| UTHEALTH HOME ABOUT SBMI A-Z WEBMAIL INSIDE THE UNIVERSITY |

|
|||||||
|
Fusion Protein:RPL7A-NTRK1 |
Fusion Protein Summary |
 Fusion gene summary Fusion gene summary |
| Fusion partner gene information | Fusion gene name: RPL7A-NTRK1 | FusionPDB ID: 76971 | FusionGDB2.0 ID: 76971 | Hgene | Tgene | Gene symbol | RPL7A | NTRK1 | Gene ID | 6130 | 4914 |
| Gene name | ribosomal protein L7a | neurotrophic receptor tyrosine kinase 1 | |
| Synonyms | L7A|SURF3|TRUP | MTC|TRK|TRK1|TRKA|Trk-A|p140-TrkA | |
| Cytomap | 9q34.2 | 1q23.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | 60S ribosomal protein L7aPLA-X polypeptidelarge ribosomal subunit protein eL8surfeit 3surfeit locus protein 3thyroid hormone receptor uncoupling protein | high affinity nerve growth factor receptorOncogene TRKTRK1-transforming tyrosine kinase proteingp140trkneurotrophic tyrosine kinase, receptor, type 1tropomyosin-related kinase Atyrosine kinase receptor A | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | P04629 | |
| Ensembl transtripts involved in fusion gene | ENST ids | ENST00000463740, ENST00000323345, ENST00000315731, | ENST00000358660, ENST00000368196, ENST00000392302, ENST00000524377, ENST00000531606, |
| Fusion gene scores for assessment (based on all fusion genes of FusionGDB 2.0) | * DoF score | 9 X 9 X 2=162 | 25 X 24 X 13=7800 |
| # samples | 18 | 33 | |
| ** MAII score | log2(18/162*10)=0.15200309344505 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | log2(33/7800*10)=-4.56293619439116 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context (manual curation of fusion genes in FusionPDB) | PubMed: RPL7A [Title/Abstract] AND NTRK1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint (based on all fusion genes of FusionGDB 2.0) | RPL7A(136215897)-NTRK1(156844362), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | RPL7A-NTRK1 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. RPL7A-NTRK1 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. RPL7A-NTRK1 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. RPL7A-NTRK1 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | NTRK1 | GO:0006468 | protein phosphorylation | 15488758 |
| Tgene | NTRK1 | GO:0008285 | negative regulation of cell proliferation | 15488758 |
| Tgene | NTRK1 | GO:0010976 | positive regulation of neuron projection development | 15488758 |
| Tgene | NTRK1 | GO:0018108 | peptidyl-tyrosine phosphorylation | 2927393 |
| Tgene | NTRK1 | GO:0043547 | positive regulation of GTPase activity | 15488758 |
| Tgene | NTRK1 | GO:0046579 | positive regulation of Ras protein signal transduction | 15488758 |
| Tgene | NTRK1 | GO:0046777 | protein autophosphorylation | 15488758 |
| Tgene | NTRK1 | GO:0048011 | neurotrophin TRK receptor signaling pathway | 15488758 |
| Tgene | NTRK1 | GO:0051092 | positive regulation of NF-kappaB transcription factor activity | 15488758 |
| Tgene | NTRK1 | GO:0070374 | positive regulation of ERK1 and ERK2 cascade | 15488758 |
| Tgene | NTRK1 | GO:1904646 | cellular response to amyloid-beta | 11927634 |
 Fusion gene breakpoints across RPL7A (5'-gene) Fusion gene breakpoints across RPL7A (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
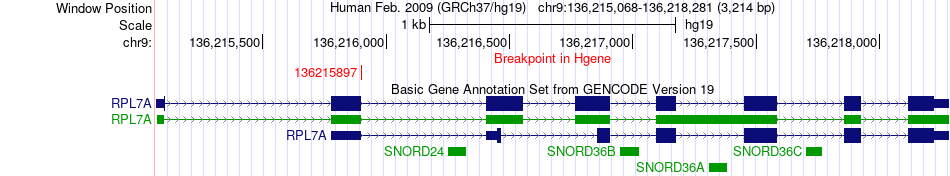 |
 Fusion gene breakpoints across NTRK1 (3'-gene) Fusion gene breakpoints across NTRK1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
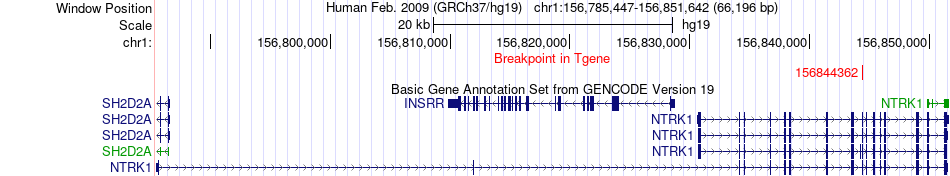 |
Top |
Fusion Gene Sample Information |
 Fusion gene information from FusionGDB2.0. Fusion gene information from FusionGDB2.0. |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChiTaRS5.0 | N/A | X06704 | RPL7A | chr9 | 136215897 | + | NTRK1 | chr1 | 156844362 | + |
Top |
Fusion ORF Analysis |
 Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. Fusion information from ORFfinder translation from full-length transcript sequence from FusionPDB. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000323345 | RPL7A | chr9 | 136215897 | + | ENST00000392302 | NTRK1 | chr1 | 156844362 | + | 1502 | 154 | 30 | 1349 | 439 |
