
|
||||||
|
Translation Factor: TNRC6A (NCBI Gene ID:27327) |
|
 Gene Summary Gene Summary |
| Gene Information | Gene Name: TNRC6A | Gene ID: 27327 | Gene Symbol | TNRC6A | Gene ID | 27327 |
| Gene Name | trinucleotide repeat containing adaptor 6A | |
| Synonyms | CAGH26|FAME6|GW1|GW182|TNRC6 | |
| Cytomap | 16p12.1 | |
| Type of Gene | protein-coding | |
| Description | trinucleotide repeat-containing gene 6A proteinCAG repeat protein 26CTD-2540M10.1EDIEEMSY interactor proteinGW182 autoantigenglycine-tryptophan protein of 182 kDatrinucleotide repeat containing 6A | |
| Modification date | 20200313 | |
| UniProtAcc | Q8NDV7 | |
 Child GO biological process term(s) under GO:0006412 Child GO biological process term(s) under GO:0006412 |
| GO ID | GO term |
| GO:0017148 | Negative regulation of translation |
| GO:0006417 | Regulation of translation |
| GO:0006412 | Translation |
 Gene ontology of translaction factor with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of translaction factor with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | TNRC6A | GO:0035278 | miRNA mediated inhibition of translation | 17671087 |
 Inferred gene age of translation factor. Inferred gene age of translation factor. |
| Gene | Inferred gene age group among (0 - 67.6], (67.6 - 355.7], (355.7 - 733], (733 - 1119.25], >1119.25 |
| TNRC6A | (355.7 - 733] |
Top |
|
 We searched PubMed using 'TNRC6A[title] AND translation [title] AND human.' We searched PubMed using 'TNRC6A[title] AND translation [title] AND human.' |
| Gene | Title | PMID |
| TNRC6A | . | . |
Top |
|
 Skipped exons in TCGA and GTEx based on Ensembl gene isoform structure. Skipped exons in TCGA and GTEx based on Ensembl gene isoform structure. * Click on the image to open the UCSC genome browser with custom track showing this image in a new window. For more annotations, please visit our ExonSkipDB. |
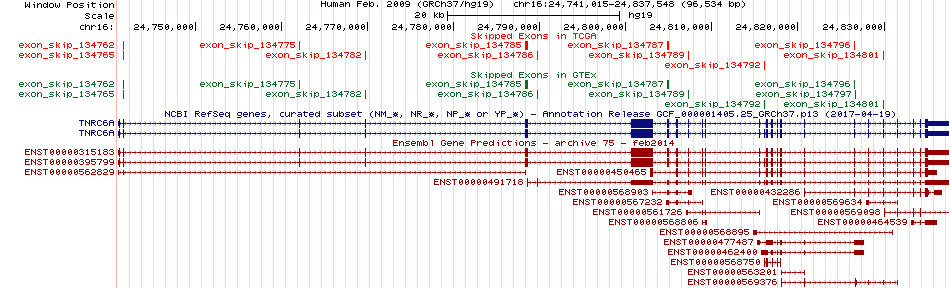 |
 Open reading frame (ORF) analsis of exon skipping events based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of exon skipping events based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ENST | Exon skip start (DNA) | Exon Skip end (DNA) | ORF |
| ENST00000395799 | 24741573 | 24741621 | In-frame |
| ENST00000395799 | 24762046 | 24762134 | Frame-shift |
| ENST00000395799 | 24769659 | 24769681 | Frame-shift |
| ENST00000395799 | 24788253 | 24788679 | In-frame |
| ENST00000395799 | 24804793 | 24804970 | In-frame |
| ENST00000395799 | 24807227 | 24807260 | In-frame |
| ENST00000395799 | 24816025 | 24816172 | In-frame |
| ENST00000395799 | 24826467 | 24826626 | In-frame |
| ENST00000395799 | 24829913 | 24830021 | In-frame |
 Exon skipping position in the amino acid sequence. Exon skipping position in the amino acid sequence. |
| ENST | Exon skip start (DNA) | Exon Skip end (DNA) | Len(transcript seq) | Exon skip start (mRNA) | Exon Skip end (mRNA) | Len(amino acid seq) | Exon skip start (AA) | Exon Skip end (AA) |
| ENST00000395799 | 24741573 | 24741621 | 8455 | 135 | 182 | 1962 | 2 | 17 |
| ENST00000395799 | 24788253 | 24788679 | 8455 | 293 | 718 | 1962 | 54 | 196 |
| ENST00000395799 | 24804793 | 24804970 | 8455 | 3305 | 3481 | 1962 | 1058 | 1117 |
| ENST00000395799 | 24816025 | 24816172 | 8455 | 3967 | 4113 | 1962 | 1279 | 1328 |
| ENST00000395799 | 24826467 | 24826626 | 8455 | 4802 | 4960 | 1962 | 1557 | 1610 |
| ENST00000395799 | 24829913 | 24830021 | 8455 | 5102 | 5209 | 1962 | 1657 | 1693 |
 Potentially (partially) lost protein functional features of UniProt. Potentially (partially) lost protein functional features of UniProt. |
| UniProtAcc | Exon skip start (AA) | Exon Skip end (AA) | Function feature start (AA) | Function feature end (AA) | Functional feature type | Functional feature desc. |
Top |
|
 Gene expression level across TCGA pancancer Gene expression level across TCGA pancancer |
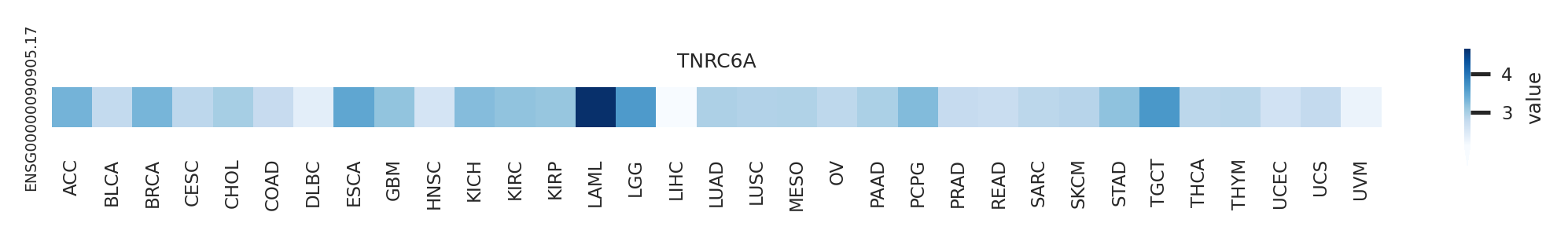 |
 Gene expression level across GTEx pantissue Gene expression level across GTEx pantissue |
 |
 Expression level of gene isoforms across TCGA pancancer Expression level of gene isoforms across TCGA pancancer |
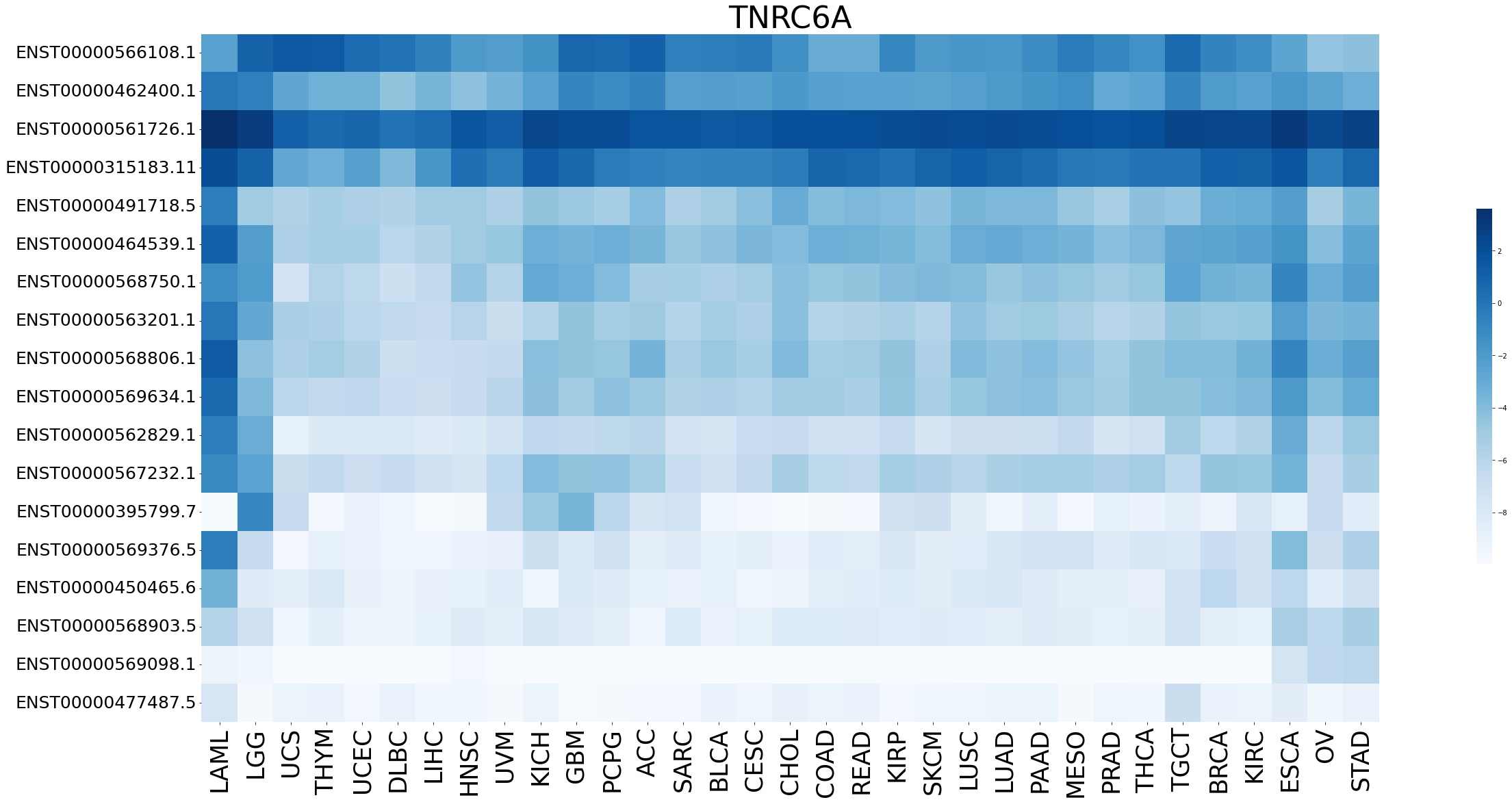 |
 Expression level of gene isoforms across GTEx pantissue Expression level of gene isoforms across GTEx pantissue |
 |
 Cancer(tissue) type-specific expression level of Translation factor using z-score distriution Cancer(tissue) type-specific expression level of Translation factor using z-score distriution |
 |
 Differential expression between tumor and matched normal (in the cancer types with more than 10 matched samples) Differential expression between tumor and matched normal (in the cancer types with more than 10 matched samples) |
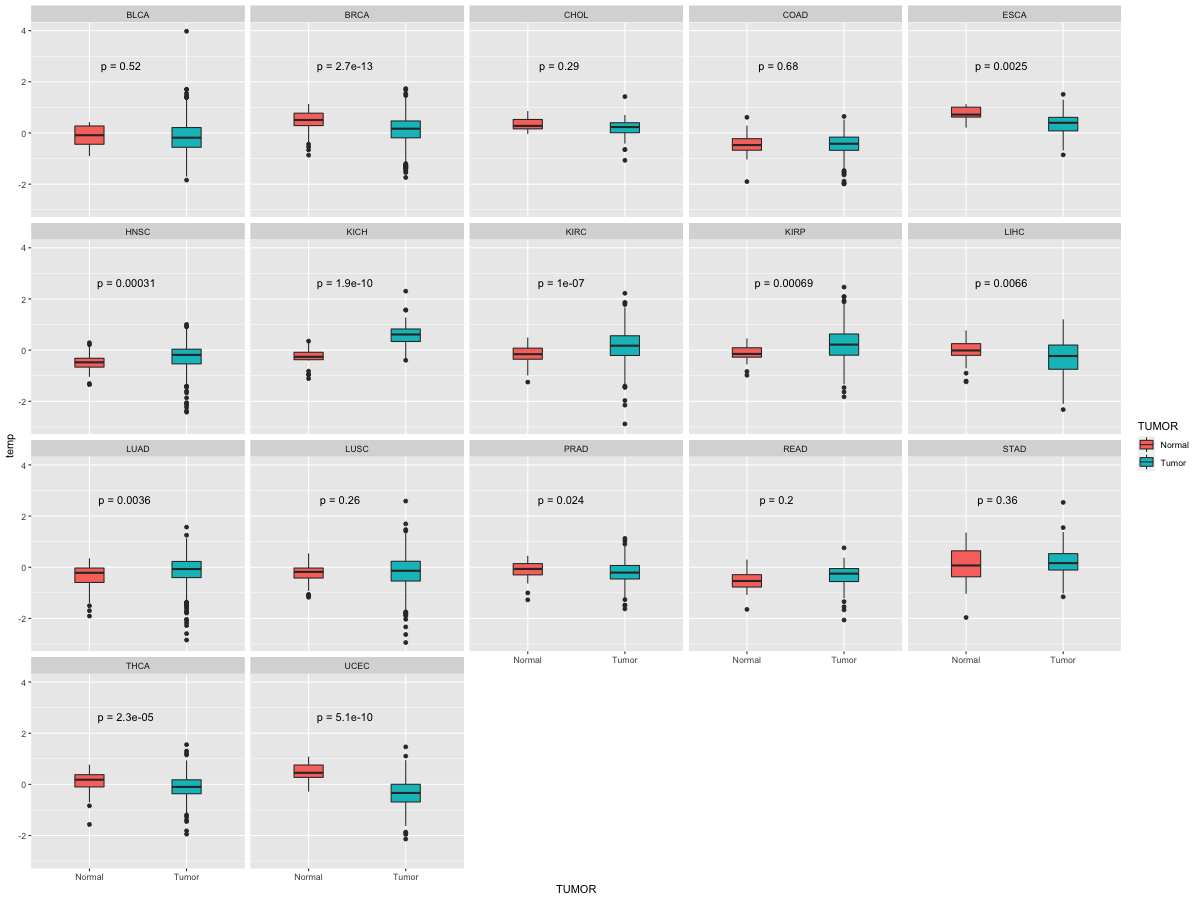 |
| Cancer type | Translation factor | FC | adj.pval |
| BRCA | TNRC6A | -1.24656754002514 | 3.76762765813611e-06 |
Top |
|
 Translation factor expression regulation through miRNA binding Translation factor expression regulation through miRNA binding |
| Cancer type | Gene | miRNA | TargetScan binding score (Context++ score percentile) | Coefficient | Pvalue |
 Translation factor expression regulation through methylation in the promoter of Translation factor Translation factor expression regulation through methylation in the promoter of Translation factor |
 |
| Cancer type | Gene | methyl group b | methyl group a | DEG pval | avg methyl in b | avg methyl in a | avg exp in b | avg exp in a |
 Translation factor expression regulation through methylation in the gene body of Translation factor (positive regulation) Translation factor expression regulation through methylation in the gene body of Translation factor (positive regulation) |
 |
| Cancer type | Gene | methyl group b | methyl group a | DEG pval | avg methyl in b | avg methyl in a | avg exp in b | avg exp in a |
| LUAD | TNRC6A | 3 | 2 | 0.0321023617316507 | 0.615448125 | 0.521751562500001 | -0.257838531018361 | -0.439792001167661 |
 Translation factor expression regulation through copy number variation of Translation factor Translation factor expression regulation through copy number variation of Translation factor |
 |
| Cancer type | Gene | Coefficient | Pvalue |
Top |
|
 Strongly correlated genes belong to cellular important gene groups with TNRC6A (coefficient>0.8, pval<0.05, node color based on FC between tumor and matched normal). Significantly associated important genes in the individual cancer types. * Cell metabolism gene: cell metabolism genes from REACTOME (black edge), IUPHAR: drug target genes from IUPHAR (blue edge), Kinase: human kinase genes (brown edge), CGC: cancer gene census genes (orange edge), TSG: tumor suppresor genes (purple edge), Epifactor: epigenetic factors (light blue edge), TF: transcription factors (green) Strongly correlated genes belong to cellular important gene groups with TNRC6A (coefficient>0.8, pval<0.05, node color based on FC between tumor and matched normal). Significantly associated important genes in the individual cancer types. * Cell metabolism gene: cell metabolism genes from REACTOME (black edge), IUPHAR: drug target genes from IUPHAR (blue edge), Kinase: human kinase genes (brown edge), CGC: cancer gene census genes (orange edge), TSG: tumor suppresor genes (purple edge), Epifactor: epigenetic factors (light blue edge), TF: transcription factors (green) |
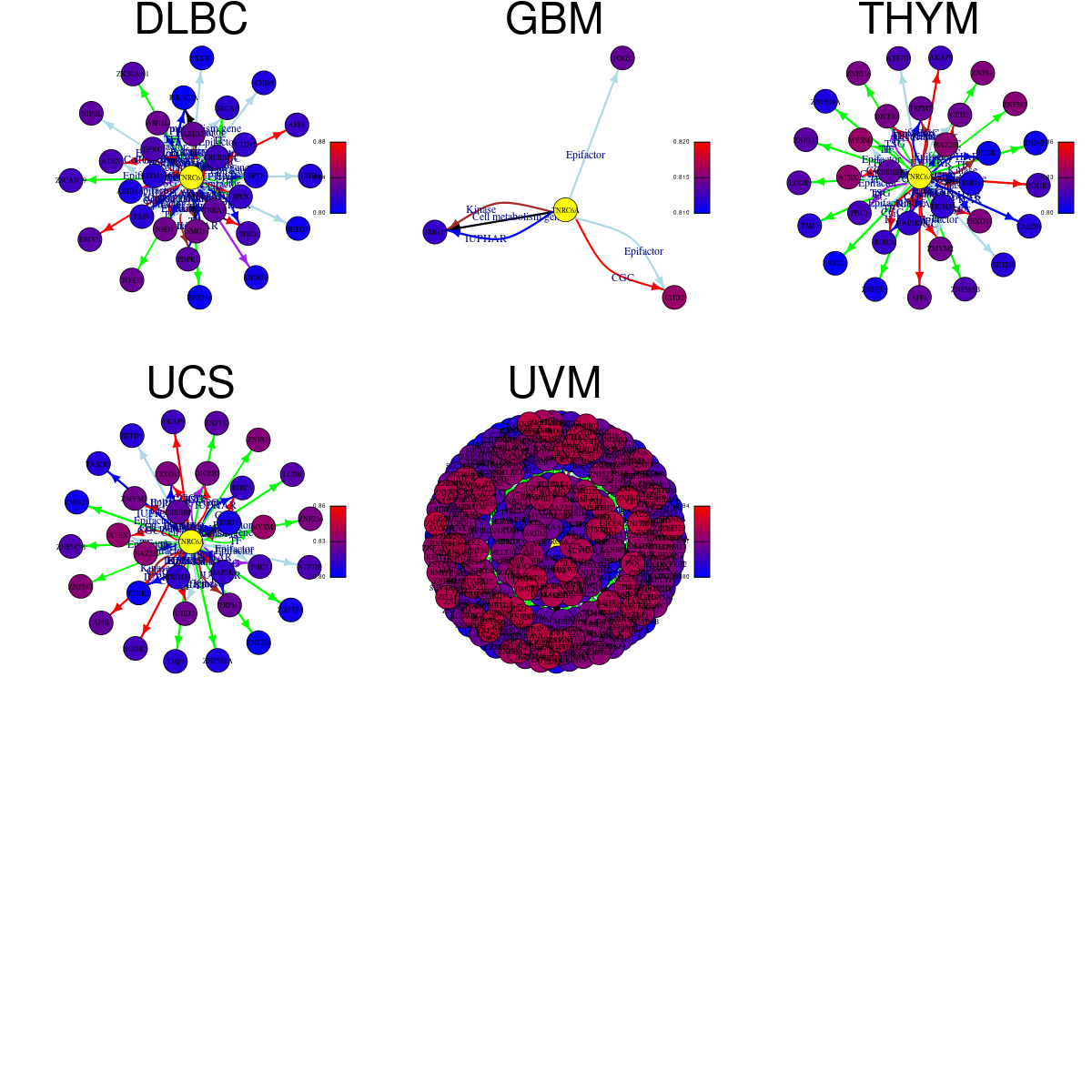 |
| Cancer type | Gene group | Translation factor | Correlated gene | Coefficient | Pvalue |
| DLBC | Cell metabolism gene | TNRC6A | PIK3C2A | 0.800229619 | 8.81E-12 |
| DLBC | Cell metabolism gene | TNRC6A | CREBBP | 0.822793756 | 7.26E-13 |
| DLBC | Cell metabolism gene | TNRC6A | EP300 | 0.829429005 | 3.25E-13 |
| DLBC | Cell metabolism gene | TNRC6A | SMG1 | 0.876499018 | 3.30E-16 |
| DLBC | CGC | TNRC6A | ARID1A | 0.806135372 | 4.73E-12 |
| DLBC | CGC | TNRC6A | TRIM33 | 0.812088013 | 2.47E-12 |
| DLBC | CGC | TNRC6A | SETD1B | 0.817001747 | 1.42E-12 |
| DLBC | CGC | TNRC6A | SPEN | 0.819464916 | 1.07E-12 |
| DLBC | CGC | TNRC6A | AFF4 | 0.82271246 | 7.33E-13 |
| DLBC | CGC | TNRC6A | CREBBP | 0.822793756 | 7.26E-13 |
| DLBC | CGC | TNRC6A | BIRC6 | 0.826088049 | 4.89E-13 |
| DLBC | CGC | TNRC6A | ATRX | 0.827454702 | 4.15E-13 |
| DLBC | CGC | TNRC6A | EP300 | 0.829429005 | 3.25E-13 |
| DLBC | CGC | TNRC6A | ERCC4 | 0.831814679 | 2.42E-13 |
| DLBC | CGC | TNRC6A | TRRAP | 0.837147312 | 1.23E-13 |
| DLBC | CGC | TNRC6A | ARID2 | 0.838706481 | 1.00E-13 |
| DLBC | CGC | TNRC6A | NSD1 | 0.875424882 | 3.98E-16 |
| DLBC | Epifactor | TNRC6A | ATXN7 | 0.803585515 | 6.20E-12 |
| DLBC | Epifactor | TNRC6A | SETD5 | 0.803967219 | 5.96E-12 |
| DLBC | Epifactor | TNRC6A | ARID1A | 0.806135372 | 4.73E-12 |
| DLBC | Epifactor | TNRC6A | BPTF | 0.806813692 | 4.40E-12 |
| DLBC | Epifactor | TNRC6A | CHD9 | 0.8095644 | 3.26E-12 |
| DLBC | Epifactor | TNRC6A | CHD6 | 0.810656266 | 2.90E-12 |
| DLBC | Epifactor | TNRC6A | TRIM33 | 0.812088013 | 2.47E-12 |
| DLBC | Epifactor | TNRC6A | TAF1 | 0.812357282 | 2.40E-12 |
| DLBC | Epifactor | TNRC6A | SRCAP | 0.814472353 | 1.89E-12 |
| DLBC | Epifactor | TNRC6A | SETD1B | 0.817001747 | 1.42E-12 |
| DLBC | Epifactor | TNRC6A | SPEN | 0.819464916 | 1.07E-12 |
| DLBC | Epifactor | TNRC6A | CREBBP | 0.822793756 | 7.26E-13 |
| DLBC | Epifactor | TNRC6A | ATRX | 0.827454702 | 4.15E-13 |
| DLBC | Epifactor | TNRC6A | EP300 | 0.829429005 | 3.25E-13 |
| DLBC | Epifactor | TNRC6A | NIPBL | 0.835862097 | 1.45E-13 |
| DLBC | Epifactor | TNRC6A | TRRAP | 0.837147312 | 1.23E-13 |
| DLBC | Epifactor | TNRC6A | ARID2 | 0.838706481 | 1.00E-13 |
| DLBC | Epifactor | TNRC6A | ASH1L | 0.866027606 | 1.91E-15 |
| DLBC | Epifactor | TNRC6A | NSD1 | 0.875424882 | 3.98E-16 |
| DLBC | IUPHAR | TNRC6A | PIK3C2A | 0.800229619 | 8.81E-12 |
| DLBC | IUPHAR | TNRC6A | BPTF | 0.806813692 | 4.40E-12 |
| DLBC | IUPHAR | TNRC6A | TRIM33 | 0.812088013 | 2.47E-12 |
| DLBC | IUPHAR | TNRC6A | TAF1 | 0.812357282 | 2.40E-12 |
| DLBC | IUPHAR | TNRC6A | SETD1B | 0.817001747 | 1.42E-12 |
| DLBC | IUPHAR | TNRC6A | CREBBP | 0.822793756 | 7.26E-13 |
| DLBC | IUPHAR | TNRC6A | BIRC6 | 0.826088049 | 4.89E-13 |
| DLBC | IUPHAR | TNRC6A | EP300 | 0.829429005 | 3.25E-13 |
| DLBC | IUPHAR | TNRC6A | PDPK1 | 0.834023405 | 1.83E-13 |
| DLBC | IUPHAR | TNRC6A | TRRAP | 0.837147312 | 1.23E-13 |
| DLBC | IUPHAR | TNRC6A | ASH1L | 0.866027606 | 1.91E-15 |
| DLBC | IUPHAR | TNRC6A | NSD1 | 0.875424882 | 3.98E-16 |
| DLBC | IUPHAR | TNRC6A | SMG1 | 0.876499018 | 3.30E-16 |
| DLBC | Kinase | TNRC6A | TRIM33 | 0.812088013 | 2.47E-12 |
| DLBC | Kinase | TNRC6A | TAF1 | 0.812357282 | 2.40E-12 |
| DLBC | Kinase | TNRC6A | PDPK1 | 0.834023405 | 1.83E-13 |
| DLBC | Kinase | TNRC6A | TRRAP | 0.837147312 | 1.23E-13 |
| DLBC | Kinase | TNRC6A | SMG1 | 0.876499018 | 3.30E-16 |
| DLBC | TF | TNRC6A | ZNF236 | 0.801984529 | 7.34E-12 |
| DLBC | TF | TNRC6A | BPTF | 0.806813692 | 4.40E-12 |
| DLBC | TF | TNRC6A | SRCAP | 0.814472353 | 1.89E-12 |
| DLBC | TF | TNRC6A | SPEN | 0.819464916 | 1.07E-12 |
| DLBC | TF | TNRC6A | ZSCAN29 | 0.823089025 | 7.01E-13 |
| DLBC | TF | TNRC6A | ZKSCAN1 | 0.830360068 | 2.90E-13 |
| DLBC | TF | TNRC6A | ARID2 | 0.838706481 | 1.00E-13 |
| DLBC | TF | TNRC6A | FOXJ3 | 0.850938561 | 1.87E-14 |
| DLBC | TF | TNRC6A | ASH1L | 0.866027606 | 1.91E-15 |
| DLBC | TSG | TNRC6A | DIDO1 | 0.804879764 | 5.41E-12 |
| DLBC | TSG | TNRC6A | ARID1A | 0.806135372 | 4.73E-12 |
| DLBC | TSG | TNRC6A | CREBBP | 0.822793756 | 7.26E-13 |
| DLBC | TSG | TNRC6A | ARID2 | 0.838706481 | 1.00E-13 |
| GBM | Cell metabolism gene | TNRC6A | SMG1 | 0.810845446 | 2.11E-41 |
| GBM | CGC | TNRC6A | CHD2 | 0.821418074 | 2.58E-43 |
| GBM | Epifactor | TNRC6A | POGZ | 0.818262998 | 9.90E-43 |
| GBM | Epifactor | TNRC6A | CHD2 | 0.821418074 | 2.58E-43 |
| GBM | IUPHAR | TNRC6A | SMG1 | 0.810845446 | 2.11E-41 |
| GBM | Kinase | TNRC6A | SMG1 | 0.810845446 | 2.11E-41 |
| THYM | Cell metabolism gene | TNRC6A | SMG1 | 0.801403786 | 1.48E-28 |
| THYM | Cell metabolism gene | TNRC6A | CREBBP | 0.81732804 | 1.64E-30 |
| THYM | CGC | TNRC6A | TRIM33 | 0.803276955 | 8.91E-29 |
