISA | WPSA | volume | donorHB | accptHB | IP | Human Oral Absorption | Percent Human Oral Absorption | Rule Of Five | Rule Of Three |
Top |
Drug Toxicity Information |
 Toxicity information of individual drugs using eToxPred Toxicity information of individual drugs using eToxPred |
| ZINC ID | Smile | Surface Accessibility | Toxicity |
Top |
Fusion Protein-Protein Interaction |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type from validated records (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type from validated records (BIOGRID-3.4.160) |
| Gene | PPI interactors |
| BRAF | YWHAB, YWHAG, YWHAQ, YWHAZ, SFN, HRAS, AKT1, MAPK3, RAP1GAP, RAF1, MRAS, RAP1A, PAK2, TERF1, CCDC88A, NEDD4L, Nedd4, MAP2K1, RNF149, KSR1, BRAP, PRKCE, RPS6KB2, HSP90AA1, BRAF, YWHAE, HSPA5, MAP2K2, HSPA1A, HSPA8, YWHAH, HSPA9, ARAF, CDC37, HSP90AB1, PHKB, LIMK1, IQGAP1, MAPK1, BAD, LIPF, MUS81, Vps4b, FBXW7, FGFR2, BRCA2, VHL, FNTA, HDAC2, PIK3CA, EGFR, PTEN, FNIP1, FNIP2, RAB3GAP1, KRAS, KIAA0141, FKBPL, ARMCX3, KCNC4, RPTOR, CYLD, NRAS, HSPA4, DNAJB6, PDCD11, PIP5K1A, DNAJC15, VANGL1, DNAJC11, FKBP5, HSPA4L, HSP90B1, GNAI2, DNAJC13, GNAS, PPP2CB, HSD11B2, DNAJB11, PLD2, RAP1B, WDR6, CPNE3, MYOF, COPA, UBLCP1, PPP1CA, RAD50, PIP4K2C, PHB, VIM, PGAM1, DNAJA1, MAP2K7, SPRY2, ALDOA, AP2B1, ATP5A1, SSB, IGF1R, JUP, PPP6C, PARP1, HSPB1, NME2, PRDX2, CCT7, RAB1A, FARSA, FASN, EPRS, TRAF2, REST, KIAA1429, NANOG, ITCH, SMURF2, WWP1, WWP2, PPP2CA, PPP2R2A, AURKA, LATS2, MAP2K3, MAP2K6, RASSF1, STK11, TERT, PEBP1, CRBN, FAR1, PSMC4, UBA52, UBB, UBC, RPS27A, HSPA6, HSP90AB3P, DSP, ATAD3A, ATAD3B, P4HB, SDF2L1, SLC25A22, SPTBN4, TMEM33, CTSB, NCL, HPX, TXNDC12, SLC25A11, NDUFA4, CTSV, FBP1, HSD17B3, ZNF189, ZNF510, KIF14, SRC, TRAP1, JTB, S100P, USP28, |
