
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:PLEKHA7-MMP9 (FusionGDB2 ID:HG144100TG4318) |
Fusion Gene Summary for PLEKHA7-MMP9 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: PLEKHA7-MMP9 | Fusion gene ID: hg144100tg4318 | Hgene | Tgene | Gene symbol | PLEKHA7 | MMP9 | Gene ID | 144100 | 4318 |
| Gene name | pleckstrin homology domain containing A7 | matrix metallopeptidase 9 | |
| Synonyms | - | CLG4B|GELB|MANDP2|MMP-9 | |
| Cytomap | ('PLEKHA7')('MMP9') 11p15.2-p15.1 | 20q13.12 | |
| Type of gene | protein-coding | protein-coding | |
| Description | pleckstrin homology domain-containing family A member 7PH domain-containing family A member 7pleckstrin homology domain containing, family A member 2pleckstrin homology domain containing, family A member 7pleckstrin homology domain-containing family A | matrix metalloproteinase-9macrophage gelatinasematrix metallopeptidase 9 (gelatinase B, 92kDa gelatinase, 92kDa type IV collagenase)matrix metalloproteinase 9 (gelatinase B, 92kDa gelatinase, 92kDa type IV collagenase)type V collagenase | |
| Modification date | 20200313 | 20200329 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000355661, ENST00000448080, ENST00000531066, ENST00000332954, ENST00000532079, | ||
| Fusion gene scores | * DoF score | 18 X 12 X 7=1512 | 5 X 4 X 4=80 |
| # samples | 22 | 5 | |
| ** MAII score | log2(22/1512*10)=-2.78088271069641 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(5/80*10)=-0.678071905112638 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: PLEKHA7 [Title/Abstract] AND MMP9 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | PLEKHA7(16872738)-MMP9(44638505), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | PLEKHA7-MMP9 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. PLEKHA7-MMP9 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. PLEKHA7-MMP9 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. PLEKHA7-MMP9 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | MMP9 | GO:0006508 | proteolysis | 2551898|15863497|19022250|24164424 |
| Tgene | MMP9 | GO:0034614 | cellular response to reactive oxygen species | 26514923 |
| Tgene | MMP9 | GO:0043388 | positive regulation of DNA binding | 22984561 |
| Tgene | MMP9 | GO:0071276 | cellular response to cadmium ion | 26514923 |
| Tgene | MMP9 | GO:1900122 | positive regulation of receptor binding | 24164424 |
| Tgene | MMP9 | GO:2001258 | negative regulation of cation channel activity | 24164424 |
 Fusion gene breakpoints across PLEKHA7 (5'-gene) Fusion gene breakpoints across PLEKHA7 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
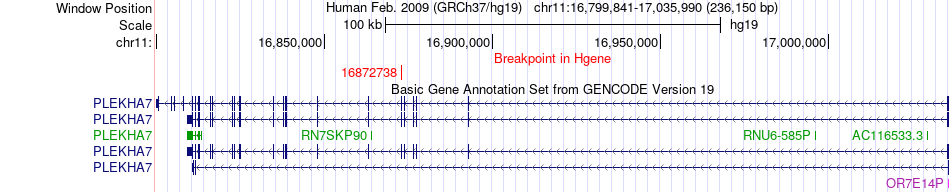 |
 Fusion gene breakpoints across MMP9 (3'-gene) Fusion gene breakpoints across MMP9 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
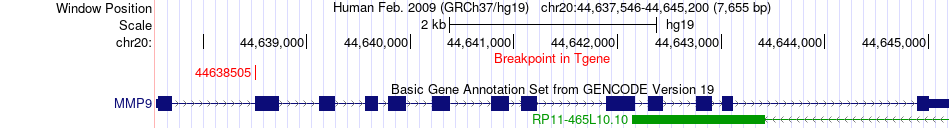 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | PRAD | TCGA-EJ-7785-01A | PLEKHA7 | chr11 | 16872738 | - | MMP9 | chr20 | 44638505 | + |
Top |
Fusion Gene ORF analysis for PLEKHA7-MMP9 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| In-frame | ENST00000355661 | ENST00000372330 | PLEKHA7 | chr11 | 16872738 | - | MMP9 | chr20 | 44638505 | + |
