| UTHEALTH HOME ABOUT SBMI A-Z WEBMAIL INSIDE THE UNIVERSITY |

|
|||||||
|
Fusion Protein:LETM1-MAEA |
Fusion Protein Summary |
 Fusion gene summary Fusion gene summary |
| Fusion partner gene information | Fusion gene name: LETM1-MAEA | FusionPDB ID: 44485 | FusionGDB2.0 ID: 44485 | Hgene | Tgene | Gene symbol | LETM1 | MAEA | Gene ID | 3954 | 10296 |
| Gene name | leucine zipper and EF-hand containing transmembrane protein 1 | macrophage erythroblast attacher, E3 ubiquitin ligase | |
| Synonyms | SLC55A1 | EMLP|EMP|GID9|HLC-10|P44EMLP|PIG5 | |
| Cytomap | 4p16.3 | 4p16.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | mitochondrial proton/calcium exchanger proteinLETM1 and EF-hand domain-containing protein 1, mitochondrialMdm38 homologleucine zipper-EF-hand containing transmembrane protein 1 | E3 ubiquitin-protein transferase MAEAGID complex subunit 9, FYV10 homologcell proliferation-inducing gene 5 proteinerythroblast macrophage proteinhuman lung cancer oncogene 10 proteinlung cancer-related protein 10macrophage erythroblast attacher | |
| Modification date | 20200328 | 20200314 | |
| UniProtAcc | O95202 | Q7L5Y9 | |
| Ensembl transtripts involved in fusion gene | ENST ids | ENST00000302787, ENST00000512189, | ENST00000512289, ENST00000264750, ENST00000303400, ENST00000452175, ENST00000505177, ENST00000505839, ENST00000510794, ENST00000514708, |
| Fusion gene scores for assessment (based on all fusion genes of FusionGDB 2.0) | * DoF score | 10 X 13 X 7=910 | 8 X 9 X 5=360 |
| # samples | 12 | 11 | |
| ** MAII score | log2(12/910*10)=-2.92283213947754 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(11/360*10)=-1.71049338280502 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context (manual curation of fusion genes in FusionPDB) | PubMed: LETM1 [Title/Abstract] AND MAEA [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint (based on all fusion genes of FusionGDB 2.0) | LETM1(1857595)-MAEA(1326544), # samples:1 LETM1(1817390)-MAEA(1332861), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | LETM1-MAEA seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. LETM1-MAEA seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. LETM1-MAEA seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. LETM1-MAEA seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. LETM1-MAEA seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. LETM1-MAEA seems lost the major protein functional domain in Hgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. LETM1-MAEA seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | LETM1 | GO:0006851 | mitochondrial calcium ion transmembrane transport | 19797662 |