| GNAI1 | GPSM3, GPSM1, NGB, CNR1, RGS19, ADCY5, RGS18, EPOR, GPR143, CRHR1, RGS4, RGS16, RASD1, RGS14, RGS12, RGS5, IGF1R, RGS10, RIC8A, S1PR1, PGR, Haus4, Cbx1, Haus1, Recql4, Cep76, Trim69, PCK1, PTH1R, GNB1, GNB2, GNB4, GNAI3, GNAI2, GPR50, NCF2, NCF1, IQCB1, MTNR1A, MTNR1B, SVIL, RAD52, ATP4A, NUCB1, RANGAP1, THAP7, RGS17, RGS1, GNG4, GNAT3, GNAO1, MTHFR, GNG10, RAP1GDS1, GNA13, GNG5, PDE3B, ABCA3, DDRGK1, ATP6V0A2, GNG7, CTDNEP1, ESR1, PSMD3, PTEN, FLNA, MYH9, MYO19, Actb, Tpm1, Dctn3, Lima1, Myh10, GLI1, DUSP22, GNG3, COX15, BCL2L1, ARNT, RNF138, PPP5C, CD81, EGFR, PDGFRA, FGFR1, DFNB26, DUSP6, EYA4, SMO, DRD4, GNG2, ADRB2, PCP2, YAP1, TFCP2, RGS20, PRKD1, EMC1, MMGT1, PKD1, ANLN, CHMP4B, ECT2, KIF14, KIF20A, KIF23, PRC1, NMRAL1, AR, GNG8, SNUPN, BTF3, NRP1, TOP3B, FURIN, DPP4, BSG, IFITM3, CLEC4D, ACE2, CCNF, |
 Protein-protein interactors based on sequence similarity (STRING) Protein-protein interactors based on sequence similarity (STRING) |
| Gene | STRING network |
| GNAI1 | 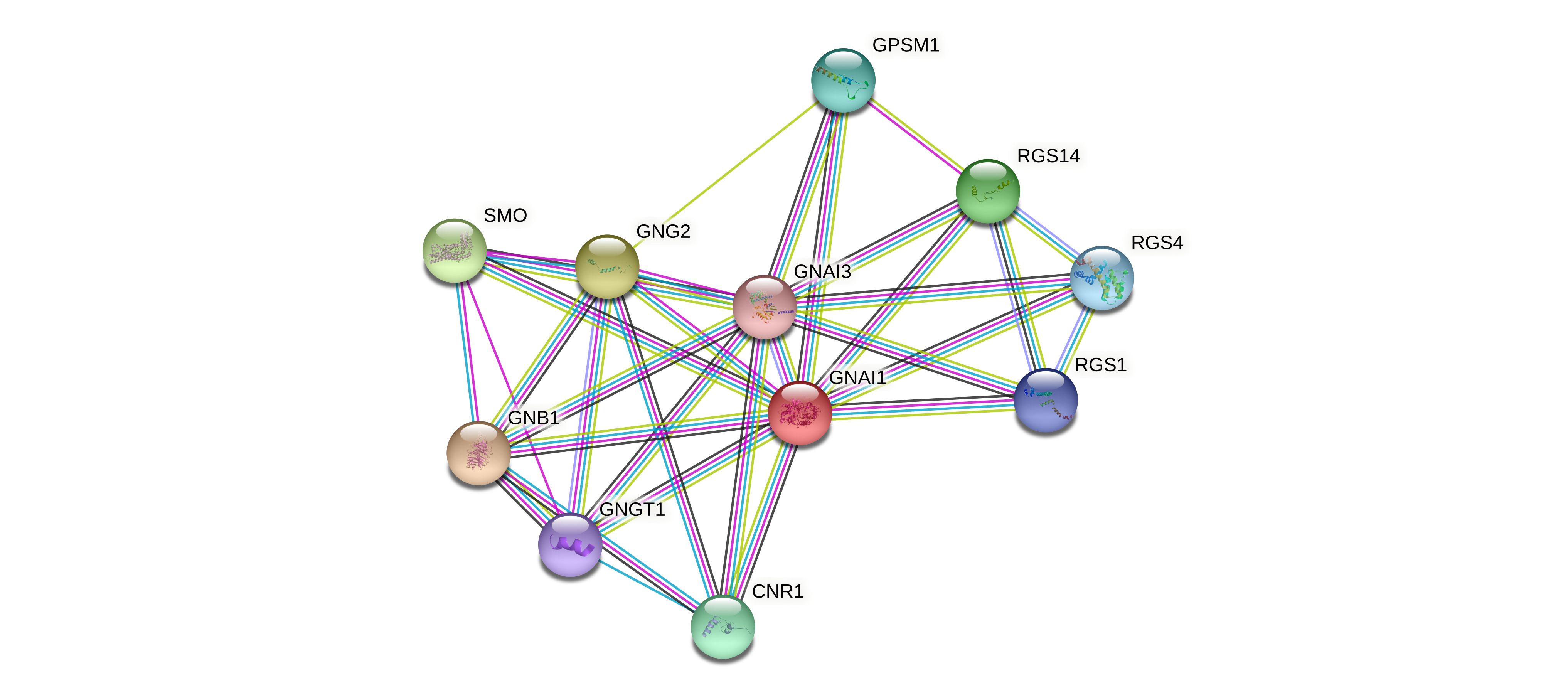 |
| BRAF | 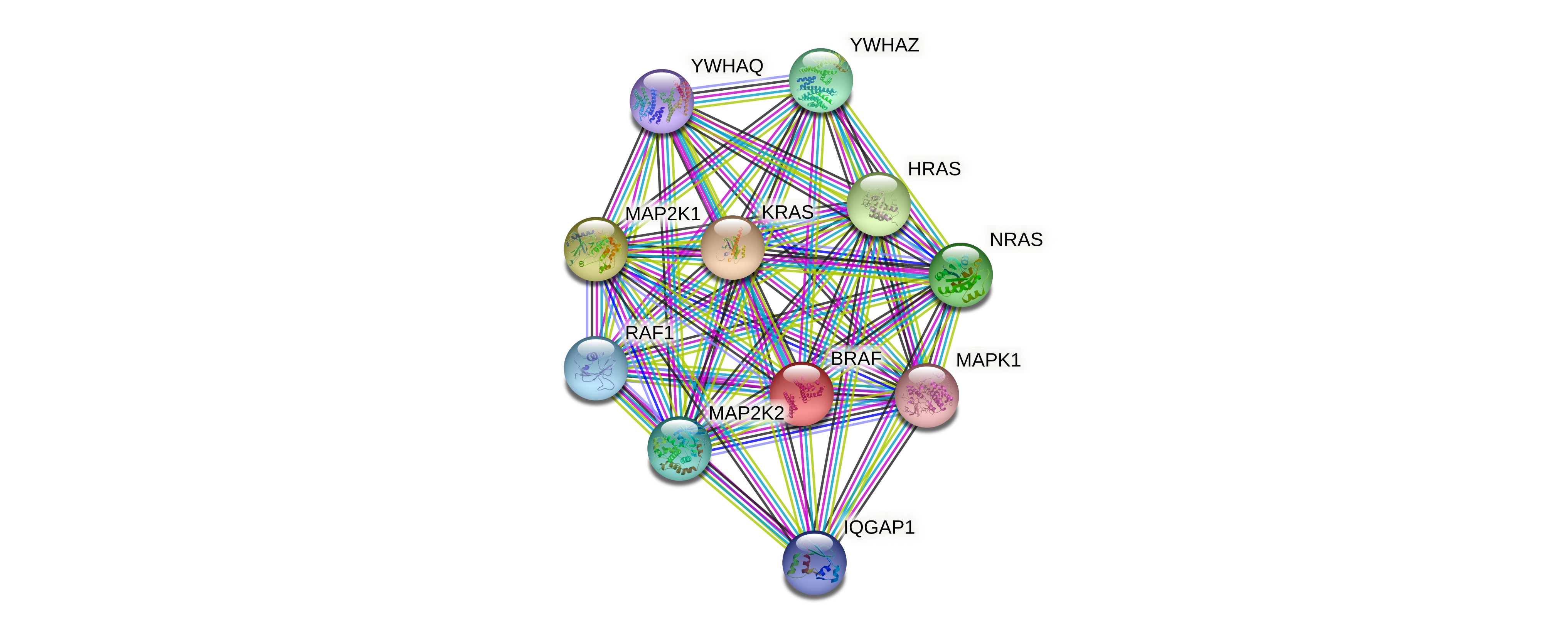 |
 - Retained interactions in fusion protein (protein functional feature from UniProt). - Retained interactions in fusion protein (protein functional feature from UniProt). |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost interactions due to fusion (protein functional feature from UniProt). - Lost interactions due to fusion (protein functional feature from UniProt). |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs to GNAI1-BRAF |
 Drugs used for this fusion-positive patient. Drugs used for this fusion-positive patient. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Drug | Source | PMID |
Top |
Related Diseases to GNAI1-BRAF |
 Diseases that have this fusion gene. Diseases that have this fusion gene. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Disease | Source | PMID |
| GNAI1 | BRAF | Pilocytic Astrocytoma | MyCancerGenome | |
| GNAI1 | BRAF | Low-Grade Glioma | MyCancerGenome | |
| GNAI1 | BRAF | Nos | MyCancerGenome | |
| GNAI1 | BRAF | Pancreatic Adenocarcinoma | MyCancerGenome | |
| GNAI1 | BRAF | Thyroid Gland Papillary Carcinoma | MyCancerGenome | |
| GNAI1 | BRAF | Astrocytoma | MyCancerGenome |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