| THYM | CGC | TNRC6A | MAP3K1 | 0.80653169 | 3.63E-29 |
| THYM | CGC | TNRC6A | BIRC6 | 0.808120468 | 2.33E-29 |
| THYM | CGC | TNRC6A | AKAP9 | 0.809114526 | 1.76E-29 |
| THYM | CGC | TNRC6A | HOOK3 | 0.809502383 | 1.58E-29 |
| THYM | CGC | TNRC6A | CREBBP | 0.81732804 | 1.64E-30 |
| THYM | CGC | TNRC6A | AFF4 | 0.819372588 | 8.88E-31 |
| THYM | CGC | TNRC6A | CHD2 | 0.823961125 | 2.19E-31 |
| THYM | CGC | TNRC6A | ZMYM2 | 0.827246408 | 7.86E-32 |
| THYM | CGC | TNRC6A | DICER1 | 0.828668929 | 5.01E-32 |
| THYM | CGC | TNRC6A | FBXO11 | 0.834988544 | 6.41E-33 |
| THYM | CGC | TNRC6A | ATRX | 0.854890453 | 5.36E-36 |
| THYM | Epifactor | TNRC6A | SETD5 | 0.802121691 | 1.22E-28 |
| THYM | Epifactor | TNRC6A | TRIM33 | 0.803276955 | 8.91E-29 |
| THYM | Epifactor | TNRC6A | PHC3 | 0.810539677 | 1.18E-29 |
| THYM | Epifactor | TNRC6A | ATF7IP | 0.8121065 | 7.51E-30 |
| THYM | Epifactor | TNRC6A | CREBBP | 0.81732804 | 1.64E-30 |
| THYM | Epifactor | TNRC6A | CHD2 | 0.823961125 | 2.19E-31 |
| THYM | Epifactor | TNRC6A | ZMYM2 | 0.827246408 | 7.86E-32 |
| THYM | Epifactor | TNRC6A | BAZ2B | 0.844301464 | 2.63E-34 |
| THYM | Epifactor | TNRC6A | ATRX | 0.854890453 | 5.36E-36 |
| THYM | Epifactor | TNRC6A | MYSM1 | 0.864473218 | 1.20E-37 |
| THYM | IUPHAR | TNRC6A | TTBK2 | 0.800458733 | 1.91E-28 |
| THYM | IUPHAR | TNRC6A | SMG1 | 0.801403786 | 1.48E-28 |
| THYM | IUPHAR | TNRC6A | TAS2R5 | 0.801870471 | 1.31E-28 |
| THYM | IUPHAR | TNRC6A | TRIM33 | 0.803276955 | 8.91E-29 |
| THYM | IUPHAR | TNRC6A | MAP3K1 | 0.80653169 | 3.63E-29 |
| THYM | IUPHAR | TNRC6A | BIRC6 | 0.808120468 | 2.33E-29 |
| THYM | IUPHAR | TNRC6A | CREBBP | 0.81732804 | 1.64E-30 |
| THYM | IUPHAR | TNRC6A | TRPM7 | 0.822484698 | 3.46E-31 |
| THYM | IUPHAR | TNRC6A | FBXO11 | 0.834988544 | 6.41E-33 |
| THYM | IUPHAR | TNRC6A | BAZ2B | 0.844301464 | 2.63E-34 |
| THYM | Kinase | TNRC6A | TTBK2 | 0.800458733 | 1.91E-28 |
| THYM | Kinase | TNRC6A | SMG1 | 0.801403786 | 1.48E-28 |
| THYM | Kinase | TNRC6A | TRIM33 | 0.803276955 | 8.91E-29 |
| THYM | Kinase | TNRC6A | MAP3K1 | 0.80653169 | 3.63E-29 |
| THYM | Kinase | TNRC6A | TRPM7 | 0.822484698 | 3.46E-31 |
| THYM | TF | TNRC6A | ZNF224 | 0.800045904 | 2.13E-28 |
| THYM | TF | TNRC6A | ZNF621 | 0.800472553 | 1.90E-28 |
| THYM | TF | TNRC6A | ZNF37A | 0.800727418 | 1.78E-28 |
| THYM | TF | TNRC6A | ZNF518A | 0.801530166 | 1.43E-28 |
| THYM | TF | TNRC6A | TMF1 | 0.802873406 | 9.94E-29 |
| THYM | TF | TNRC6A | ZNF585B | 0.814768436 | 3.47E-30 |
| THYM | TF | TNRC6A | LCOR | 0.814932145 | 3.31E-30 |
| THYM | TF | TNRC6A | ZNF333 | 0.816156828 | 2.31E-30 |
| THYM | TF | TNRC6A | ZNF236 | 0.831689018 | 1.90E-32 |
| THYM | TF | TNRC6A | ZNF81 | 0.84134804 | 7.41E-34 |
| THYM | TF | TNRC6A | ZNF587 | 0.843216298 | 3.86E-34 |
| THYM | TF | TNRC6A | BAZ2B | 0.844301464 | 2.63E-34 |
| THYM | TF | TNRC6A | MYSM1 | 0.864473218 | 1.20E-37 |
| THYM | TSG | TNRC6A | PHC3 | 0.810539677 | 1.18E-29 |
| THYM | TSG | TNRC6A | CREBBP | 0.81732804 | 1.64E-30 |
| THYM | TSG | TNRC6A | DICER1 | 0.828668929 | 5.01E-32 |
| UCS | Cell metabolism gene | TNRC6A | SMG1 | 0.801403786 | 1.48E-28 |
| UCS | Cell metabolism gene | TNRC6A | CREBBP | 0.81732804 | 1.64E-30 |
| UCS | CGC | TNRC6A | TRIM33 | 0.803276955 | 8.91E-29 |
| UCS | CGC | TNRC6A | MAP3K1 | 0.80653169 | 3.63E-29 |
| UCS | CGC | TNRC6A | BIRC6 | 0.808120468 | 2.33E-29 |
| UCS | CGC | TNRC6A | AKAP9 | 0.809114526 | 1.76E-29 |
| UCS | CGC | TNRC6A | HOOK3 | 0.809502383 | 1.58E-29 |
| UCS | CGC | TNRC6A | CREBBP | 0.81732804 | 1.64E-30 |
| UCS | CGC | TNRC6A | AFF4 | 0.819372588 | 8.88E-31 |
| UCS | CGC | TNRC6A | CHD2 | 0.823961125 | 2.19E-31 |
| UCS | CGC | TNRC6A | ZMYM2 | 0.827246408 | 7.86E-32 |
| UCS | CGC | TNRC6A | DICER1 | 0.828668929 | 5.01E-32 |
| UCS | CGC | TNRC6A | FBXO11 | 0.834988544 | 6.41E-33 |
| UCS | CGC | TNRC6A | ATRX | 0.854890453 | 5.36E-36 |
| UCS | Epifactor | TNRC6A | SETD5 | 0.802121691 | 1.22E-28 |
| UCS | Epifactor | TNRC6A | TRIM33 | 0.803276955 | 8.91E-29 |
| UCS | Epifactor | TNRC6A | PHC3 | 0.810539677 | 1.18E-29 |
| UCS | Epifactor | TNRC6A | ATF7IP | 0.8121065 | 7.51E-30 |
| UCS | Epifactor | TNRC6A | CREBBP | 0.81732804 | 1.64E-30 |
| UCS | Epifactor | TNRC6A | CHD2 | 0.823961125 | 2.19E-31 |
| UCS | Epifactor | TNRC6A | ZMYM2 | 0.827246408 | 7.86E-32 |
| UCS | Epifactor | TNRC6A | BAZ2B | 0.844301464 | 2.63E-34 |
| UCS | Epifactor | TNRC6A | ATRX | 0.854890453 | 5.36E-36 |
| UCS | Epifactor | TNRC6A | MYSM1 | 0.864473218 | 1.20E-37 |
| UCS | IUPHAR | TNRC6A | TTBK2 | 0.800458733 | 1.91E-28 |
| UCS | IUPHAR | TNRC6A | SMG1 | 0.801403786 | 1.48E-28 |
| UCS | IUPHAR | TNRC6A | TAS2R5 | 0.801870471 | 1.31E-28 |
| UCS | IUPHAR | TNRC6A | TRIM33 | 0.803276955 | 8.91E-29 |
| UCS | IUPHAR | TNRC6A | MAP3K1 | 0.80653169 | 3.63E-29 |
| UCS | IUPHAR | TNRC6A | BIRC6 | 0.808120468 | 2.33E-29 |
| UCS | IUPHAR | TNRC6A | CREBBP | 0.81732804 | 1.64E-30 |
| UCS | IUPHAR | TNRC6A | TRPM7 | 0.822484698 | 3.46E-31 |
| UCS | IUPHAR | TNRC6A | FBXO11 | 0.834988544 | 6.41E-33 |
| UCS | IUPHAR | TNRC6A | BAZ2B | 0.844301464 | 2.63E-34 |
| UCS | Kinase | TNRC6A | TTBK2 | 0.800458733 | 1.91E-28 |
| UCS | Kinase | TNRC6A | SMG1 | 0.801403786 | 1.48E-28 |
| UCS | Kinase | TNRC6A | TRIM33 | 0.803276955 | 8.91E-29 |
| UCS | Kinase | TNRC6A | MAP3K1 | 0.80653169 | 3.63E-29 |
| UCS | Kinase | TNRC6A | TRPM7 | 0.822484698 | 3.46E-31 |
| UCS | TF | TNRC6A | ZNF224 | 0.800045904 | 2.13E-28 |
| UCS | TF | TNRC6A | ZNF621 | 0.800472553 | 1.90E-28 |
| UCS | TF | TNRC6A | ZNF37A | 0.800727418 | 1.78E-28 |
| UCS | TF | TNRC6A | ZNF518A | 0.801530166 | 1.43E-28 |
| UCS | TF | TNRC6A | TMF1 | 0.802873406 | 9.94E-29 |
| UCS | TF | TNRC6A | ZNF585B | 0.814768436 | 3.47E-30 |
| UCS | TF | TNRC6A | LCOR | 0.814932145 | 3.31E-30 |
| UCS | TF | TNRC6A | ZNF333 | 0.816156828 | 2.31E-30 |
| UCS | TF | TNRC6A | ZNF236 | 0.831689018 | 1.90E-32 |
| UCS | TF | TNRC6A | ZNF81 | 0.84134804 | 7.41E-34 |
| UCS | TF | TNRC6A | ZNF587 | 0.843216298 | 3.86E-34 |
| UCS | TF | TNRC6A | BAZ2B | 0.844301464 | 2.63E-34 |
| UCS | TF | TNRC6A | MYSM1 | 0.864473218 | 1.20E-37 |
| UCS | TSG | TNRC6A | PHC3 | 0.810539677 | 1.18E-29 |
| UCS | TSG | TNRC6A | CREBBP | 0.81732804 | 1.64E-30 |
| UCS | TSG | TNRC6A | DICER1 | 0.828668929 | 5.01E-32 |
| UVM | Cell metabolism gene | TNRC6A | IDI1 | 0.800207244 | 5.35E-19 |
| UVM | Cell metabolism gene | TNRC6A | MED17 | 0.801449394 | 4.30E-19 |
| UVM | Cell metabolism gene | TNRC6A | PSMC6 | 0.801570885 | 4.21E-19 |
| UVM | Cell metabolism gene | TNRC6A | GPAM | 0.802602405 | 3.51E-19 |
| UVM | Cell metabolism gene | TNRC6A | XPO1 | 0.803671757 | 2.90E-19 |
| UVM | Cell metabolism gene | TNRC6A | ACADSB | 0.804489906 | 2.51E-19 |
| UVM | Cell metabolism gene | TNRC6A | MED4 | 0.805276796 | 2.18E-19 |
| UVM | Cell metabolism gene | TNRC6A | ENOPH1 | 0.807905282 | 1.35E-19 |
| UVM | Cell metabolism gene | TNRC6A | EDEM3 | 0.809293402 | 1.05E-19 |
| UVM | Cell metabolism gene | TNRC6A | TPH1 | 0.809452398 | 1.02E-19 |
| UVM | Cell metabolism gene | TNRC6A | TBL1XR1 | 0.810667778 | 8.13E-20 |
| UVM | Cell metabolism gene | TNRC6A | NUDT12 | 0.810895563 | 7.80E-20 |
| UVM | Cell metabolism gene | TNRC6A | PIK3R1 | 0.811059153 | 7.56E-20 |
| UVM | Cell metabolism gene | TNRC6A | GRPEL2 | 0.811690572 | 6.72E-20 |
| UVM | Cell metabolism gene | TNRC6A | MTMR6 | 0.813402719 | 4.88E-20 |
| UVM | Cell metabolism gene | TNRC6A | NFYB | 0.813584098 | 4.71E-20 |
| UVM | Cell metabolism gene | TNRC6A | DCP2 | 0.815113961 | 3.52E-20 |
| UVM | Cell metabolism gene | TNRC6A | DIS3 | 0.816740373 | 2.58E-20 |
| UVM | Cell metabolism gene | TNRC6A | RORA | 0.818294738 | 1.91E-20 |
| UVM | Cell metabolism gene | TNRC6A | DCK | 0.819559106 | 1.49E-20 |
| UVM | Cell metabolism gene | TNRC6A | AMY2B | 0.820193516 | 1.32E-20 |
| UVM | Cell metabolism gene | TNRC6A | NUP107 | 0.820600339 | 1.22E-20 |
| UVM | Cell metabolism gene | TNRC6A | PNPLA8 | 0.820819027 | 1.17E-20 |
| UVM | Cell metabolism gene | TNRC6A | PPP1CB | 0.821004942 | 1.12E-20 |
| UVM | Cell metabolism gene | TNRC6A | TRMT11 | 0.821847924 | 9.51E-21 |
| UVM | Cell metabolism gene | TNRC6A | GNAQ | 0.823861408 | 6.36E-21 |
| UVM | Cell metabolism gene | TNRC6A | MANEA | 0.826539618 | 3.69E-21 |
| UVM | Cell metabolism gene | TNRC6A | DBT | 0.827672114 | 2.92E-21 |
| UVM | Cell metabolism gene | TNRC6A | SLC19A2 | 0.828327347 | 2.55E-21 |
| UVM | Cell metabolism gene | TNRC6A | SACM1L | 0.828908361 | 2.26E-21 |
| UVM | Cell metabolism gene | TNRC6A | PDE8A | 0.83010515 | 1.76E-21 |
| UVM | Cell metabolism gene | TNRC6A | PLA2G12A | 0.83143557 | 1.33E-21 |
| UVM | Cell metabolism gene | TNRC6A | ALG13 | 0.837419172 | 3.67E-22 |
| UVM | Cell metabolism gene | TNRC6A | GNPDA2 | 0.846117229 | 5.12E-23 |
| UVM | Cell metabolism gene | TNRC6A | NUPL2 | 0.846938332 | 4.23E-23 |
| UVM | Cell metabolism gene | TNRC6A | NUPL1 | 0.847201498 | 3.97E-23 |
| UVM | Cell metabolism gene | TNRC6A | CSGALNACT2 | 0.849098335 | 2.53E-23 |
| UVM | Cell metabolism gene | TNRC6A | TRDMT1 | 0.853189648 | 9.42E-24 |
| UVM | Cell metabolism gene | TNRC6A | MTR | 0.853654954 | 8.40E-24 |
| UVM | Cell metabolism gene | TNRC6A | PRKAA1 | 0.85386135 | 7.98E-24 |
| UVM | Cell metabolism gene | TNRC6A | ALG10B | 0.853908579 | 7.89E-24 |
| UVM | Cell metabolism gene | TNRC6A | LPGAT1 | 0.854101081 | 7.52E-24 |
| UVM | Cell metabolism gene | TNRC6A | PIKFYVE | 0.855993087 | 4.69E-24 |
| UVM | Cell metabolism gene | TNRC6A | NUP54 | 0.857926937 | 2.88E-24 |
| UVM | Cell metabolism gene | TNRC6A | FBXL3 | 0.867469962 | 2.31E-25 |
| UVM | Cell metabolism gene | TNRC6A | FAR1 | 0.870413039 | 1.02E-25 |
| UVM | Cell metabolism gene | TNRC6A | RANBP2 | 0.870755995 | 9.26E-26 |
| UVM | Cell metabolism gene | TNRC6A | SMG1 | 0.882136551 | 3.18E-27 |
| UVM | Cell metabolism gene | TNRC6A | ETNK1 | 0.894776197 | 4.88E-29 |
| UVM | Cell metabolism gene | TNRC6A | FBXW7 | 0.901496005 | 4.24E-30 |
| UVM | Cell metabolism gene | TNRC6A | GXYLT1 | 0.907527858 | 4.06E-31 |
| UVM | CGC | TNRC6A | AKAP9 | 0.801666811 | 4.14E-19 |
| UVM | CGC | TNRC6A | XPO1 | 0.803671757 | 2.90E-19 |
| UVM | CGC | TNRC6A | USP8 | 0.804169908 | 2.66E-19 |
| UVM | CGC | TNRC6A | PALB2 | 0.806446497 | 1.76E-19 |
| UVM | CGC | TNRC6A | JAK2 | 0.809804717 | 9.54E-20 |
| UVM | CGC | TNRC6A | TBL1XR1 | 0.810667778 | 8.13E-20 |
| UVM | CGC | TNRC6A | PIK3R1 | 0.811059153 | 7.56E-20 |
| UVM | CGC | TNRC6A | ATF1 | 0.812180872 | 6.13E-20 |
| UVM | CGC | TNRC6A | KIF5B | 0.813426847 | 4.85E-20 |
| UVM | CGC | TNRC6A | ZMYM2 | 0.814576838 | 3.90E-20 |
| UVM | CGC | TNRC6A | RB1 | 0.818827398 | 1.72E-20 |
| UVM | CGC | TNRC6A | BMPR1A | 0.823051043 | 7.48E-21 |
| UVM | CGC | TNRC6A | GNAQ | 0.823861408 | 6.36E-21 |
| UVM | CGC | TNRC6A | KTN1 | 0.824873929 | 5.18E-21 |
| UVM | CGC | TNRC6A | ATR | 0.831166016 | 1.41E-21 |
| UVM | CGC | TNRC6A | APC | 0.832526314 | 1.06E-21 |
| UVM | CGC | TNRC6A | PWWP2A | 0.833246976 | 9.08E-22 |
| UVM | CGC | TNRC6A | ERCC4 | 0.833899565 | 7.89E-22 |
| UVM | CGC | TNRC6A | FBXO11 | 0.834737529 | 6.59E-22 |
| UVM | CGC | TNRC6A | KRAS | 0.835128604 | 6.05E-22 |
| UVM | CGC | TNRC6A | TRIM33 | 0.838415898 | 2.95E-22 |
| UVM | CGC | TNRC6A | PSIP1 | 0.840075742 | 2.04E-22 |
| UVM | CGC | TNRC6A | ARHGAP5 | 0.841111187 | 1.62E-22 |
| UVM | CGC | TNRC6A | MAP3K1 | 0.848033032 | 3.27E-23 |
| UVM | CGC | TNRC6A | SUZ12 | 0.849782215 | 2.15E-23 |
| UVM | CGC | TNRC6A | DDX5 | 0.852086159 | 1.23E-23 |
| UVM | CGC | TNRC6A | ATRX | 0.85241368 | 1.14E-23 |
| UVM | CGC | TNRC6A | CREB1 | 0.854597438 | 6.65E-24 |
| UVM | CGC | TNRC6A | MDM4 | 0.859241003 | 2.06E-24 |
| UVM | CGC | TNRC6A | BCLAF1 | 0.860075581 | 1.66E-24 |
| UVM | CGC | TNRC6A | SMAD4 | 0.860834021 | 1.36E-24 |
| UVM | CGC | TNRC6A | MLLT10 | 0.865386067 | 4.07E-25 |
| UVM | CGC | TNRC6A | GOPC | 0.868209673 | 1.88E-25 |
| UVM | CGC | TNRC6A | ARID2 | 0.8697744 | 1.22E-25 |
| UVM | CGC | TNRC6A | RANBP2 | 0.870755995 | 9.26E-26 |
| UVM | CGC | TNRC6A | PMS1 | 0.875688665 | 2.24E-26 |
| UVM | CGC | TNRC6A | TET2 | 0.887917165 | 5.01E-28 |