| Hgene | LETM1 | GO:0051560 | mitochondrial calcium ion homeostasis | 19797662 |
| Hgene | LETM1 | GO:0099093 | calcium export from the mitochondrion | 19797662 |
| Tgene | MAEA | GO:0007155 | cell adhesion | 9763581 |
| Tgene | MAEA | GO:0033033 | negative regulation of myeloid cell apoptotic process | 9763581 |
 Fusion gene breakpoints across LETM1 (5'-gene) Fusion gene breakpoints across LETM1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
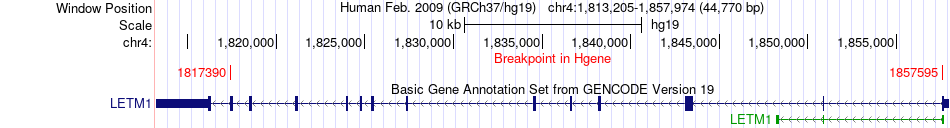 |
 Fusion gene breakpoints across MAEA (3'-gene) Fusion gene breakpoints across MAEA (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
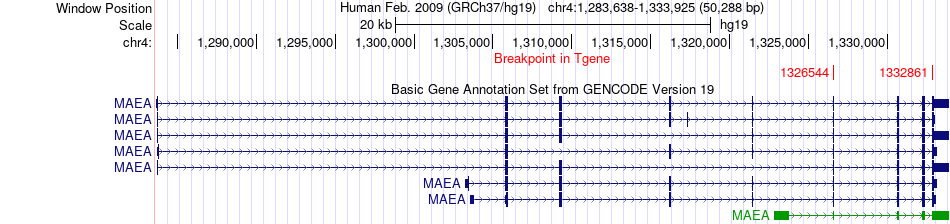 |
Top |
Fusion Gene Sample Information |
 Fusion gene information from FusionGDB2.0. Fusion gene information from FusionGDB2.0. |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | ESCA | TCGA-LN-A7HZ | LETM1 | chr4 | 1857595 | - | MAEA | chr4 | 1326544 | + |
| ChimerDB4 | STAD | TCGA-HU-A4GP | LETM1 | chr4 | 1817390 | - | MAEA | chr4 | 1332861 | + |
Top |
Fusion ORF Analysis |
 Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000302787 | LETM1 | chr4 | 1857595 | - | ENST00000514708 | MAEA | chr4 | 1326544 | + | 1849 | 379 | 543 | 1 | 181 |
| ENST00000302787 | LETM1 | chr4 | 1817390 | - | ENST00000452175 | MAEA | chr4 | 1332861 | + | 2658 | 2367 | 297 | 2462 | 721 |
| ENST00000302787 | LETM1 | chr4 | 1817390 | - | ENST00000514708 | MAEA | chr4 | 1332861 | + | 3402 | 2367 | 297 | 2462 | 721 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000302787 | ENST00000514708 | LETM1 | chr4 | 1857595 | - | MAEA | chr4 | 1326544 | + | 0.28035444 | 0.7196456 |
| ENST00000302787 | ENST00000452175 | LETM1 | chr4 | 1817390 | - | MAEA | chr4 | 1332861 | + | 0.019727325 | 0.9802727 |
| ENST00000302787 | ENST00000514708 | LETM1 | chr4 | 1817390 | - | MAEA | chr4 | 1332861 | + | 0.012407264 | 0.98759276 |
Top |
Fusion Amino Acid Sequences |
 For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >FusionGDB ID_FusionGDB isoform ID_FGname_Hgene_Hchr_Hbp_Henst_Tgene_Tchr_Tbp_Tenst_length(fusion AA) seq_BP >44485_44485_1_LETM1-MAEA_LETM1_chr4_1817390_ENST00000302787_MAEA_chr4_1332861_ENST00000452175_length(amino acids)=721AA_BP=690 MASILLRSCRGRAPARLPPPPRYTVPRGSPGDPAHLSCASTLGLRNCLNVPFGCCTPIHPVYTSSRGDHLGCWALRPECLRIVSRAPWTS TSVGFVAVGPQCLPVRGWHSSRPVRDDSVVEKSLKSLKDKNKKLEEGGPVYSPPAEVVVKKSLGQRVLDELKHYYHGFRLLWIDTKIAAR MLWRILNGHSLTRRERRQFLRICADLFRLVPFLVFVVVPFMEFLLPVAVKLFPNMLPSTFETQSLKEERLKKELRVKLELAKFLQDTIEE MALKNKAAKGSATKDFSVFFQKIRETGERPSNEEIMRFSKLFEDELTLDNLTRPQLVALCKLLELQSIGTNNFLRFQLTMRLRSIKADDK LIAEEGVDSLNVKELQAACRARGMRALGVTEDRLRGQLKQWLDLHLHQEIPTSLLILSRAMYLPDTLSPADQLKSTLQTLPEIVAKEAQV