| Hgene | GNAI1 | C0002622 | Amnesia | 1 | CTD_human |
| Hgene | GNAI1 | C0038587 | Substance Withdrawal Syndrome | 1 | CTD_human |
| Hgene | GNAI1 | C0086189 | Drug Withdrawal Symptoms | 1 | CTD_human |
| Hgene | GNAI1 | C0087169 | Withdrawal Symptoms | 1 | CTD_human |
| Hgene | GNAI1 | C0233750 | Hysterical amnesia | 1 | CTD_human |
| Hgene | GNAI1 | C0233796 | Temporary Amnesia | 1 | CTD_human |
| Hgene | GNAI1 | C0236795 | Dissociative Amnesia | 1 | CTD_human |
| Hgene | GNAI1 | C0262497 | Global Amnesia | 1 | CTD_human |
| Hgene | GNAI1 | C0750906 | Tactile Amnesia | 1 | CTD_human |
| Hgene | GNAI1 | C0750907 | Amnestic State | 1 | CTD_human |
| Tgene | BRAF | C0025202 | melanoma | 24 | CGI;CTD_human;UNIPROT |
| Tgene | BRAF | C1275081 | Cardio-facio-cutaneous syndrome | 14 | CLINGEN;CTD_human;GENOMICS_ENGLAND;ORPHANET;UNIPROT |
| Tgene | BRAF | C0009402 | Colorectal Carcinoma | 8 | CTD_human;UNIPROT |
| Tgene | BRAF | C0028326 | Noonan Syndrome | 8 | CLINGEN;CTD_human;GENOMICS_ENGLAND;ORPHANET |
| Tgene | BRAF | C0238463 | Papillary thyroid carcinoma | 8 | CTD_human;ORPHANET |
| Tgene | BRAF | C0040136 | Thyroid Neoplasm | 6 | CGI;CTD_human |
| Tgene | BRAF | C0151468 | Thyroid Gland Follicular Adenoma | 6 | CTD_human |
| Tgene | BRAF | C0175704 | LEOPARD Syndrome | 6 | CLINGEN;GENOMICS_ENGLAND |
| Tgene | BRAF | C0549473 | Thyroid carcinoma | 6 | CGI;CTD_human |
| Tgene | BRAF | C3150970 | NOONAN SYNDROME 7 | 5 | CTD_human;GENOMICS_ENGLAND;UNIPROT |
| Tgene | BRAF | C0009404 | Colorectal Neoplasms | 4 | CTD_human |
| Tgene | BRAF | C3150971 | LEOPARD SYNDROME 3 | 4 | CTD_human;GENOMICS_ENGLAND;UNIPROT |
| Tgene | BRAF | C1519086 | Pilomyxoid astrocytoma | 3 | ORPHANET |
| Tgene | BRAF | C0004565 | Melanoma, B16 | 2 | CTD_human |
| Tgene | BRAF | C0009075 | Melanoma, Cloudman S91 | 2 | CTD_human |
| Tgene | BRAF | C0018598 | Melanoma, Harding-Passey | 2 | CTD_human |
| Tgene | BRAF | C0023443 | Hairy Cell Leukemia | 2 | CGI;ORPHANET |
| Tgene | BRAF | C0025205 | Melanoma, Experimental | 2 | CTD_human |