| In-frame | ENST00000448080 | ENST00000372330 | PLEKHA7 | chr11 | 16872738 | - | MMP9 | chr20 | 44638505 | + |
| In-frame | ENST00000531066 | ENST00000372330 | PLEKHA7 | chr11 | 16872738 | - | MMP9 | chr20 | 44638505 | + |
| intron-3CDS | ENST00000332954 | ENST00000372330 | PLEKHA7 | chr11 | 16872738 | - | MMP9 | chr20 | 44638505 | + |
| intron-3CDS | ENST00000532079 | ENST00000372330 | PLEKHA7 | chr11 | 16872738 | - | MMP9 | chr20 | 44638505 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000531066 | PLEKHA7 | chr11 | 16872738 | - | ENST00000372330 | MMP9 | chr20 | 44638505 | + | 2917 | 738 | 42 | 2723 | 893 |
| ENST00000355661 | PLEKHA7 | chr11 | 16872738 | - | ENST00000372330 | MMP9 | chr20 | 44638505 | + | 2886 | 707 | 11 | 2692 | 893 |
| ENST00000448080 | PLEKHA7 | chr11 | 16872738 | - | ENST00000372330 | MMP9 | chr20 | 44638505 | + | 2890 | 711 | 15 | 2696 | 893 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000531066 | ENST00000372330 | PLEKHA7 | chr11 | 16872738 | - | MMP9 | chr20 | 44638505 | + | 0.002338196 | 0.9976618 |
| ENST00000355661 | ENST00000372330 | PLEKHA7 | chr11 | 16872738 | - | MMP9 | chr20 | 44638505 | + | 0.002376735 | 0.99762326 |
| ENST00000448080 | ENST00000372330 | PLEKHA7 | chr11 | 16872738 | - | MMP9 | chr20 | 44638505 | + | 0.002354079 | 0.9976459 |
Top |
Fusion Genomic Features for PLEKHA7-MMP9 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| PLEKHA7 | chr11 | 16872737 | - | MMP9 | chr20 | 44638504 | + | 0.01460192 | 0.98539805 |
| PLEKHA7 | chr11 | 16872737 | - | MMP9 | chr20 | 44638504 | + | 0.01460192 | 0.98539805 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
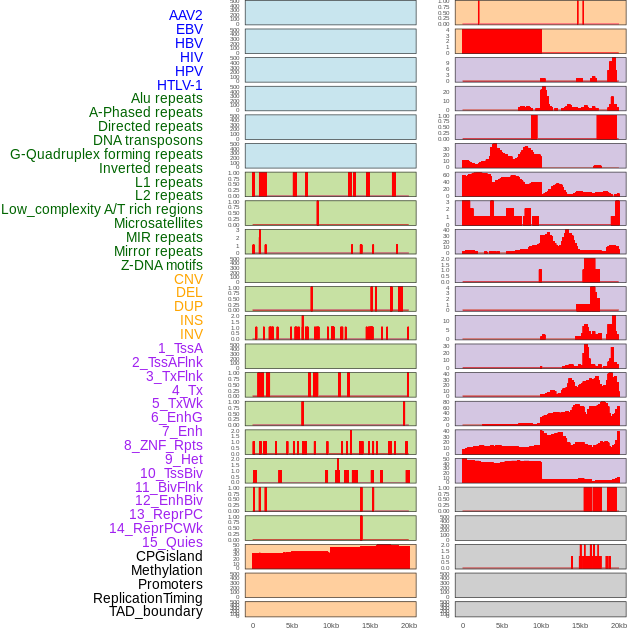 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
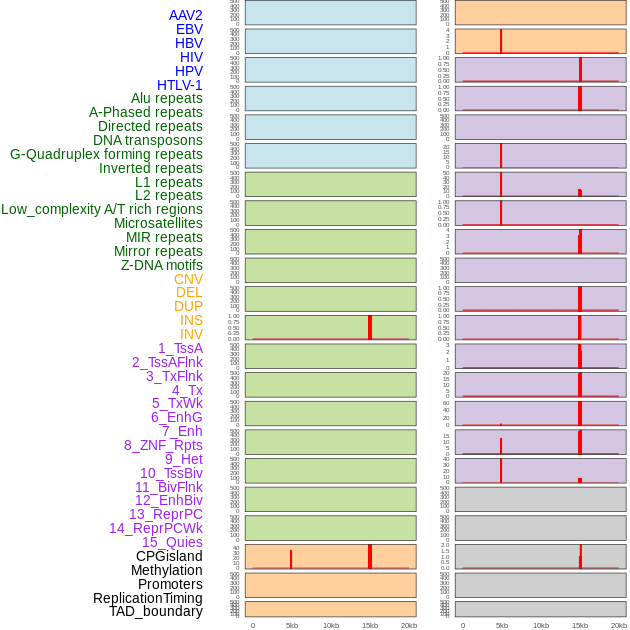 |
Top |
Fusion Protein Features for PLEKHA7-MMP9 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr11:16872738/chr20:44638505) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | PLEKHA7 | chr11:16872738 | chr20:44638505 | ENST00000355661 | - | 8 | 23 | 54_87 | 232 | 1122.0 | Domain | WW 2 |
| Hgene | PLEKHA7 | chr11:16872738 | chr20:44638505 | ENST00000355661 | - | 8 | 23 | 9_42 | 232 | 1122.0 | Domain | WW 1 |
| Hgene | PLEKHA7 | chr11:16872738 | chr20:44638505 | ENST00000448080 | - | 8 | 23 | 54_87 | 232 | 1123.0 | Domain | WW 2 |
| Hgene | PLEKHA7 | chr11:16872738 | chr20:44638505 | ENST00000448080 | - | 8 | 23 | 9_42 | 232 | 1123.0 | Domain | WW 1 |
| Tgene | MMP9 | chr11:16872738 | chr20:44638505 | ENST00000372330 | 0 | 13 | 225_273 | 46 | 708.0 | Domain | Fibronectin type-II 1 | |
| Tgene | MMP9 | chr11:16872738 | chr20:44638505 | ENST00000372330 | 0 | 13 | 283_331 | 46 | 708.0 | Domain | Fibronectin type-II 2 | |
| Tgene | MMP9 | chr11:16872738 | chr20:44638505 | ENST00000372330 | 0 | 13 | 342_390 | 46 | 708.0 | Domain | Fibronectin type-II 3 | |
| Tgene | MMP9 | chr11:16872738 | chr20:44638505 | ENST00000372330 | 0 | 13 | 97_104 | 46 | 708.0 | Motif | Cysteine switch | |
| Tgene | MMP9 | chr11:16872738 | chr20:44638505 | ENST00000372330 | 0 | 13 | 518_563 | 46 | 708.0 | Repeat | Note=Hemopexin 1 | |