| UVM | CGC | TNRC6A | PHF6 | 0.890363678 | 2.22E-28 |
| UVM | CGC | TNRC6A | STAG2 | 0.891610803 | 1.46E-28 |
| UVM | CGC | TNRC6A | ACVR2A | 0.892063563 | 1.25E-28 |
| UVM | CGC | TNRC6A | SF3B1 | 0.892957616 | 9.19E-29 |
| UVM | CGC | TNRC6A | MALT1 | 0.893977387 | 6.46E-29 |
| UVM | CGC | TNRC6A | ETNK1 | 0.894776197 | 4.88E-29 |
| UVM | CGC | TNRC6A | RGPD3 | 0.900112395 | 7.12E-30 |
| UVM | CGC | TNRC6A | FBXW7 | 0.901496005 | 4.24E-30 |
| UVM | CGC | TNRC6A | DICER1 | 0.914037635 | 2.67E-32 |
| UVM | Epifactor | TNRC6A | DDX50 | 0.800767335 | 4.85E-19 |
| UVM | Epifactor | TNRC6A | NSL1 | 0.801390158 | 4.35E-19 |
| UVM | Epifactor | TNRC6A | CTBP2 | 0.802394583 | 3.64E-19 |
| UVM | Epifactor | TNRC6A | AEBP2 | 0.802709547 | 3.45E-19 |
| UVM | Epifactor | TNRC6A | TDG | 0.805167834 | 2.22E-19 |
| UVM | Epifactor | TNRC6A | SUV420H1 | 0.806099092 | 1.88E-19 |
| UVM | Epifactor | TNRC6A | EID1 | 0.806488747 | 1.75E-19 |
| UVM | Epifactor | TNRC6A | PHF14 | 0.808905678 | 1.13E-19 |
| UVM | Epifactor | TNRC6A | JAK2 | 0.809804717 | 9.54E-20 |
| UVM | Epifactor | TNRC6A | EPC1 | 0.809894691 | 9.39E-20 |
| UVM | Epifactor | TNRC6A | TBL1XR1 | 0.810667778 | 8.13E-20 |
| UVM | Epifactor | TNRC6A | RSF1 | 0.811139395 | 7.45E-20 |
| UVM | Epifactor | TNRC6A | PCGF6 | 0.811762724 | 6.63E-20 |
| UVM | Epifactor | TNRC6A | NFYB | 0.813584098 | 4.71E-20 |
| UVM | Epifactor | TNRC6A | ANKRD32 | 0.813988299 | 4.36E-20 |
| UVM | Epifactor | TNRC6A | TAF7 | 0.814040108 | 4.32E-20 |
| UVM | Epifactor | TNRC6A | ZMYM2 | 0.814576838 | 3.90E-20 |
| UVM | Epifactor | TNRC6A | SHPRH | 0.816003907 | 2.97E-20 |
| UVM | Epifactor | TNRC6A | SUDS3 | 0.816778717 | 2.56E-20 |
| UVM | Epifactor | TNRC6A | MYSM1 | 0.818773438 | 1.74E-20 |
| UVM | Epifactor | TNRC6A | RB1 | 0.818827398 | 1.72E-20 |
| UVM | Epifactor | TNRC6A | CDK17 | 0.818936915 | 1.69E-20 |
| UVM | Epifactor | TNRC6A | SIRT1 | 0.821099081 | 1.10E-20 |
| UVM | Epifactor | TNRC6A | MPHOSPH8 | 0.821425662 | 1.03E-20 |
| UVM | Epifactor | TNRC6A | SENP1 | 0.822173978 | 8.91E-21 |
| UVM | Epifactor | TNRC6A | DNAJC2 | 0.822450833 | 8.43E-21 |
| UVM | Epifactor | TNRC6A | MGEA5 | 0.823210678 | 7.24E-21 |
| UVM | Epifactor | TNRC6A | BRCC3 | 0.825770436 | 4.32E-21 |
| UVM | Epifactor | TNRC6A | TLK1 | 0.827889494 | 2.80E-21 |
| UVM | Epifactor | TNRC6A | USP15 | 0.829504135 | 2.00E-21 |
| UVM | Epifactor | TNRC6A | ATR | 0.831166016 | 1.41E-21 |
| UVM | Epifactor | TNRC6A | BRWD3 | 0.831352845 | 1.36E-21 |
| UVM | Epifactor | TNRC6A | SMARCAD1 | 0.834525471 | 6.90E-22 |
| UVM | Epifactor | TNRC6A | HMGB1 | 0.835162348 | 6.01E-22 |
| UVM | Epifactor | TNRC6A | HCFC2 | 0.835731667 | 5.31E-22 |
| UVM | Epifactor | TNRC6A | SMEK2 | 0.836993604 | 4.03E-22 |
| UVM | Epifactor | TNRC6A | TRIM33 | 0.838415898 | 2.95E-22 |
| UVM | Epifactor | TNRC6A | DNTTIP2 | 0.839914272 | 2.11E-22 |
| UVM | Epifactor | TNRC6A | PSIP1 | 0.840075742 | 2.04E-22 |
| UVM | Epifactor | TNRC6A | FAM175B | 0.840670594 | 1.78E-22 |
| UVM | Epifactor | TNRC6A | YEATS4 | 0.847203257 | 3.97E-23 |
| UVM | Epifactor | TNRC6A | CIR1 | 0.849365708 | 2.38E-23 |
| UVM | Epifactor | TNRC6A | SUZ12 | 0.849782215 | 2.15E-23 |
| UVM | Epifactor | TNRC6A | MBIP | 0.850206205 | 1.94E-23 |
| UVM | Epifactor | TNRC6A | CUL5 | 0.850582416 | 1.78E-23 |
| UVM | Epifactor | TNRC6A | CUL4B | 0.851249299 | 1.51E-23 |
| UVM | Epifactor | TNRC6A | TAF9B | 0.851822576 | 1.32E-23 |
| UVM | Epifactor | TNRC6A | ATRX | 0.85241368 | 1.14E-23 |
| UVM | Epifactor | TNRC6A | PRKAA1 | 0.85386135 | 7.98E-24 |
| UVM | Epifactor | TNRC6A | PHC3 | 0.854470811 | 6.86E-24 |
| UVM | Epifactor | TNRC6A | PCGF3 | 0.855005899 | 6.01E-24 |
| UVM | Epifactor | TNRC6A | RLIM | 0.856673416 | 3.96E-24 |
| UVM | Epifactor | TNRC6A | BRWD1 | 0.860781627 | 1.38E-24 |
| UVM | Epifactor | TNRC6A | ZMYND11 | 0.86367265 | 6.45E-25 |
| UVM | Epifactor | TNRC6A | MLLT10 | 0.865386067 | 4.07E-25 |
| UVM | Epifactor | TNRC6A | SMEK1 | 0.867822183 | 2.10E-25 |
| UVM | Epifactor | TNRC6A | ARID2 | 0.8697744 | 1.22E-25 |
| UVM | Epifactor | TNRC6A | ARID4B | 0.873170629 | 4.65E-26 |
| UVM | Epifactor | TNRC6A | EPC2 | 0.875694108 | 2.23E-26 |
| UVM | Epifactor | TNRC6A | CUL2 | 0.880611621 | 5.10E-27 |
| UVM | Epifactor | TNRC6A | CHD1 | 0.881545997 | 3.82E-27 |
| UVM | Epifactor | TNRC6A | ATAD2B | 0.8838118 | 1.88E-27 |
| UVM | Epifactor | TNRC6A | RMI1 | 0.88436771 | 1.58E-27 |
| UVM | Epifactor | TNRC6A | CHD9 | 0.884370649 | 1.58E-27 |
| UVM | Epifactor | TNRC6A | INO80D | 0.88494528 | 1.31E-27 |
| UVM | Epifactor | TNRC6A | BAZ2B | 0.885356676 | 1.15E-27 |
| UVM | Epifactor | TNRC6A | TET2 | 0.887917165 | 5.01E-28 |
| UVM | Epifactor | TNRC6A | PHIP | 0.888520008 | 4.11E-28 |
| UVM | Epifactor | TNRC6A | SF3B1 | 0.892957616 | 9.19E-29 |
| UVM | Epifactor | TNRC6A | UHRF2 | 0.893524774 | 7.56E-29 |
| UVM | Epifactor | TNRC6A | RCOR3 | 0.893596276 | 7.37E-29 |
| UVM | Epifactor | TNRC6A | NIPBL | 0.894406743 | 5.56E-29 |
| UVM | Epifactor | TNRC6A | OGT | 0.898469633 | 1.30E-29 |
| UVM | Epifactor | TNRC6A | ARID4A | 0.900174053 | 6.96E-30 |
| UVM | Epifactor | TNRC6A | JMJD1C | 0.901386301 | 4.42E-30 |
| UVM | Epifactor | TNRC6A | MBTD1 | 0.907774072 | 3.67E-31 |
| UVM | IUPHAR | TNRC6A | IDI1 | 0.800207244 | 5.35E-19 |
| UVM | IUPHAR | TNRC6A | TTBK2 | 0.80030867 | 5.26E-19 |
| UVM | IUPHAR | TNRC6A | MAP4K3 | 0.801942027 | 3.95E-19 |
| UVM | IUPHAR | TNRC6A | XPO1 | 0.803671757 | 2.90E-19 |
| UVM | IUPHAR | TNRC6A | NIPA1 | 0.804441857 | 2.53E-19 |
| UVM | IUPHAR | TNRC6A | TPH1 | 0.809452398 | 1.02E-19 |
| UVM | IUPHAR | TNRC6A | JAK2 | 0.809804717 | 9.54E-20 |
| UVM | IUPHAR | TNRC6A | PIK3R1 | 0.811059153 | 7.56E-20 |
| UVM | IUPHAR | TNRC6A | ATP11B | 0.813162262 | 5.10E-20 |
| UVM | IUPHAR | TNRC6A | TAS2R14 | 0.813448387 | 4.83E-20 |
| UVM | IUPHAR | TNRC6A | MAP3K2 | 0.813537548 | 4.75E-20 |
| UVM | IUPHAR | TNRC6A | PKN2 | 0.817372251 | 2.28E-20 |
| UVM | IUPHAR | TNRC6A | NEK7 | 0.818010902 | 2.02E-20 |
| UVM | IUPHAR | TNRC6A | RORA | 0.818294738 | 1.91E-20 |
| UVM | IUPHAR | TNRC6A | CDK17 | 0.818936915 | 1.69E-20 |
| UVM | IUPHAR | TNRC6A | SIRT1 | 0.821099081 | 1.10E-20 |
| UVM | IUPHAR | TNRC6A | SENP1 | 0.822173978 | 8.91E-21 |
| UVM | IUPHAR | TNRC6A | BMPR1A | 0.823051043 | 7.48E-21 |
| UVM | IUPHAR | TNRC6A | SLC25A27 | 0.82309933 | 7.41E-21 |
| UVM | IUPHAR | TNRC6A | ABCA5 | 0.823128778 | 7.36E-21 |
| UVM | IUPHAR | TNRC6A | GNAQ | 0.823861408 | 6.36E-21 |
| UVM | IUPHAR | TNRC6A | CYP2R1 | 0.824499439 | 5.59E-21 |
| UVM | IUPHAR | TNRC6A | CRBN | 0.826228153 | 3.93E-21 |
| UVM | IUPHAR | TNRC6A | TLK1 | 0.827889494 | 2.80E-21 |
| UVM | IUPHAR | TNRC6A | SLC19A2 | 0.828327347 | 2.55E-21 |
| UVM | IUPHAR | TNRC6A | PDE8A | 0.83010515 | 1.76E-21 |
| UVM | IUPHAR | TNRC6A | ATR | 0.831166016 | 1.41E-21 |
| UVM | IUPHAR | TNRC6A | YES1 | 0.831312778 | 1.37E-21 |
| UVM | IUPHAR | TNRC6A | BRWD3 | 0.831352845 | 1.36E-21 |
| UVM | IUPHAR | TNRC6A | PLA2G12A | 0.83143557 | 1.33E-21 |
| UVM | IUPHAR | TNRC6A | NAPEPLD | 0.832724251 | 1.01E-21 |
| UVM | IUPHAR | TNRC6A | IRAK4 | 0.833081562 | 9.40E-22 |
| UVM | IUPHAR | TNRC6A | PRPF4B | 0.834495226 | 6.94E-22 |
| UVM | IUPHAR | TNRC6A | FBXO11 | 0.834737529 | 6.59E-22 |
| UVM | IUPHAR | TNRC6A | KRAS | 0.835128604 | 6.05E-22 |
| UVM | IUPHAR | TNRC6A | TAS2R5 | 0.835874518 | 5.15E-22 |
| UVM | IUPHAR | TNRC6A | SLK | 0.837746948 | 3.42E-22 |
| UVM | IUPHAR | TNRC6A | TRIM33 | 0.838415898 | 2.95E-22 |
| UVM | IUPHAR | TNRC6A | SLC4A7 | 0.845731158 | 5.60E-23 |
| UVM | IUPHAR | TNRC6A | STK38L | 0.846095433 | 5.15E-23 |
| UVM | IUPHAR | TNRC6A | MAP3K1 | 0.848033032 | 3.27E-23 |
| UVM | IUPHAR | TNRC6A | MTR | 0.853654954 | 8.40E-24 |
| UVM | IUPHAR | TNRC6A | PRKAA1 | 0.85386135 | 7.98E-24 |
| UVM | IUPHAR | TNRC6A | SLC25A36 | 0.855959484 | 4.73E-24 |
| UVM | IUPHAR | TNRC6A | PIKFYVE | 0.855993087 | 4.69E-24 |
| UVM | IUPHAR | TNRC6A | TNKS2 | 0.856469004 | 4.16E-24 |
| UVM | IUPHAR | TNRC6A | BRWD1 | 0.860781627 | 1.38E-24 |
| UVM | IUPHAR | TNRC6A | ABCA10 | 0.861593313 | 1.12E-24 |
| UVM | IUPHAR | TNRC6A | CSNK1G3 | 0.86315845 | 7.39E-25 |
| UVM | IUPHAR | TNRC6A | ZMYND11 | 0.86367265 | 6.45E-25 |
| UVM | IUPHAR | TNRC6A | NEK1 | 0.864957851 | 4.57E-25 |
| UVM | IUPHAR | TNRC6A | RIOK3 | 0.865938847 | 3.51E-25 |
| UVM | IUPHAR | TNRC6A | BIRC2 | 0.866525187 | 2.99E-25 |
| UVM | IUPHAR | TNRC6A | NR2C1 | 0.872206434 | 6.14E-26 |
| UVM | IUPHAR | TNRC6A | XIAP | 0.872389796 | 5.82E-26 |
| UVM | IUPHAR | TNRC6A | TRPM7 | 0.882124342 | 3.19E-27 |
| UVM | IUPHAR | TNRC6A | SMG1 | 0.882136551 | 3.18E-27 |
| UVM | IUPHAR | TNRC6A | ATAD2B | 0.8838118 | 1.88E-27 |
| UVM | IUPHAR | TNRC6A | BAZ2B | 0.885356676 | 1.15E-27 |
| UVM | IUPHAR | TNRC6A | PHIP | 0.888520008 | 4.11E-28 |
| UVM | IUPHAR | TNRC6A | NR3C1 | 0.890494816 | 2.13E-28 |
| UVM | IUPHAR | TNRC6A | ACVR2A | 0.892063563 | 1.25E-28 |
| UVM | IUPHAR | TNRC6A | SENP6 | 0.893549166 | 7.49E-29 |
| UVM | IUPHAR | TNRC6A | MALT1 | 0.893977387 | 6.46E-29 |
| UVM | IUPHAR | TNRC6A | SENP7 | 0.899635538 | 8.49E-30 |
| UVM | IUPHAR | TNRC6A | CLK1 | 0.900799137 | 5.51E-30 |
| UVM | IUPHAR | TNRC6A | JMJD1C | 0.901386301 | 4.42E-30 |
| UVM | IUPHAR | TNRC6A | CLK4 | 0.92313933 | 4.05E-34 |
| UVM | Kinase | TNRC6A | TTBK2 | 0.80030867 | 5.26E-19 |
| UVM | Kinase | TNRC6A | MAP4K3 | 0.801942027 | 3.95E-19 |
| UVM | Kinase | TNRC6A | JAK2 | 0.809804717 | 9.54E-20 |
| UVM | Kinase | TNRC6A | MAP3K2 | 0.813537548 | 4.75E-20 |
| UVM | Kinase | TNRC6A | PKN2 | 0.817372251 | 2.28E-20 |
| UVM | Kinase | TNRC6A | NEK7 | 0.818010902 | 2.02E-20 |
| UVM | Kinase | TNRC6A | CDK17 | 0.818936915 | 1.69E-20 |
| UVM | Kinase | TNRC6A | BMPR1A | 0.823051043 | 7.48E-21 |
| UVM | Kinase | TNRC6A | TLK1 | 0.827889494 | 2.80E-21 |
| UVM | Kinase | TNRC6A | ATR | 0.831166016 | 1.41E-21 |
| UVM | Kinase | TNRC6A | YES1 | 0.831312778 | 1.37E-21 |
| UVM | Kinase | TNRC6A | IRAK4 | 0.833081562 | 9.40E-22 |
| UVM | Kinase | TNRC6A | PRPF4B | 0.834495226 | 6.94E-22 |
| UVM | Kinase | TNRC6A | SLK | 0.837746948 | 3.42E-22 |
| UVM | Kinase | TNRC6A | TRIM33 | 0.838415898 | 2.95E-22 |
| UVM | Kinase | TNRC6A | STK38L | 0.846095433 | 5.15E-23 |
| UVM | Kinase | TNRC6A | MAP3K1 | 0.848033032 | 3.27E-23 |
| UVM | Kinase | TNRC6A | PRKAA1 | 0.85386135 | 7.98E-24 |
| UVM | Kinase | TNRC6A | CSNK1G3 | 0.86315845 | 7.39E-25 |
| UVM | Kinase | TNRC6A | NEK1 | 0.864957851 | 4.57E-25 |
| UVM | Kinase | TNRC6A | RIOK3 | 0.865938847 | 3.51E-25 |
| UVM | Kinase | TNRC6A | TRPM7 | 0.882124342 | 3.19E-27 |
| UVM | Kinase | TNRC6A | SMG1 | 0.882136551 | 3.18E-27 |
| UVM | Kinase | TNRC6A | ACVR2A | 0.892063563 | 1.25E-28 |
| UVM | Kinase | TNRC6A | CLK1 | 0.900799137 | 5.51E-30 |
| UVM | Kinase | TNRC6A | PAN3 | 0.91206192 | 6.24E-32 |
| UVM | Kinase | TNRC6A | CLK4 | 0.92313933 | 4.05E-34 |
| UVM | TF | TNRC6A | ZNF506 | 0.801260149 | 4.45E-19 |
| UVM | TF | TNRC6A | ZNF600 | 0.801348248 | 4.38E-19 |
| UVM | TF | TNRC6A | ZNF502 | 0.801930884 | 3.95E-19 |
| UVM | TF | TNRC6A | AEBP2 | 0.802709547 | 3.45E-19 |
| UVM | TF | TNRC6A | ZNF709 | 0.80322878 | 3.14E-19 |
| UVM | TF | TNRC6A | ZNF286A | 0.803592566 | 2.95E-19 |
| UVM | TF | TNRC6A | TIGD4 | 0.804598733 | 2.46E-19 |
| UVM | TF | TNRC6A | ZBTB44 | 0.805295817 | 2.17E-19 |
| UVM | TF | TNRC6A | ZNF879 | 0.806269723 | 1.82E-19 |
| UVM | TF | TNRC6A | ZNF222 | 0.806402088 | 1.78E-19 |
| UVM | TF | TNRC6A | ZNF587 | 0.807172025 | 1.55E-19 |
| UVM | TF | TNRC6A | ZNF223 | 0.810456448 | 8.46E-20 |
| UVM | TF | TNRC6A | ZNF420 | 0.810660808 | 8.15E-20 |
| UVM | TF | TNRC6A | PCGF6 | 0.811762724 | 6.63E-20 |
| UVM | TF | TNRC6A | ZNF627 | 0.812099487 | 6.23E-20 |
| UVM | TF | TNRC6A | ATF1 | 0.812180872 | 6.13E-20 |
| UVM | TF | TNRC6A | VEZF1 | 0.813008564 | 5.25E-20 |
| UVM | TF | TNRC6A | ZNF326 | 0.813050667 | 5.21E-20 |
| UVM | TF | TNRC6A | NFYB | 0.813584098 | 4.71E-20 |
| UVM | TF | TNRC6A | ZNF480 | 0.814505859 | 3.96E-20 |
| UVM | TF | TNRC6A | ZNF564 | 0.814527654 | 3.94E-20 |
| UVM | TF | TNRC6A | ZNF267 | 0.814905135 | 3.67E-20 |