| ENST00000323345 | RPL7A | chr9 | 136215897 | + | ENST00000368196 | NTRK1 | chr1 | 156844362 | + | 1558 | 154 | 30 | 1349 | 439 |
| ENST00000323345 | RPL7A | chr9 | 136215897 | + | ENST00000524377 | NTRK1 | chr1 | 156844362 | + | 1350 | 154 | 30 | 1349 | 439 |
| ENST00000323345 | RPL7A | chr9 | 136215897 | + | ENST00000358660 | NTRK1 | chr1 | 156844362 | + | 1428 | 154 | 30 | 1358 | 442 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000323345 | ENST00000392302 | RPL7A | chr9 | 136215897 | + | NTRK1 | chr1 | 156844362 | + | 0.021238165 | 0.9787618 |
| ENST00000323345 | ENST00000368196 | RPL7A | chr9 | 136215897 | + | NTRK1 | chr1 | 156844362 | + | 0.019550415 | 0.98044956 |
| ENST00000323345 | ENST00000524377 | RPL7A | chr9 | 136215897 | + | NTRK1 | chr1 | 156844362 | + | 0.024425851 | 0.97557414 |
| ENST00000323345 | ENST00000358660 | RPL7A | chr9 | 136215897 | + | NTRK1 | chr1 | 156844362 | + | 0.01601388 | 0.98398614 |
Top |
Fusion Amino Acid Sequences |
 For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. For individual full-length fusion transcript sequence from FusionPDB, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >FusionGDB ID_FusionGDB isoform ID_FGname_Hgene_Hchr_Hbp_Henst_Tgene_Tchr_Tbp_Tenst_length(fusion AA) seq_BP >76971_76971_1_RPL7A-NTRK1_RPL7A_chr9_136215897_ENST00000323345_NTRK1_chr1_156844362_ENST00000358660_length(amino acids)=442AA_BP=41 MPKGKKAKGKKVAPAPAVVKKQEAKKVVNPLFEKRPKNFGIDTNSTSGDPVEKKDETPFGVSVAVGLAVFACLFLSTLLLVLNKCGRRNK FGINRPAVLAPEDGLAMSLHFMTLGGSSLSPTEGKGSGLQGHIIENPQYFSDASPSGVHHIKRRDIVLKWELGEGAFGKVFLAECHNLLP EQDKMLVAVKALKEASESARQDFQREAELLTMLQHQHIVRFFGVCTEGRPLLMVFEYMRHGDLNRFLRSHGPDAKLLAGGEDVAPGPLGL GQLLAVASQVAAGMVYLAGLHFVHRDLATRNCLVGQGLVVKIGDFGMSRDIYSTDYYRVGGRTMLPIRWMPPESILYRKFTTESDVWSFG -------------------------------------------------------------- >76971_76971_2_RPL7A-NTRK1_RPL7A_chr9_136215897_ENST00000323345_NTRK1_chr1_156844362_ENST00000368196_length(amino acids)=439AA_BP=41 MPKGKKAKGKKVAPAPAVVKKQEAKKVVNPLFEKRPKNFGIDTNSTSGDPVEKKDETPFGVSVAVGLAVFACLFLSTLLLVLNKCGRRNK FGINRPAVLAPEDGLAMSLHFMTLGGSSLSPTEGKGSGLQGHIIENPQYFSDACVHHIKRRDIVLKWELGEGAFGKVFLAECHNLLPEQD KMLVAVKALKEASESARQDFQREAELLTMLQHQHIVRFFGVCTEGRPLLMVFEYMRHGDLNRFLRSHGPDAKLLAGGEDVAPGPLGLGQL LAVASQVAAGMVYLAGLHFVHRDLATRNCLVGQGLVVKIGDFGMSRDIYSTDYYRVGGRTMLPIRWMPPESILYRKFTTESDVWSFGVVL -------------------------------------------------------------- >76971_76971_3_RPL7A-NTRK1_RPL7A_chr9_136215897_ENST00000323345_NTRK1_chr1_156844362_ENST00000392302_length(amino acids)=439AA_BP=41 MPKGKKAKGKKVAPAPAVVKKQEAKKVVNPLFEKRPKNFGIDTNSTSGDPVEKKDETPFGVSVAVGLAVFACLFLSTLLLVLNKCGRRNK FGINRPAVLAPEDGLAMSLHFMTLGGSSLSPTEGKGSGLQGHIIENPQYFSDACVHHIKRRDIVLKWELGEGAFGKVFLAECHNLLPEQD KMLVAVKALKEASESARQDFQREAELLTMLQHQHIVRFFGVCTEGRPLLMVFEYMRHGDLNRFLRSHGPDAKLLAGGEDVAPGPLGLGQL LAVASQVAAGMVYLAGLHFVHRDLATRNCLVGQGLVVKIGDFGMSRDIYSTDYYRVGGRTMLPIRWMPPESILYRKFTTESDVWSFGVVL -------------------------------------------------------------- >76971_76971_4_RPL7A-NTRK1_RPL7A_chr9_136215897_ENST00000323345_NTRK1_chr1_156844362_ENST00000524377_length(amino acids)=439AA_BP=41 MPKGKKAKGKKVAPAPAVVKKQEAKKVVNPLFEKRPKNFGIDTNSTSGDPVEKKDETPFGVSVAVGLAVFACLFLSTLLLVLNKCGRRNK FGINRPAVLAPEDGLAMSLHFMTLGGSSLSPTEGKGSGLQGHIIENPQYFSDACVHHIKRRDIVLKWELGEGAFGKVFLAECHNLLPEQD KMLVAVKALKEASESARQDFQREAELLTMLQHQHIVRFFGVCTEGRPLLMVFEYMRHGDLNRFLRSHGPDAKLLAGGEDVAPGPLGLGQL LAVASQVAAGMVYLAGLHFVHRDLATRNCLVGQGLVVKIGDFGMSRDIYSTDYYRVGGRTMLPIRWMPPESILYRKFTTESDVWSFGVVL -------------------------------------------------------------- |
Top |
Fusion Protein Functional Features |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr9:136215897/chr1:156844362) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
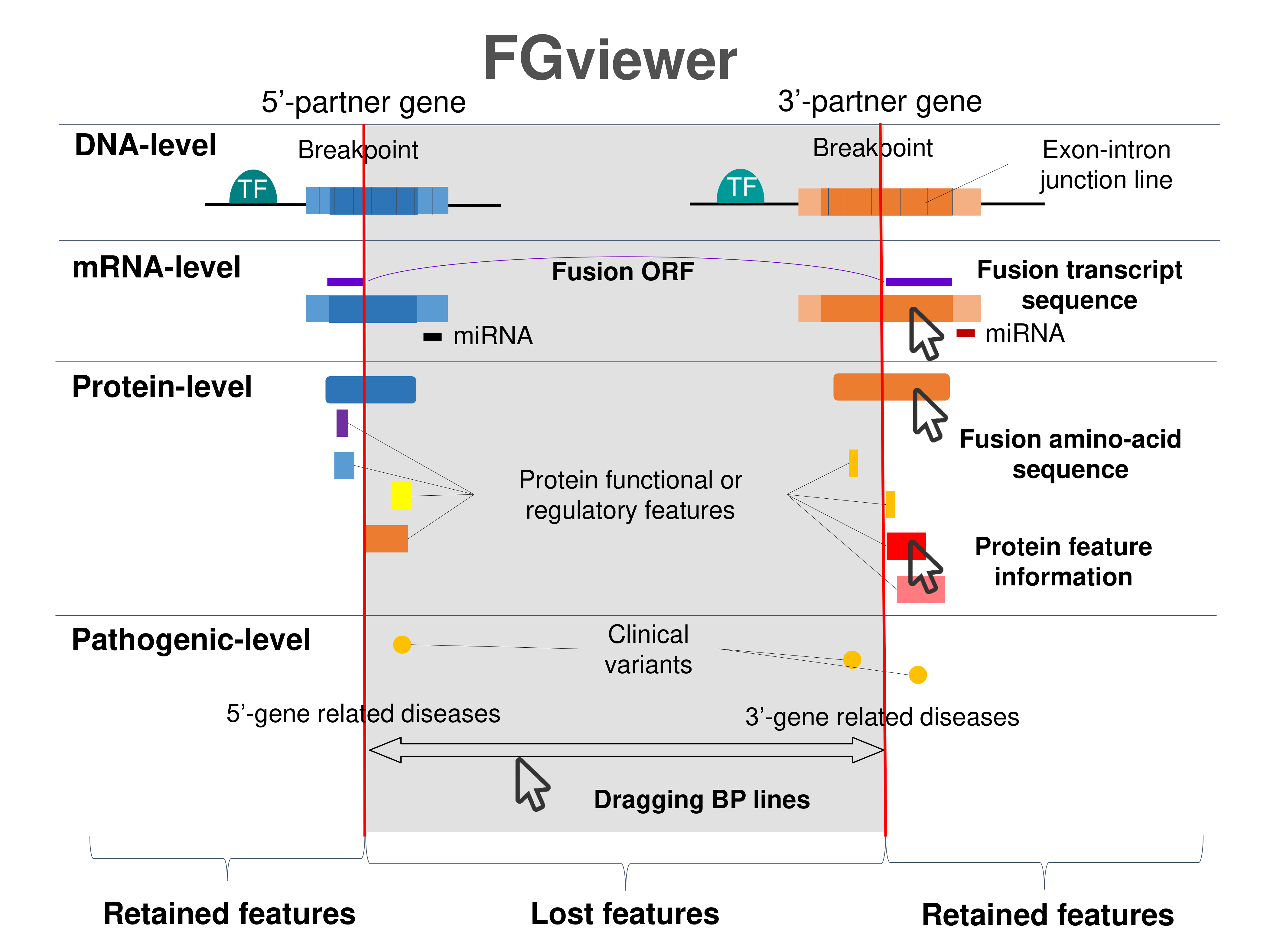 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | NTRK1 |