| Tgene | MMP9 | chr11:16872738 | chr20:44638505 | ENST00000372330 | 0 | 13 | 564_608 | 46 | 708.0 | Repeat | Note=Hemopexin 2 | |
| Tgene | MMP9 | chr11:16872738 | chr20:44638505 | ENST00000372330 | 0 | 13 | 610_657 | 46 | 708.0 | Repeat | Note=Hemopexin 3 | |
| Tgene | MMP9 | chr11:16872738 | chr20:44638505 | ENST00000372330 | 0 | 13 | 658_704 | 46 | 708.0 | Repeat | Note=Hemopexin 4 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | PLEKHA7 | chr11:16872738 | chr20:44638505 | ENST00000355661 | - | 8 | 23 | 1067_1094 | 232 | 1122.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | PLEKHA7 | chr11:16872738 | chr20:44638505 | ENST00000355661 | - | 8 | 23 | 700_801 | 232 | 1122.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | PLEKHA7 | chr11:16872738 | chr20:44638505 | ENST00000448080 | - | 8 | 23 | 1067_1094 | 232 | 1123.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | PLEKHA7 | chr11:16872738 | chr20:44638505 | ENST00000448080 | - | 8 | 23 | 700_801 | 232 | 1123.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | PLEKHA7 | chr11:16872738 | chr20:44638505 | ENST00000355661 | - | 8 | 23 | 567_599 | 232 | 1122.0 | Compositional bias | Note=Pro-rich |
| Hgene | PLEKHA7 | chr11:16872738 | chr20:44638505 | ENST00000355661 | - | 8 | 23 | 845_942 | 232 | 1122.0 | Compositional bias | Note=Pro-rich |
| Hgene | PLEKHA7 | chr11:16872738 | chr20:44638505 | ENST00000448080 | - | 8 | 23 | 567_599 | 232 | 1123.0 | Compositional bias | Note=Pro-rich |
| Hgene | PLEKHA7 | chr11:16872738 | chr20:44638505 | ENST00000448080 | - | 8 | 23 | 845_942 | 232 | 1123.0 | Compositional bias | Note=Pro-rich |
| Hgene | PLEKHA7 | chr11:16872738 | chr20:44638505 | ENST00000355661 | - | 8 | 23 | 164_282 | 232 | 1122.0 | Domain | PH |
| Hgene | PLEKHA7 | chr11:16872738 | chr20:44638505 | ENST00000448080 | - | 8 | 23 | 164_282 | 232 | 1123.0 | Domain | PH |
Top |
Fusion Gene Sequence for PLEKHA7-MMP9 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >66222_66222_1_PLEKHA7-MMP9_PLEKHA7_chr11_16872738_ENST00000355661_MMP9_chr20_44638505_ENST00000372330_length(transcript)=2886nt_BP=707nt GCTCGGCGAACATGGCGGCGGCGACGGTCGGGCGGGACACTTTACCTGAGCATTGGTCCTACGGGGTGTGCCGGGATGGCCGCGTCTTCT TCATCAATGACCAGCTCCGCTGCACGACCTGGCTGCATCCGCGCACCGGGGAGCCCGTCAACTCGGGCCACATGATCCGCTCAGACCTGC CCCGCGGCTGGGAGGAGGGCTTCACGGAGGAGGGCGCCAGCTACTTCATCGACCATAACCAGCAGACCACAGCATTCAGGCATCCTGTGA CGGGACAGTTTTCTCCAGAAAATAGTGAATTCATTCTTCAAGAAGAGCCGAATCCACATATGTCGAAGCAAGACAGAAACCAAAGACCGT CCAGCATGGTCAGTGAAACATCCACGGCTGGGACCGCCTCCACCCTGGAGGCCAAGCCTGGACCCAAGATCATAAAGTCCAGCAGTAAAG TCCACAGCTTTGGGAAGAGAGACCAGGCCATTCGGAGGAACCCCAATGTTCCCGTGGTGGTGAGGGGCTGGCTGCACAAGCAGGACAGTT CTGGGATGAGGCTGTGGAAAAGGAGGTGGTTTGTGCTTGCTGATTACTGCTTATTTTACTATAAAGACAGCCGAGAAGAAGCGGTCCTCG GGAGCATCCCCTTGCCCAGCTACGTGATCTCTCCTGTGGCCCCTGAGGATCGCATAAGCCGCAAATATTCCTTTAAGGAATACCTGTACC GCTATGGTTACACTCGGGTGGCAGAGATGCGTGGAGAGTCGAAATCTCTGGGGCCTGCGCTGCTGCTTCTCCAGAAGCAACTGTCCCTGC CCGAGACCGGTGAGCTGGATAGCGCCACGCTGAAGGCCATGCGAACCCCACGGTGCGGGGTCCCAGACCTGGGCAGATTCCAAACCTTTG AGGGCGACCTCAAGTGGCACCACCACAACATCACCTATTGGATCCAAAACTACTCGGAAGACTTGCCGCGGGCGGTGATTGACGACGCCT