| UVM | TF | TNRC6A | ZNF236 | 0.815645671 | 3.18E-20 |
| UVM | TF | TNRC6A | ZFP62 | 0.817736143 | 2.13E-20 |
| UVM | TF | TNRC6A | RORA | 0.818294738 | 1.91E-20 |
| UVM | TF | TNRC6A | ZNF546 | 0.818411009 | 1.87E-20 |
| UVM | TF | TNRC6A | MYSM1 | 0.818773438 | 1.74E-20 |
| UVM | TF | TNRC6A | ZNF547 | 0.819042735 | 1.65E-20 |
| UVM | TF | TNRC6A | ZNF654 | 0.819160812 | 1.61E-20 |
| UVM | TF | TNRC6A | ZNF397 | 0.819476671 | 1.52E-20 |
| UVM | TF | TNRC6A | ZNF230 | 0.819596824 | 1.48E-20 |
| UVM | TF | TNRC6A | TTF1 | 0.819606074 | 1.48E-20 |
| UVM | TF | TNRC6A | ZNF594 | 0.820084343 | 1.35E-20 |
| UVM | TF | TNRC6A | ZNF81 | 0.820225389 | 1.31E-20 |
| UVM | TF | TNRC6A | ZNF714 | 0.820288807 | 1.29E-20 |
| UVM | TF | TNRC6A | ZNF514 | 0.82095766 | 1.13E-20 |
| UVM | TF | TNRC6A | ZFP30 | 0.821539506 | 1.01E-20 |
| UVM | TF | TNRC6A | FOXN2 | 0.822859002 | 7.77E-21 |
| UVM | TF | TNRC6A | ZNF440 | 0.823592306 | 6.71E-21 |
| UVM | TF | TNRC6A | TOPORS | 0.825338032 | 4.72E-21 |
| UVM | TF | TNRC6A | ZNF570 | 0.826825942 | 3.48E-21 |
| UVM | TF | TNRC6A | MYNN | 0.827232513 | 3.20E-21 |
| UVM | TF | TNRC6A | ZNF235 | 0.828673772 | 2.38E-21 |
| UVM | TF | TNRC6A | ZNF765 | 0.829068238 | 2.19E-21 |
| UVM | TF | TNRC6A | ZNF484 | 0.829119557 | 2.17E-21 |
| UVM | TF | TNRC6A | ZNF443 | 0.830024522 | 1.79E-21 |
| UVM | TF | TNRC6A | ZNF799 | 0.830880334 | 1.50E-21 |
| UVM | TF | TNRC6A | ZNF131 | 0.831432058 | 1.33E-21 |
| UVM | TF | TNRC6A | ZBTB6 | 0.83325453 | 9.06E-22 |
| UVM | TF | TNRC6A | ZNF791 | 0.834745353 | 6.58E-22 |
| UVM | TF | TNRC6A | TIGD7 | 0.834763047 | 6.55E-22 |
| UVM | TF | TNRC6A | PRDM10 | 0.834788548 | 6.52E-22 |
| UVM | TF | TNRC6A | ZNF585B | 0.835320108 | 5.81E-22 |
| UVM | TF | TNRC6A | ZNF675 | 0.835335743 | 5.79E-22 |
| UVM | TF | TNRC6A | ZNF569 | 0.836487326 | 4.51E-22 |
| UVM | TF | TNRC6A | GABPA | 0.836813238 | 4.20E-22 |
| UVM | TF | TNRC6A | ZNF770 | 0.837733343 | 3.43E-22 |
| UVM | TF | TNRC6A | ZNF780A | 0.838405214 | 2.96E-22 |
| UVM | TF | TNRC6A | ZBTB41 | 0.838586573 | 2.84E-22 |
| UVM | TF | TNRC6A | ZNF383 | 0.839756022 | 2.19E-22 |
| UVM | TF | TNRC6A | ZBTB34 | 0.841908517 | 1.35E-22 |
| UVM | TF | TNRC6A | ZNF721 | 0.842350754 | 1.22E-22 |
| UVM | TF | TNRC6A | ZNF354A | 0.842537906 | 1.17E-22 |
| UVM | TF | TNRC6A | MEF2A | 0.842547555 | 1.17E-22 |
| UVM | TF | TNRC6A | ZNF461 | 0.842736725 | 1.12E-22 |
| UVM | TF | TNRC6A | ZSCAN29 | 0.843104731 | 1.03E-22 |
| UVM | TF | TNRC6A | ZNF808 | 0.843406763 | 9.59E-23 |
| UVM | TF | TNRC6A | SKIL | 0.843675658 | 9.01E-23 |
| UVM | TF | TNRC6A | ZNF143 | 0.843792962 | 8.77E-23 |
| UVM | TF | TNRC6A | ZNF655 | 0.844060744 | 8.25E-23 |
| UVM | TF | TNRC6A | ZNF345 | 0.844520065 | 7.42E-23 |
| UVM | TF | TNRC6A | SP3 | 0.844609457 | 7.27E-23 |
| UVM | TF | TNRC6A | ZNF169 | 0.84535164 | 6.12E-23 |
| UVM | TF | TNRC6A | ZNF800 | 0.845924349 | 5.36E-23 |
| UVM | TF | TNRC6A | ZNF227 | 0.846240447 | 4.98E-23 |
| UVM | TF | TNRC6A | THAP5 | 0.846631 | 4.54E-23 |
| UVM | TF | TNRC6A | ZNF519 | 0.846803695 | 4.36E-23 |
| UVM | TF | TNRC6A | ZNF280C | 0.848205917 | 3.13E-23 |
| UVM | TF | TNRC6A | ZNF845 | 0.84896841 | 2.61E-23 |
| UVM | TF | TNRC6A | RBAK | 0.849081332 | 2.55E-23 |
| UVM | TF | TNRC6A | ZNF430 | 0.849510257 | 2.30E-23 |
| UVM | TF | TNRC6A | ZNF260 | 0.8496476 | 2.22E-23 |
| UVM | TF | TNRC6A | ZNF43 | 0.850043442 | 2.02E-23 |
| UVM | TF | TNRC6A | ZNF615 | 0.85136526 | 1.47E-23 |
| UVM | TF | TNRC6A | ZNF624 | 0.851454485 | 1.44E-23 |
| UVM | TF | TNRC6A | ZNF148 | 0.851705365 | 1.35E-23 |
| UVM | TF | TNRC6A | ZNF28 | 0.85190422 | 1.29E-23 |
| UVM | TF | TNRC6A | JRKL | 0.852351236 | 1.16E-23 |
| UVM | TF | TNRC6A | ZNF347 | 0.852365057 | 1.15E-23 |
| UVM | TF | TNRC6A | ZNF558 | 0.853910905 | 7.88E-24 |
| UVM | TF | TNRC6A | ZNF253 | 0.854465529 | 6.87E-24 |
| UVM | TF | TNRC6A | IKZF5 | 0.854574895 | 6.69E-24 |
| UVM | TF | TNRC6A | CREB1 | 0.854597438 | 6.65E-24 |
| UVM | TF | TNRC6A | ZBTB49 | 0.854763865 | 6.38E-24 |
| UVM | TF | TNRC6A | EEA1 | 0.854945551 | 6.10E-24 |
| UVM | TF | TNRC6A | ZNF644 | 0.855073845 | 5.91E-24 |
| UVM | TF | TNRC6A | ZNF189 | 0.857093966 | 3.56E-24 |
| UVM | TF | TNRC6A | ZNF449 | 0.85823576 | 2.66E-24 |
| UVM | TF | TNRC6A | ZNF585A | 0.858319982 | 2.60E-24 |
| UVM | TF | TNRC6A | ZNF701 | 0.859114973 | 2.12E-24 |
| UVM | TF | TNRC6A | PURB | 0.85928054 | 2.04E-24 |
| UVM | TF | TNRC6A | ZNF470 | 0.859295169 | 2.03E-24 |
| UVM | TF | TNRC6A | ZNF12 | 0.859823101 | 1.77E-24 |
| UVM | TF | TNRC6A | ZNF417 | 0.859823357 | 1.77E-24 |
| UVM | TF | TNRC6A | ZNF226 | 0.860357025 | 1.54E-24 |
| UVM | TF | TNRC6A | ZFP28 | 0.860591573 | 1.45E-24 |
| UVM | TF | TNRC6A | SMAD4 | 0.860834021 | 1.36E-24 |
| UVM | TF | TNRC6A | ZNF268 | 0.861968016 | 1.01E-24 |
| UVM | TF | TNRC6A | ZFX | 0.862656486 | 8.44E-25 |
| UVM | TF | TNRC6A | AHCTF1 | 0.862678286 | 8.39E-25 |
| UVM | TF | TNRC6A | ZNF24 | 0.862747681 | 8.24E-25 |
| UVM | TF | TNRC6A | ZNF566 | 0.863746571 | 6.32E-25 |
| UVM | TF | TNRC6A | ZNF41 | 0.865318795 | 4.15E-25 |
| UVM | TF | TNRC6A | ZNF254 | 0.865412314 | 4.04E-25 |
| UVM | TF | TNRC6A | LIN54 | 0.865412487 | 4.04E-25 |
| UVM | TF | TNRC6A | NFXL1 | 0.866473351 | 3.03E-25 |
| UVM | TF | TNRC6A | ZNF507 | 0.86849044 | 1.74E-25 |
| UVM | TF | TNRC6A | ZNF182 | 0.86957679 | 1.29E-25 |
| UVM | TF | TNRC6A | ZNF266 | 0.869622904 | 1.27E-25 |
| UVM | TF | TNRC6A | ARID2 | 0.8697744 | 1.22E-25 |
| UVM | TF | TNRC6A | ZNF761 | 0.86999385 | 1.15E-25 |
| UVM | TF | TNRC6A | ZNF44 | 0.870751601 | 9.27E-26 |
| UVM | TF | TNRC6A | NR2C1 | 0.872206434 | 6.14E-26 |
| UVM | TF | TNRC6A | ZNF140 | 0.8737778 | 3.91E-26 |
| UVM | TF | TNRC6A | SMAD5 | 0.874382716 | 3.28E-26 |
| UVM | TF | TNRC6A | ZNF674 | 0.875359555 | 2.46E-26 |
| UVM | TF | TNRC6A | ZNF527 | 0.875965281 | 2.06E-26 |
| UVM | TF | TNRC6A | ZNF320 | 0.87608258 | 1.99E-26 |
| UVM | TF | TNRC6A | ZBTB11 | 0.876612445 | 1.70E-26 |
| UVM | TF | TNRC6A | ZNF37A | 0.876659522 | 1.68E-26 |
| UVM | TF | TNRC6A | ZNF141 | 0.877155166 | 1.45E-26 |
| UVM | TF | TNRC6A | ZNF708 | 0.877939863 | 1.15E-26 |
| UVM | TF | TNRC6A | ZNF441 | 0.878862147 | 8.69E-27 |
| UVM | TF | TNRC6A | ZBTB1 | 0.879493406 | 7.17E-27 |
| UVM | TF | TNRC6A | ZNF432 | 0.879585584 | 6.97E-27 |
| UVM | TF | TNRC6A | ZNF83 | 0.879699029 | 6.74E-27 |
| UVM | TF | TNRC6A | ZNF75D | 0.879917477 | 6.30E-27 |
| UVM | TF | TNRC6A | ZNF354B | 0.880153936 | 5.86E-27 |
| UVM | TF | TNRC6A | ZNF23 | 0.881080537 | 4.41E-27 |
| UVM | TF | TNRC6A | TMF1 | 0.881551786 | 3.81E-27 |
| UVM | TF | TNRC6A | ZNF91 | 0.881975683 | 3.34E-27 |
| UVM | TF | TNRC6A | ZNF14 | 0.882024538 | 3.29E-27 |
| UVM | TF | TNRC6A | ZNF195 | 0.884327837 | 1.60E-27 |
| UVM | TF | TNRC6A | THAP6 | 0.884466474 | 1.53E-27 |
| UVM | TF | TNRC6A | ZBED5 | 0.884503107 | 1.51E-27 |
| UVM | TF | TNRC6A | ZNF700 | 0.884958889 | 1.31E-27 |
| UVM | TF | TNRC6A | ZNF136 | 0.88506139 | 1.26E-27 |
| UVM | TF | TNRC6A | BAZ2B | 0.885356676 | 1.15E-27 |
| UVM | TF | TNRC6A | ELF2 | 0.885386806 | 1.14E-27 |
| UVM | TF | TNRC6A | ZNF181 | 0.88586966 | 9.76E-28 |
| UVM | TF | TNRC6A | ZNF26 | 0.885995553 | 9.37E-28 |
| UVM | TF | TNRC6A | ZNF33A | 0.886494986 | 7.97E-28 |
| UVM | TF | TNRC6A | ZNF611 | 0.886610955 | 7.68E-28 |
| UVM | TF | TNRC6A | ZNF234 | 0.887145051 | 6.45E-28 |
| UVM | TF | TNRC6A | TET2 | 0.887917165 | 5.01E-28 |
| UVM | TF | TNRC6A | ZNF180 | 0.88800929 | 4.86E-28 |
| UVM | TF | TNRC6A | GPBP1 | 0.889272385 | 3.20E-28 |
| UVM | TF | TNRC6A | ZNF280D | 0.889773364 | 2.71E-28 |
| UVM | TF | TNRC6A | NR3C1 | 0.890494816 | 2.13E-28 |
| UVM | TF | TNRC6A | ZNF658 | 0.890495583 | 2.13E-28 |
| UVM | TF | TNRC6A | ZNF292 | 0.890558119 | 2.08E-28 |
| UVM | TF | TNRC6A | ZMAT1 | 0.892067839 | 1.25E-28 |
| UVM | TF | TNRC6A | ZNF25 | 0.896548279 | 2.61E-29 |
| UVM | TF | TNRC6A | ZNF567 | 0.89672971 | 2.44E-29 |
| UVM | TF | TNRC6A | ZFP14 | 0.89684784 | 2.34E-29 |
| UVM | TF | TNRC6A | ZNF224 | 0.897405268 | 1.92E-29 |
| UVM | TF | TNRC6A | CREBZF | 0.898059069 | 1.51E-29 |
| UVM | TF | TNRC6A | ZNF302 | 0.901028893 | 5.06E-30 |
| UVM | TF | TNRC6A | ZNF510 | 0.902005047 | 3.50E-30 |
| UVM | TF | TNRC6A | THAP9 | 0.904908509 | 1.15E-30 |
| UVM | TF | TNRC6A | ZNF431 | 0.907438209 | 4.21E-31 |
| UVM | TF | TNRC6A | ZNF160 | 0.907709399 | 3.77E-31 |
| UVM | TF | TNRC6A | ZNF529 | 0.908462757 | 2.78E-31 |
| UVM | TF | TNRC6A | ZNF493 | 0.908756872 | 2.47E-31 |
| UVM | TF | TNRC6A | ZNF84 | 0.909429764 | 1.87E-31 |
| UVM | TF | TNRC6A | ZNF782 | 0.913777169 | 2.99E-32 |
| UVM | TF | TNRC6A | ZNF518A | 0.916065321 | 1.09E-32 |
| UVM | TF | TNRC6A | ZNF780B | 0.916392176 | 9.46E-33 |
| UVM | TF | TNRC6A | ZNF606 | 0.918686358 | 3.34E-33 |
| UVM | TF | TNRC6A | ZNF10 | 0.920593566 | 1.37E-33 |
| UVM | TF | TNRC6A | DMTF1 | 0.933103269 | 2.18E-36 |
| UVM | TF | TNRC6A | ZNF248 | 0.935170288 | 6.67E-37 |
| UVM | TSG | TNRC6A | WDR11 | 0.804081413 | 2.70E-19 |
| UVM | TSG | TNRC6A | NEDD4 | 0.806312548 | 1.81E-19 |
| UVM | TSG | TNRC6A | PALB2 | 0.806446497 | 1.76E-19 |
| UVM | TSG | TNRC6A | RASSF8 | 0.808255768 | 1.27E-19 |
| UVM | TSG | TNRC6A | LIN9 | 0.812898912 | 5.36E-20 |
| UVM | TSG | TNRC6A | FAM188A | 0.81455974 | 3.92E-20 |
| UVM | TSG | TNRC6A | SHPRH | 0.816003907 | 2.97E-20 |
| UVM | TSG | TNRC6A | MCM9 | 0.816416311 | 2.75E-20 |
| UVM | TSG | TNRC6A | RB1 | 0.818827398 | 1.72E-20 |
| UVM | TSG | TNRC6A | TTF1 | 0.819606074 | 1.48E-20 |
| UVM | TSG | TNRC6A | USP33 | 0.820844959 | 1.16E-20 |
| UVM | TSG | TNRC6A | SIRT1 | 0.821099081 | 1.10E-20 |
| UVM | TSG | TNRC6A | BMPR1A | 0.823051043 | 7.48E-21 |
| UVM | TSG | TNRC6A | TOPORS | 0.825338032 | 4.72E-21 |
| UVM | TSG | TNRC6A | COPS2 | 0.828113974 | 2.67E-21 |
| UVM | TSG | TNRC6A | SMCHD1 | 0.828885982 | 2.28E-21 |
| UVM | TSG | TNRC6A | INTS6 | 0.83098399 | 1.47E-21 |
| UVM | TSG | TNRC6A | ATR | 0.831166016 | 1.41E-21 |
| UVM | TSG | TNRC6A | APC | 0.832526314 | 1.06E-21 |
| UVM | TSG | TNRC6A | NAPEPLD | 0.832724251 | 1.01E-21 |
| UVM | TSG | TNRC6A | MLH3 | 0.842544679 | 1.17E-22 |
| UVM | TSG | TNRC6A | SKIL | 0.843675658 | 9.01E-23 |
| UVM | TSG | TNRC6A | PPM1A | 0.848702075 | 2.79E-23 |
| UVM | TSG | TNRC6A | SUZ12 | 0.849782215 | 2.15E-23 |
| UVM | TSG | TNRC6A | CUL5 | 0.850582416 | 1.78E-23 |
| UVM | TSG | TNRC6A | CCAR1 | 0.85226499 | 1.18E-23 |
| UVM | TSG | TNRC6A | VEZT | 0.853856043 | 7.99E-24 |
| UVM | TSG | TNRC6A | PRKAA1 | 0.85386135 | 7.98E-24 |
| UVM | TSG | TNRC6A | PHC3 | 0.854470811 | 6.86E-24 |
| UVM | TSG | TNRC6A | SMAD4 | 0.860834021 | 1.36E-24 |
| UVM | TSG | TNRC6A | GORAB | 0.862680345 | 8.39E-25 |
| UVM | TSG | TNRC6A | ZMYND11 | 0.86367265 | 6.45E-25 |
| UVM | TSG | TNRC6A | APAF1 | 0.868842172 | 1.58E-25 |
| UVM | TSG | TNRC6A | ARID2 | 0.8697744 | 1.22E-25 |
| UVM | TSG | TNRC6A | DCLRE1A | 0.876540263 | 1.74E-26 |
| UVM | TSG | TNRC6A | KRIT1 | 0.879193159 | 7.86E-27 |
| UVM | TSG | TNRC6A | CUL2 | 0.880611621 | 5.10E-27 |
| UVM | TSG | TNRC6A | CHD1 | 0.881545997 | 3.82E-27 |
| UVM | TSG | TNRC6A | TET2 | 0.887917165 | 5.01E-28 |
| UVM | TSG | TNRC6A | PHF6 | 0.890363678 | 2.22E-28 |
| UVM | TSG | TNRC6A | ZNF292 | 0.890558119 | 2.08E-28 |
| UVM | TSG | TNRC6A | TANK | 0.89187271 | 1.33E-28 |
| UVM | TSG | TNRC6A | UHRF2 | 0.893524774 | 7.56E-29 |
| UVM | TSG | TNRC6A | PNN | 0.895928763 | 3.25E-29 |
| UVM | TSG | TNRC6A | FBXW7 | 0.901496005 | 4.24E-30 |
| UVM | TSG | TNRC6A | DICER1 | 0.914037635 | 2.67E-32 |
| UVM | TSG | TNRC6A | DMTF1 | 0.933103269 | 2.18E-36 |
Top |
|
 Protein 3D structure Protein 3D structureVisit iCn3D. |
Top |
|
 Protein-protein interaction networks Protein-protein interaction networks * Overlap between up-regulated DEGs (log2FC<-1 and adj.P<0.05) and STRING PPI network (center: Translation factor, node: DEGs, edges: weighted by -log2(adj.P)) |
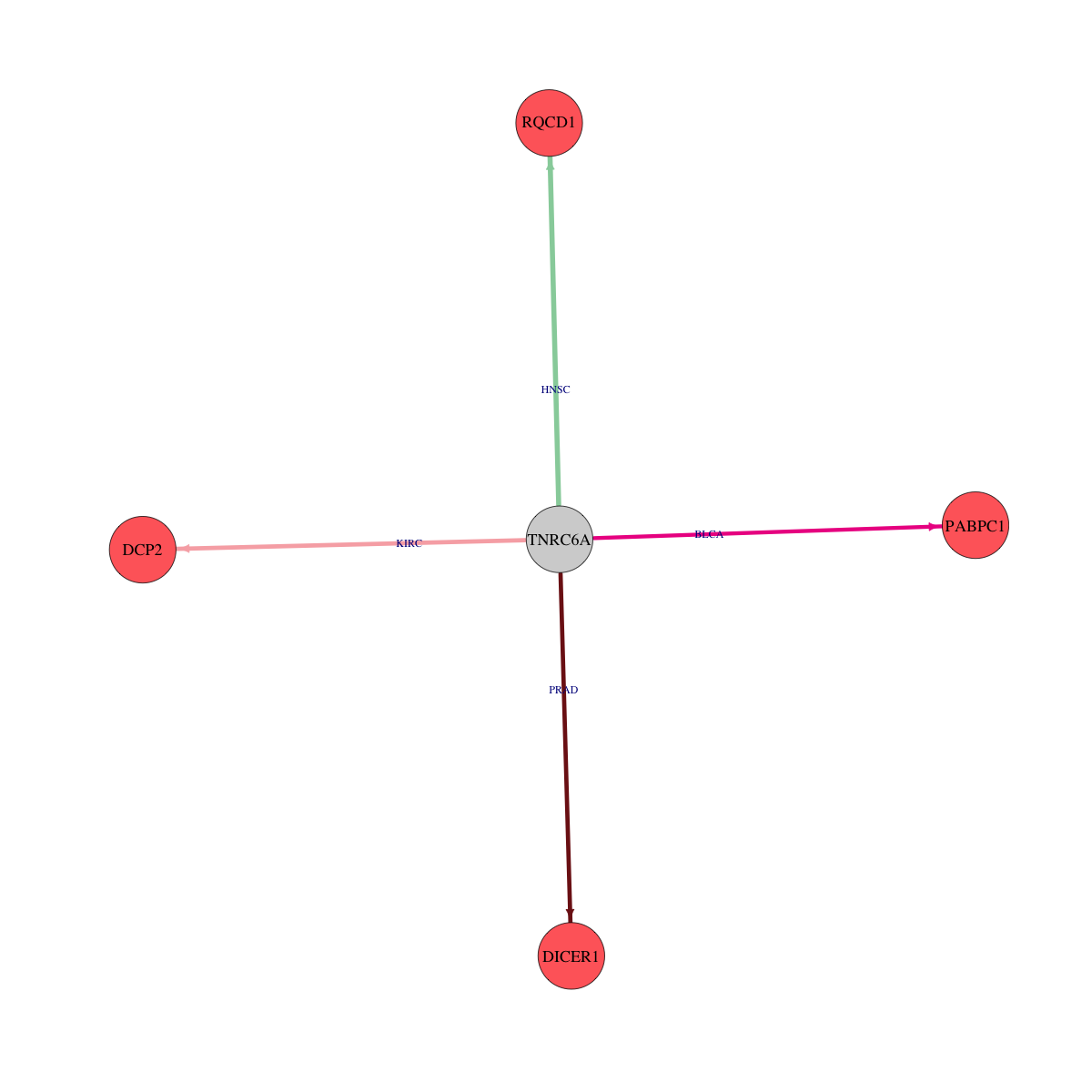 |
 Overlap between down-regulated DEGs (log2FC>1 and adj.P<0.05) and STRING PPI network (center: Translation factor, node: DEGs, edges: weighted by -log2(adj.P)) Overlap between down-regulated DEGs (log2FC>1 and adj.P<0.05) and STRING PPI network (center: Translation factor, node: DEGs, edges: weighted by -log2(adj.P)) |