| Tgene | BRAF | C0033578 | Prostatic Neoplasms | 2 | CTD_human |
| Tgene | BRAF | C0152013 | Adenocarcinoma of lung (disorder) | 2 | CGI;CTD_human |
| Tgene | BRAF | C0376358 | Malignant neoplasm of prostate | 2 | CTD_human |
| Tgene | BRAF | C0587248 | Costello syndrome (disorder) | 2 | CLINGEN;CTD_human |
| Tgene | BRAF | C3501843 | Nonmedullary Thyroid Carcinoma | 2 | CTD_human |
| Tgene | BRAF | C3501844 | Familial Nonmedullary Thyroid Cancer | 2 | CTD_human |
| Tgene | BRAF | C0002448 | Ameloblastoma | 1 | CTD_human |
| Tgene | BRAF | C0004114 | Astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0010276 | Craniopharyngioma | 1 | CTD_human;ORPHANET |
| Tgene | BRAF | C0011860 | Diabetes Mellitus, Non-Insulin-Dependent | 1 | CTD_human |
| Tgene | BRAF | C0017638 | Glioma | 1 | CGI;CTD_human |
| Tgene | BRAF | C0019621 | Histiocytosis, Langerhans-Cell | 1 | CGI;ORPHANET |
| Tgene | BRAF | C0022665 | Kidney Neoplasm | 1 | CTD_human |
| Tgene | BRAF | C0023903 | Liver neoplasms | 1 | CTD_human |
| Tgene | BRAF | C0024232 | Lymphatic Metastasis | 1 | CTD_human |
| Tgene | BRAF | C0024694 | Mandibular Neoplasms | 1 | CTD_human |
| Tgene | BRAF | C0027659 | Neoplasms, Experimental | 1 | CTD_human |
| Tgene | BRAF | C0027962 | Melanocytic nevus | 1 | GENOMICS_ENGLAND |
| Tgene | BRAF | C0036920 | Sezary Syndrome | 1 | CTD_human |
| Tgene | BRAF | C0041409 | Turner Syndrome, Male | 1 | CTD_human |
| Tgene | BRAF | C0079773 | Lymphoma, T-Cell, Cutaneous | 1 | CTD_human |
| Tgene | BRAF | C0205768 | Subependymal Giant Cell Astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0206686 | Adrenocortical carcinoma | 1 | CTD_human |
| Tgene | BRAF | C0206754 | Neuroendocrine Tumors | 1 | CTD_human |
| Tgene | BRAF | C0259783 | mixed gliomas | 1 | CTD_human |
| Tgene | BRAF | C0278875 | Adult Craniopharyngioma | 1 | CTD_human |
| Tgene | BRAF | C0280783 | Juvenile Pilocytic Astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0280785 | Diffuse Astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0334579 | Anaplastic astrocytoma | 1 | CGI;CTD_human |