KVAEVEGEQVDNKAKLEATLQEEAAIQQEHREKELQKRSEVAKDFEPERVVAAPQRPGTEPQPEMPDTVLQSETLKDTAPVLEGLKEEEI TKEEIDILSDACSKLQEQKKSLTKEKEELELLKEDVQDYSEDLQEIKKELSKTGEEKYVEESKASKRLTKRVQQMIGQIDGLISQLEMDQ QAGKLAPANGMPTGENVISVAELINAMKQVKHIPESKLTSLAAALDENKDGKVNIDDLVKSLLSIRQDDKVVCPRTKEVFHFSQAEKVYI -------------------------------------------------------------- >44485_44485_2_LETM1-MAEA_LETM1_chr4_1817390_ENST00000302787_MAEA_chr4_1332861_ENST00000514708_length(amino acids)=721AA_BP=690 MASILLRSCRGRAPARLPPPPRYTVPRGSPGDPAHLSCASTLGLRNCLNVPFGCCTPIHPVYTSSRGDHLGCWALRPECLRIVSRAPWTS TSVGFVAVGPQCLPVRGWHSSRPVRDDSVVEKSLKSLKDKNKKLEEGGPVYSPPAEVVVKKSLGQRVLDELKHYYHGFRLLWIDTKIAAR MLWRILNGHSLTRRERRQFLRICADLFRLVPFLVFVVVPFMEFLLPVAVKLFPNMLPSTFETQSLKEERLKKELRVKLELAKFLQDTIEE MALKNKAAKGSATKDFSVFFQKIRETGERPSNEEIMRFSKLFEDELTLDNLTRPQLVALCKLLELQSIGTNNFLRFQLTMRLRSIKADDK LIAEEGVDSLNVKELQAACRARGMRALGVTEDRLRGQLKQWLDLHLHQEIPTSLLILSRAMYLPDTLSPADQLKSTLQTLPEIVAKEAQV KVAEVEGEQVDNKAKLEATLQEEAAIQQEHREKELQKRSEVAKDFEPERVVAAPQRPGTEPQPEMPDTVLQSETLKDTAPVLEGLKEEEI TKEEIDILSDACSKLQEQKKSLTKEKEELELLKEDVQDYSEDLQEIKKELSKTGEEKYVEESKASKRLTKRVQQMIGQIDGLISQLEMDQ QAGKLAPANGMPTGENVISVAELINAMKQVKHIPESKLTSLAAALDENKDGKVNIDDLVKSLLSIRQDDKVVCPRTKEVFHFSQAEKVYI -------------------------------------------------------------- >44485_44485_3_LETM1-MAEA_LETM1_chr4_1857595_ENST00000302787_MAEA_chr4_1326544_ENST00000514708_length(amino acids)=181AA_BP=1 MSYRNCWISIRHRAGSRRSLYGEMCVSGGKASMPMAWRTSSSWLPSAWLKCFLACPAGRCTEAAAGGGRAPGRGSSSVRWTPCARARRPL RPPALPPSPRRPRLFAVPAALQTRPARTADRGGWPRTGGALLKDPAPAAALSRHRRPEARGGAARPRLPLGPSAPAAGQVRGGPRHRGLP -------------------------------------------------------------- |
Top |
Fusion Protein Functional Features |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr4:1857595/chr4:1326544) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
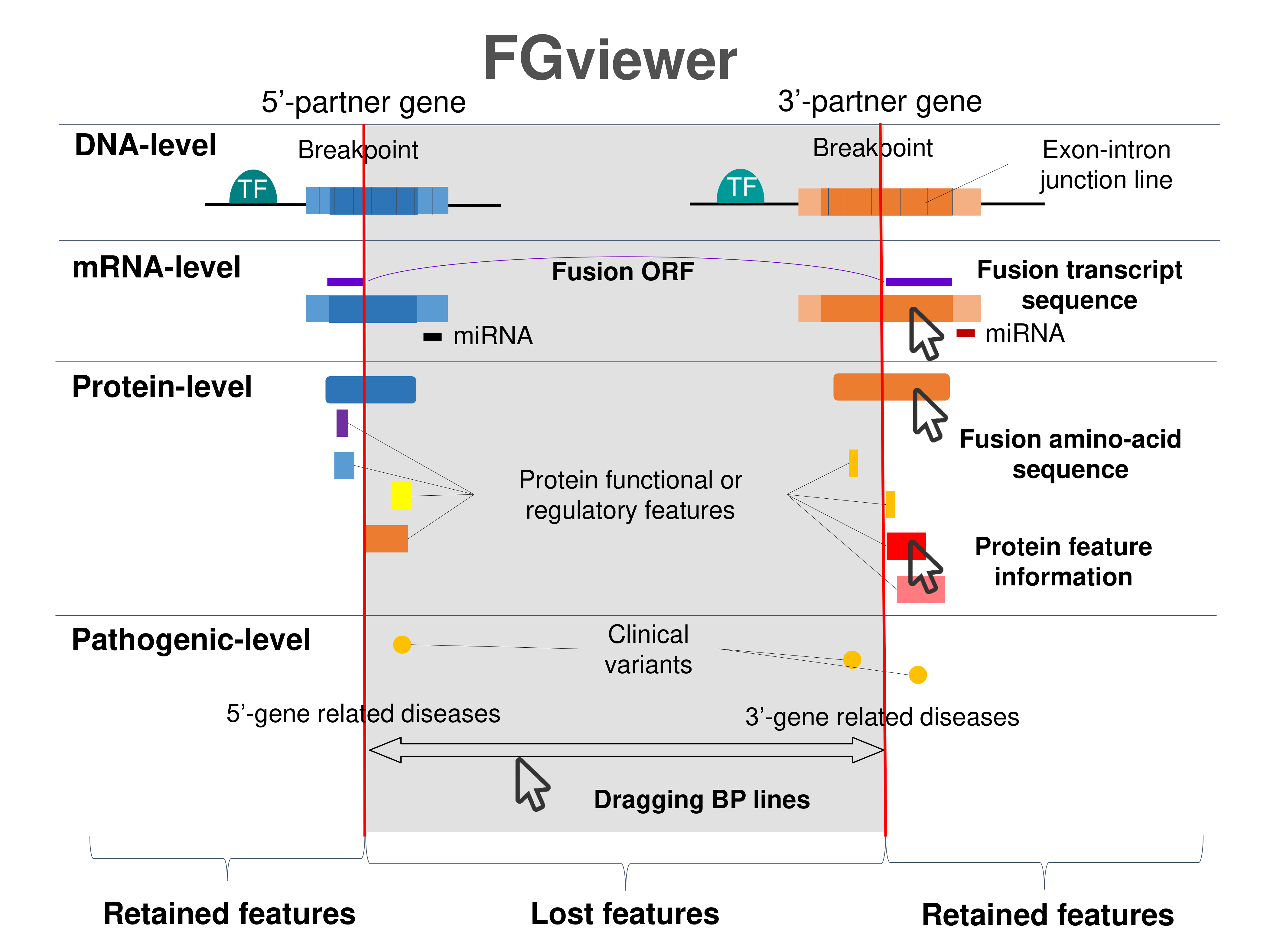 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| LETM1 | MAEA |
| FUNCTION: Mitochondrial proton/calcium antiporter that mediates proton-dependent calcium efflux from mitochondrion (PubMed:19797662). Crucial for the maintenance of mitochondrial tubular networks and for the assembly of the supercomplexes of the respiratory chain (PubMed:18628306). Required for the maintenance of the tubular shape and cristae organization (PubMed:18628306). In contrast to SLC8B1/NCLX, does not constitute the major factor for mitochondrial calcium extrusion (PubMed:24898248). {ECO:0000269|PubMed:18628306, ECO:0000269|PubMed:19797662, ECO:0000269|PubMed:24898248}. | FUNCTION: Core component of the CTLH E3 ubiquitin-protein ligase complex that selectively accepts ubiquitin from UBE2H and mediates ubiquitination and subsequent proteasomal