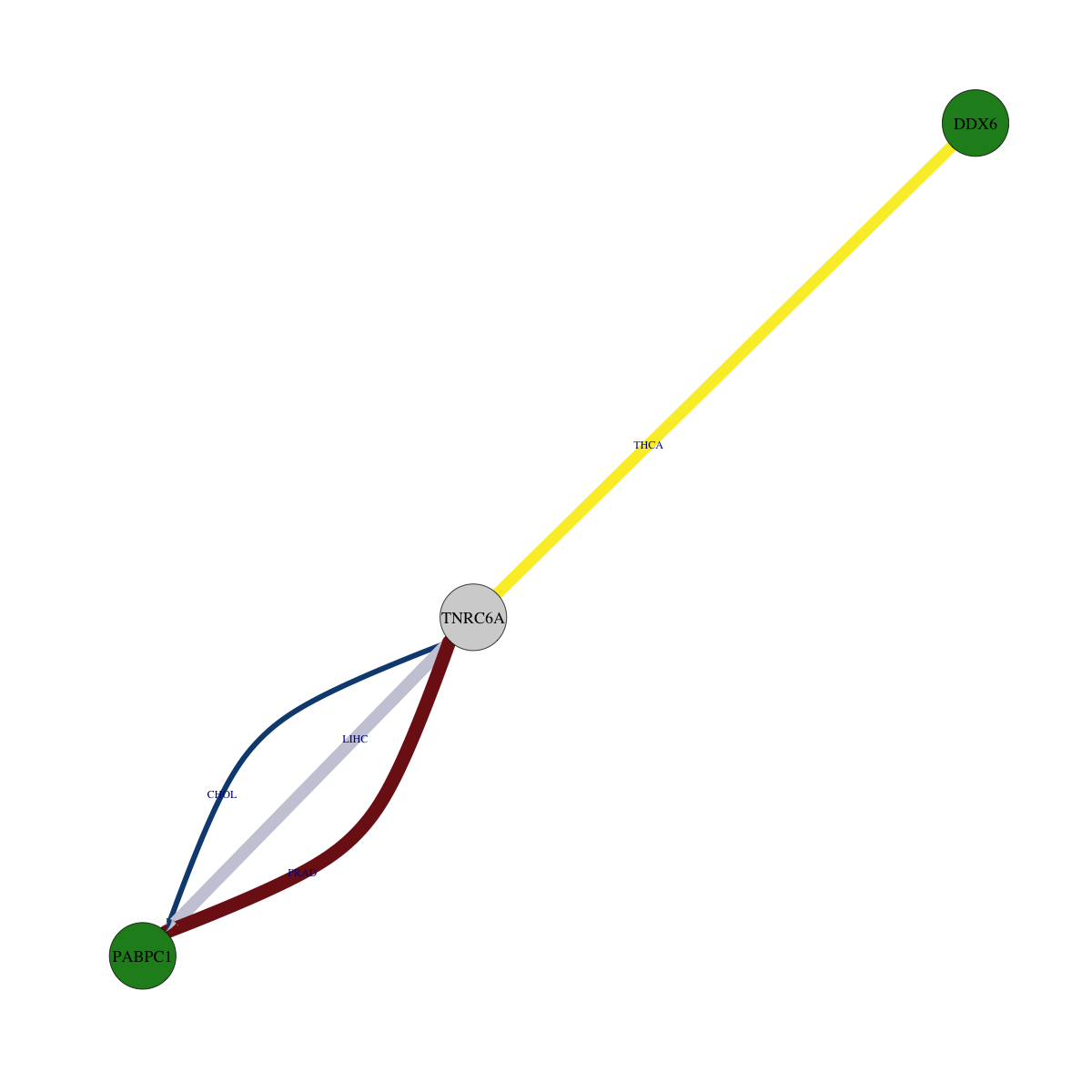 |
 * Edge colors based on TCGA cancer types. |
 * Overlap between DEGs (log2FC>1 and adj.P<0.05) and STRING PPI network per cancer (center: Translation factor, node: DEGs, node color: log2FC, edges: weighted by -log2(adj.P)) * Overlap between DEGs (log2FC>1 and adj.P<0.05) and STRING PPI network per cancer (center: Translation factor, node: DEGs, node color: log2FC, edges: weighted by -log2(adj.P)) |
 |
| Cancer type | Translation factor | Interacting protein coding gene | FC | adj.pval |
| CHOL | TNRC6A | PABPC1 | -4.56272528815353 | 0.00390625 |
| HNSC | TNRC6A | RQCD1 | 1.97297540658491 | 0.00585215220849023 |
| KIRC | TNRC6A | DCP2 | 1.26475649682567 | 0.0124189994647713 |
| PRAD | TNRC6A | DICER1 | 1.54768558737379 | 0.0130781232419534 |
| BLCA | TNRC6A | PABPC1 | 1.31485302792746 | 0.0180816650390625 |
| LIHC | TNRC6A | PABPC1 | -1.72076961362188 | 2.98009622125841e-06 |
| THCA | TNRC6A | DDX6 | -4.18593714762416 | 8.3083675979354e-06 |
| PRAD | TNRC6A | PABPC1 | -2.10200318363476 | 8.55379952759014e-07 |
 Protein-protein interactors with this translation factor (BIOGRID-3.4.160) Protein-protein interactors with this translation factor (BIOGRID-3.4.160) |
| PPI interactors with TNRC6A |
| AGO4, AGO2, WNK1, UBC, PABPC1, UBR5, CUL3, AGO1, AGO3, SAV1, TRIM65, CEP250, TRIP13, CNOT1, CNOT3, CNOT6L, CNOT7, CNOT10, PAN2, CNOT2, CNOT6, CNOT8, PAN3, BTRC, SSX2IP, CEP152, CNTROB, CEP135, CNTRL, NINL, ODF2, RPGRIP1L, CENPJ, ITGB4, CLTB, CLTC, PICALM, GTSE1, FANCM, KIAA2013, RQCD1, Ubr5, EYA2, SMARCB1, BRCA1, ESR2, EZH2, SUZ12, SFPQ, TRIM24, MED12, AHNAK2, PSPC1, PCF11, MED1, KMT2B, EIF2AK4, NCOA6, CPSF7, MED14, CLP1, MED17, MED13, TJP2, CDK5RAP3, CNOT11, ANAPC4, SRSF6, SEC63, TAP1, MED23, MED4, MED6, SCD, TP53BP1, CCAR2, ANAPC1, ARID1A, TNKS1BP1, GATAD2A, AFF4, CDC23, MDC1, ANAPC7, ANAPC5, CDC16, CDC27, NAT10, NES, SMARCD1, WDR5, KMT2C, ASH2L, RBBP5, ZNF24, MED27, HIST3H3, TOMM20, ZC3H7A, IKBKG, TNIP2, NR2C2, PATL1, DDX6, ZBTB10, BICD2, BICD1, NIN, ARIH2, MIB1, HCVgp1, HIST2H2BC, nsp12, nsp7, ESR1, ANKRD17, CELF1, CEP85, CPEB4, DAZL, FAM120C, FUBP3, KIAA0355, MEX3B, PRRC2A, PUM1, R3HDM2, ANKHD1-EIF4EBP3, DCP1B, DLG5, EIF4E2, EIF4ENIF1, GIGYF1, GIGYF2, HELZ, KIAA0430, PRRC2B, RC3H1, RC3H2, RNF219, SMG5, SMG7, TBK1, TBKBP1, TNRC6B, TNRC6C, AP2A1, AP2B1, AP2M1, CEP192, CYLD, DCP1A, DCP2, EDC3, EDC4, EDRF1, FAM175B, FAM184A, FAM193A, FBF1, GPBP1, GPBP1L1, ITSN2, KIZ, KLHL15, LZTS2, MIB2, MPHOSPH9, NME7, SCLT1, SEC23B, SPATA2, TCHP, TRIP11, USP54, UBAP2L, UNK, YTHDF1, YTHDF2, YTHDF3, ZFP36, MOV10, ALG13, KANK2, SYNM, TNPO1, BTG3, SEC16A, AP2A2, SEC24B, TXLNG, ZC3HAV1, MKRN2, RBMS1, CNOT4, OTUD4, TNPO2, CPVL, DZIP3, KIF14, KIF20A, PHIP, TRIM66, TRIM37, ACTR3, AKAP1, AMOT, ANAPC2, CLTA, EMD, GFAP, IMPDH2, KDM1A, KRT18, KRT19, KRT8, LATS1, MLLT4, NUP155, PRPH, PXN, RPS20, SASS6, SERBP1, SQSTM1, STIL, STX4, SYNE3, VASP, VIM, ZYX, FKBP9, AK4, QKI, DMRTB1, TXNL4A, RBFOX2, Ssbp1, N, CPEB1, GLI3, ISX, RB1CC1, |
Top |
|
 Clinically associated variants from ClinVar. Clinically associated variants from ClinVar. |
| Gene | Chr | Position | RefSeq | VarSeq | RefSeeq | VarType | Pathogenic | Disease | VarInfo |
| TNRC6A | chr16 | 24788422 | AGCAGCCACAGCC | A | Deletion | Benign | not_provided | SO:0001822|inframe_deletion | SO:0001822|inframe_deletion |
| TNRC6A | chr16 | 24788549 | G | A | single_nucleotide_variant | Benign | not_provided | SO:0001819|synonymous_variant | SO:0001819|synonymous_variant |
| TNRC6A | chr16 | 24788645 | T | A | single_nucleotide_variant | Benign | not_provided | SO:0001583|missense_variant | SO:0001583|missense_variant |
| TNRC6A | chr16 | 24801145 | G | A | single_nucleotide_variant | Uncertain_significance | not_provided | SO:0001819|synonymous_variant | SO:0001819|synonymous_variant |
| TNRC6A | chr16 | 24801638 | A | G | single_nucleotide_variant | Benign | not_provided | SO:0001583|missense_variant | SO:0001583|missense_variant |
| TNRC6A | chr16 | 24802366 | G | A | single_nucleotide_variant | Benign | not_provided | SO:0001819|synonymous_variant | SO:0001819|synonymous_variant |
| TNRC6A | chr16 | 24802572 | A | G | single_nucleotide_variant | Likely_benign | not_provided | SO:0001583|missense_variant | SO:0001583|missense_variant |
| TNRC6A | chr16 | 24802972 | T | C | single_nucleotide_variant | Benign | not_provided | SO:0001819|synonymous_variant | SO:0001819|synonymous_variant |
| TNRC6A | chr16 | 24817614 | G | A | single_nucleotide_variant | Benign | not_provided | SO:0001627|intron_variant | SO:0001627|intron_variant |
| TNRC6A | chr16 | 24818114 | C | G | single_nucleotide_variant | Likely_benign | not_provided | SO:0001627|intron_variant | SO:0001627|intron_variant |
| TNRC6A | chr16 | 24831640 | T | C | single_nucleotide_variant | Uncertain_significance | not_provided | SO:0001583|missense_variant | SO:0001583|missense_variant |
| TNRC6A | chr16 | 24834950 | G | A | single_nucleotide_variant | Likely_benign | not_provided | SO:0001583|missense_variant | SO:0001583|missense_variant |
 nsSNVs with sample frequency (size of circle) from TCGA 33 cancers. nsSNVs with sample frequency (size of circle) from TCGA 33 cancers. |
 SNVs and Indels SNVs and Indels |
| Gene | Cancer type | Chromosome | Start | End | RefSeeq | MutSeq | Mutation type | AAchange | # samples |
| TNRC6A | BLCA | chr16 | 24826541 | 24826541 | A | C | Silent | 7 | |
| TNRC6A | COAD | chr16 | 24788423 | 24788434 | GCAGCCACAGCC | - | In_Frame_Del | p.111_115del | 7 |
| TNRC6A | KIRC | chr16 | 24802365 | 24802366 | - | G | Frame_Shift_Ins | p.R801fs | 5 |
| TNRC6A | CESC | chr16 | 24788506 | 24788506 | G | A | Missense_Mutation | p.R139Q | 5 |
| TNRC6A | LUAD | chr16 | 24817548 | 24817548 | A | G | Missense_Mutation | p.R1425G | 5 |
| TNRC6A | BLCA | chr16 | 24807240 | 24807240 | A | - | Frame_Shift_Del | p.K1182fs | 4 |
| TNRC6A | SKCM | chr16 | 24828142 | 24828142 | C | T | Missense_Mutation | p.R1613C | 4 |
| TNRC6A | HNSC | chr16 | 24826581 | 24826581 | C | T | Missense_Mutation | p.R1596C | 4 |
| TNRC6A | KIRP | chr16 | 24826534 | 24826534 | G | A | Missense_Mutation | p.S1580N | 4 |
| TNRC6A | PCPG | chr16 | 24817584 | 24817584 | C | T | Missense_Mutation | p.R1437W | 4 |
| TNRC6A | BRCA | chr16 | 24807329 | 24807332 | ATTT | - | Frame_Shift_Del | p.I192fs | 4 |
| TNRC6A | LGG | chr16 | 24817937 | 24817937 | C | T | Missense_Mutation | p.R1458C | 4 |
| TNRC6A | SKCM | chr16 | 24801035 | 24801035 | C | T | Missense_Mutation | p.P358S | 4 |
| TNRC6A | UCEC | chr16 | 24826565 | 24826565 | A | G | Missense_Mutation | p.I1590M | 3 |
| TNRC6A | LUAD | chr16 | 24834793 | 24834793 | A | T | Missense_Mutation | p.S1852C | 3 |
| TNRC6A | BLCA | chr16 | 24807240 | 24807240 | A | - | Frame_Shift_Del | p.K1181fs | 3 |
| TNRC6A | SARC | chr16 | 24815560 | 24815560 | C | T | Nonsense_Mutation | p.R1253* | 3 |
| TNRC6A | LIHC | chr16 | 24762087 | 24762087 | A | - | Frame_Shift_Del | p.K35fs | 3 |
| TNRC6A | BRCA | chr16 | 24801088 | 24801088 | C | T | Silent | p.N375 | 3 |
| TNRC6A | PCPG | chr16 | 24817584 | 24817584 | C | T | Missense_Mutation | 3 | |
| TNRC6A | STAD | chr16 | 24802366 | 24802366 | G | - | Frame_Shift_Del | p.Q801fs | 3 |
| TNRC6A | UCEC | chr16 | 24802677 | 24802677 | A | G | Missense_Mutation | p.N905S | 3 |
| TNRC6A | BLCA | chr16 | 24788506 | 24788506 | G | T | Missense_Mutation | 3 | |
| TNRC6A | BRCA | chr16 | 24801817 | 24801817 | C | A | Missense_Mutation | p.N618K | 3 |
| TNRC6A | UCEC | chr16 | 24831626 | 24831626 | T | A | Missense_Mutation | p.N1749K | 3 |
| TNRC6A | LUAD | chr16 | 24801933 | 24801933 | A | T | Missense_Mutation | p.N657I | 3 |
| TNRC6A | LIHC | chr16 | 24816089 | 24816089 | C | - | Frame_Shift_Del | p.P1302fs | 3 |
| TNRC6A | HNSC | chr16 | 24802394 | 24802394 | A | G | Missense_Mutation | p.T811A | 3 |
| TNRC6A | LIHC | chr16 | 24834929 | 24834929 | T | G | Missense_Mutation | p.L1897R | 3 |
| TNRC6A | LUAD | chr16 | 24762097 | 24762097 | A | C | Missense_Mutation | p.K35T | 3 |
| TNRC6A | LIHC | chr16 | 24818061 | 24818061 | A | - | Frame_Shift_Del | p.Q1499fs | 3 |
| TNRC6A | ESCA | chr16 | 24818001 | 24818001 | A | G | Missense_Mutation | p.H1479R | 3 |
| TNRC6A | LIHC | chr16 | 24826473 | 24826473 | A | C | Missense_Mutation | 3 | |
| TNRC6A | BRCA | chr16 | 24831678 | 24831678 | C | T | Missense_Mutation | p.P1767S | 3 |
| TNRC6A | BLCA | chr16 | 24802570 | 24802570 | C | T | Silent | p.A869A | 3 |
| TNRC6A | KIRP | chr16 | 24801321 | 24801321 | C | G | Missense_Mutation | p.P453R | 3 |
| TNRC6A | BRCA | chr16 | 24833445 | 24833445 | G | A | Missense_Mutation | p.V1784I | 3 |
| TNRC6A | HNSC | chr16 | 24826581 | 24826581 | C | T | Missense_Mutation | 3 | |
| TNRC6A | HNSC | chr16 | 24835045 | 24835045 | G | A | Missense_Mutation | p.D1936N | 3 |
| TNRC6A | UCEC | chr16 | 24802899 | 24802899 | C | T | Missense_Mutation | p.A979V | 3 |
| TNRC6A | LIHC | chr16 | 24831552 | 24831552 | A | - | Frame_Shift_Del | p.K1725fs | 3 |
| TNRC6A | LIHC | chr16 | 24834929 | 24834929 | T | G | Missense_Mutation | 3 | |
| TNRC6A | COAD | chr16 | 24835087 | 24835087 | C | A | Missense_Mutation | p.L1950I | 3 |
| TNRC6A | KIRP | chr16 | 24788620 | 24788620 | C | G | Missense_Mutation | p.A177G | 3 |
| TNRC6A | SKCM | chr16 | 24802755 | 24802755 | C | T | Missense_Mutation | p.S931F | 3 |
| TNRC6A | UCEC | chr16 | 24788584 | 24788584 | C | A | Missense_Mutation | p.A165D | 3 |
| TNRC6A | CHOL | chr16 | 24788389 | 24788389 | A | C | Missense_Mutation | 3 | |
| TNRC6A | SKCM | chr16 | 24801572 | 24801572 | C | T | Missense_Mutation | p.P537S | 2 |
| TNRC6A | KIRP | chr16 | 24802526 | 24802526 | G | - | Frame_Shift_Del | p.D855fs | 2 |
| TNRC6A | STAD | chr16 | 24803029 | 24803029 | G | T | Missense_Mutation | p.W1022C | 2 |
| TNRC6A | BLCA | chr16 | 24815594 | 24815594 | A | G | Missense_Mutation | p.Q1264R | 2 |
| TNRC6A | CESC | chr16 | 24800641 | 24800641 | G | A | Silent | 2 | |
| TNRC6A | LIHC | chr16 | 24816121 | 24816121 | C | A | Silent | p.A1311A | 2 |
| TNRC6A | UCEC | chr16 | 24808845 | 24808845 | C | T | Missense_Mutation | p.A1199V | 2 |
| TNRC6A | HNSC | chr16 | 24800997 | 24800997 | G | C | Missense_Mutation | p.W345S | 2 |
| TNRC6A | STAD | chr16 | 24834270 | 24834270 | G | A | Missense_Mutation | p.G1817R | 2 |
| TNRC6A | BLCA | chr16 | 24803020 | 24803020 | G | A | Silent | p.V1019V | 2 |
| TNRC6A | STAD | chr16 | 24815625 | 24815625 | T | C | Silent | p.N1274N | 2 |
| TNRC6A | LGG | chr16 | 24817937 | 24817937 | C | T | Missense_Mutation | 2 | |
| TNRC6A | UCEC | chr16 | 24801346 | 24801346 | C | A | Silent | p.T461 | 2 |
| TNRC6A | CHOL | chr16 | 24826538 | 24826538 | T | G | Silent | 2 | |
| TNRC6A | ESCA | chr16 | 24818001 | 24818001 | A | G | Missense_Mutation | 2 | |
| TNRC6A | STAD | chr16 | 24816169 | 24816169 | A | T | Missense_Mutation | p.R1327S | 2 |
| TNRC6A | BLCA | chr16 | 24815516 | 24815516 | G | A | Missense_Mutation | p.R1238Q | 2 |
| TNRC6A | CESC | chr16 | 24805874 | 24805874 | C | G | Missense_Mutation | 2 | |
| TNRC6A | LIHC | chr16 | 24788429 | 24788429 | A | G | Silent | p.P113P | 2 |
| TNRC6A | UCEC | chr16 | 24815565 | 24815565 | G | A | Silent | p.G194S | 2 |
| TNRC6A | LUAD | chr16 | 24801038 | 24801038 | A | T | Missense_Mutation | p.S359C | 2 |
| TNRC6A | LGG | chr16 | 24788441 | 24788441 | G | A | Silent | p.P117P | 2 |
| TNRC6A | BLCA | chr16 | 24816470 | 24816470 | C | T | Nonsense_Mutation | p.Q1374* | 2 |
| TNRC6A | STAD | chr16 | 24820702 | 24820702 | T | A | Missense_Mutation | p.D1524E | 2 |
| TNRC6A | LIHC | chr16 | 24816374 | 24816374 | G | T | Missense_Mutation | 2 | |
| TNRC6A | UCEC | chr16 | 24801529 | 24801529 | G | A | Nonsense_Mutation | p.W522* | 2 |
| TNRC6A | CHOL | chr16 | 24826538 | 24826538 | T | G | Silent | p.T1581T | 2 |
| TNRC6A | ESCA | chr16 | 24788375 | 24788375 | A | G | Silent | 2 | |
| TNRC6A | SKCM | chr16 | 24826599 | 24826599 | G | A | Missense_Mutation | p.G1602S | 2 |
| TNRC6A | STAD | chr16 | 24788506 | 24788506 | G | T | Missense_Mutation | p.R139L | 2 |