| FUNCTION: Might normally function as a transcriptional repressor. EWS-fusion-proteins (EFPS) may play a role in the tumorigenic process. They may disturb gene expression by mimicking, or interfering with the normal function of CTD-POLII within the transcription initiation complex. They may also contribute to an aberrant activation of the fusion protein target genes. | FUNCTION: Receptor tyrosine kinase involved in the development and the maturation of the central and peripheral nervous systems through regulation of proliferation, differentiation and survival of sympathetic and nervous neurons. High affinity receptor for NGF which is its primary ligand (PubMed:1850821, PubMed:1849459, PubMed:1281417, PubMed:8325889, PubMed:15488758, PubMed:22649032, PubMed:17196528, PubMed:27445338). Can also bind and be activated by NTF3/neurotrophin-3. However, NTF3 only supports axonal extension through NTRK1 but has no effect on neuron survival (By similarity). Upon dimeric NGF ligand-binding, undergoes homodimerization, autophosphorylation and activation (PubMed:1281417). Recruits, phosphorylates and/or activates several downstream effectors including SHC1, FRS2, SH2B1, SH2B2 and PLCG1 that regulate distinct overlapping signaling cascades driving cell survival and differentiation. Through SHC1 and FRS2 activates a GRB2-Ras-MAPK cascade that regulates cell differentiation and survival. Through PLCG1 controls NF-Kappa-B activation and the transcription of genes involved in cell survival. Through SHC1 and SH2B1 controls a Ras-PI3 kinase-AKT1 signaling cascade that is also regulating survival. In absence of ligand and activation, may promote cell death, making the survival of neurons dependent on trophic factors. {ECO:0000250|UniProtKB:P35739, ECO:0000250|UniProtKB:Q3UFB7, ECO:0000269|PubMed:11244088, ECO:0000269|PubMed:1281417, ECO:0000269|PubMed:15488758, ECO:0000269|PubMed:17196528, ECO:0000269|PubMed:1849459, ECO:0000269|PubMed:1850821, ECO:0000269|PubMed:22649032, ECO:0000269|PubMed:27445338, ECO:0000269|PubMed:27676246, ECO:0000269|PubMed:8155326, ECO:0000269|PubMed:8325889}.; FUNCTION: [Isoform TrkA-III]: Resistant to NGF, it constitutively activates AKT1 and NF-kappa-B and is unable to activate the Ras-MAPK signaling cascade. Antagonizes the anti-proliferative NGF-NTRK1 signaling that promotes neuronal precursors differentiation. Isoform TrkA-III promotes angiogenesis and has oncogenic activity when overexpressed. {ECO:0000269|PubMed:15488758}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - Retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000368196 | 7 | 16 | 510_781 | 392.3333333333333 | 791.0 | Domain | Protein kinase | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000392302 | 8 | 17 | 510_781 | 362.3333333333333 | 761.0 | Domain | Protein kinase | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000524377 | 8 | 17 | 510_781 | 398.3333333333333 | 797.0 | Domain | Protein kinase | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000368196 | 7 | 16 | 537_541 | 392.3333333333333 | 791.0 | Motif | DXXLL | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000368196 | 7 | 16 | 607_611 | 392.3333333333333 | 791.0 | Motif | DXXLL | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000392302 | 8 | 17 | 537_541 | 362.3333333333333 | 761.0 | Motif | DXXLL | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000392302 | 8 | 17 | 607_611 | 362.3333333333333 | 761.0 | Motif | DXXLL | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000524377 | 8 | 17 | 537_541 | 398.3333333333333 | 797.0 | Motif | DXXLL | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000524377 | 8 | 17 | 607_611 | 398.3333333333333 | 797.0 | Motif | DXXLL | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000368196 | 7 | 16 | 516_524 | 392.3333333333333 | 791.0 | Nucleotide binding | ATP | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000392302 | 8 | 17 | 516_524 | 362.3333333333333 | 761.0 | Nucleotide binding | ATP | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000524377 | 8 | 17 | 516_524 | 398.3333333333333 | 797.0 | Nucleotide binding | ATP | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000368196 | 7 | 16 | 440_796 | 392.3333333333333 | 791.0 | Topological domain | Cytoplasmic | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000392302 | 8 | 17 | 440_796 | 362.3333333333333 | 761.0 | Topological domain | Cytoplasmic | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000524377 | 8 | 17 | 440_796 | 398.3333333333333 | 797.0 | Topological domain | Cytoplasmic | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000368196 | 7 | 16 | 424_439 | 392.3333333333333 | 791.0 | Transmembrane | Helical | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000392302 | 8 | 17 | 424_439 | 362.3333333333333 | 761.0 | Transmembrane | Helical | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000524377 | 8 | 17 | 424_439 | 398.3333333333333 | 797.0 | Transmembrane | Helical |