degradation of the transcription factor HBP1. MAEA and RMND5A are both required for catalytic activity of the CTLH E3 ubiquitin-protein ligase complex (PubMed:29911972). MAEA is required for normal cell proliferation (PubMed:29911972). The CTLH E3 ubiquitin-protein ligase complex is not required for the degradation of enzymes involved in gluconeogenesis, such as FBP1 (PubMed:29911972). Plays a role in erythroblast enucleation during erythrocyte maturation and in the development of mature macrophages (By similarity). Mediates the attachment of erythroid cell to mature macrophages; this MAEA-mediated contact inhibits erythroid cell apoptosis (PubMed:9763581). Participates in erythroblastic island formation, which is the functional unit of definitive erythropoiesis. Associates with F-actin to regulate actin distribution in erythroblasts and macrophages (By similarity). May contribute to nuclear architecture and cells division events (Probable). {ECO:0000250|UniProtKB:Q4VC33, ECO:0000269|PubMed:29911972, ECO:0000269|PubMed:9763581, ECO:0000305|PubMed:16510120}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - Retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | LETM1 | chr4:1817390 | chr4:1332861 | ENST00000302787 | - | 13 | 14 | 676_688 | 690.0 | 740.0 | Calcium binding | Ontology_term=ECO:0000255 |
| Hgene | LETM1 | chr4:1817390 | chr4:1332861 | ENST00000302787 | - | 13 | 14 | 115_136 | 690.0 | 740.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | LETM1 | chr4:1817390 | chr4:1332861 | ENST00000302787 | - | 13 | 14 | 462_490 | 690.0 | 740.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | LETM1 | chr4:1817390 | chr4:1332861 | ENST00000302787 | - | 13 | 14 | 537_627 | 690.0 | 740.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | LETM1 | chr4:1817390 | chr4:1332861 | ENST00000302787 | - | 13 | 14 | 18_21 | 690.0 | 740.0 | Compositional bias | Note=Poly-Pro |
| Hgene | LETM1 | chr4:1857595 | chr4:1326544 | ENST00000302787 | - | 1 | 14 | 18_21 | 27.333333333333332 | 740.0 | Compositional bias | Note=Poly-Pro |
| Hgene | LETM1 | chr4:1817390 | chr4:1332861 | ENST00000302787 | - | 13 | 14 | 252_537 | 690.0 | 740.0 | Domain | Letm1 RBD |
| Hgene | LETM1 | chr4:1817390 | chr4:1332861 | ENST00000302787 | - | 13 | 14 | 116_208 | 690.0 | 740.0 | Topological domain | Mitochondrial intermembrane |
| Hgene | LETM1 | chr4:1817390 | chr4:1332861 | ENST00000302787 | - | 13 | 14 | 209_229 | 690.0 | 740.0 | Transmembrane | Helical |
| Tgene | MAEA | chr4:1857595 | chr4:1326544 | ENST00000264750 | 3 | 8 | 314_381 | 177.66666666666666 | 356.0 | Zinc finger | RING-Gid-type | |