| TNRC6A | BLCA | chr16 | 24817006 | 24817006 | G | A | Silent | p.L1401L | 2 |
| TNRC6A | LIHC | chr16 | 24831569 | 24831569 | G | A | Silent | p.P1730P | 2 |
| TNRC6A | UCEC | chr16 | 24816352 | 24816352 | G | A | Missense_Mutation | p.M1334I | 2 |
| TNRC6A | UCEC | chr16 | 24741605 | 24741605 | G | T | Nonsense_Mutation | p.E13* | 2 |
| TNRC6A | LUAD | chr16 | 24804935 | 24804935 | C | G | Missense_Mutation | p.S1106C | 2 |
| TNRC6A | UCS | chr16 | 24834300 | 24834300 | G | T | Nonsense_Mutation | p.E1827* | 2 |
| TNRC6A | STAD | chr16 | 24817959 | 24817959 | T | C | Missense_Mutation | p.L1465S | 2 |
| TNRC6A | BLCA | chr16 | 24802034 | 24802034 | C | T | Nonsense_Mutation | p.Q691* | 2 |
| TNRC6A | STAD | chr16 | 24828182 | 24828182 | C | A | Missense_Mutation | p.P1626H | 2 |
| TNRC6A | THYM | chr16 | 24788444 | 24788444 | G | A | Silent | p.Q118Q | 2 |
| TNRC6A | UCEC | chr16 | 24802006 | 24802006 | C | T | Silent | p.R681 | 2 |
| TNRC6A | SKCM | chr16 | 24826600 | 24826600 | G | A | Missense_Mutation | p.G1602D | 2 |
| TNRC6A | HNSC | chr16 | 24801674 | 24801674 | G | A | Missense_Mutation | p.G571S | 2 |
| TNRC6A | BLCA | chr16 | 24802299 | 24802299 | C | T | Missense_Mutation | p.S779F | 2 |
| TNRC6A | LUAD | chr16 | 24803129 | 24803129 | A | T | Missense_Mutation | p.S1056C | 2 |
| TNRC6A | UCEC | chr16 | 24818078 | 24818078 | A | G | Missense_Mutation | p.I1505V | 2 |
| TNRC6A | LIHC | chr16 | 24800799 | 24800799 | A | - | Frame_Shift_Del | p.E279fs | 2 |
| TNRC6A | UCEC | chr16 | 24762107 | 24762107 | G | T | Missense_Mutation | p.K38N | 2 |
| TNRC6A | LUAD | chr16 | 24826516 | 24826516 | A | T | Missense_Mutation | p.Y1574F | 2 |
| TNRC6A | HNSC | chr16 | 24800705 | 24800705 | G | C | Missense_Mutation | p.E248Q | 2 |
| TNRC6A | STAD | chr16 | 24802783 | 24802783 | A | T | Silent | p.P940P | 2 |
| TNRC6A | BLCA | chr16 | 24802570 | 24802570 | C | T | Silent | 2 | |
| TNRC6A | BLCA | chr16 | 24801938 | 24801938 | C | G | Missense_Mutation | p.Q659E | 2 |
| TNRC6A | STAD | chr16 | 24803058 | 24803058 | G | A | Missense_Mutation | p.R1032H | 2 |
| TNRC6A | UCEC | chr16 | 24802265 | 24802265 | G | T | Missense_Mutation | p.D768Y | 2 |
| TNRC6A | COAD | chr16 | 24788434 | 24788434 | C | A | Missense_Mutation | p.P115Q | 2 |
| TNRC6A | SKCM | chr16 | 24817563 | 24817563 | A | G | Missense_Mutation | p.R1430G | 2 |
| TNRC6A | HNSC | chr16 | 24834933 | 24834933 | T | A | Missense_Mutation | p.N1898K | 2 |
| TNRC6A | UCEC | chr16 | 24826566 | 24826566 | G | T | Nonsense_Mutation | p.G1591* | 2 |
| TNRC6A | UCEC | chr16 | 24788401 | 24788401 | C | T | Missense_Mutation | p.P104L | 2 |
| TNRC6A | ESCA | chr16 | 24815534 | 24815534 | A | T | Missense_Mutation | p.H1244L | 2 |
| TNRC6A | HNSC | chr16 | 24800634 | 24800634 | A | T | Missense_Mutation | p.N224I | 2 |
| TNRC6A | STAD | chr16 | 24802619 | 24802619 | G | A | Missense_Mutation | p.G886S | 2 |
| TNRC6A | BLCA | chr16 | 24802435 | 24802435 | G | C | Missense_Mutation | p.E824D | 2 |
| TNRC6A | STAD | chr16 | 24802947 | 24802947 | T | C | Missense_Mutation | p.I995T | 2 |
| TNRC6A | TGCT | chr16 | 24826529 | 24826529 | C | T | Silent | 2 | |
| TNRC6A | UCEC | chr16 | 24802307 | 24802307 | G | A | Missense_Mutation | p.G782R | 2 |
| TNRC6A | SKCM | chr16 | 24817561 | 24817561 | A | G | Missense_Mutation | p.Q1429R | 2 |
| TNRC6A | LGG | chr16 | 24788378 | 24788378 | G | T | Missense_Mutation | p.Q96H | 2 |
| TNRC6A | STAD | chr16 | 24803039 | 24803039 | G | A | Missense_Mutation | p.V1026I | 2 |
| TNRC6A | UCEC | chr16 | 24829957 | 24829957 | C | A | Silent | p.L563M | 2 |
| TNRC6A | UCEC | chr16 | 24788441 | 24788441 | G | A | Silent | p.P117 | 2 |
| TNRC6A | CESC | chr16 | 24831599 | 24831599 | G | T | Missense_Mutation | p.Q1740H | 2 |
| TNRC6A | STAD | chr16 | 24802952 | 24802952 | C | T | Missense_Mutation | p.R997C | 2 |
| TNRC6A | BLCA | chr16 | 24826603 | 24826603 | C | G | Missense_Mutation | p.S1603C | 2 |
| TNRC6A | BLCA | chr16 | 24801369 | 24801369 | C | T | Missense_Mutation | 2 | |
| TNRC6A | UCEC | chr16 | 24802415 | 24802415 | C | T | Nonsense_Mutation | p.Q818* | 2 |
| TNRC6A | SKCM | chr16 | 24831623 | 24831623 | T | A | Missense_Mutation | p.D1748E | 2 |
| TNRC6A | SKCM | chr16 | 24831582 | 24831582 | C | A | Missense_Mutation | p.P1735T | 2 |
| TNRC6A | STAD | chr16 | 24802647 | 24802647 | C | A | Missense_Mutation | p.S895Y | 2 |
| TNRC6A | UCEC | chr16 | 24829988 | 24829988 | G | A | Missense_Mutation | p.V1683I | 2 |
| TNRC6A | LIHC | chr16 | 24835083 | 24835083 | T | - | Frame_Shift_Del | p.A1948fs | 2 |
| TNRC6A | UCEC | chr16 | 24788596 | 24788596 | T | C | Missense_Mutation | p.L169S | 2 |
| TNRC6A | LGG | chr16 | 24802981 | 24802981 | A | C | Silent | 2 | |
| TNRC6A | STAD | chr16 | 24834864 | 24834864 | C | T | Silent | p.L1875L | 2 |
| TNRC6A | BLCA | chr16 | 24808841 | 24808841 | G | C | Missense_Mutation | p.E1198Q | 2 |
| TNRC6A | STAD | chr16 | 24805922 | 24805922 | G | T | Missense_Mutation | p.R1137L | 2 |
| TNRC6A | UCEC | chr16 | 24802458 | 24802458 | C | T | Missense_Mutation | p.S832L | 2 |
| TNRC6A | SKCM | chr16 | 24831624 | 24831624 | A | C | Missense_Mutation | p.N1749H | 2 |
| TNRC6A | SKCM | chr16 | 24816999 | 24816999 | C | T | Missense_Mutation | p.A1399V | 2 |
| TNRC6A | STAD | chr16 | 24817979 | 24817979 | C | T | Missense_Mutation | p.P1472S | 2 |
| TNRC6A | BLCA | chr16 | 24800967 | 24800967 | C | T | Missense_Mutation | p.S335F | 2 |
| TNRC6A | UCEC | chr16 | 24831637 | 24831637 | G | A | Missense_Mutation | p.R1753H | 2 |
| TNRC6A | UCEC | chr16 | 24788637 | 24788637 | C | T | Missense_Mutation | p.P183S | 2 |
| TNRC6A | ESCA | chr16 | 24788428 | 24788428 | C | A | Missense_Mutation | p.P113Q | 2 |
| TNRC6A | STAD | chr16 | 24820702 | 24820702 | T | A | Missense_Mutation | 2 | |
| TNRC6A | LGG | chr16 | 24802982 | 24802982 | G | C | Missense_Mutation | 2 | |
| TNRC6A | STAD | chr16 | 24788440 | 24788440 | C | A | Missense_Mutation | p.P117Q | 2 |
| TNRC6A | CESC | chr16 | 24831599 | 24831599 | G | T | Missense_Mutation | 2 | |
| TNRC6A | LUAD | chr16 | 24801619 | 24801619 | G | T | Missense_Mutation | p.Q552H | 2 |
| TNRC6A | UCEC | chr16 | 24802773 | 24802773 | C | T | Missense_Mutation | p.S937L | 2 |
| TNRC6A | SKCM | chr16 | 24801472 | 24801472 | C | T | Silent | p.S503S | 2 |
| TNRC6A | HNSC | chr16 | 24829935 | 24829935 | C | T | Missense_Mutation | p.T1665I | 2 |
| TNRC6A | LGG | chr16 | 24802267 | 24802267 | T | C | Silent | p.D768D | 2 |
| TNRC6A | STAD | chr16 | 24806029 | 24806029 | A | G | Missense_Mutation | p.M1173V | 2 |
| TNRC6A | LIHC | chr16 | 24802369 | 24802369 | G | A | Silent | p.G802G | 2 |
| TNRC6A | UCEC | chr16 | 24834197 | 24834197 | C | T | Silent | p.R683* | 2 |
| TNRC6A | BLCA | chr16 | 24801369 | 24801369 | C | T | Missense_Mutation | p.S469L | 2 |
| TNRC6A | UCEC | chr16 | 24800899 | 24800899 | A | G | Silent | p.P312 | 2 |
| TNRC6A | ESCA | chr16 | 24802699 | 24802699 | A | T | Missense_Mutation | p.E912D | 2 |
| TNRC6A | STAD | chr16 | 24815625 | 24815625 | T | C | Silent | 2 | |
| TNRC6A | LGG | chr16 | 24802267 | 24802267 | T | C | Silent | 2 | |
| TNRC6A | SKCM | chr16 | 24802347 | 24802347 | C | T | Missense_Mutation | p.S795F | 2 |
| TNRC6A | KIRP | chr16 | 24802526 | 24802526 | G | - | Frame_Shift_Del | p.S854fs | 2 |
| TNRC6A | LIHC | chr16 | 24816121 | 24816121 | C | A | Silent | 2 | |
| TNRC6A | BLCA | chr16 | 24818089 | 24818089 | C | T | Silent | p.F1508F | 2 |
| TNRC6A | CESC | chr16 | 24835021 | 24835021 | C | A | Missense_Mutation | 2 | |
| TNRC6A | UCEC | chr16 | 24802902 | 24802904 | AAG | - | In_Frame_Del | p.E983in_frame_del | 2 |
| TNRC6A | HNSC | chr16 | 24816384 | 24816384 | G | A | Missense_Mutation | p.R1345Q | 2 |
| TNRC6A | LGG | chr16 | 24802981 | 24802981 | A | C | Silent | p.S1006S | 2 |
| TNRC6A | STAD | chr16 | 24801521 | 24801521 | A | G | Missense_Mutation | p.T520A | 2 |
| TNRC6A | BLCA | chr16 | 24815516 | 24815516 | G | A | Missense_Mutation | 2 | |
| TNRC6A | UCEC | chr16 | 24834277 | 24834277 | C | T | Missense_Mutation | p.A1819V | 2 |
| TNRC6A | BLCA | chr16 | 24801938 | 24801938 | C | G | Missense_Mutation | 2 | |
| TNRC6A | UCEC | chr16 | 24801003 | 24801003 | C | G | Missense_Mutation | p.T347S | 2 |
| TNRC6A | KIRC | chr16 | 24788568 | 24788568 | G | A | Missense_Mutation | p.A160T | 2 |
| TNRC6A | STAD | chr16 | 24816407 | 24816407 | C | G | Missense_Mutation | p.Q1353E | 2 |
| TNRC6A | LGG | chr16 | 24788378 | 24788378 | G | T | Missense_Mutation | 2 | |
| TNRC6A | STAD | chr16 | 24817552 | 24817552 | C | T | Missense_Mutation | p.A1426V | 2 |
| TNRC6A | LIHC | chr16 | 24831569 | 24831569 | G | A | Silent | 2 | |
| TNRC6A | CESC | chr16 | 24826503 | 24826503 | C | T | Missense_Mutation | 2 | |
| TNRC6A | LIHC | chr16 | 24802560 | 24802560 | G | - | Frame_Shift_Del | p.W866fs | 2 |
| TNRC6A | LUAD | chr16 | 24802953 | 24802953 | G | T | Missense_Mutation | p.R997L | 2 |
| TNRC6A | UCEC | chr16 | 24805947 | 24805947 | G | T | Nonsense_Mutation | p.G85* | 2 |
| TNRC6A | HNSC | chr16 | 24801451 | 24801451 | A | T | Silent | p.P496P | 2 |
| TNRC6A | LGG | chr16 | 24802982 | 24802982 | G | C | Missense_Mutation | p.A1007P | 2 |
| TNRC6A | LUAD | chr16 | 24834870 | 24834870 | C | T | Silent | p.S1877S | 2 |
| TNRC6A | UCEC | chr16 | 24835005 | 24835005 | G | A | Silent | p.A813T | 2 |
| TNRC6A | THYM | chr16 | 24816122 | 24816122 | C | A | Missense_Mutation | 2 | |
| TNRC6A | UCEC | chr16 | 24801151 | 24801151 | T | C | Silent | p.S396 | 2 |
| TNRC6A | ESCA | chr16 | 24815534 | 24815534 | A | T | Missense_Mutation | 2 | |
| TNRC6A | KIRC | chr16 | 24816069 | 24816069 | C | T | Missense_Mutation | p.S1294F | 2 |
| TNRC6A | LGG | chr16 | 24788379 | 24788379 | C | T | Nonsense_Mutation | 2 | |
| TNRC6A | PAAD | chr16 | 24831545 | 24831545 | G | A | Silent | 1 | |
| TNRC6A | HNSC | chr16 | 24800705 | 24800705 | G | C | Missense_Mutation | 1 | |
| TNRC6A | SKCM | chr16 | 24804818 | 24804818 | C | T | Missense_Mutation | p.P1067L | 1 |
| TNRC6A | UCEC | chr16 | 24835005 | 24835005 | G | A | Silent | p.P1922P | 1 |
| TNRC6A | LUAD | chr16 | 24802584 | 24802584 | G | T | Missense_Mutation | p.S874I | 1 |
| TNRC6A | COAD | chr16 | 24802313 | 24802313 | G | T | Missense_Mutation | p.G784C | 1 |
| TNRC6A | LIHC | chr16 | 24802674 | 24802675 | - | A | Frame_Shift_Ins | p.Q904fs | 1 |
| TNRC6A | UCEC | chr16 | 24802902 | 24802904 | AAG | - | In_Frame_Del | p.E983del | 1 |
| TNRC6A | LUAD | chr16 | 24828156 | 24828156 | A | G | Silent | p.P1617P | 1 |
| TNRC6A | UCS | chr16 | 24802366 | 24802366 | G | - | Frame_Shift_Del | 1 | |
| TNRC6A | LUAD | chr16 | 24801401 | 24801401 | A | T | Missense_Mutation | p.T480S | 1 |
| TNRC6A | DLBC | chr16 | 24802245 | 24802245 | T | G | Missense_Mutation | p.F761C | 1 |
| TNRC6A | READ | chr16 | 24801675 | 24801675 | G | A | Missense_Mutation | p.G571D | 1 |
| TNRC6A | HNSC | chr16 | 24818089 | 24818089 | C | T | Silent | 1 | |
| TNRC6A | SKCM | chr16 | 24803121 | 24803121 | A | G | Missense_Mutation | p.E1053G | 1 |
| TNRC6A | LGG | chr16 | 24800571 | 24800571 | C | G | Nonsense_Mutation | p.S203* | 1 |
| TNRC6A | BLCA | chr16 | 24817006 | 24817006 | G | A | Silent | 1 | |
| TNRC6A | KIRC | chr16 | 24802866 | 24802866 | C | A | Missense_Mutation | p.T968K | 1 |
| TNRC6A | STAD | chr16 | 24802194 | 24802194 | G | A | Missense_Mutation | p.R744Q | 1 |
| TNRC6A | LIHC | chr16 | 24834848 | 24834848 | C | A | Missense_Mutation | 1 | |
| TNRC6A | THYM | chr16 | 24818020 | 24818021 | TT | - | Frame_Shift_Del | 1 | |
| TNRC6A | LIHC | chr16 | 24788537 | 24788537 | A | - | Frame_Shift_Del | p.L149fs | 1 |
| TNRC6A | MESO | chr16 | 24788482 | 24788482 | G | A | Missense_Mutation | 1 | |
| TNRC6A | SARC | chr16 | 24817054 | 24817054 | G | T | Splice_Site | p.Q1417_splice | 1 |
| TNRC6A | COAD | chr16 | 24802325 | 24802325 | C | T | Missense_Mutation | p.P788S | 1 |
| TNRC6A | PAAD | chr16 | 24788479 | 24788479 | C | A | Missense_Mutation | p.P130H | 1 |
| TNRC6A | HNSC | chr16 | 24801451 | 24801451 | A | T | Silent | 1 | |