| - Not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000368196 | 7 | 16 | 148_193 | 392.3333333333333 | 791.0 | Domain | Note=LRRCT | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000368196 | 7 | 16 | 194_283 | 392.3333333333333 | 791.0 | Domain | Ig-like C2-type 1 | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000368196 | 7 | 16 | 299_365 | 392.3333333333333 | 791.0 | Domain | Ig-like C2-type 2 | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000392302 | 8 | 17 | 148_193 | 362.3333333333333 | 761.0 | Domain | Note=LRRCT | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000392302 | 8 | 17 | 194_283 | 362.3333333333333 | 761.0 | Domain | Ig-like C2-type 1 | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000392302 | 8 | 17 | 299_365 | 362.3333333333333 | 761.0 | Domain | Ig-like C2-type 2 | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000524377 | 8 | 17 | 148_193 | 398.3333333333333 | 797.0 | Domain | Note=LRRCT | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000524377 | 8 | 17 | 194_283 | 398.3333333333333 | 797.0 | Domain | Ig-like C2-type 1 | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000524377 | 8 | 17 | 299_365 | 398.3333333333333 | 797.0 | Domain | Ig-like C2-type 2 | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000368196 | 7 | 16 | 116_137 | 392.3333333333333 | 791.0 | Repeat | Note=LRR 2 | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000368196 | 7 | 16 | 90_113 | 392.3333333333333 | 791.0 | Repeat | Note=LRR 1 | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000392302 | 8 | 17 | 116_137 | 362.3333333333333 | 761.0 | Repeat | Note=LRR 2 | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000392302 | 8 | 17 | 90_113 | 362.3333333333333 | 761.0 | Repeat | Note=LRR 1 | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000524377 | 8 | 17 | 116_137 | 398.3333333333333 | 797.0 | Repeat | Note=LRR 2 | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000524377 | 8 | 17 | 90_113 | 398.3333333333333 | 797.0 | Repeat | Note=LRR 1 | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000368196 | 7 | 16 | 33_423 | 392.3333333333333 | 791.0 | Topological domain | Extracellular | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000392302 | 8 | 17 | 33_423 | 362.3333333333333 | 761.0 | Topological domain | Extracellular | |
| Tgene | NTRK1 | chr9:136215897 | chr1:156844362 | ENST00000524377 | 8 | 17 | 33_423 | 398.3333333333333 | 797.0 | Topological domain | Extracellular |
Top |
Fusion Protein-Protein Interaction |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type from validated records (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type from validated records (BIOGRID-3.4.160) |
| Gene | PPI interactors |
| NTRK1 | ABL1, NGF, MATK, PLCG1, GRB2, CAV1, NTF3, SQSTM1, SH2B1, SHC1, SH2B2, ARHGAP32, FRS2, NTRK1, RAP1A, SHC3, GIPC1, RGS19, PTPN1, TRAF6, NEDD4, CBL, HSP90AA1, HSPA4, CDC37, NEDD4L, MYC, SGSM3, SORBS1, TNK2, SNX17, SNX27, GAREML, SOS2, PIK3CA, GAREM, PIK3C2B, ARHGEF40, PIK3R1, PIK3R2, SOS1, PIK3CB, PML, UBC, AMFR, CBLB, GAB1, EPN1, PIK3C2A, ANKRD13A, DOCK4, TRIO, LGR5, WRNIP1, RABEP1, PDGFRA, KANK2, RABGEF1, VAV2, TBC1D15, PTPN11, GAK, IRS2, PCDHGC3, EPS15, LRRN3, HUWE1, ACVR1B, MIB1, OFD1, PIBF1, ZNF497, TXN, CAT, FABP3, JUP, DSG1, TGM1, CFL1, LCN1, HIST2H4A, DSP, PFN1, LGALS3BP, HIST1H2BL, LTF, S100A9, IGLL5, TGM3, DSC1, LYZ, DCD, LEKR1, ENO2, PPIA, BTN1A1, XRCC5, XRCC6, HLA-A, GSTP1, ABCB7, PA2G4, PRDX1, HDGF, HSPA4L, TBC1D10B, HP1BP3, KIF26B, SYT4, RHOBTB3, NOTCH3, GAB2, ATAD3A, RBM15B, KCNH2, USP15, PDCD11, RBFOX2, RBM14, RNF31, ADNP, ZC3HAV