| Tgene | MAEA | chr4:1857595 | chr4:1326544 | ENST00000303400 | 4 | 9 | 314_381 | 218.66666666666666 | 397.0 | Zinc finger | RING-Gid-type |
| - Not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | LETM1 | chr4:1857595 | chr4:1326544 | ENST00000302787 | - | 1 | 14 | 676_688 | 27.333333333333332 | 740.0 | Calcium binding | Ontology_term=ECO:0000255 |
| Hgene | LETM1 | chr4:1817390 | chr4:1332861 | ENST00000302787 | - | 13 | 14 | 708_739 | 690.0 | 740.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | LETM1 | chr4:1857595 | chr4:1326544 | ENST00000302787 | - | 1 | 14 | 115_136 | 27.333333333333332 | 740.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | LETM1 | chr4:1857595 | chr4:1326544 | ENST00000302787 | - | 1 | 14 | 462_490 | 27.333333333333332 | 740.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | LETM1 | chr4:1857595 | chr4:1326544 | ENST00000302787 | - | 1 | 14 | 537_627 | 27.333333333333332 | 740.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | LETM1 | chr4:1857595 | chr4:1326544 | ENST00000302787 | - | 1 | 14 | 708_739 | 27.333333333333332 | 740.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | LETM1 | chr4:1817390 | chr4:1332861 | ENST00000302787 | - | 13 | 14 | 663_698 | 690.0 | 740.0 | Domain | EF-hand |
| Hgene | LETM1 | chr4:1857595 | chr4:1326544 | ENST00000302787 | - | 1 | 14 | 252_537 | 27.333333333333332 | 740.0 | Domain | Letm1 RBD |
| Hgene | LETM1 | chr4:1857595 | chr4:1326544 | ENST00000302787 | - | 1 | 14 | 663_698 | 27.333333333333332 | 740.0 | Domain | EF-hand |
| Hgene | LETM1 | chr4:1817390 | chr4:1332861 | ENST00000302787 | - | 13 | 14 | 230_739 | 690.0 | 740.0 | Topological domain | Mitochondrial matrix |
| Hgene | LETM1 | chr4:1857595 | chr4:1326544 | ENST00000302787 | - | 1 | 14 | 116_208 | 27.333333333333332 | 740.0 | Topological domain | Mitochondrial intermembrane |
| Hgene | LETM1 | chr4:1857595 | chr4:1326544 | ENST00000302787 | - | 1 | 14 | 230_739 | 27.333333333333332 | 740.0 | Topological domain | Mitochondrial matrix |
| Hgene | LETM1 | chr4:1857595 | chr4:1326544 | ENST00000302787 | - | 1 | 14 | 209_229 | 27.333333333333332 | 740.0 | Transmembrane | Helical |
| Tgene | MAEA | chr4:1817390 | chr4:1332861 | ENST00000264750 | 6 | 8 | 121_153 | 324.0 | 356.0 | Domain | LisH | |
| Tgene | MAEA | chr4:1817390 | chr4:1332861 | ENST00000264750 | 6 | 8 | 159_216 | 324.0 | 356.0 | Domain | CTLH | |
| Tgene | MAEA | chr4:1817390 | chr4:1332861 | ENST00000303400 | 7 | 9 | 121_153 | 365.0 | 397.0 | Domain | LisH | |
| Tgene | MAEA | chr4:1817390 | chr4:1332861 | ENST00000303400 | 7 | 9 | 159_216 | 365.0 | 397.0 | Domain | CTLH | |
| Tgene | MAEA | chr4:1857595 | chr4:1326544 | ENST00000264750 | 3 | 8 | 121_153 | 177.66666666666666 | 356.0 | Domain | LisH | |