| TNRC6A | SKCM | chr16 | 24834251 | 24834251 | C | T | Silent | p.F1810F | 1 |
| TNRC6A | KIRP | chr16 | 24801321 | 24801321 | C | G | Missense_Mutation | 1 | |
| TNRC6A | LIHC | chr16 | 24820735 | 24820735 | A | - | Frame_Shift_Del | p.L1535fs | 1 |
| TNRC6A | UCEC | chr16 | 24802656 | 24802656 | G | - | Frame_Shift_Del | p.W898fs | 1 |
| TNRC6A | UCS | chr16 | 24834300 | 24834300 | G | T | Nonsense_Mutation | p.E1827X | 1 |
| TNRC6A | LUAD | chr16 | 24808843 | 24808843 | A | C | Missense_Mutation | p.E1198D | 1 |
| TNRC6A | DLBC | chr16 | 24802204 | 24802204 | C | T | Silent | p.D747D | 1 |
| TNRC6A | READ | chr16 | 24801755 | 24801755 | C | T | Nonsense_Mutation | p.R598X | 1 |
| TNRC6A | SKCM | chr16 | 24815594 | 24815594 | A | T | Missense_Mutation | p.Q1264L | 1 |
| TNRC6A | STAD | chr16 | 24762058 | 24762060 | AAG | - | In_Frame_Del | p.22_22del | 1 |
| TNRC6A | BLCA | chr16 | 24808846 | 24808846 | G | A | Silent | 1 | |
| TNRC6A | LGG | chr16 | 24801111 | 24801111 | C | A | Missense_Mutation | 1 | |
| TNRC6A | STAD | chr16 | 24806025 | 24806025 | G | T | Missense_Mutation | p.K1171N | 1 |
| TNRC6A | BLCA | chr16 | 24826603 | 24826603 | C | G | Missense_Mutation | 1 | |
| TNRC6A | THYM | chr16 | 24834851 | 24834851 | G | A | Missense_Mutation | p.G1871D | 1 |
| TNRC6A | LIHC | chr16 | 24835047 | 24835047 | C | - | Frame_Shift_Del | p.D1936fs | 1 |
| TNRC6A | MESO | chr16 | 24788482 | 24788482 | G | A | Missense_Mutation | p.R131Q | 1 |
| TNRC6A | COAD | chr16 | 24802967 | 24802967 | G | A | Missense_Mutation | p.D1002N | 1 |
| TNRC6A | PAAD | chr16 | 24831545 | 24831545 | G | A | Silent | p.L1722L | 1 |
| TNRC6A | HNSC | chr16 | 24831573 | 24831573 | C | A | Missense_Mutation | 1 | |
| TNRC6A | HNSC | chr16 | 24834300 | 24834300 | G | C | Missense_Mutation | 1 | |
| TNRC6A | SKCM | chr16 | 24834197 | 24834197 | C | T | Silent | p.I1792I | 1 |
| TNRC6A | KIRP | chr16 | 24826534 | 24826534 | G | A | Missense_Mutation | 1 | |
| TNRC6A | CESC | chr16 | 24828149 | 24828149 | G | T | Missense_Mutation | 1 | |
| TNRC6A | UCEC | chr16 | 24801151 | 24801151 | T | C | Silent | p.S396S | 1 |
| TNRC6A | LUAD | chr16 | 24801439 | 24801439 | G | T | Missense_Mutation | p.Q492H | 1 |
| TNRC6A | LUAD | chr16 | 24804889 | 24804890 | - | G | Frame_Shift_Ins | p.G1091fs | 1 |
| TNRC6A | DLBC | chr16 | 24804956 | 24804956 | C | A | Missense_Mutation | p.A1113E | 1 |
| TNRC6A | READ | chr16 | 24816116 | 24816116 | C | G | Missense_Mutation | p.Q1310E | 1 |
| TNRC6A | HNSC | chr16 | 24834300 | 24834300 | G | C | Missense_Mutation | p.E1827Q | 1 |
| TNRC6A | SKCM | chr16 | 24831632 | 24831632 | C | T | Silent | p.P1751P | 1 |
| TNRC6A | LGG | chr16 | 24828136 | 24828136 | G | A | Splice_Site | 1 | |
| TNRC6A | BLCA | chr16 | 24802526 | 24802526 | G | A | Missense_Mutation | 1 | |
| TNRC6A | KIRP | chr16 | 24801625 | 24801625 | C | G | Silent | p.G554G | 1 |
| TNRC6A | LGG | chr16 | 24815620 | 24815620 | C | T | Missense_Mutation | 1 | |
| TNRC6A | STAD | chr16 | 24762058 | 24762060 | AAG | - | In_Frame_Del | p.Q22del | 1 |
| TNRC6A | BLCA | chr16 | 24808841 | 24808841 | G | C | Missense_Mutation | 1 | |
| TNRC6A | LIHC | chr16 | 24801289 | 24801289 | A | G | Silent | 1 | |
| TNRC6A | LIHC | chr16 | 24762082 | 24762082 | A | - | Frame_Shift_Del | p.E30fs | 1 |
| TNRC6A | OV | chr16 | 24709047 | 24709047 | A | G | Missense_Mutation | 1 | |
| TNRC6A | ESCA | chr16 | 24815566 | 24815566 | G | T | Missense_Mutation | p.V1255F | 1 |
| TNRC6A | SARC | chr16 | 24815560 | 24815560 | C | T | Nonsense_Mutation | 1 | |
| TNRC6A | COAD | chr16 | 24804909 | 24804909 | G | T | Silent | p.A1097A | 1 |
| TNRC6A | HNSC | chr16 | 24816384 | 24816384 | G | A | Missense_Mutation | 1 | |
| TNRC6A | SKCM | chr16 | 24802246 | 24802246 | T | G | Missense_Mutation | p.F761L | 1 |
| TNRC6A | KIRP | chr16 | 24788620 | 24788620 | C | G | Missense_Mutation | 1 | |
| TNRC6A | CESC | chr16 | 24802308 | 24802308 | G | T | Missense_Mutation | 1 | |
| TNRC6A | LIHC | chr16 | 24801289 | 24801289 | A | G | Silent | p.Q442Q | 1 |
| TNRC6A | CESC | chr16 | 24809285 | 24809285 | G | T | Missense_Mutation | 1 | |
| TNRC6A | LUAD | chr16 | 24800787 | 24800787 | A | C | Missense_Mutation | p.N275T | 1 |
| TNRC6A | ESCA | chr16 | 24788375 | 24788375 | A | G | Silent | p.P95 | 1 |
| TNRC6A | READ | chr16 | 24826591 | 24826591 | C | T | Missense_Mutation | p.S1599L | 1 |
| TNRC6A | SKCM | chr16 | 24803029 | 24803029 | G | A | Nonsense_Mutation | p.W1022* | 1 |
| TNRC6A | LGG | chr16 | 24788379 | 24788379 | C | T | Nonsense_Mutation | p.Q97* | 1 |
| TNRC6A | LIHC | chr16 | 24801737 | 24801737 | G | A | Missense_Mutation | 1 | |
| TNRC6A | STAD | chr16 | 24788376 | 24788381 | CAGCAG | - | In_Frame_Del | p.QQ102del | 1 |
| TNRC6A | LIHC | chr16 | 24802369 | 24802369 | G | A | Silent | 1 | |
| TNRC6A | THYM | chr16 | 24818020 | 24818021 | TT | - | Frame_Shift_Del | p.1485_1485del | 1 |
| TNRC6A | LIHC | chr16 | 24818039 | 24818039 | C | - | Frame_Shift_Del | p.P1492fs | 1 |
| TNRC6A | OV | chr16 | 24725471 | 24725471 | C | T | Nonsense_Mutation | p.Q1469* | 1 |
| TNRC6A | ESCA | chr16 | 24802699 | 24802699 | A | T | Missense_Mutation | 1 | |
| TNRC6A | SARC | chr16 | 24788423 | 24788434 | GCAGCCACAGCC | - | In_Frame_Del | p.PQPQ115in_frame_del | 1 |
| TNRC6A | COAD | chr16 | 24828176 | 24828176 | T | C | Missense_Mutation | p.I1624T | 1 |
| TNRC6A | HNSC | chr16 | 24802886 | 24802887 | - | - | Frame_Shift_Ins | 1 | |
| TNRC6A | SKCM | chr16 | 24834902 | 24834902 | C | T | Missense_Mutation | p.S1888F | 1 |
| TNRC6A | LGG | chr16 | 24800571 | 24800571 | C | G | Nonsense_Mutation | p.S203X | 1 |
| TNRC6A | STAD | chr16 | 24815592 | 24815592 | T | C | Silent | p.P1263P | 1 |
| TNRC6A | BLCA | chr16 | 24808846 | 24808846 | G | A | Silent | p.A1199A | 1 |
| TNRC6A | LIHC | chr16 | 24801444 | 24801444 | A | G | Missense_Mutation | p.Q494R | 1 |
| TNRC6A | LUAD | chr16 | 24835027 | 24835027 | G | T | Missense_Mutation | p.G1930C | 1 |
| TNRC6A | LUAD | chr16 | 24804889 | 24804890 | - | G | Frame_Shift_Ins | p.W1091fs | 1 |
| TNRC6A | READ | chr16 | 24820776 | 24820776 | C | A | Missense_Mutation | p.S1549Y | 1 |
| TNRC6A | STAD | chr16 | 24807240 | 24807240 | A | - | Frame_Shift_Del | 1 | |
| TNRC6A | LGG | chr16 | 24801111 | 24801111 | C | A | Missense_Mutation | p.P383H | 1 |
| TNRC6A | BLCA | chr16 | 24820766 | 24820766 | G | T | Missense_Mutation | 1 | |
| TNRC6A | KIRP | chr16 | 24801182 | 24801182 | A | G | Missense_Mutation | p.I407V | 1 |
| TNRC6A | LIHC | chr16 | 24801508 | 24801508 | A | G | Silent | 1 | |
| TNRC6A | BLCA | chr16 | 24805971 | 24805971 | T | C | Silent | 1 | |
| TNRC6A | LIHC | chr16 | 24826592 | 24826592 | G | A | Silent | p.S1599S | 1 |
| TNRC6A | THYM | chr16 | 24834791 | 24834791 | C | T | Missense_Mutation | p.A1851V | 1 |
| TNRC6A | LIHC | chr16 | 24834206 | 24834206 | A | - | Frame_Shift_Del | p.S1795fs | 1 |
| TNRC6A | COAD | chr16 | 24788590 | 24788590 | A | C | Missense_Mutation | p.K167T | 1 |
| TNRC6A | OV | chr16 | 24725061 | 24725061 | C | T | Nonsense_Mutation | p.Q1429* | 1 |
| TNRC6A | GBM | chr16 | 24802455 | 24802455 | A | C | Missense_Mutation | 1 | |
| TNRC6A | SKCM | chr16 | 24803029 | 24803029 | G | A | Nonsense_Mutation | p.W1022X | 1 |
| TNRC6A | COAD | chr16 | 24829970 | 24829970 | C | T | Missense_Mutation | p.R1677C | 1 |
| TNRC6A | PRAD | chr16 | 24808870 | 24808870 | G | A | Silent | p.Q1207Q | 1 |
| TNRC6A | HNSC | chr16 | 24800997 | 24800997 | G | C | Missense_Mutation | 1 | |
| TNRC6A | SKCM | chr16 | 24834284 | 24834284 | C | T | Silent | p.V1821V | 1 |
| TNRC6A | BLCA | chr16 | 24802526 | 24802526 | G | A | Missense_Mutation | p.D855N | 1 |
| TNRC6A | UCEC | chr16 | 24800695 | 24800695 | C | T | Silent | p.G244G | 1 |
| TNRC6A | LUAD | chr16 | 24834925 | 24834925 | G | A | Missense_Mutation | p.D1896N | 1 |
| TNRC6A | LUSC | chr16 | 24831651 | 24831651 | T | A | Missense_Mutation | p.W1758R | 1 |
| TNRC6A | READ | chr16 | 24831490 | 24831490 | C | T | Missense_Mutation | p.S1704F | 1 |
| TNRC6A | HNSC | chr16 | 24802886 | 24802887 | - | T | Frame_Shift_Ins | p.T975fs | 1 |
| TNRC6A | STAD | chr16 | 24802902 | 24802904 | AAG | - | In_Frame_Del | 1 | |
| TNRC6A | LGG | chr16 | 24815620 | 24815620 | C | T | Missense_Mutation | p.R1273C | 1 |
| TNRC6A | BLCA | chr16 | 24800967 | 24800967 | C | T | Missense_Mutation | 1 | |
| TNRC6A | KIRP | chr16 | 24788401 | 24788401 | C | A | Missense_Mutation | p.P104Q | 1 |
| TNRC6A | STAD | chr16 | 24831653 | 24831653 | G | A | Nonsense_Mutation | p.W1758X | 1 |
| TNRC6A | TGCT | chr16 | 24801979 | 24801979 | C | T | Silent | 1 | |
| TNRC6A | LIHC | chr16 | 24834981 | 24834981 | C | T | Silent | p.H1914H | 1 |
| TNRC6A | THYM | chr16 | 24788471 | 24788471 | G | T | Missense_Mutation | p.Q127H | 1 |
| TNRC6A | LIHC | chr16 | 24835104 | 24835104 | G | - | Frame_Shift_Del | p.L1955fs | 1 |
| TNRC6A | COAD | chr16 | 24788645 | 24788645 | T | A | Missense_Mutation | p.N185K | 1 |
| TNRC6A | OV | chr16 | 24710436 | 24710436 | C | A | Missense_Mutation | p.S991Y | 1 |
| TNRC6A | HNSC | chr16 | 24800634 | 24800634 | A | T | Missense_Mutation | 1 | |
| TNRC6A | COAD | chr16 | 24831665 | 24831665 | C | T | Silent | p.D1762D | 1 |
| TNRC6A | PRAD | chr16 | 24801364 | 24801364 | C | T | Silent | p.S467S | 1 |
| TNRC6A | HNSC | chr16 | 24802394 | 24802394 | A | G | Missense_Mutation | 1 | |
| TNRC6A | HNSC | chr16 | 24833454 | 24833454 | A | G | Missense_Mutation | p.N1787D | 1 |
| TNRC6A | SKCM | chr16 | 24802845 | 24802845 | C | T | Missense_Mutation | p.A961V | 1 |
| TNRC6A | LGG | chr16 | 24788379 | 24788379 | C | T | Nonsense_Mutation | p.Q97X | 1 |
| TNRC6A | BLCA | chr16 | 24788347 | 24788347 | A | G | Missense_Mutation | 1 | |
| TNRC6A | LIHC | chr16 | 24834848 | 24834848 | C | A | Missense_Mutation | p.P1870H | 1 |
| TNRC6A | UCEC | chr16 | 24802006 | 24802006 | C | T | Silent | p.R681R | 1 |
| TNRC6A | LUAD | chr16 | 24802810 | 24802810 | A | T | Missense_Mutation | p.Q949H | 1 |
| TNRC6A | LUAD | chr16 | 24800859 | 24800859 | G | T | Missense_Mutation | p.S299I | 1 |
| TNRC6A | CESC | chr16 | 24805874 | 24805874 | C | G | Missense_Mutation | p.S1121C | 1 |
| TNRC6A | LUSC | chr16 | 24788444 | 24788444 | G | T | Missense_Mutation | p.Q118H | 1 |
| TNRC6A | ESCA | chr16 | 24834982 | 24834982 | G | T | Missense_Mutation | p.G1915C | 1 |
| TNRC6A | SARC | chr16 | 24802185 | 24802185 | G | T | Missense_Mutation | 1 | |
| TNRC6A | HNSC | chr16 | 24801106 | 24801106 | A | G | Silent | p.K381K | 1 |
| TNRC6A | STAD | chr16 | 24816407 | 24816407 | C | G | Missense_Mutation | 1 | |
| TNRC6A | BLCA | chr16 | 24802802 | 24802802 | A | G | Missense_Mutation | 1 | |
| TNRC6A | LIHC | chr16 | 24805973 | 24805973 | G | A | Missense_Mutation | 1 | |
| TNRC6A | BLCA | chr16 | 24816096 | 24816096 | C | T | Missense_Mutation | 1 | |
| TNRC6A | LIHC | chr16 | 24826473 | 24826473 | A | C | Missense_Mutation | p.I1560L | 1 |
| TNRC6A | THYM | chr16 | 24806011 | 24806011 | C | T | Missense_Mutation | p.P1167S | 1 |
| TNRC6A | LUAD | chr16 | 24788604 | 24788604 | C | G | Missense_Mutation | p.Q172E | 1 |
| TNRC6A | COAD | chr16 | 24800755 | 24800756 | AG | - | Frame_Shift_Del | p.264_264del | 1 |
| TNRC6A | OV | chr16 | 24801413 | 24801413 | C | T | Missense_Mutation | p.R484C | 1 |
| TNRC6A | HNSC | chr16 | 24804954 | 24804954 | A | T | Missense_Mutation | 1 | |
| TNRC6A | COAD | chr16 | 24834233 | 24834233 | C | T | Silent | p.H1804H | 1 |
| TNRC6A | PRAD | chr16 | 24801953 | 24801953 | G | A | Missense_Mutation | p.A664T | 1 |
| TNRC6A | SKCM | chr16 | 24808856 | 24808856 | C | T | Missense_Mutation | p.P1203S | 1 |
| TNRC6A | BLCA | chr16 | 24788645 | 24788645 | T | A | Missense_Mutation | 1 | |
| TNRC6A | LIHC | chr16 | 24788418 | 24788441 | CAGCAGCAGCCACAGCCGCAGCCG | - | In_Frame_Del | p.109_117del | 1 |
| TNRC6A | LUAD | chr16 | 24815581 | 24815581 | G | T | Missense_Mutation | p.G1260C | 1 |
| TNRC6A | BLCA | chr16 | 24801809 | 24801809 | G | C | Missense_Mutation | 1 | |
| TNRC6A | BLCA | chr16 | 24805971 | 24805971 | T | C | Silent | p.N1153N | 1 |