| Tgene | BRAF | C0334580 | Protoplasmic astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0334581 | Gemistocytic astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0334582 | Fibrillary Astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0334583 | Pilocytic Astrocytoma | 1 | CGI;CTD_human |
| Tgene | BRAF | C0338070 | Childhood Cerebral Astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0345904 | Malignant neoplasm of liver | 1 | CTD_human |
| Tgene | BRAF | C0376407 | Granulomatous Slack Skin | 1 | CTD_human |
| Tgene | BRAF | C0406803 | Syringocystadenoma Papilliferum | 1 | GENOMICS_ENGLAND |
| Tgene | BRAF | C0431128 | Papillary craniopharyngioma | 1 | CTD_human |
| Tgene | BRAF | C0431129 | Adamantinous Craniopharyngioma | 1 | CTD_human |
| Tgene | BRAF | C0547065 | Mixed oligoastrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0555198 | Malignant Glioma | 1 | CTD_human |
| Tgene | BRAF | C0596263 | Carcinogenesis | 1 | CTD_human |
| Tgene | BRAF | C0684249 | Carcinoma of lung | 1 | CGI;UNIPROT |
| Tgene | BRAF | C0740457 | Malignant neoplasm of kidney | 1 | CTD_human |
| Tgene | BRAF | C0750935 | Cerebral Astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0750936 | Intracranial Astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C0751061 | Craniopharyngioma, Child | 1 | CTD_human |
| Tgene | BRAF | C0920269 | Microsatellite Instability | 1 | CTD_human |
| Tgene | BRAF | C1527404 | Female Pseudo-Turner Syndrome | 1 | CTD_human |
| Tgene | BRAF | C1704230 | Grade I Astrocytoma | 1 | CTD_human |
| Tgene | BRAF | C1721098 | Replication Error Phenotype | 1 | CTD_human |
| Tgene | BRAF | C2239176 | Liver carcinoma | 1 | CTD_human |
| Tgene | BRAF | C4551484 | Leopard Syndrome 1 | 1 | GENOMICS_ENGLAND |
| Tgene | BRAF | C4551602 | Noonan Syndrome 1 | 1 | CTD_human |
| Tgene | BRAF | C4721532 | Lymphoma, Non-Hodgkin, Familial | 1 | UNIPROT |
| Tgene | BRAF | C4733333 | familial non-medullary thyroid cancer | 1 | GENOMICS_ENGLAND |