| Tgene | MAEA | chr4:1857595 | chr4:1326544 | ENST00000264750 | 3 | 8 | 159_216 | 177.66666666666666 | 356.0 | Domain | CTLH | |
| Tgene | MAEA | chr4:1857595 | chr4:1326544 | ENST00000303400 | 4 | 9 | 121_153 | 218.66666666666666 | 397.0 | Domain | LisH | |
| Tgene | MAEA | chr4:1857595 | chr4:1326544 | ENST00000303400 | 4 | 9 | 159_216 | 218.66666666666666 | 397.0 | Domain | CTLH | |
| Tgene | MAEA | chr4:1817390 | chr4:1332861 | ENST00000264750 | 6 | 8 | 1_124 | 324.0 | 356.0 | Region | Extracellular and involved in cell to cell contact | |
| Tgene | MAEA | chr4:1817390 | chr4:1332861 | ENST00000303400 | 7 | 9 | 1_124 | 365.0 | 397.0 | Region | Extracellular and involved in cell to cell contact | |
| Tgene | MAEA | chr4:1857595 | chr4:1326544 | ENST00000264750 | 3 | 8 | 1_124 | 177.66666666666666 | 356.0 | Region | Extracellular and involved in cell to cell contact | |
| Tgene | MAEA | chr4:1857595 | chr4:1326544 | ENST00000303400 | 4 | 9 | 1_124 | 218.66666666666666 | 397.0 | Region | Extracellular and involved in cell to cell contact | |
| Tgene | MAEA | chr4:1817390 | chr4:1332861 | ENST00000264750 | 6 | 8 | 314_381 | 324.0 | 356.0 | Zinc finger | RING-Gid-type | |
| Tgene | MAEA | chr4:1817390 | chr4:1332861 | ENST00000303400 | 7 | 9 | 314_381 | 365.0 | 397.0 | Zinc finger | RING-Gid-type |
Top |
Fusion Protein-Protein Interaction |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type from validated records (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type from validated records (BIOGRID-3.4.160) |
| Gene | PPI interactors |
 Protein-protein interactors based on sequence similarity (STRING) Protein-protein interactors based on sequence similarity (STRING) |
| Gene | STRING network |
| LETM1 | |
| MAEA |
 - Retained interactions in fusion protein (protein functional feature from UniProt). - Retained interactions in fusion protein (protein functional feature from UniProt). |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost interactions due to fusion (protein functional feature from UniProt). - Lost interactions due to fusion (protein functional feature from UniProt). |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs to LETM1-MAEA |
 Drugs used for this fusion-positive patient. Drugs used for this fusion-positive patient. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Drug | Source | PMID |
Top |
Related Diseases to LETM1-MAEA |
 Diseases that have this fusion gene. Diseases that have this fusion gene. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Disease | Source | PMID |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |

