
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:SLX4IP-CDS2 (FusionGDB2 ID:HG128710TG8760) |
Fusion Gene Summary for SLX4IP-CDS2 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: SLX4IP-CDS2 | Fusion gene ID: hg128710tg8760 | Hgene | Tgene | Gene symbol | SLX4IP | CDS2 | Gene ID | 128710 | 8760 |
| Gene name | SLX4 interacting protein | CDP-diacylglycerol synthase 2 | |
| Synonyms | C20orf94|bA204H22.1|bA254M13.1|dJ1099D15.3 | - | |
| Cytomap | ('C20orf94','SLX4IP')('CDS2','CDS2') 20p12.2 | 20p12.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | protein SLX4IP | phosphatidate cytidylyltransferase 2CDP-DAG synthase 2CDP-DG synthase 2CDP-DG synthetase 2CDP-diacylglycerol synthase (phosphatidate cytidylyltransferase) 2CDP-diglyceride diphosphorylase 2CDP-diglyceride pyrophosphorylase 2CDP-diglyceride synthase | |
| Modification date | 20200313 | 20200327 | |
| UniProtAcc | . | O95674 | |
| Ensembl transtripts involved in fusion gene | ENST00000334534, | ||
| Fusion gene scores | * DoF score | 13 X 9 X 5=585 | 10 X 11 X 7=770 |
| # samples | 13 | 13 | |
| ** MAII score | log2(13/585*10)=-2.16992500144231 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(13/770*10)=-2.56634682255381 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: SLX4IP [Title/Abstract] AND CDS2 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | C20orf94(10582467)-CDS2(5154169), # samples:1 SLX4IP(10582467)-CDS2(5154169), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across SLX4IP (5'-gene) Fusion gene breakpoints across SLX4IP (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
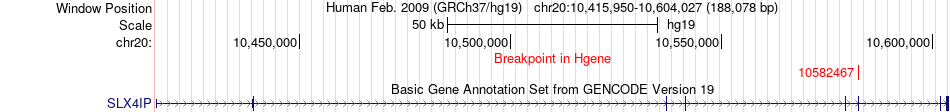 |
 Fusion gene breakpoints across CDS2 (3'-gene) Fusion gene breakpoints across CDS2 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
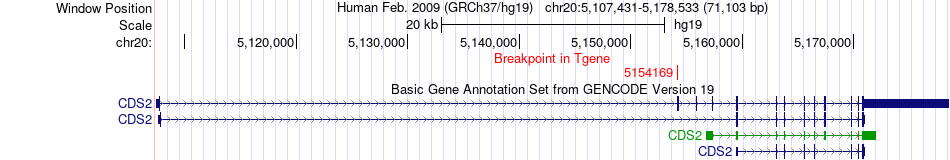 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | HNSC | TCGA-CX-7082-01A | C20orf94 | chr20 | 10582467 | - | CDS2 | chr20 | 5154169 | + |
| ChimerDB4 | HNSC | TCGA-CX-7082-01A | SLX4IP | chr20 | 10582467 | + | CDS2 | chr20 | 5154169 | + |
Top |
Fusion Gene ORF analysis for SLX4IP-CDS2 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000334534 | ENST00000379062 | SLX4IP | chr20 | 10582467 | + | CDS2 | chr20 | 5154169 | + |
| 5CDS-intron | ENST00000334534 | ENST00000379070 | SLX4IP | chr20 | 10582467 | + | CDS2 | chr20 | 5154169 | + |
| 5CDS-intron | ENST00000334534 | ENST00000535100 | SLX4IP | chr20 | 10582467 | + | CDS2 | chr20 | 5154169 | + |
| In-frame | ENST00000334534 | ENST00000460006 | SLX4IP | chr20 | 10582467 | + | CDS2 | chr20 | 5154169 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000334534 | SLX4IP | chr20 | 10582467 | + | ENST00000460006 | CDS2 | chr20 | 5154169 | + | 9519 | 585 | 180 | 1865 | 561 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000334534 | ENST00000460006 | SLX4IP | chr20 | 10582467 | + | CDS2 | chr20 | 5154169 | + | 0.000611893 | 0.9993881 |
Top |
Fusion Genomic Features for SLX4IP-CDS2 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| SLX4IP | chr20 | 10582467 | + | CDS2 | chr20 | 5154168 | + | 0.007618686 | 0.9923813 |
| SLX4IP | chr20 | 10582467 | + | CDS2 | chr20 | 5154168 | + | 0.007618686 | 0.9923813 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
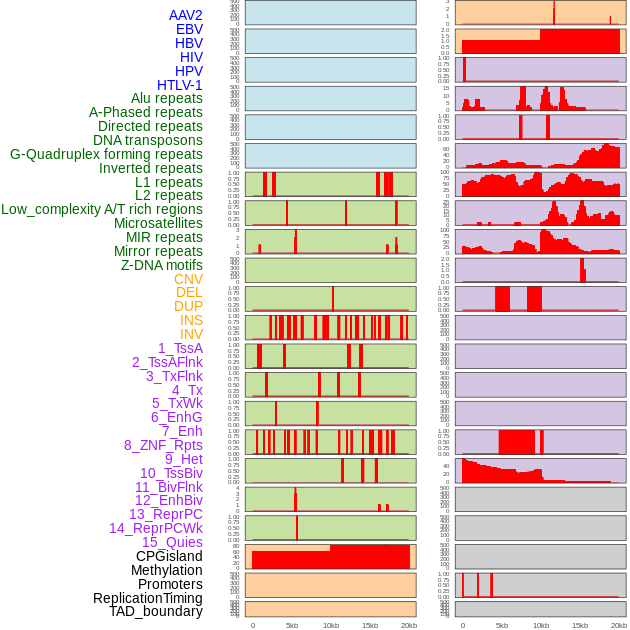 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
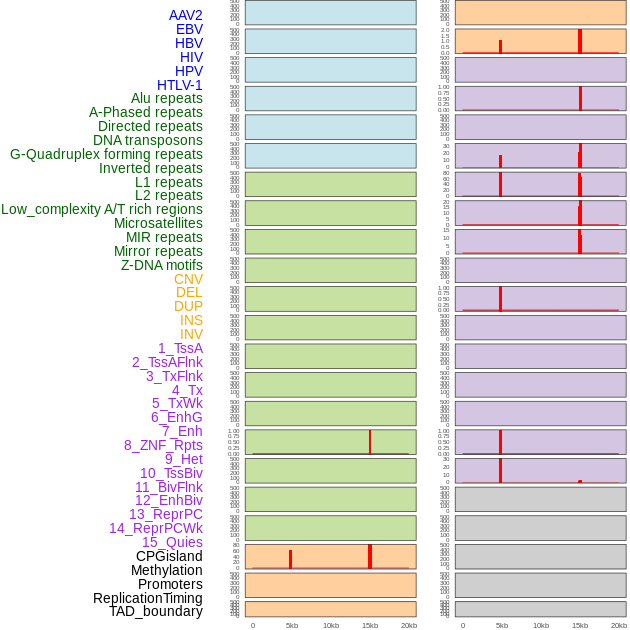 |
Top |
Fusion Protein Features for SLX4IP-CDS2 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr20:10582467/chr20:5154169) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | CDS2 |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Catalyzes the conversion of phosphatidic acid (PA) to CDP-diacylglycerol (CDP-DAG), an essential intermediate in the synthesis of phosphatidylglycerol, cardiolipin and phosphatidylinositol (PubMed:25375833). Exhibits specificity for the nature of the acyl chains at the sn-1 and sn-2 positions in the substrate, PA and the preferred acyl