| TNRC6A | TGCT | chr16 | 24826529 | 24826529 | C | T | Silent | p.N1578N | 1 |
| TNRC6A | LUAD | chr16 | 24788543 | 24788543 | G | C | Missense_Mutation | p.R151S | 1 |
| TNRC6A | CESC | chr16 | 24826503 | 24826503 | C | T | Missense_Mutation | p.P1570S | 1 |
| TNRC6A | LUSC | chr16 | 24801321 | 24801321 | C | A | Missense_Mutation | p.P453Q | 1 |
| TNRC6A | SARC | chr16 | 24831519 | 24831519 | G | T | Missense_Mutation | 1 | |
| TNRC6A | HNSC | chr16 | 24802886 | 24802887 | - | T | Frame_Shift_Ins | p.I975fs | 1 |
| TNRC6A | STAD | chr16 | 24802902 | 24802904 | AAG | - | In_Frame_Del | p.KE980del | 1 |
| TNRC6A | KIRP | chr16 | 24801496 | 24801496 | G | A | Silent | p.E511E | 1 |
| TNRC6A | BLCA | chr16 | 24818054 | 24818054 | G | A | Missense_Mutation | 1 | |
| TNRC6A | LIHC | chr16 | 24818081 | 24818081 | A | G | Missense_Mutation | p.N1506D | 1 |
| TNRC6A | THYM | chr16 | 24818020 | 24818021 | TT | - | Frame_Shift_Del | p.SF1485fs | 1 |
| TNRC6A | COAD | chr16 | 24801465 | 24801465 | G | T | Missense_Mutation | p.G501V | 1 |
| TNRC6A | OV | chr16 | 24803028 | 24803029 | GG | TT | Missense_Mutation | p.W1022F | 1 |
| TNRC6A | HNSC | chr16 | 24834933 | 24834933 | T | A | Missense_Mutation | 1 | |
| TNRC6A | COAD | chr16 | 24835018 | 24835018 | A | G | Missense_Mutation | p.S1927G | 1 |
| TNRC6A | READ | chr16 | 24788505 | 24788505 | C | T | Nonsense_Mutation | p.R139X | 1 |
| TNRC6A | HNSC | chr16 | 24801106 | 24801106 | A | G | Silent | 1 | |
| TNRC6A | SKCM | chr16 | 24802181 | 24802181 | C | T | Missense_Mutation | p.P740S | 1 |
| TNRC6A | BLCA | chr16 | 24815594 | 24815594 | A | G | Missense_Mutation | 1 | |
| TNRC6A | BLCA | chr16 | 24820728 | 24820728 | C | G | Nonsense_Mutation | p.S1533* | 1 |
| TNRC6A | UCEC | chr16 | 24800814 | 24800814 | G | A | Missense_Mutation | p.R284Q | 1 |
| TNRC6A | LUAD | chr16 | 24831611 | 24831611 | G | T | Missense_Mutation | p.L1744F | 1 |
| TNRC6A | BLCA | chr16 | 24803020 | 24803020 | G | A | Silent | 1 | |
| TNRC6A | THCA | chr16 | 24834233 | 24834233 | C | T | Silent | 1 | |
| TNRC6A | LIHC | chr16 | 24769670 | 24769670 | A | - | Frame_Shift_Del | p.Q51fs | 1 |
| TNRC6A | LUAD | chr16 | 24805965 | 24805965 | G | A | Missense_Mutation | p.M1151I | 1 |
| TNRC6A | CESC | chr16 | 24800641 | 24800641 | G | A | Silent | p.K226 | 1 |
| TNRC6A | LUSC | chr16 | 24802560 | 24802560 | G | T | Missense_Mutation | p.W866L | 1 |
| TNRC6A | SARC | chr16 | 24831527 | 24831527 | G | T | Missense_Mutation | 1 | |
| TNRC6A | KICH | chr16 | 24817998 | 24817998 | T | C | Missense_Mutation | p.L1478P | 1 |
| TNRC6A | STAD | chr16 | 24801392 | 24801392 | A | G | Missense_Mutation | p.S477G | 1 |
| TNRC6A | STAD | chr16 | 24818069 | 24818069 | C | T | Missense_Mutation | p.P1502S | 1 |
| TNRC6A | LIHC | chr16 | 24834278 | 24834278 | T | C | Silent | p.A1819A | 1 |
| TNRC6A | THYM | chr16 | 24788444 | 24788444 | G | A | Silent | p.Q118 | 1 |
| TNRC6A | LUAD | chr16 | 24802765 | 24802765 | G | T | Missense_Mutation | p.W934C | 1 |
| TNRC6A | COAD | chr16 | 24801737 | 24801737 | G | A | Missense_Mutation | p.A592T | 1 |
| TNRC6A | OV | chr16 | 24804800 | 24804800 | G | T | Missense_Mutation | p.G1061V | 1 |
| TNRC6A | HNSC | chr16 | 24801674 | 24801674 | G | A | Missense_Mutation | 1 | |
| TNRC6A | READ | chr16 | 24788511 | 24788511 | C | T | Missense_Mutation | p.R141C | 1 |
| TNRC6A | HNSC | chr16 | 24833454 | 24833454 | A | G | Missense_Mutation | 1 | |
| TNRC6A | SKCM | chr16 | 24802182 | 24802182 | C | T | Missense_Mutation | p.P740L | 1 |
| TNRC6A | BLCA | chr16 | 24802802 | 24802802 | A | G | Missense_Mutation | p.N947D | 1 |
| TNRC6A | LIHC | chr16 | 24815534 | 24815534 | A | G | Missense_Mutation | p.H1244R | 1 |
| TNRC6A | UCEC | chr16 | 24800899 | 24800899 | A | G | Silent | p.P312P | 1 |
| TNRC6A | BLCA | chr16 | 24816096 | 24816096 | C | T | Missense_Mutation | p.S1303L | 1 |
| TNRC6A | THYM | chr16 | 24801109 | 24801109 | G | T | Silent | 1 | |
| TNRC6A | LIHC | chr16 | 24800905 | 24800905 | T | - | Frame_Shift_Del | p.G314fs | 1 |
| TNRC6A | CESC | chr16 | 24835021 | 24835021 | C | A | Missense_Mutation | p.L1928M | 1 |
| TNRC6A | LUSC | chr16 | 24801959 | 24801959 | A | G | Missense_Mutation | p.T666A | 1 |
| TNRC6A | ESCA | chr16 | 24834982 | 24834982 | G | T | Missense_Mutation | 1 | |
| TNRC6A | SARC | chr16 | 24801720 | 24801720 | G | T | Missense_Mutation | 1 | |
| TNRC6A | STAD | chr16 | 24831653 | 24831653 | G | A | Nonsense_Mutation | p.W1758* | 1 |
| TNRC6A | SKCM | chr16 | 24831512 | 24831512 | C | T | Silent | p.T1711T | 1 |
| TNRC6A | BLCA | chr16 | 24788436 | 24788436 | C | T | Nonsense_Mutation | p.Q116* | 1 |
| TNRC6A | UCEC | chr16 | 24801085 | 24801085 | G | A | Silent | p.E374E | 1 |
| TNRC6A | COAD | chr16 | 24802132 | 24802132 | G | T | Missense_Mutation | p.W723C | 1 |
| TNRC6A | PAAD | chr16 | 24788408 | 24788408 | G | T | Missense_Mutation | 1 | |
| TNRC6A | HNSC | chr16 | 24802355 | 24802355 | T | C | Missense_Mutation | 1 | |
| TNRC6A | LUAD | chr16 | 24802337 | 24802337 | T | C | Missense_Mutation | p.W792R | 1 |
| TNRC6A | DLBC | chr16 | 24788401 | 24788402 | - | GCA | In_Frame_Ins | p.P104delinsPQ | 1 |
| TNRC6A | READ | chr16 | 24788528 | 24788528 | A | C | Missense_Mutation | p.K146N | 1 |
| TNRC6A | HNSC | chr16 | 24835045 | 24835045 | G | A | Missense_Mutation | 1 | |
| TNRC6A | SKCM | chr16 | 24829927 | 24829927 | C | T | Silent | p.S1662S | 1 |
| TNRC6A | STAD | chr16 | 24788376 | 24788381 | CAGCAG | - | In_Frame_Del | p.95_97del | 1 |
| TNRC6A | BLCA | chr16 | 24802299 | 24802299 | C | T | Missense_Mutation | 1 | |
| TNRC6A | BLCA | chr16 | 24801809 | 24801809 | G | C | Missense_Mutation | p.E616Q | 1 |
| TNRC6A | LIHC | chr16 | 24802005 | 24802005 | G | T | Missense_Mutation | p.R681L | 1 |
| TNRC6A | STAD | chr16 | 24815588 | 24815588 | G | A | Missense_Mutation | p.R1262Q | 1 |
| TNRC6A | BLCA | chr16 | 24802435 | 24802435 | G | C | Missense_Mutation | 1 | |
| TNRC6A | BLCA | chr16 | 24818054 | 24818054 | G | A | Missense_Mutation | p.E1497K | 1 |
| TNRC6A | LIHC | chr16 | 24762093 | 24762093 | A | - | Frame_Shift_Del | p.K35fs | 1 |
| TNRC6A | LUSC | chr16 | 24788259 | 24788259 | G | C | Missense_Mutation | p.E57Q | 1 |
| TNRC6A | SARC | chr16 | 24833408 | 24833408 | G | T | Missense_Mutation | 1 | |
| TNRC6A | STAD | chr16 | 24802902 | 24802904 | AAG | - | In_Frame_Del | p.980_980del | 1 |
 Copy number variation (CNV) of TNRC6A Copy number variation (CNV) of TNRC6A * Click on the image to open the original image in a new window. |
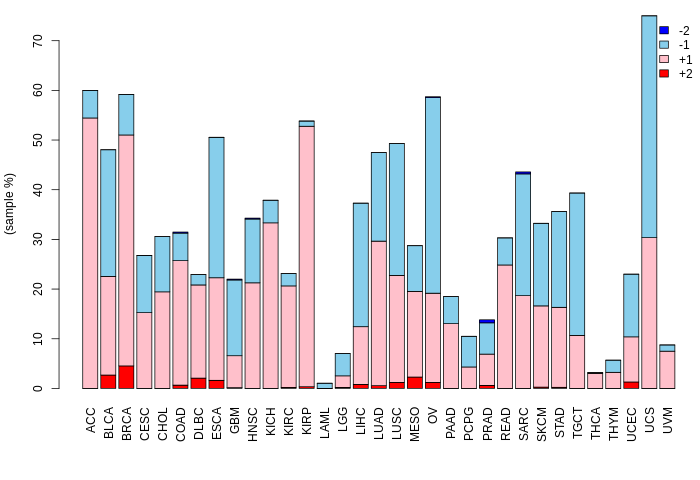 |
 Fusion gene breakpoints (product of the structural variants (SVs)) across TNRC6A Fusion gene breakpoints (product of the structural variants (SVs)) across TNRC6A * Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
 Fusion genes with this translation factor from FusionGDB2.0. Fusion genes with this translation factor from FusionGDB2.0. |
| FusionGDB2 ID | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 98852 | BRCA | TCGA-S3-AA0Z-01A | ARHGAP17 | chr16 | 24953308 | - | TNRC6A | chr16 | 24762047 | + |
| 98852 | N/A | AA507348 | ATF2 | chr2 | 175936987 | - | TNRC6A | chr16 | 24730973 | + |
| 98852 | N/A | N22604 | ATXN2 | chr12 | 112010838 | - | TNRC6A | chr16 | 24837543 | - |
| 98852 | N/A | EC527969 | CACNG3 | chr16 | 24346112 | + | TNRC6A | chr16 | 24644105 | + |
| 98852 | N/A | BF437094 | CLN8 | chr8 | 1711096 | + | TNRC6A | chr16 | 24802241 | - |
| 98852 | GBM | TCGA-06-0211-01B | CSNK2A1 | chr20 | 485762 | - | TNRC6A | chr16 | 24788254 | + |
| 98852 | N/A | AA531082 | GALNTL6 | chr4 | 172884246 | + | TNRC6A | chr16 | 24741835 | - |
| 98852 | HNSC | TCGA-BA-6869-01A | PDLIM1 | chr10 | 97023621 | - | TNRC6A | chr16 | 24800553 | + |
| 98852 | N/A | BF920959 | RNF112 | chr17 | 19320049 | - | TNRC6A | chr16 | 24790369 | + |
| 98852 | N/A | CV353496 | SLC9A9 | chr3 | 143030963 | + | TNRC6A | chr16 | 24769851 | - |
| 98852 | N/A | BF733944 | SMG6 | chr17 | 2203379 | + | TNRC6A | chr16 | 24802489 | - |
| 98852 | N/A | AK025277 | SORBS2 | chr4 | 186610266 | - | TNRC6A | chr16 | 24828954 | + |
| 98852 | N/A | AA962358 | STRN4 | chr19 | 47222828 | + | TNRC6A | chr16 | 24756977 | - |
| 98852 | N/A | BI491476 | TCF7L1 | chr2 | 85446844 | + | TNRC6A | chr16 | 24734039 | - |
| 98852 | N/A | BF839733 | TG | chr8 | 134025682 | - | TNRC6A | chr16 | 24831390 | - |
| 92857 | N/A | BI046776 | TNRC6A | chr16 | 24770977 | + | ACTG2 | chr2 | 74129762 | - |
| 95742 | N/A | AA653866 | TNRC6A | chr16 | 24746555 | - | ADAMTS9-AS2 | chr3 | 64827552 | - |
| 92857 | LUAD | TCGA-69-A59K-01A | TNRC6A | chr16 | 24802355 | + | ANKS4B | chr16 | 21262929 | + |
| 92857 | N/A | N64162 | TNRC6A | chr16 | 24643092 | - | FTH1P10 | chr5 | 17353804 | + |
| 92857 | OV | TCGA-13-0765 | TNRC6A | chr16 | 24741621 | + | LCMT1 | chr16 | 25139795 | + |
| 92857 | OV | TCGA-13-0765 | TNRC6A | chr16 | 24762134 | + | LCMT1 | chr16 | 25139795 | + |
| 92857 | OV | TCGA-13-0765-01A | TNRC6A | chr16 | 24741621 | + | LCMT1 | chr16 | 25139796 | + |
| 92857 | OV | TCGA-13-0765-01A | TNRC6A | chr16 | 24762134 | + | LCMT1 | chr16 | 25066140 | + |
| 92857 | OV | TCGA-13-0765-01A | TNRC6A | chr16 | 24762134 | + | LCMT1 | chr16 | 25137285 | + |
| 92857 | OV | TCGA-13-0765-01A | TNRC6A | chr16 | 24762134 | + | LCMT1 | chr16 | 25139796 | + |
| 95786 | PCPG | TCGA-TT-A6YN | TNRC6A | chr16 | 24809287 | + | LIX1L | chr1 | 145492234 | + |
| 95786 | PCPG | TCGA-TT-A6YN-01A | TNRC6A | chr16 | 24809287 | + | LIX1L | chr1 | 145492235 | - |
| 97011 | LUSC | TCGA-O2-A52S-01A | TNRC6A | chr16 | 24741167 | + | METTL9 | chr16 | 21666548 | + |
| 97631 | GBM | TCGA-06-0882-01A | TNRC6A | chr16 | 24741621 | + | PRKCB | chr16 | 24192111 | + |
| 92857 | N/A | BE175576 | TNRC6A | chr16 | 24813992 | - | RPA1 | chr17 | 1782975 | - |
| 98852 | N/A | AA501799 | TNRC6A | chr16 | 24780527 | + | TNRC6A | chr16 | 24780356 | - |
| 98852 | N/A | AV730574 | WDFY4 | chr10 | 50000063 | - | TNRC6A | chr16 | 24816958 | - |
Top |
|
 Kaplan-Meier plots with logrank tests of overall survival (OS) Kaplan-Meier plots with logrank tests of overall survival (OS) |
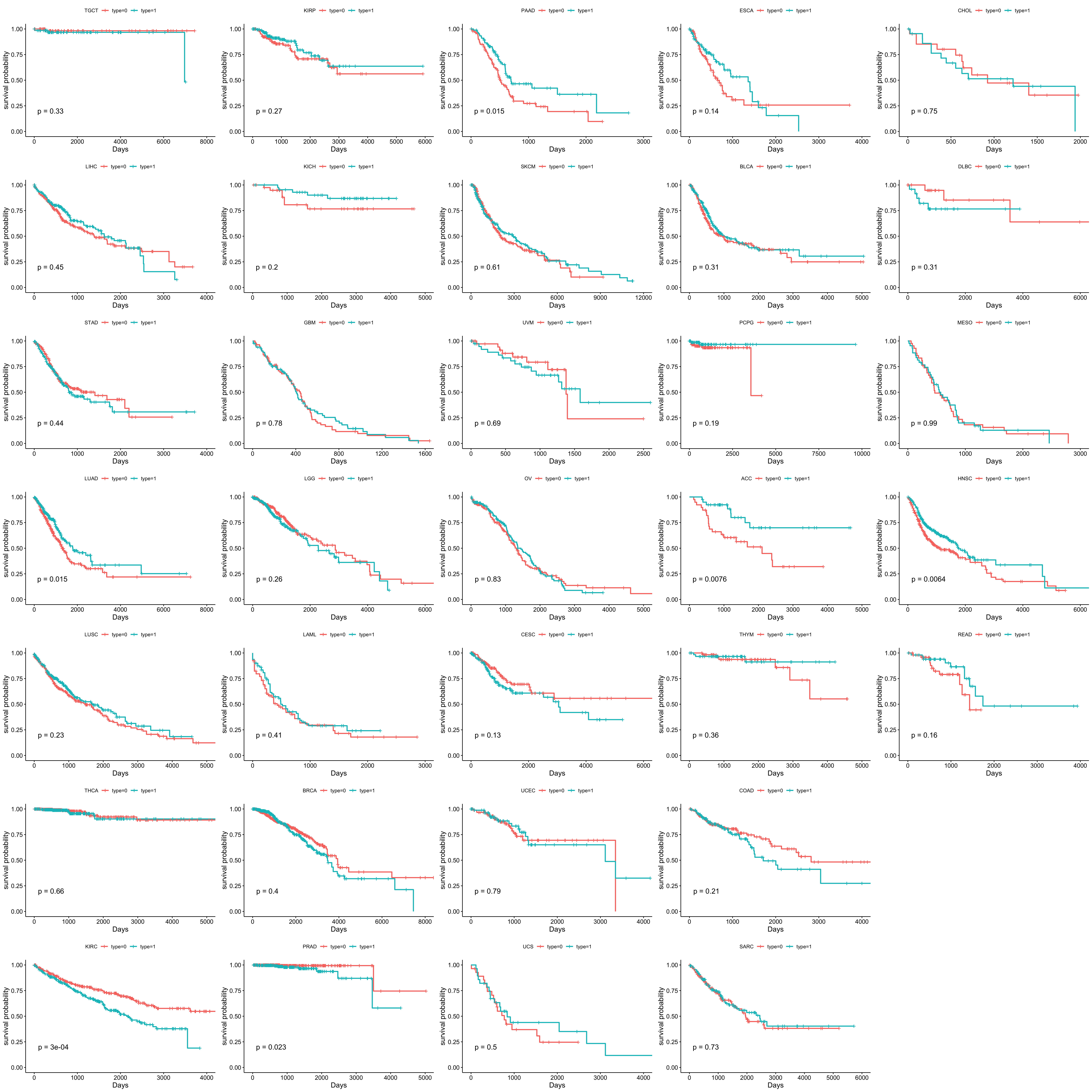 |
| Cancer type | Translation factor | Coefficent | Hazard ratio | Wald test pval | Likelihool ratio pval | Logrank test pval | # samples |
Top |
|
 Differential gene expression between female and male. (Wilcoxon test, pval<0.05) Differential gene expression between female and male. (Wilcoxon test, pval<0.05) |
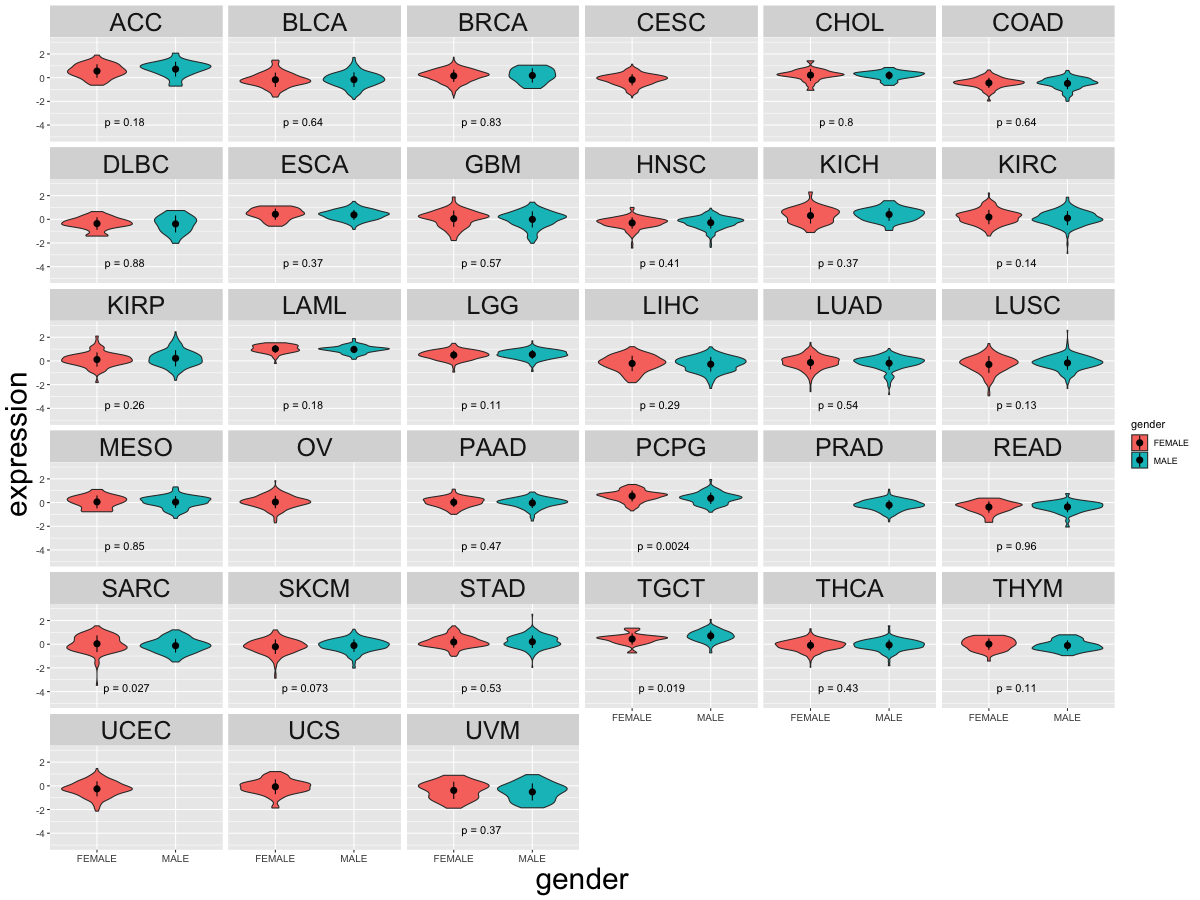 |
| Cancer type | Translation factor | pval | adj.p |
| PCPG | TNRC6A | 0.00238143665653013 | 0.067 |
| TGCT | TNRC6A | 0.0185447807146848 | 0.5 |
| SARC | TNRC6A | 0.0273874462783429 | 0.71 |
Top |
|
 Differential gene expression between young and old age groups (Wilcoxon test, pval<0.05) Differential gene expression between young and old age groups (Wilcoxon test, pval<0.05) |
 |
| Cancer type | Translation factor | pval | adj.p |
| KIRC | TNRC6A | 0.0323646821186705 | 1 |
| BLCA | TNRC6A | 0.0226469949338202 | 0.72 |
| SARC | TNRC6A | 0.0059899877017452 | 0.2 |
Top |
|
 Drugs targeting genes involved in this translation factor. Drugs targeting genes involved in this translation factor. (DrugBank Version 5.1.8 2021-05-08) |
| UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
|
 Diseases associated with this translation factor. Diseases associated with this translation factor. (DisGeNet 4.0) |
| Disease ID | Disease Name | # PubMeds | Disease source |
| C0014550 | Myoclonic Epilepsy | 1 | CTD_human |
| C0338478 | Idiopathic Myoclonic Epilepsy | 1 | CTD_human |
| C0338479 | Symptomatic Myoclonic Epilepsy | 1 | CTD_human |
| C0393695 | Early Childhood Epilepsy, Myoclonic | 1 | CTD_human |
| C0393702 | Myoclonic Astatic Epilepsy | 1 | CTD_human |
| C0393703 | Myoclonic Absence Epilepsy | 1 | CTD_human |
| C0438414 | Myoclonic Encephalopathy | 1 | CTD_human |
| C0751120 | Benign Infantile Myoclonic Epilepsy | 1 | CTD_human |
| C0751122 | Infantile Severe Myoclonic Epilepsy | 1 | CTD_human |
| C0917800 | Epilepsy, Myoclonic, Infantile | 1 | CTD_human |