1, MAP7D1, USP4, JAK1, PTPRS, PDGFRB, HIST1H1C, USP11, PPFIBP1, INTS6, VGF, ARHGEF11, SMPD4, SYNPO2, HNRNPM, SLC27A4, DDX41, EGFR, H1FX, ASAP1, HIST1H1B, RRBP1, EPHA2, KIDINS220, CPT1A, UNC5B, IGF1R, DDR2, AUP1, XPR1, RP9, MARK2, REPS1, EMD, SMARCA4, DDX21, DAB2IP, C7, MSTO1, CDC27, ZNF629, ASAP2, NCAPH2, SRBD1, ATAD3B, TCF12, PLEKHA6, INPPL1, PRPF4B, DPM1, SMARCA1, FIP1L1, DLGAP5, PLEKHH3, TRA2A, ACIN1, DDX10, TAX1BP1, SMARCA5, RABEP2, DDX5, MYO1C, SRSF10, DNAJA2, RET, TRA2B, MYEF2, PNPLA6, SLC12A2, TAF6L, RSL1D1, ARAP3, DDX18, ABL2, KHDRBS1, CERK, SRSF7, SPTLC2, PBXIP1, DNAJA1, IQGAP3, DDX31, ZC3H11A, PLRG1, CNP, FMR1, HIP1R, KIAA1429, CDC5L, ARAP1, ZNF316, SMARCC2, EML5, VANGL2, SLC25A1, UNC5C, SLC16A1, RFC1, ZKSCAN1, CHD1, PTK7, SMARCD2, AP3B1, YTHDC2, AKAP8L, RPL15, CTNND1, PHLDB1, ZNF574, DDX27, TOM1, RASA1, MORC2, MTCH2, IMMT, SMARCC1, ROR2, CAPN2, DNAJC11, SYT11, AP2A1, LBR, MEPCE, NCAPD3, CEP97, QKI, LRRC41, LPCAT2, EPS15L1, LAS1L, TMEM131, CWF19L2, NCAPG2, MAP4K5, CCP110, ARHGEF2, ITGAV, ZC3H18, CWC22, VIM, FAM171A1, TMEM132A, SRSF9, SOGA1, NKRF, RBL1, FMNL3, TOP1, PIGT, RBM15, GBAS, EDC3, WDR5, NOP2, GNL2, SLC25A11, RRP1B, FAM115A, INTS5, SLC25A5, AAK1, HERC5, CCNT1, FERMT2, CDC42BPA, MRPL28, UBAP2, WWP2, SEC24B, HNRNPU, GPD2, FXR2, DHX15, SLC25A22, SSR1, PLXNB2, PROX1, TEX10, USP19, FTSJ3, TMEM59, GTF3C2, PGAM5, CPSF1, KLHL22, TMX3, SIN3A, NOP58, PTCD3, RPRD2, DHX9, PICALM, FARP1, SLC25A6, ERLIN2, SMARCD1, RHOT1, ALDH1B1, SGPL1, USP9X, HNRNPF, KIAA1211, MAPK7, DDX17, GSE1, DVL3, MRPL44, DNAJB6, GTPBP4, HNRNPA0, PLXNA4, SFXN3, NAT10, DDX28, SNW1, MYO6, MRPL24, PRRT4, FAM98A, PURA, DHX29, C1QBP, TRIP4, TBK1, RBM28, ZNF609, POLRMT, PCDHB3, XRN1, HSD17B11, DKC1, DHX30, CDK13, SENP3, LPHN2, GPATCH4, NDUFA9, RASA4, MCCC1, DDX24, RPL14, RPS8, ATG2B, MYO1B, PTPN12, CEBPZ, ILVBL, RALY, KIF14, ZC3H7A, MOV10, MAP4K4, DHX57, RBMX, FXR1, MRPS35, SART1, MARK3, DECR1, CCDC88A, ANKLE2, PRPF31, LARP7, RPL10, PACS1, RPL13, FAM120A, SLC25A13, ADAR, CDK1, HNRNPR, CSNK1E, DLG1, LEMD3, NOL9, IFT122, NCLN, STK11IP, GATAD2B, CHERP, LARP4, MAP3K1, HNRNPUL2, PPIL2, ESYT2, MYH10, RPL8, UQCRC2, NDUFA10, HAX1, HELLS, MED14, RBM42, CHCHD3, RPL7, ABCD3, TUBGCP6, MRPS9, SNRNP200, IGF2BP3, LRCH3, MRPL3, RSBN1L, SAFB, TUBG1, ALDH16A1, ARHGEF12, EPB41L5, AP2A2, SRSF6, HLTF, PTPLAD1, TMPO, AGO2, POP1, RPS9, HDAC6, ILF3, VWA8, ABCB1, SCRIB, GTF3C5, YBX1, RBM4, PTPN9, PRPF8, BRD3, TRIM24, CLASP2, MADD, TCF4, MAP4K3, TSC2, DDX23, RPL7A, RRP12, PRKD2, TRMT1L, EDC4, CSTF3, RPS2, NDUFS2, RPL13A, PTPRR, ATP7A, EXD2, IK, HIP1, MATR3, IGSF3, PTPRF, SPATA5L1, EFTUD2, RPS3A, TUBGCP5, TTN, GIT1, PATL1, NUP160, PDE4B, SUN2, PTPN23, RPS6, DCP1A, CDC73, MEX3A, EIF2AK2, RPL3, PPP1R10, SRP72, ELAVL3, NOM1, RPL18, CEP170, THRAP3, MED17, MRPS7, DDX20, PDPR, SH3PXD2A, DBN1, DAP3, OAS3, STT3B, COPE, MED12, AP3D1, CAPRIN1, PHB2, U2SURP, BSG, TECR, FARSA, HSD17B12, RPL4, HNRNPUL1, PPFIA1, MRPS22, TBL3, GNL3, MTA2, CPSF2, ATP2B1, AP3B2, PPIP5K2, DDX56, ASCC3, FMNL2, SMCHD1, MYBBP1A, SRPK2, RHOT2, SRP68, PUM1, SLC25A3, SRPK1, RPL19, BEND3, SEC61A1, LSG1, NOC3L, TNKS1BP1, GTF3C3, TUBGCP3, PRPF40A, TRMT10C, URGCP, XRN2, MSH3, XRCC1, GEMIN4, DIS3L2, BCLAF1, SBF2, DHTKD1, ATXN2, ARID4B, TOR1AIP1, RPL29, NAA16, PRPF6, KIFAP3, TMEM33, WASF2, NACAD, LIG3, POLE, CKAP4, TCEB3, MTA1, GNA13, DHX38, PHF6, ELAVL4, DDX3X, CTR9, PRPF19, CAMSAP2, PRMT3, SH3PXD2B, SCAF8, MAGED1, ILF2, TONSL, PPP1R12A, SRSF5, KIAA0020, PTBP1, CAD, ACSL3, CHD1L, STAU1, MRPL38, DVL2, GNAI3, RPL10A, RPL6, MRPS5, CDK11B, MRPS34, PES1, ATP5C1, STXBP3, HNRNPLL, CHTF18, VPS39, SART3, COPA, PRPF3, UBE2O, ABCF2, HNRNPH3, EARS2, DHX40, ARHGEF7, EMC1, CASK, PTK2, SORT1, OGT, ATP12A, LMNB1, SLC4A7, PFKP, VDAC2, RBM25, ATL3, DHX36, TFIP11, EXOSC10, GTF3C1, SRSF1, PPP6C, TRAPPC9, RAD21, DNAJC13, ACOT1, ZC3H4, SRRM1, NDUFS3, DYRK1A, INTS1, SF3B2, EHHADH, MYO9B, ASCC2, WDR11, RPN1, EHD4, WDR91, SMARCAL1, NBAS, PI4KA, PAF1, EIF2B4, NUP107, EMC2, SYNE1, KLHL13, TBL1XR1, DNAJC10, SEC16A, METTL13, C7orf50, CDC42BPB, CLCN5, PGM3, DOCK7, ALYREF, POLR3A, TOP3A, WDR48, WDR81, DOCK1, CSNK2A2, CYC1, TLE3, THOC2, EIF2S3, MYO5A, SNX9, TARDBP, NAP1L4, CRKL, YTHDF2, PYGB, PABPC1, LTV1, PSMD4, KTN1, UBTF, WDR6, FUBP3, UBXN7, PHB, PDXP, CNOT3, TBL2, LLGL1, PLOD1, AHSA1, AAAS, ERGIC1, LARP4B, CYFIP1, DNM1, TARS2, ACSL1, ATXN1L, WDR59, SUCLA2, MLH1, CSNK2B, SYNCRIP, WDR36, FAM91A1, MTDH, UBE3C, CAPN1, BRIX1, DNAJB11, EDEM3, PLOD3, CLASP1, DPYSL3, MED16, NONO, EPM2AIP1, NT5DC2, ADRBK2, RPTOR, TELO2, MRPL37, PRKACB, ELMO1, TRAPPC8, ANAPC1, NIPSNAP1, EWSR1, PIK3R4, POLD1, IQGAP1, PRRC2A, COG1, ANKRD44, TFRC, DIEXF, FHOD1, SERPINH1, SPATA5, MED1, ABCF3, ZSWIM8, LMNA, HAT1, PABPC4, SMC4, HADH, RNGTT, TUBGCP2, ZNF598, SKIV2L2, GNB2L1, LARP1, DIP2B, BCR, IDH3B, LSM14A, FBL, NDUFS1, APPL2, USP39, SMC2, HCFC1, LANCL1, POLR2B, UPF2, CCBL2, MRPL15, MAGED2, AGK, GNPAT, AIMP1, GTF3C4, KDM1A, SCAMP3, RDH11, SRGAP2, CRNKL1, RFC5, PFKL, SPTBN1, PHOX2A, VDAC3, HNRNPA1, WDR26, DNM2, HAUS5, LCMT2, MCM7, CLINT1, UGDH, ACO1, TRIM46, PHKB, USP10, PCF11, YTHDF3, AARS2, FIGNL1, SFPQ, ATP5A1, NUP133, CPSF3, ANKRD28, PNN, C9orf64, ABCF1, ATL1, NAP1L1, HSPA9, MCCC2, FKBP5, CNOT1, TAF6, NEMF, UHRF1, SDHA, PELP1, HADHB, OAT, NEK9, AP3M1, PRKCA, LPCAT1, MRPL39, AFG3L2, WASF1, HNRNPL, RALGAPB, HK1, GBA, PLD3, CPSF7, HADHA, PDE12, DCX, LIMA1, HNRNPA2B1, POLR2A, MYH9, ZNF207, DDX55, ELP3, EIF2B5, MICAL1, TTI1, NES, HSDL1, GPSM1, PPM1F, C4orf27, SRRM2, OSBPL9, STAT3, IRF2BPL, KLC2, P3H1, STAT5B, UQCRC1, PCK2, CCAR2, TBC1D5, SACM1L, QARS, XPO4, CHD4, KIF2A, TACC3, NCL, BTAF1, RARS2, NCKAP1, EIF3D, RPS3, YME1L1, EIF4A3, HAUS6, HNRNPDL, MON2, G3BP2, WDHD1, CSNK1A1, SDAD1, RPS25, LSM14B, AP1M1, CCDC47, HNRNPK, DNMT1, NUP85, CUL5, TANC1, STAT1, COG5, STOML2, CDC45, OSBPL11, CIC, ABCD2, ARFGEF1, TROVE2, GBF1, SRPRB, UBE3A, GTPBP1, SCCPDH, RPS4X, PRPF4, ACTL6A, CCAR1, KIAA1033, TRAPPC12, HNRNPA3, FASTKD5, LYN, VPS18, PGRMC2, NUP214, AIFM1, TSG101, PPP6R3, OCRL, GANAB, IARS, SARS2, ATP2A2, EXOC1, NCAPH, TTC27, GNE, SRC, MIOS, ZCCHC8, GOLGA2, CARM1, ABCC1, PRMT5, LEO1, LMNB2, USP8, DARS, GYS1, APEH, ZYG11B, AKAP1, ELAVL1, MCM5, LRWD1, PSMC1, MOGS, TRIM28, EXOC6, LUC7L3, ANKFY1, NSUN2, HDAC2, G3BP1, QRICH1, INTS3, UBR1, XAB2, DYNC1LI1, TUFM, GAA, EXOC8, DHX8, LIMCH1, CLTC, SRPR, RNH1, ABI2, AQR, PREP, PPM1E, COG2, SEPT9, IDH2, MFN2, EXOC4, VPS33B, INTS4, GNB1, MSH6, HNRNPC, ERLIN1, EPRS, LRRC49, RPAP1, MCM3, PSMC4, PSMD3, GNL1, TRIP13, CUX1, PFAS, MAP1S, LRCH1, MRPL19, ABCD1, EIF3F, PDS5B, SMC3, KIF4A, FANCD2, IRF2BP1, SMC1A, RNF20, CLPX, NOC2L, DLAT, CTBP1, APPL1, NAA35, VPS45, PUM2, RELA, NUP210, RANBP9, DNAJC9, WRN, PRKAA1, AIMP2, VPS8, MAD1L1, MRE11A, ATP13A1, ATG7, CEP131, TRIM33, SLC25A10, HERC4, LRRFIP1, POLR1C, IPO9, GEMIN5, METAP2, SMG8, WASL, PPP1CB, SCFD1, RANBP3, RB1, CLTA, RNF40, SNRNP70, NCSTN, MGEA5, NARS, EIF4E, INPP4A, TFB2M, SEL1L, ARGLU1, PPP4R1, ANKRD17, TTK, RAD50, KIF2C, GOLGA3, TRAPPC11, GIGYF2, MSI2, CYFIP2, CDK5, HBS1L, IFT81, UFL1, PFKM, SMARCAD1, WDR35, PRIM2, NAMPT, PSMD2, MSH2, SOGA3, HEATR6, SFXN1, ARCN1, OGDH, FDPS, PSMC5, INSR, IRF2BP2, DDOST, ATXN2L, CSNK2A1, EIF2S2, MAP4, LTN1, COPB1, BRAT1, SMC6, SNRPN, RIC8A, SKIV2L, EIF4A1, KIF15, CLPTM1L, KPNA1, VPS16, VPS26B, RNPEP, RBM39, PMS1, FAM129B, RFC3, NUP155, RBM26, KIF21A, GAPVD1, C2CD5, WAPAL, RPS13, ARHGEF1, ACTR1A, KLC4, ISLR2, DCK, LRRC8A, STT3A, BOP1, RPN2, AFTPH, ORC3, FOXK1, CD2BP2, PDHA1, PPP6R1, PANK4, KIAA0196, WDR77, CHUK, DDX1, COLGALT1, VAPA, PPME1, CAMSAP1, RABL6, RTCB, SRRT, PIK3C3, NUMA1, STXBP1, OTUB1, UBAP2L, EXOC5, LRRC59, EXOC3, RQCD1, FLII, COPG2, USP5, XPOT, TIMM44, ATP1A1, SBF1, ARPC1B, PLK1, OSBPL8, BAG6, PPP6R2, FAM98B, VPS52, FADS2, EIF4G2, EIF4G3, HSPA8, POLR1B, PSMC6, SFSWAP, VPS33A, MCM10, AVL9, RINT1, PPIB, FBXL18, MFAP1, KIF13B, ATP2C1, ERCC2, CTTNBP2NL, WDR82, VARS, KPNA6, MIA3, HEATR1, SEC23B, AASDHPPT, POLR3B, ALDH18A1, PIKFYVE, MPP6, MRI1, CAPN7, RAB3GAP2, NCAPD2, UGGT1, IKBKAP, KDM3B, CLNS1A, FEN1, ECH1, AP2B1, HNRNPH1, RARS, AGPS, FANCI, PYCR2, EHD1, BUB3, KIF5B, SYMPK, PC, ACAA2, NDC80, PRRC2C, MTHFD1L, FASN, RANBP6, VPS11, NUP93, STRN, SCYL1, DDX19A, GTF2I, SEC23A, CTNNB1, STAM, VPRBP, VCL, PDHB, SEC24D, ERCC4, GRN, ACADM, HSPA6, ERCC6L, RPL5, ARHGAP17, DDX6, MAPK8IP3, TJP1, RIOK1, ERP29, PARG, NUP205, PALLD, COG4, TPP2, EPB41, VAC14, FUS, ZW10, KIF11, GET4, MCM6, EIF2S1, CSDE1, DHX16, GFM1, RABGAP1, TTC28, XPO1, EXOC2, DCTN1, ACBD3, RFC4, PARP1, COPG1, GPRIN1, EIF3C, CUL1, C14orf166, RTL1, STRIP1, MRPS2, PPP2R5D, EIF4G1, EXOSC4, MCM4, SEC63, WDR1, CUL4B, CRMP1, NSF, RNMT, BLVRA, UBE4A, PCBP1, MLLT4, RNF214, ARVCF, CKAP5, NUP188, SMCR8, CTNNA1, SYT1, PCM1, KNTC1, GFPT1, THYN1, SARM1, LPIN1, TIA1, NUP54, PKN1, SUMO4, DICER1, KIF3A, IPO8, CLIC4, PSMD14, GLG1, ABCE1, SMEK1, NCAPG, PPP5C, RTF1, IARS2, PDS5A, PHGDH, RPS6KA3, LONP1, ROCK1, CDK6, LARS, KPNA3, HNRNPAB, MCM2, PPP2R2A, HSP90AB2P, VCPIP1, PCBP2, RPA1, SEC24A, HECTD3, RCC2, DIS3, SMS, KHSRP, CLGN, GNAS, FOCAD, SEC23IP, ACOT7, HK2, CCDC6, MFN1, WDR7, CNN3, KIAA1524, RAB3GAP1, OXCT1, SBNO1, PRPSAP2, ACTR3, CTPS2, TMEM43, FAM21A, LRRC47, VAPB, SCYL2, STRN4, NAE1, SREK1, TFG, SF3A1, CDK2, GPKOW, GPHN, PDCD4, U2AF2, UNC45A, VPS35, NAA15, ASNS, TOP2B, ACO2, AGAP3, ROCK2, NUDT21, NPEPPS, ADD1, AP1G1, CPSF6, DNAJC2, RPS5, CTPS1, IDH1, SEC24C, KIF3B, ACTR2, CPD, EEF1D, SAMHD1, ETFB, DNM1L, LRPPRC, MCMBP, RBM12, OPA1, FARSB, UPF1, PCMT1, USO1, COG3, CLPTM1, TLK2, PRUNE, ASPSCR1, MARS, PYCR1, TCERG1, MED23, PITPNB, ARPC2, UBE4B, DBR1, ETFA, BICD1, KLC1, WDR70, PXDN, DPYSL4, USP7, CLUH, TBRG4, OGFR, SV2A, PDCD6IP, PDLIM5, PSMC2, TRAP1, DRG1, SMEK2, TCP1, TNPO2, CALD1, ARIH2, POR, FKBP3, DCTN2, NPM1, USP48, IPO13, NUP88, NNT, TAOK1, PPWD1, PSPC1, PABPN1, POLA1, MTR, BCCIP, RUVBL2, C16orf62, SORD, PPP2CA, STRN3, RUVBL1, PPP1CA, CACTIN, RPAP3, VPS51, TUBB3, PRKDC, HSPA5, CCDC132, MRPS27, RBBP4, CAPZA1, SLC4A1AP, THUMPD3, FAF1, CDK4, GSTO1, NCBP1, CAPZB, SNRPA, TXLNG, NELFE, MUT, DYNC1LI2, CTTN, DYNC1I2, TIMELESS, CBS, TTLL12, ESYT1, LAMB1, KARS, PPIL4, LRRC40, TXLNA, RAD23B, TBCE, ACAD9, THADA, GCN1L1, SLK, PNKP, ACLY, ATP6V0A1, VTA1, TXNDC5, OXSR1, ACACA, TNPO3, CCDC93, TIPRL, RAN, HOOK3, PSME4, YWHAQ, PPM1G, PSMC3, SCARB2, EXOC7, DFNA5, TRMT6, CPNE3, SLC3A2, G6PD, EIF2B1, SF3B1, EXOC6B, HGS, DYNC1H1, DNAJC7, CANX, TSR1, TTC37, DARS2, ZPR1, SERBP1, HSP90AB1, TBCB, UCHL5, IMPDH2, PUF60, RANGAP1, PDXDC1, YWHAH, LARS2, ZC3H15, GMPS, YWHAG, SUPV3L1, SPAG9, IMPDH1, HSD17B10, GTF2F1, EXOSC9, CUL3, ARHGAP35, TUBB4B, EIF5B, KIF5C, PDIA6, LDHA, ATL2, SNX1, SF3B3, SRM, MAP2, DIAPH1, DBH, NHLRC2, ELP2, GRPEL1, TWF1, WBP11, ERP44, SERPINB6, VCP, HSPH1, CMTR1, KIF1A, USP47, EML4, PRMT1, ATP6V1B2, GPS1, CAND1, CLPB, PCNA, RPLP0, CORO7, TUBB, ANXA2, API5, SUPT16H, RUFY1, KIAA1279, PDIA3, ACTA1, SF3A3, DCTN4, CUL2, WARS, PGM2, NELFB, XPO5, UGP2, HSP90B1, GSR, SHMT2, IPO11, EIF3E, RDX, DNAJB1, KIAA1598, AARS, EFTUD1, HEATR5B, ENAH, TNPO1, LIG1, NRD1, DLD, STRA6, PAFAH1B1, NT5C2, SSRP1, NTRK3, HDGFRP2, EIF3G, CSE1L, ACAT1, TBCD, NAA10, MTHFD1, RALA, EIF3A, PRPS1, ISOC1, IPO7, HNRNPD, GAPDH, U2AF1, SUPT5H, PSMD8, RPS6KA1, NUF2, GDAP1, VPS26A, TUBB2B, CDH2, TMOD1, CBR1, HDLBP, CS, HSPD1, LMAN1, EIF3B, DDB1, PSMD1, DPYSL2, STRAP, RTN4, FLNA, AHCY, HSD17B4, GOLIM4, SPTAN1, IPO4, PKM, EIF3I, SF1, BZW2, XPO7, ACADVL, STAG2, DPP9, RECQL, GDI1, GARS, MMS19, EIF2A, IPO5, TCOF1, RAB5C, SAP30BP, VDAC1, TUBA1A, SEPT2, DBNL, PLAA, TOMM70A, NOMO1, ISYNA1, CTNNA2, EEF1B2, PAK4, NELFCD, GSPT1, KPNB1, MAP1A, COPS4, YWHAZ, KPNA2, EIF4B, PLS3, COPS3, HSPBP1, AP1B1, NASP, DDX39A, LUC7L2, APMAP, SEC31A, SRSF2, TXNRD1, ATP1A3, RRM2, COG7, FSD1, PRDX4, DCPS, NCAM1, NUDCD1, NAA25, RPL9, NPLOC4, YWHAB, ATP1B1, LSS, DNAAF5, METTL16, COPS6, XPNPEP1, ELAC2, PUS7, CCT3, SFN, AGL, PAPOLA, SRSF11, ALDH2, PSME2, CKMT1B, LDHB, PSMA7, RAB11A, PPP2R1A, GNAO1, LUC7L, COPS5, IMPA1, RRM1, PPID, DPP3, KIAA0368, SLC1A5, YARS, NUP50, TPM4, ITGB1, SARS, SEZ6L2, GTF2F2, GART, ALDOC, PSMD11, EIF6, DFFA, RAP1GDS1, NELFA, CARS, CCT4, P4HB, FSCN1, PSMD13, PSMA5, CCT5, PAPSS1, OSBP, SAE1, ATP6V1A, LMAN2, YWHAE, SRP54, IDE, SNX2, RPSA, BZW1, SND1, UBQLN1, MAPK1, CALR, MARCKSL1, SUGT1, TBC1D4, GLRX3, LETM1, HMGB1, EIF3H, SLC9A3R1, DDX42, LAP3, PRKCSH, EEF2, PSMD6, BCAP31, ANXA11, ACTN1, CCT6A, PSMA4, ANXA5, CCT7, EIF3L, FUBP1, SSB, PGM1, MAPRE1, UBA2, AMPD2, PPAT, CLIC1, CCT2, ECHS1, LTA4H, USP14, CCT8, ANK2, PDIA4, TPR, NRCAM, MAT2A, HTATSF1, THOP1, RANBP1, RSRC2, NMT1, AK2, EEF1G, TPM3, PSMA3, EIF4H, GLUD1, CAP1, TKT, ATP5B, TXNL1, STIP1, HMGB2, GGH, HSPB1, CALU, RPL17, PSMA1, RSRC1, ST13, PRDX6, EIF3J, PNPT1, APEX1, TARS, MARCKS, PSMA6, EEF1A1, COPB2, NUDT5, HSPA1A, ERO1L, ME2, RPH3A, ESD, EIF5, ADSL, RUFY3, DPYSL5, ANXA6, NUDC, NANS, GOT2, GLDC, PSAT1, SET, PAK2, UBA6, UCHL1, L1CAM, PGLS, MAP1B, CCDC124, EIF3M, VAT1, PAFAH1B3, ALDOA, IRGQ, UBQLN2, NACA, OLA1, ACTN4, ARHGDIA, PSMD7, CACYBP, PSMA2, CPNE1, MDH2, PGAM1, DDX46, ACAT2, ALCAM, HARS, CKB, FKBP4, ASNA1, GPI, INA, HYOU1, CHGA, MDH1, EZR, ANP32A, UBA1, ARG1, ANP32B, PAICS, ATIC, TPI1, PPA1, TALDO1, PGK1, DDX39B, TLN1, AKR1B1, ENO1, NGFR, TPTE, GGA1, GGA2, GGA3, USP36, NDFIP1, SOCS2, CDK12, EPHB4, NDUFA4L2, AKT1, SLC5A5, DOT1L, AKT2, SHC4, TRAF4, PTPN6, ERBB2, EGR1, HIPK2, POU4F2, |
 Protein-protein interactors based on sequence similarity (STRING) Protein-protein interactors based on sequence similarity (STRING) |
| Gene | STRING network |
| RPL7A | |
| NTRK1 |  |
 - Retained interactions in fusion protein (protein functional feature from UniProt). - Retained interactions in fusion protein (protein functional feature from UniProt). |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost interactions due to fusion (protein functional feature from UniProt). - Lost interactions due to fusion (protein functional feature from UniProt). |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs to RPL7A-NTRK1 |
 Drugs used for this fusion-positive patient. Drugs used for this fusion-positive patient. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Drug | Source | PMID |
Top |
Related Diseases to RPL7A-NTRK1 |
 Diseases that have this fusion gene. Diseases that have this fusion gene. (Manual curation of PubMed, 04-30-2022 + MyCancerGenome) |
| Hgene | Tgene | Disease | Source | PMID |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
| Tgene | NTRK1 | C0020074 | HSAN Type IV | 17 | CTD_human;GENOMICS_ENGLAND;UNIPROT |
| Tgene | NTRK1 | C0238463 | Papillary thyroid carcinoma | 3 | ORPHANET |
| Tgene | NTRK1 | C0002768 | Congenital Pain Insensitivity | 1 | ORPHANET |
| Tgene | NTRK1 | C0005586 | Bipolar Disorder | 1 | CTD_human |
| Tgene | NTRK1 | C0005587 | Depression, Bipolar | 1 | CTD_human |
| Tgene | NTRK1 | C0017638 | Glioma | 1 | CTD_human |
| Tgene | NTRK1 | C0020075 | Hereditary Sensory Autonomic Neuropathy, Type 5 | 1 | CTD_human;ORPHANET |
| Tgene | NTRK1 | C0024713 | Manic Disorder | 1 | CTD_human |
| Tgene | NTRK1 | C0027796 | Neuralgia | 1 | CTD_human |
| Tgene | NTRK1 | C0027819 | Neuroblastoma | 1 | CTD_human |
| Tgene | NTRK1 | C0033958 | Psychosis, Brief Reactive | 1 | CTD_human |
| Tgene | NTRK1 | C0033975 | Psychotic Disorders | 1 | CTD_human |
| Tgene | NTRK1 | C0036337 | Schizoaffective Disorder | 1 | CTD_human |
| Tgene | NTRK1 | C0036341 | Schizophrenia | 1 | CTD_human |
| Tgene | NTRK1 | C0036358 | Schizophreniform Disorders | 1 | CTD_human |
| Tgene | NTRK1 | C0038870 | Neuralgia, Supraorbital | 1 | CTD_human |
| Tgene | NTRK1 | C0042656 | Neuralgia, Vidian | 1 | CTD_human |
| Tgene | NTRK1 | C0234247 | Neuralgia, Atypical | 1 | CTD_human |
| Tgene | NTRK1 | C0234249 | Neuralgia, Stump | 1 | CTD_human |
| Tgene | NTRK1 | C0259783 | mixed gliomas | 1 | CTD_human |
| Tgene | NTRK1 | C0273115 | Lung Injury | 1 | CTD_human |
| Tgene | NTRK1 | C0338831 | Manic | 1 | CTD_human |
| Tgene | NTRK1 | C0423711 | Neuralgia, Perineal | 1 | CTD_human |
| Tgene | NTRK1 | C0423712 | Neuralgia, Iliohypogastric Nerve | 1 | CTD_human |
| Tgene | NTRK1 | C0555198 | Malignant Glioma | 1 | CTD_human |
| Tgene | NTRK1 | C0598589 | Inherited neuropathies | 1 | GENOMICS_ENGLAND |
| Tgene | NTRK1 | C0751371 | Neuralgia, Ilioinguinal | 1 | CTD_human |
| Tgene | NTRK1 | C0751372 | Nerve Pain | 1 | CTD_human |
| Tgene | NTRK1 | C0751373 | Paroxysmal Nerve Pain | 1 | CTD_human |
| Tgene | NTRK1 | C0752347 | Lewy Body Disease | 1 | CTD_human |
| Tgene | NTRK1 | C1833921 | Familial medullary thyroid carcinoma | 1 | CTD_human;GENOMICS_ENGLAND;ORPHANET |
| Tgene | NTRK1 | C2350344 | Chronic Lung Injury | 1 | CTD_human |