chain composition is 1-stearoyl-2-arachidonoyl-sn-phosphatidic acid (PubMed:25375833). Plays an important role in regulating the growth and maturation of lipid droplets which are storage organelles at the center of lipid and energy homeostasis (PubMed:26946540, PubMed:31548309). {ECO:0000269|PubMed:25375833, ECO:0000269|PubMed:26946540, ECO:0000269|PubMed:31548309}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | CDS2 | chr20:10582467 | chr20:5154169 | ENST00000460006 | 0 | 13 | 132_152 | 19 | 446.0 | Transmembrane | Helical | |
| Tgene | CDS2 | chr20:10582467 | chr20:5154169 | ENST00000460006 | 0 | 13 | 166_186 | 19 | 446.0 | Transmembrane | Helical | |
| Tgene | CDS2 | chr20:10582467 | chr20:5154169 | ENST00000460006 | 0 | 13 | 213_233 | 19 | 446.0 | Transmembrane | Helical | |
| Tgene | CDS2 | chr20:10582467 | chr20:5154169 | ENST00000460006 | 0 | 13 | 262_282 | 19 | 446.0 | Transmembrane | Helical | |
| Tgene | CDS2 | chr20:10582467 | chr20:5154169 | ENST00000460006 | 0 | 13 | 340_360 | 19 | 446.0 | Transmembrane | Helical | |
| Tgene | CDS2 | chr20:10582467 | chr20:5154169 | ENST00000460006 | 0 | 13 | 79_99 | 19 | 446.0 | Transmembrane | Helical |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
Top |
Fusion Gene Sequence for SLX4IP-CDS2 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >83746_83746_1_SLX4IP-CDS2_SLX4IP_chr20_10582467_ENST00000334534_CDS2_chr20_5154169_ENST00000460006_length(transcript)=9519nt_BP=585nt GCGAAAGTGCTTCCCTTAAGCTTCTGAAGGTTGGCTGCAGTTCCGGCTACCTGTGTAGTCCGAGTTTCCACAGCCAGGTACTACTCCGCC AGTGACCCTGGACAGTAACAAAACATATAAAGCCCGAGCCCAAACCCCGCCACCATCATAGGTCTGTAGTTACTGTGGAATCAATAAGCC ATGGCATCTAAGAAATTTGCTGTTAAATGTGGGAATTTTGCTGTCCTCGTGGATCTTCATATCTTGCCACAAGGTTCAAACAAAGATACA AGCTGGTTTTCTGAACAGAAGAAAGAGGAAGTCTGTTTACTGTTAAAAGAAACCATTGATTCAAGAGTTCAGGAGTACTTGGAAGTTCGC AAACAGCACAGGCCATCAAATGCAGAATTCACAAGATCCAATCCCTTGTCCTTAAAAGGTTATGGCTTTCAAATCACAGCCTATTTCCTC AAGAGAGGGATACGCCTTCGCTGCATCAGGAGCACACAGAATGCTGAACTCTGTGTATTCCCTGACAGATTTGTGGTTTGTGTCAGTCAG CTTGCATTCAGTCGTGATCTTTTAGCAAGTCAGAATGAAGATTTGGAGTCAGAGTCAGAAGCAAAGGTAGATGGAGAGACTGCATCGGAC AGTGAGAGCCGGGCAGAATCCGCACCCCTGCCAGTCTCTGCAGATGATACCCCGGAGGTCCTCAATAGGGCCCTTTCCAACTTGTCTTCA AGATGGAAGAACTGGTGGGTGAGAGGCATCCTGACTTTGGCCATGATTGCATTTTTCTTCATCATCATTTACCTGGGACCAATGGTTTTG ATGATAATCGTGATGTGCGTTCAGATTAAGTGTTTCCATGAGATAATCACTATTGGCTACAACGTCTACCACTCATATGATCTGCCCTGG TTCAGGACGCTCAGCTGGTACTTTCTCCTGTGTGTAAACTATTTCTTCTATGGTGAGACAGTGACGGATTACTTCTTCACCCTGGTCCAG AGAGAAGAGCCTTTGCGGATTCTCAGTAAATACCACCGGTTCATTTCCTTTACTCTCTATCTAATAGGATTCTGCATGTTTGTACTGAGT CTGGTCAAGAAGCATTATCGACTGCAGTTCTACATGTTTGGCTGGACCCATGTGACATTGCTGATTGTTGTAACACAGTCACATCTTGTT ATCCACAACCTATTTGAAGGAATGATCTGGTTCATTGTCCCCATATCTTGTGTGATCTGTAATGACATCATGGCCTATATGTTTGGCTTT TTCTTTGGTCGGACCCCACTCATCAAGCTGTCCCCGAAGAAGACCTGGGAAGGCTTCATTGGGGGCTTCTTTGCTACTGTGGTGTTTGGC CTTCTGCTGTCCTATGTGATGTCCGGGTACAGATGCTTTGTCTGCCCTGTGGAGTACAACAATGACACCAACAGCTTCACTGTGGACTGT GAGCCCTCGGACCTGTTTCGCCTGCAGGAGTACAACATTCCTGGGGTGATCCAGTCAGTCATTGGCTGGAAAACGGTCCGGATGTACCCC TTCCAGATTCACAGCATCGCTCTCTCCACCTTTGCCTCGCTCATTGGCCCCTTTGGAGGATTCTTCGCAAGTGGATTCAAACGAGCCTTT AAAATCAAAGACTTTGCCAATACCATTCCTGGCCATGGAGGCATCATGGATCGCTTTGACTGCCAGTATCTGATGGCCACCTTTGTCAAT GTATACATCGCCAGTTTTATCAGAGGCCCTAACCCAAGCAAACTGATTCAGCAGTTCCTGACTTTACGGCCAGATCAGCAGCTCCACATC TTCAACACGCTGCGGTCTCATCTGATCGACAAAGGGATGCTGACATCCACCACAGAGGACGAGTAGGGGCCACCCAGGGCCAGGAGAACA GGAACAGAACTGAGCAGGGGCAGGTCTCCAAGGCAAGCCCAGCTGGTGTGACTTAGACAATGACGAGGCTTCAACTCACTGTCTTTTTTT TTTTTTTTTGGAGGGTATTTTTTATTTGTGGGTTCAAAAAATCTGTATATACAGTCTATGTGTTTAGAATTTGTGTTGTAAGTAAACTAC AGCTTTGAGTTGGAAAGAAGTCACGGGTTGTAAAACCATTTGGATTTTTTTAAAACAAAAGTATTAATAATCTGGAAGACAGTGTTGCCC AGGTCAGGAGTGTTTTCTTGGTGGTTCCAGCCCCCATCAATTGAACTGTTTCTGGGCTCAGTCAGACACAGACATTCATCTGTGTCTGAC CAAATCAGGGGACTTCCCCACCTGTGGTGGGAGGCACAGCTTAGATGTTTTGTACACCTGGTCTTTTCTAGAAATCCCTGCTTGGAGCTG CAGAAGGGTTGCCTTCTGTAGGTCGGAGGAATGGAGGCTTACTAACCAGGTAAGCCTTCTATGCATCCACACCAAAATCCTGCAGAATGT AAGTAAGCTCTGCTTTATAAGATGGGTTCACCTTCATCGCAGACTGAAAGTTTCAGTTTTTATTTTTTTCAGAAAGCACGAAAAATTATT TATAATAGTCTGGAGAAAAAACACACTGTAATATTTCAAGTGTATGCAGTAGAATGTACTGTAACTGAGCCCTTTCCCACATGTCTAGGC TCCAATGTCTCCTGTAGGTCCACCTAACTGTGTGTTTTCAGGGACAATGCCATCCATGTTTGTGCTGTAGACTTGCTGCTGCTGAATCCT TTCTGGGGACTTTCTCATCGGGCAGGGAGCAGAGGGCTTCTCGTTCATGCACCCTTTGCCTGAACACCCATGTAGCTGCTGTGTTGTGTA TATATTACTCTTAAGAGGAGTGTGTGTGTCTGTGTTTGTTTTAAAAGTCACTTATTTCTTACAGTGATTTCAATTGCACCATGACTTCTT CACTAAAACCACAAAGTCCTGCTTAAAACTATGGAAAACCTAACCTGATTAGAGCCTTGACTATTTTGAAGATTAAATGCACACTTTTTA TATAATGTGACCAGTTTAAATGTAGTTTGTATTGTACTGGGGGACCTTTCGTTGTTGTTGTTTGCTTAAACTGTGATTTTTTTTCCCCTC CCTAATTTCAGGGGTGAGATTGACTTTGGGAAGACAGATTAGTTCTTTGTCAGGCCAACAAGTGATGGAGTGCGGGAGAGAGAACTTGGC ACCCAAATATATCAAACTATTCCGTGCCGTGGATGTTTTCATTGCCAACGAGGGTGTAATGATTTGCTTCTGCACCTTGGTCTAGTGCTG GTTTGTGGTGTTTTTGTTTGTAAATTAGCCTCACTGCCTCCCTGAAAGTGCACAGTCAGCCCAGGTCTCTAGAGTGTCCAGCAGAGGATG GAGCCCTCGGGATGTTTCAGCTCACACATCCTTGAGTGACTTCCCATATGTGTGTGGGGCTGAAGCGGCTTCTTTCCGCTGAACTTTTAT TTCCATTTCAGCCTCGTGGCTTCTCTTATTCAGGTCCAAGGCCCCTTAAAAACAAAACATATTATCGCAAGGGCTTTCATTTCAGTATTG GCTGTGTCACTGGCAGGATGCATCACTCTCCAGGGCCTCTTTTTCGGAGCTACTCCCAATTCCACTCTTCTGTGATTCTGGCTGAGCAAG TTCATGCCGGTAGAGGCTCATTAAGTGAGGTCCTGGGCACCCCCCAAACAGGAAGTGTTTTCAGGCCCTTACATGCTTATCCTTAAGGTG AGTAGGTATCTCAGCAGCACACCCGAACAGTAAAGGTGATCAATAACGAAACTGTAGTTTTCAGATTGAAGAATGAACCTGGATCACCAA ACTAGATACCGAAAGGGCTCCTCATTTGTCTCTTTTGTCCCATTGTGGGCATATGGTGGGCTCCTTGAGAGGAGGGAGTCTCAGCTGGGG CCTGGCTGCAGAGACCTGCTCTTGAATAGGGCTGCCTTTGGGATCAGGGAGCCGAGCATACTGGGCAAGGCTCATCATGTTCCTTTGTGG AAGTGGTTACTTGGTGGGGGTGGGGTGGGGAAGGGGGGTCTAGGAAGCCGAAGCTGAGTGCGCACTAGAGCTGCTAGCAGGCAGGCCAGA GGGTCAGCTTGTGTTTTATTTTCTGATTTCTCCTGCATGCTTTCTGGAGATGAGGATTTTTTGTCAGGTAGCTAGGAGAGCACTGCCACC TTGTGTCCATATAGCACAATGTTTCTCTTTCCTATTCACAGACAAGATTCAAGAAACCATTTTTACGTGTGTACATTCAAATCTTCATGA ATGGCATCACTTGCTTTAGACCCATTTTTTTTTTATTGGTCAGAAATAAAATGTGACTGCAACTGTGTGCCCTCCCTTGTCGTGAATTCC CTGTTTATCACTTCCATGCATGGTTTTATGCTTCTCATTTGTTTATGTATTCATGAATACTGCATAGTCCTGATTCAGAGACTTTTGCTT TGCTTTCCAACTTGTATCCCTACATTCTCACAACAGCCTTCTTACCTCGGGTGAGAGTCATGAGCACCTTACCAGGCAGGGAGCTTCCCC AGGATAGTGTGGTGGAGTCAGGACTCACTCCATCCTGCACTTTACCCACAGGACACCTGGAGGGTCCCCCAGCCCTGCTTGCTTTTGTGA ACGTCGTGTGAGTACACCGAGGTGTTGCAGCTTTCCTTCCCTCTAGCTCAGGAGGCCTGGCCTGAGCCAGGTTGGAGCTTAAATTGCTGT TGGCTCCTGCCTGGTGGAACAAGGGGGTCATAGAAACCAGGGCTGATGGGCACCCCCCAGCCCCTGGGAGCTCTGCTCATGACCGCTCAC AGGGTACCGAGTCTTTGCCTTTTGTGTGTGCGCATGCTGCCCTCTTAACTGCTTTTAGTATTCAGAAAGGTTCCATTTATGTTTTACTCC TGAAAGTGAAACCCTCTTAGGCAACTTGTTTCATATTCTGGGCATTTTAGTAGCTACAGTATTTTGCATTTAAAATGTAATTACTCTTAG TATTGTACAGCCCTATAATGTATGAGAACTAGTTTCACATCCTTTGAAACTAATCTCTTTCATTGGGGAGGATACAACTTTTGACTTTAC ACAGGTGGACTTTGAATCCATTTTTATAGAATTTTTTTCCACTGTAACTTGACCTATATTATGCAAGAAAATCCCAAAAATATGTTTTAA GGGAGACTACACAGACCTTGTGATTTGGTCTGAGCTAGTTGCTGAGTTGACTGGAGCCTTATTTGAATGGATCTTTGAGTTGAGATCTTC ACACTGTCAGTGAGCCCTTGGCTCAGCCCTTGTCACCTGTTTGTATGGGGACCATGTTTCCCAAAGCTCAGGTTCATCTCCACTATTGAC AGAGTCGGGTAGTGGGAGAGTATAAGTGACTTGAGACAGAGTGTCCTTGCTCAGAAAACTGTGAAGCAGGGGTCTTGTTAGTTACAAAGC CAGTCCTAGTTGCATGTCCAAGGGGGTAGATTTTCATCACCCTACTAAATGTGGTATGTCACTGCTGAAGCCTTTTCCTGTTTTAGTAAT ATCTTAGGCACTATTGTCTTTTGTCTTCCAGAGTAACATATCTGGGCCCACCTTTCTTGCACACTGTGATTATGTGCTAAGAAAAGGGTG GAGTTGGGAGGACAATTTCACTTGAACAGGTTTCTGTTCTTCTGAGGTCAGGGTGGGGAGGATGTCCAGGTGACTTTTATGGACACATTG CAAGACAGTAAGCCGCGGATGGACTCTCACTTGTGCTTTGGAACACTGGACTCCAAAGAGCAGGTGGAACAAACTCTGGCCTGAAGATGA TGGCTAATACTCAGGCAGAGCCCCGTATTCTTCCCTGACTTGGTCAAAACAAACTTCCTTTTCCCTTGATCACCCTGCACGCAGCTGCCC AGCTTCCAGTTTTTAAAGAGATGAACAGCTCTCACTTCCTGACTCAGTTTCTCTTGTTGGGTGGGGGCCTTGTGCCACAACCTGCATGGG GTGCTAGGGTTTAAACCAAGTGGTCACGAAGGATTCTTATCTCAGGGCTCTGCCTGGCCCACCTTCTCTTGGGCCTGGACTCTCCTTATC CACATCCCTGCAGAAGGTGGGCCCTATTCTCAGAAGAGTCCTGGCTGCTGTTGCCATCTGAATGCGCCCTTCCCTTTCAGGTTGACCAGC AGTGGCCTTACCTGGCCTCTCAGGTGGATGGAGGCTGTTCTCATCTGGGGCTGTCTCTTGATTAAGGCTGTCACCTGTTTGTATAGGGAC CATGTTTCCCAACGCACTTTGAGTGCAGTTTAGGAGGTATCCTTTGGATGCTGAAGTGTTGGGATTTTGAGGACATGCGGATGATTTATG AGAATAGAGGGGCAGAGTTTGTGAAGTCTAGGGTTTTTTGTCCAGTGGGTTTCAAATTTGTTGGCTGTTTAAAGACCCTGTCTCCTCCCC CACCCTCCCCTTCCTTTTCCCATCAAATGCCTACCTACTGAGGCGCCTTCTCTTTTTATTCTTGAGCCAATGAGATATCAGTGTGTTTAT TAATAAGGATTGCAATAGTATAGTGCTTAATATGTGCCAGGCATAGTGCTAAGGGTTTACATACGTTATCCCTTTTAATCCAGAAAACAG CCTTTGATTAGGTATTGTTGTCTCCATTTTATAGATGAGGAAACTGAGGCAGTATGACTGAGGCCGAGAGCAGTGGTTAAGACAGACTTT GGGATCAAATCACCCTAGGTGCGGGCAGGTTTCTAGGCTGCTCTAAGCTTCTACCTTGAAGGCAGTTGGGTCTGATAATAGTACCTTCTT CCTGGGGTGGTTGTGAGGATATAACGAGGTAATGCATGTAAGCATTTAGTACCATGCCTGGCACATAGTAAGCACTCAATAAATGGTAGT TACTTTTGTTGAGCTCAGCTGGTAGTTTGACCTCCGTTGGTGCAATGCCAGAGATGGGATTGTCAGCCACCACACTGCCACCACCCTGCA CACAGTCTGGGTCCCTCCGAGAGAAGCACGCCAACCTTAACATCCCCTACAGCATCTTGGTGCTTCACACACCATTTAGGAGGCCACCCT AACGAGGCAAGAGGACATGGGGGTGTGCAGAAGTTCTGTTCTTAGTGAGAATGGCTGGCCTGAGAATCCAGCTCACCTCCTCTGCCCACA GTCTGGACTGCTTGTGGGAAGGCCATCTCCTCCTGTCTGGTGGTACCTCCACTCTGGGCAGGGAGGGCCTGTCTGTGACTCGCTCTGGAC CCCACTTCCCGCTCTGGCCAAGGGCTGTCTTCTGTGCACATAGCTCAGGAGCCTGCAGCAGTTCCCTGTATGGGTGGATGTCCTCAGGGG CCAAGAAGAGAAGAATGGGGACAACAGAAACCTGCTTTGTAGGGGATTATGTAGACTAAAAGCCTAGAAGTTCTGCCCACATCCTTATGA CGCAGCCAATCTGGAGCTTCATGTTTCCTTCTAAAAGTAGTAACCGGGTAGAAGTGGGTGACCCATTGAGGACACATGATGTGAGGTATT TCTGCCGGATTCTAGGTTAGCAAAGACTCAAATGGGTGACTCATGAATGAAGGGAGAACAGGCACAGGACTTAATGAAAAGAGTTTCTTG AACTTAGTTTTCTGAATGGGGGTGGGGAGTTGGGGCACGTGTGTGTAAGGGGAGGCTGCCTCACTTTGAAGGAATGGTGTTGGCTAACAG GATCCATGAAATTCCTAGGTCAGTTGAATAATAACTGAACTACTGGTTTGACTGTGGATTATTTCCTGGGTATTAGTGGGACCCAACAGC ACATTGCTATTAATAGTATGGAAATAATGGTCCCTCTGTTTCATTGAATTGCTGAAGTAATGATAATGCCATGAACCAGGAGAAGATGGT TGTGGGGGTTTCTAGGAAGCAGCTGCTGTGACAGGCCAATCCCAGGCTTGCCCCCGAGCCCTCCTGTCCTTCTGGGTTTGTGTTTGCCTT GCCTGGTGCTTCCTCCCTCATTGTGTTGTTGAATAGACCTAGGCAGCCTTGTATCCATAGAAAGCCTCCACTGCTGGGCCGGGGAGGGGG TTGTGGTAAGGGCCCTCTGGAAATCAAGCAGAGAATGCCTTTGCAGTGGTCTCCCGGGCATTGGGCAGGGCCTAGCCTTTTGCAGCCCCC AGAGGACCTTGCCTGCATAAAGCACTCAGAAGGGTTAGCCACCTGGTGATTTTGTCTCTAGTGAAACTGTGTTTGCCGATAATCATCCAC ATAGCCACCTCTGTCCTCGCAGTTTGGATGCCTTTTTTTGACACCTGCGCTAGGTTCTTGTATGCTGGCTTTTCTTAAAGGAACAGCAGA GTAGAAAAGAGGTTGAGAATGTTTTCTAGCAGGCAGAATGTGCATACATGTTTTCATGAGTGTCCTTTGGGTGCTGTTTCTTTTAAATCC TCTGTGCACAGGGCTCTGGCCTTTAGTAAACTGTTTTTCTGTCTTACGTCATGCTGACTGGGTGCTAGGGGCTGATTACAAAGGGGAAGA GTTGAACAGACATCAGGGGCCGATGAAACCAAAGGACTAGGAGTCAGGAGAACAAGTCAGGGATTAGGAGACAGCGGTTTGGTTTATTGT TATCCAGCTGGAGGACTCCTAGGGGCAGCAGCAGGAGGAATACCAGGGCCACGGAGGGGCCAGGAGTCTCACAGTGGAGGGCAGACTCTA ACAGATGCCAGCTGAACGCTCGCTGGCCCTGGATGTCATACGAGTTGGGGACCAGAAATCTGGGCTCAGAGAACCCGTCCAGGGAGATTT GAAGCCATGGGTTATCTTCTAGAGTTGATACTGATAATATATTTTAATTTTTATTGATGTTTAATACCTTCTGAAACAGGAGGGTAAGAT CAGATGGGAAGCCCCTCTGTTGAAGGATCTTGGGAACCTTGGTGGTTTTTTTTTTTTTTTTGGTTTTTTTTTTTTGATCGAGCTGTGGAC ATCCTTCTTAATTCGATTCTGAGGATTTGTTTAACTAAAAAGTTCCCAAACACAGAAAGGGCCTCCCCACCTGCTTTGGGGAGCTGTCTG TGCTGGGAGTGCCAGGCATCCCATGGGACCCATCACTGCCAGTGTCTGTGCCTCCCAGAGGTCAGCCGTGTCTGCCCTGGCTCTGTCTCC TCTGTGACAGGGCAGAGCATTTCTGGTCAGTTTCTCCATGGTGCCTCCCACCCCTTTGTAAAGTGGATGGACATGATGGAATTCAGTTGT CTCACCCTGATAGCCTGGGTGTTGATATTCACTTTACCCGCACTCAGACACAGGCGACCTTGAAGCAGTTCTCGGTGTGTAGAGTCCACG TGACAGTCCCCACAGCCTCCCCAGATAGCTGTGTGCCTGTGCGCTACTGCTGTGCCATTTTCCCAACTTGGCGTTTCACTAAATGCAGCT GATCTCTCTCTCTGTGCACTCGTGATCCATGTTGAACAATACATGTAGGTTCTTTTTCCACGCAATGTAAGAACATGATATACTGTACGT >83746_83746_1_SLX4IP-CDS2_SLX4IP_chr20_10582467_ENST00000334534_CDS2_chr20_5154169_ENST00000460006_length(amino acids)=561AA_BP=135 MASKKFAVKCGNFAVLVDLHILPQGSNKDTSWFSEQKKEEVCLLLKETIDSRVQEYLEVRKQHRPSNAEFTRSNPLSLKGYGFQITAYFL KRGIRLRCIRSTQNAELCVFPDRFVVCVSQLAFSRDLLASQNEDLESESEAKVDGETASDSESRAESAPLPVSADDTPEVLNRALSNLSS RWKNWWVRGILTLAMIAFFFIIIYLGPMVLMIIVMCVQIKCFHEIITIGYNVYHSYDLPWFRTLSWYFLLCVNYFFYGETVTDYFFTLVQ REEPLRILSKYHRFISFTLYLIGFCMFVLSLVKKHYRLQFYMFGWTHVTLLIVVTQSHLVIHNLFEGMIWFIVPISCVICNDIMAYMFGF FFGRTPLIKLSPKKTWEGFIGGFFATVVFGLLLSYVMSGYRCFVCPVEYNNDTNSFTVDCEPSDLFRLQEYNIPGVIQSVIGWKTVRMYP FQIHSIALSTFASLIGPFGGFFASGFKRAFKIKDFANTIPGHGGIMDRFDCQYLMATFVNVYIASFIRGPNPSKLIQQFLTLRPDQQLHI -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for SLX4IP-CDS2 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for SLX4IP-CDS2 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for SLX4IP-CDS2 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