TTGCCCGCGCCTTCGCACTGTGGAGCGCGGTGACGCCGCTCACCTTCACTCGCGTGTACAGCCGGGACGCAGACATCGTCATCCAGTTTG GTGTCGCGGAGCACGGAGACGGGTATCCCTTCGACGGGAAGGACGGGCTCCTGGCACACGCCTTTCCTCCTGGCCCCGGCATTCAGGGAG ACGCCCATTTCGACGATGACGAGTTGTGGTCCCTGGGCAAGGGCGTCGTGGTTCCAACTCGGTTTGGAAACGCAGATGGCGCGGCCTGCC ACTTCCCCTTCATCTTCGAGGGCCGCTCCTACTCTGCCTGCACCACCGACGGTCGCTCCGACGGCTTGCCCTGGTGCAGTACCACGGCCA ACTACGACACCGACGACCGGTTTGGCTTCTGCCCCAGCGAGAGACTCTACACCCAGGACGGCAATGCTGATGGGAAACCCTGCCAGTTTC CATTCATCTTCCAAGGCCAATCCTACTCCGCCTGCACCACGGACGGTCGCTCCGACGGCTACCGCTGGTGCGCCACCACCGCCAACTACG ACCGGGACAAGCTCTTCGGCTTCTGCCCGACCCGAGCTGACTCGACGGTGATGGGGGGCAACTCGGCGGGGGAGCTGTGCGTCTTCCCCT TCACTTTCCTGGGTAAGGAGTACTCGACCTGTACCAGCGAGGGCCGCGGAGATGGGCGCCTCTGGTGCGCTACCACCTCGAACTTTGACA GCGACAAGAAGTGGGGCTTCTGCCCGGACCAAGGATACAGTTTGTTCCTCGTGGCGGCGCATGAGTTCGGCCACGCGCTGGGCTTAGATC ATTCCTCAGTGCCGGAGGCGCTCATGTACCCTATGTACCGCTTCACTGAGGGGCCCCCCTTGCATAAGGACGACGTGAATGGCATCCGGC ACCTCTATGGTCCTCGCCCTGAACCTGAGCCACGGCCTCCAACCACCACCACACCGCAGCCCACGGCTCCCCCGACGGTCTGCCCCACCG GACCCCCCACTGTCCACCCCTCAGAGCGCCCCACAGCTGGCCCCACAGGTCCCCCCTCAGCTGGCCCCACAGGTCCCCCCACTGCTGGCC CTTCTACGGCCACTACTGTGCCTTTGAGTCCGGTGGACGATGCCTGCAACGTGAACATCTTCGACGCCATCGCGGAGATTGGGAACCAGC TGTATTTGTTCAAGGATGGGAAGTACTGGCGATTCTCTGAGGGCAGGGGGAGCCGGCCGCAGGGCCCCTTCCTTATCGCCGACAAGTGGC CCGCGCTGCCCCGCAAGCTGGACTCGGTCTTTGAGGAGCGGCTCTCCAAGAAGCTTTTCTTCTTCTCTGGGCGCCAGGTGTGGGTGTACA CAGGCGCGTCGGTGCTGGGCCCGAGGCGTCTGGACAAGCTGGGCCTGGGAGCCGACGTGGCCCAGGTGACCGGGGCCCTCCGGAGTGGCA GGGGGAAGATGCTGCTGTTCAGCGGGCGGCGCCTCTGGAGGTTCGACGTGAAGGCGCAGATGGTGGATCCCCGGAGCGCCAGCGAGGTGG ACCGGATGTTCCCCGGGGTGCCTTTGGACACGCACGACGTCTTCCAGTACCGAGAGAAAGCCTATTTCTGCCAGGACCGCTTCTACTGGC GCGTGAGTTCCCGGAGTGAGTTGAACCAGGTGGACCAAGTGGGCTACGTGACCTATGACATCCTGCAGTGCCCTGAGGACTAGGGCTCCC GTCCTGCTTTGGCAGTGCCATGTAAATCCCCACTGGGACCAACCCTGGGGAAGGAGCCAGTTTGCCGGATACAAACTGGTATTCTGTTCT GGAGGAAAGGGAGGAGTGGAGGTGGGCTGGGCCCTCTCTTCTCACCTTTGTTTTTTGTTGGAGTGTTTCTAATAAACTTGGATTCTCTAA >66222_66222_1_PLEKHA7-MMP9_PLEKHA7_chr11_16872738_ENST00000355661_MMP9_chr20_44638505_ENST00000372330_length(amino acids)=893AA_BP=232 MAAATVGRDTLPEHWSYGVCRDGRVFFINDQLRCTTWLHPRTGEPVNSGHMIRSDLPRGWEEGFTEEGASYFIDHNQQTTAFRHPVTGQF SPENSEFILQEEPNPHMSKQDRNQRPSSMVSETSTAGTASTLEAKPGPKIIKSSSKVHSFGKRDQAIRRNPNVPVVVRGWLHKQDSSGMR LWKRRWFVLADYCLFYYKDSREEAVLGSIPLPSYVISPVAPEDRISRKYSFKEYLYRYGYTRVAEMRGESKSLGPALLLLQKQLSLPETG ELDSATLKAMRTPRCGVPDLGRFQTFEGDLKWHHHNITYWIQNYSEDLPRAVIDDAFARAFALWSAVTPLTFTRVYSRDADIVIQFGVAE HGDGYPFDGKDGLLAHAFPPGPGIQGDAHFDDDELWSLGKGVVVPTRFGNADGAACHFPFIFEGRSYSACTTDGRSDGLPWCSTTANYDT DDRFGFCPSERLYTQDGNADGKPCQFPFIFQGQSYSACTTDGRSDGYRWCATTANYDRDKLFGFCPTRADSTVMGGNSAGELCVFPFTFL GKEYSTCTSEGRGDGRLWCATTSNFDSDKKWGFCPDQGYSLFLVAAHEFGHALGLDHSSVPEALMYPMYRFTEGPPLHKDDVNGIRHLYG PRPEPEPRPPTTTTPQPTAPPTVCPTGPPTVHPSERPTAGPTGPPSAGPTGPPTAGPSTATTVPLSPVDDACNVNIFDAIAEIGNQLYLF KDGKYWRFSEGRGSRPQGPFLIADKWPALPRKLDSVFEERLSKKLFFFSGRQVWVYTGASVLGPRRLDKLGLGADVAQVTGALRSGRGKM -------------------------------------------------------------- >66222_66222_2_PLEKHA7-MMP9_PLEKHA7_chr11_16872738_ENST00000448080_MMP9_chr20_44638505_ENST00000372330_length(transcript)=2890nt_BP=711nt CCGCGCTCGGCGAACATGGCGGCGGCGACGGTCGGGCGGGACACTTTACCTGAGCATTGGTCCTACGGGGTGTGCCGGGATGGCCGCGTC TTCTTCATCAATGACCAGCTCCGCTGCACGACCTGGCTGCATCCGCGCACCGGGGAGCCCGTCAACTCGGGCCACATGATCCGCTCAGAC CTGCCCCGCGGCTGGGAGGAGGGCTTCACGGAGGAGGGCGCCAGCTACTTCATCGACCATAACCAGCAGACCACAGCATTCAGGCATCCT GTGACGGGACAGTTTTCTCCAGAAAATAGTGAATTCATTCTTCAAGAAGAGCCGAATCCACATATGTCGAAGCAAGACAGAAACCAAAGA CCGTCCAGCATGGTCAGTGAAACATCCACGGCTGGGACCGCCTCCACCCTGGAGGCCAAGCCTGGACCCAAGATCATAAAGTCCAGCAGT AAAGTCCACAGCTTTGGGAAGAGAGACCAGGCCATTCGGAGGAACCCCAATGTTCCCGTGGTGGTGAGGGGCTGGCTGCACAAGCAGGAC AGTTCTGGGATGAGGCTGTGGAAAAGGAGGTGGTTTGTGCTTGCTGATTACTGCTTATTTTACTATAAAGACAGCCGAGAAGAAGCGGTC CTCGGGAGCATCCCCTTGCCCAGCTACGTGATCTCTCCTGTGGCCCCTGAGGATCGCATAAGCCGCAAATATTCCTTTAAGGAATACCTG TACCGCTATGGTTACACTCGGGTGGCAGAGATGCGTGGAGAGTCGAAATCTCTGGGGCCTGCGCTGCTGCTTCTCCAGAAGCAACTGTCC CTGCCCGAGACCGGTGAGCTGGATAGCGCCACGCTGAAGGCCATGCGAACCCCACGGTGCGGGGTCCCAGACCTGGGCAGATTCCAAACC TTTGAGGGCGACCTCAAGTGGCACCACCACAACATCACCTATTGGATCCAAAACTACTCGGAAGACTTGCCGCGGGCGGTGATTGACGAC GCCTTTGCCCGCGCCTTCGCACTGTGGAGCGCGGTGACGCCGCTCACCTTCACTCGCGTGTACAGCCGGGACGCAGACATCGTCATCCAG TTTGGTGTCGCGGAGCACGGAGACGGGTATCCCTTCGACGGGAAGGACGGGCTCCTGGCACACGCCTTTCCTCCTGGCCCCGGCATTCAG GGAGACGCCCATTTCGACGATGACGAGTTGTGGTCCCTGGGCAAGGGCGTCGTGGTTCCAACTCGGTTTGGAAACGCAGATGGCGCGGCC TGCCACTTCCCCTTCATCTTCGAGGGCCGCTCCTACTCTGCCTGCACCACCGACGGTCGCTCCGACGGCTTGCCCTGGTGCAGTACCACG GCCAACTACGACACCGACGACCGGTTTGGCTTCTGCCCCAGCGAGAGACTCTACACCCAGGACGGCAATGCTGATGGGAAACCCTGCCAG TTTCCATTCATCTTCCAAGGCCAATCCTACTCCGCCTGCACCACGGACGGTCGCTCCGACGGCTACCGCTGGTGCGCCACCACCGCCAAC TACGACCGGGACAAGCTCTTCGGCTTCTGCCCGACCCGAGCTGACTCGACGGTGATGGGGGGCAACTCGGCGGGGGAGCTGTGCGTCTTC CCCTTCACTTTCCTGGGTAAGGAGTACTCGACCTGTACCAGCGAGGGCCGCGGAGATGGGCGCCTCTGGTGCGCTACCACCTCGAACTTT GACAGCGACAAGAAGTGGGGCTTCTGCCCGGACCAAGGATACAGTTTGTTCCTCGTGGCGGCGCATGAGTTCGGCCACGCGCTGGGCTTA GATCATTCCTCAGTGCCGGAGGCGCTCATGTACCCTATGTACCGCTTCACTGAGGGGCCCCCCTTGCATAAGGACGACGTGAATGGCATC CGGCACCTCTATGGTCCTCGCCCTGAACCTGAGCCACGGCCTCCAACCACCACCACACCGCAGCCCACGGCTCCCCCGACGGTCTGCCCC ACCGGACCCCCCACTGTCCACCCCTCAGAGCGCCCCACAGCTGGCCCCACAGGTCCCCCCTCAGCTGGCCCCACAGGTCCCCCCACTGCT GGCCCTTCTACGGCCACTACTGTGCCTTTGAGTCCGGTGGACGATGCCTGCAACGTGAACATCTTCGACGCCATCGCGGAGATTGGGAAC CAGCTGTATTTGTTCAAGGATGGGAAGTACTGGCGATTCTCTGAGGGCAGGGGGAGCCGGCCGCAGGGCCCCTTCCTTATCGCCGACAAG TGGCCCGCGCTGCCCCGCAAGCTGGACTCGGTCTTTGAGGAGCGGCTCTCCAAGAAGCTTTTCTTCTTCTCTGGGCGCCAGGTGTGGGTG TACACAGGCGCGTCGGTGCTGGGCCCGAGGCGTCTGGACAAGCTGGGCCTGGGAGCCGACGTGGCCCAGGTGACCGGGGCCCTCCGGAGT GGCAGGGGGAAGATGCTGCTGTTCAGCGGGCGGCGCCTCTGGAGGTTCGACGTGAAGGCGCAGATGGTGGATCCCCGGAGCGCCAGCGAG GTGGACCGGATGTTCCCCGGGGTGCCTTTGGACACGCACGACGTCTTCCAGTACCGAGAGAAAGCCTATTTCTGCCAGGACCGCTTCTAC TGGCGCGTGAGTTCCCGGAGTGAGTTGAACCAGGTGGACCAAGTGGGCTACGTGACCTATGACATCCTGCAGTGCCCTGAGGACTAGGGC TCCCGTCCTGCTTTGGCAGTGCCATGTAAATCCCCACTGGGACCAACCCTGGGGAAGGAGCCAGTTTGCCGGATACAAACTGGTATTCTG TTCTGGAGGAAAGGGAGGAGTGGAGGTGGGCTGGGCCCTCTCTTCTCACCTTTGTTTTTTGTTGGAGTGTTTCTAATAAACTTGGATTCT >66222_66222_2_PLEKHA7-MMP9_PLEKHA7_chr11_16872738_ENST00000448080_MMP9_chr20_44638505_ENST00000372330_length(amino acids)=893AA_BP=232 MAAATVGRDTLPEHWSYGVCRDGRVFFINDQLRCTTWLHPRTGEPVNSGHMIRSDLPRGWEEGFTEEGASYFIDHNQQTTAFRHPVTGQF SPENSEFILQEEPNPHMSKQDRNQRPSSMVSETSTAGTASTLEAKPGPKIIKSSSKVHSFGKRDQAIRRNPNVPVVVRGWLHKQDSSGMR LWKRRWFVLADYCLFYYKDSREEAVLGSIPLPSYVISPVAPEDRISRKYSFKEYLYRYGYTRVAEMRGESKSLGPALLLLQKQLSLPETG ELDSATLKAMRTPRCGVPDLGRFQTFEGDLKWHHHNITYWIQNYSEDLPRAVIDDAFARAFALWSAVTPLTFTRVYSRDADIVIQFGVAE HGDGYPFDGKDGLLAHAFPPGPGIQGDAHFDDDELWSLGKGVVVPTRFGNADGAACHFPFIFEGRSYSACTTDGRSDGLPWCSTTANYDT DDRFGFCPSERLYTQDGNADGKPCQFPFIFQGQSYSACTTDGRSDGYRWCATTANYDRDKLFGFCPTRADSTVMGGNSAGELCVFPFTFL GKEYSTCTSEGRGDGRLWCATTSNFDSDKKWGFCPDQGYSLFLVAAHEFGHALGLDHSSVPEALMYPMYRFTEGPPLHKDDVNGIRHLYG PRPEPEPRPPTTTTPQPTAPPTVCPTGPPTVHPSERPTAGPTGPPSAGPTGPPTAGPSTATTVPLSPVDDACNVNIFDAIAEIGNQLYLF KDGKYWRFSEGRGSRPQGPFLIADKWPALPRKLDSVFEERLSKKLFFFSGRQVWVYTGASVLGPRRLDKLGLGADVAQVTGALRSGRGKM -------------------------------------------------------------- >66222_66222_3_PLEKHA7-MMP9_PLEKHA7_chr11_16872738_ENST00000531066_MMP9_chr20_44638505_ENST00000372330_length(transcript)=2917nt_BP=738nt CCCCCTCCCCCCCACCCCGCGGCGGCTCCGCGCTCGGCGAACATGGCGGCGGCGACGGTCGGGCGGGACACTTTACCTGAGCATTGGTCC TACGGGGTGTGCCGGGATGGCCGCGTCTTCTTCATCAATGACCAGCTCCGCTGCACGACCTGGCTGCATCCGCGCACCGGGGAGCCCGTC AACTCGGGCCACATGATCCGCTCAGACCTGCCCCGCGGCTGGGAGGAGGGCTTCACGGAGGAGGGCGCCAGCTACTTCATCGACCATAAC CAGCAGACCACAGCATTCAGGCATCCTGTGACGGGACAGTTTTCTCCAGAAAATAGTGAATTCATTCTTCAAGAAGAGCCGAATCCACAT ATGTCGAAGCAAGACAGAAACCAAAGACCGTCCAGCATGGTCAGTGAAACATCCACGGCTGGGACCGCCTCCACCCTGGAGGCCAAGCCT GGACCCAAGATCATAAAGTCCAGCAGTAAAGTCCACAGCTTTGGGAAGAGAGACCAGGCCATTCGGAGGAACCCCAATGTTCCCGTGGTG GTGAGGGGCTGGCTGCACAAGCAGGACAGTTCTGGGATGAGGCTGTGGAAAAGGAGGTGGTTTGTGCTTGCTGATTACTGCTTATTTTAC TATAAAGACAGCCGAGAAGAAGCGGTCCTCGGGAGCATCCCCTTGCCCAGCTACGTGATCTCTCCTGTGGCCCCTGAGGATCGCATAAGC CGCAAATATTCCTTTAAGGAATACCTGTACCGCTATGGTTACACTCGGGTGGCAGAGATGCGTGGAGAGTCGAAATCTCTGGGGCCTGCG CTGCTGCTTCTCCAGAAGCAACTGTCCCTGCCCGAGACCGGTGAGCTGGATAGCGCCACGCTGAAGGCCATGCGAACCCCACGGTGCGGG GTCCCAGACCTGGGCAGATTCCAAACCTTTGAGGGCGACCTCAAGTGGCACCACCACAACATCACCTATTGGATCCAAAACTACTCGGAA GACTTGCCGCGGGCGGTGATTGACGACGCCTTTGCCCGCGCCTTCGCACTGTGGAGCGCGGTGACGCCGCTCACCTTCACTCGCGTGTAC AGCCGGGACGCAGACATCGTCATCCAGTTTGGTGTCGCGGAGCACGGAGACGGGTATCCCTTCGACGGGAAGGACGGGCTCCTGGCACAC GCCTTTCCTCCTGGCCCCGGCATTCAGGGAGACGCCCATTTCGACGATGACGAGTTGTGGTCCCTGGGCAAGGGCGTCGTGGTTCCAACT CGGTTTGGAAACGCAGATGGCGCGGCCTGCCACTTCCCCTTCATCTTCGAGGGCCGCTCCTACTCTGCCTGCACCACCGACGGTCGCTCC GACGGCTTGCCCTGGTGCAGTACCACGGCCAACTACGACACCGACGACCGGTTTGGCTTCTGCCCCAGCGAGAGACTCTACACCCAGGAC GGCAATGCTGATGGGAAACCCTGCCAGTTTCCATTCATCTTCCAAGGCCAATCCTACTCCGCCTGCACCACGGACGGTCGCTCCGACGGC TACCGCTGGTGCGCCACCACCGCCAACTACGACCGGGACAAGCTCTTCGGCTTCTGCCCGACCCGAGCTGACTCGACGGTGATGGGGGGC AACTCGGCGGGGGAGCTGTGCGTCTTCCCCTTCACTTTCCTGGGTAAGGAGTACTCGACCTGTACCAGCGAGGGCCGCGGAGATGGGCGC CTCTGGTGCGCTACCACCTCGAACTTTGACAGCGACAAGAAGTGGGGCTTCTGCCCGGACCAAGGATACAGTTTGTTCCTCGTGGCGGCG CATGAGTTCGGCCACGCGCTGGGCTTAGATCATTCCTCAGTGCCGGAGGCGCTCATGTACCCTATGTACCGCTTCACTGAGGGGCCCCCC TTGCATAAGGACGACGTGAATGGCATCCGGCACCTCTATGGTCCTCGCCCTGAACCTGAGCCACGGCCTCCAACCACCACCACACCGCAG CCCACGGCTCCCCCGACGGTCTGCCCCACCGGACCCCCCACTGTCCACCCCTCAGAGCGCCCCACAGCTGGCCCCACAGGTCCCCCCTCA GCTGGCCCCACAGGTCCCCCCACTGCTGGCCCTTCTACGGCCACTACTGTGCCTTTGAGTCCGGTGGACGATGCCTGCAACGTGAACATC TTCGACGCCATCGCGGAGATTGGGAACCAGCTGTATTTGTTCAAGGATGGGAAGTACTGGCGATTCTCTGAGGGCAGGGGGAGCCGGCCG CAGGGCCCCTTCCTTATCGCCGACAAGTGGCCCGCGCTGCCCCGCAAGCTGGACTCGGTCTTTGAGGAGCGGCTCTCCAAGAAGCTTTTC TTCTTCTCTGGGCGCCAGGTGTGGGTGTACACAGGCGCGTCGGTGCTGGGCCCGAGGCGTCTGGACAAGCTGGGCCTGGGAGCCGACGTG GCCCAGGTGACCGGGGCCCTCCGGAGTGGCAGGGGGAAGATGCTGCTGTTCAGCGGGCGGCGCCTCTGGAGGTTCGACGTGAAGGCGCAG ATGGTGGATCCCCGGAGCGCCAGCGAGGTGGACCGGATGTTCCCCGGGGTGCCTTTGGACACGCACGACGTCTTCCAGTACCGAGAGAAA GCCTATTTCTGCCAGGACCGCTTCTACTGGCGCGTGAGTTCCCGGAGTGAGTTGAACCAGGTGGACCAAGTGGGCTACGTGACCTATGAC ATCCTGCAGTGCCCTGAGGACTAGGGCTCCCGTCCTGCTTTGGCAGTGCCATGTAAATCCCCACTGGGACCAACCCTGGGGAAGGAGCCA GTTTGCCGGATACAAACTGGTATTCTGTTCTGGAGGAAAGGGAGGAGTGGAGGTGGGCTGGGCCCTCTCTTCTCACCTTTGTTTTTTGTT >66222_66222_3_PLEKHA7-MMP9_PLEKHA7_chr11_16872738_ENST00000531066_MMP9_chr20_44638505_ENST00000372330_length(amino acids)=893AA_BP=232 MAAATVGRDTLPEHWSYGVCRDGRVFFINDQLRCTTWLHPRTGEPVNSGHMIRSDLPRGWEEGFTEEGASYFIDHNQQTTAFRHPVTGQF SPENSEFILQEEPNPHMSKQDRNQRPSSMVSETSTAGTASTLEAKPGPKIIKSSSKVHSFGKRDQAIRRNPNVPVVVRGWLHKQDSSGMR LWKRRWFVLADYCLFYYKDSREEAVLGSIPLPSYVISPVAPEDRISRKYSFKEYLYRYGYTRVAEMRGESKSLGPALLLLQKQLSLPETG ELDSATLKAMRTPRCGVPDLGRFQTFEGDLKWHHHNITYWIQNYSEDLPRAVIDDAFARAFALWSAVTPLTFTRVYSRDADIVIQFGVAE HGDGYPFDGKDGLLAHAFPPGPGIQGDAHFDDDELWSLGKGVVVPTRFGNADGAACHFPFIFEGRSYSACTTDGRSDGLPWCSTTANYDT DDRFGFCPSERLYTQDGNADGKPCQFPFIFQGQSYSACTTDGRSDGYRWCATTANYDRDKLFGFCPTRADSTVMGGNSAGELCVFPFTFL GKEYSTCTSEGRGDGRLWCATTSNFDSDKKWGFCPDQGYSLFLVAAHEFGHALGLDHSSVPEALMYPMYRFTEGPPLHKDDVNGIRHLYG PRPEPEPRPPTTTTPQPTAPPTVCPTGPPTVHPSERPTAGPTGPPSAGPTGPPTAGPSTATTVPLSPVDDACNVNIFDAIAEIGNQLYLF KDGKYWRFSEGRGSRPQGPFLIADKWPALPRKLDSVFEERLSKKLFFFSGRQVWVYTGASVLGPRRLDKLGLGADVAQVTGALRSGRGKM -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for PLEKHA7-MMP9 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | PLEKHA7 | chr11:16872738 | chr20:44638505 | ENST00000355661 | - | 8 | 23 | 538_696 | 232.0 | 1122.0 | CTNND1 |
| Hgene | PLEKHA7 | chr11:16872738 | chr20:44638505 | ENST00000448080 | - | 8 | 23 | 538_696 | 232.0 | 1123.0 | CTNND1 |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for PLEKHA7-MMP9 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for PLEKHA7-MMP9 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
| Hgene | PLEKHA7 | C0017605 | Angle Closure Glaucoma | 2 | CTD_human |
| Hgene | PLEKHA7 | C0008924 | Cleft upper lip | 1 | GENOMICS_ENGLAND |
| Tgene | C0027626 | Neoplasm Invasiveness | 7 | CTD_human | |
| Tgene | C0027627 | Neoplasm Metastasis | 5 | CTD_human | |
| Tgene | C2937358 | Cerebral Hemorrhage | 5 | CTD_human | |
| Tgene | C0001973 | Alcoholic Intoxication, Chronic | 3 | PSYGENET | |
| Tgene | C0007621 | Neoplastic Cell Transformation | 3 | CTD_human | |
| Tgene | C0002152 | Alloxan Diabetes | 2 | CTD_human | |
| Tgene | C0004096 | Asthma | 2 | CTD_human | |
| Tgene | C0005586 | Bipolar Disorder | 2 | PSYGENET | |
| Tgene | C0006663 | Calcinosis | 2 | CTD_human | |
| Tgene | C0011853 | Diabetes Mellitus, Experimental | 2 | CTD_human | |
| Tgene | C0021368 | Inflammation | 2 | CTD_human | |
| Tgene | C0027051 | Myocardial Infarction | 2 | CTD_human | |
| Tgene | C0031051 | Pericementitis | 2 | CTD_human | |
| Tgene | C0031099 | Periodontitis | 2 | CTD_human | |
| Tgene | C0036341 | Schizophrenia | 2 | PSYGENET | |
| Tgene | C0038433 | Streptozotocin Diabetes | 2 | CTD_human | |
| Tgene | C0263628 | Tumoral calcinosis | 2 | CTD_human | |
| Tgene | C0521174 | Microcalcification | 2 | CTD_human | |
| Tgene | C0003486 | Aortic Aneurysm | 1 | CTD_human | |
| Tgene | C0003493 | Aortic Diseases | 1 | CTD_human | |
| Tgene | C0003496 | Aortic Rupture | 1 | CTD_human | |
| Tgene | C0004114 | Astrocytoma | 1 | CTD_human | |
| Tgene | C0005398 | Cholestasis, Extrahepatic | 1 | CTD_human | |
| Tgene | C0005684 | Malignant neoplasm of urinary bladder | 1 | CTD_human | |
| Tgene | C0005695 | Bladder Neoplasm | 1 | CTD_human | |
| Tgene | C0005967 | Bone neoplasms | 1 | CTD_human | |
| Tgene | C0006114 | Cerebral Edema | 1 | CTD_human | |
| Tgene | C0006142 | Malignant neoplasm of breast | 1 | CTD_human | |
| Tgene | C0007102 | Malignant tumor of colon | 1 | CTD_human | |
| Tgene | C0007114 | Malignant neoplasm of skin | 1 | CTD_human | |
| Tgene | C0007131 | Non-Small Cell Lung Carcinoma | 1 | CTD_human | |
| Tgene | C0007137 | Squamous cell carcinoma | 1 | CTD_human | |
| Tgene | C0007786 | Brain Ischemia | 1 | CTD_human | |
| Tgene | C0009324 | Ulcerative Colitis | 1 | CTD_human | |
| Tgene | C0009375 | Colonic Neoplasms | 1 | CTD_human | |
| Tgene | C0011882 | Diabetic Neuropathies | 1 | CTD_human | |
| Tgene | C0017636 | Glioblastoma | 1 | CTD_human | |
| Tgene | C0019193 | Hepatitis, Toxic | 1 | CTD_human | |
| Tgene | C0020507 | Hyperplasia | 1 | CTD_human | |
| Tgene | C0020538 | Hypertensive disease | 1 | CTD_human | |
| Tgene | C0023176 | Lead Poisoning | 1 | CTD_human | |
| Tgene | C0023893 | Liver Cirrhosis, Experimental | 1 | CTD_human | |
| Tgene | C0023895 | Liver diseases | 1 | CTD_human | |
| Tgene | C0024117 | Chronic Obstructive Airway Disease | 1 | CTD_human | |
| Tgene | C0024143 | Lupus Nephritis | 1 | CTD_human | |
| Tgene | C0024668 | Mammary Neoplasms, Experimental | 1 | CTD_human | |
| Tgene | C0024796 | Marfan Syndrome | 1 | CTD_human | |
| Tgene | C0026552 | Morphine Dependence | 1 | CTD_human | |
| Tgene | C0026766 | Multiple Organ Failure | 1 | CTD_human | |
| Tgene | C0028754 | Obesity | 1 | CTD_human | |
| Tgene | C0029172 | Oral Submucous Fibrosis | 1 | CTD_human | |
| Tgene | C0030297 | Pancreatic Neoplasm | 1 | CTD_human | |
| Tgene | C0033578 | Prostatic Neoplasms | 1 | CTD_human | |
| Tgene | C0034067 | Pulmonary Emphysema | 1 | CTD_human | |
| Tgene | C0034069 | Pulmonary Fibrosis | 1 | CTD_human | |
| Tgene | C0034189 | Pyemia | 1 | CTD_human | |
| Tgene | C0035126 | Reperfusion Injury | 1 | CTD_human | |
| Tgene | C0035309 | Retinal Diseases | 1 | CTD_human | |
| Tgene | C0036690 | Septicemia | 1 | CTD_human | |
| Tgene | C0037286 | Skin Neoplasms | 1 | CTD_human | |
| Tgene | C0038358 | Gastric ulcer | 1 | CTD_human | |
| Tgene | C0038587 | Substance Withdrawal Syndrome | 1 | CTD_human | |
| Tgene | C0086189 | Drug Withdrawal Symptoms | 1 | CTD_human | |
| Tgene | C0086565 | Liver Dysfunction | 1 | CTD_human | |
| Tgene | C0087169 | Withdrawal Symptoms | 1 | CTD_human | |
| Tgene | C0151526 | Premature Birth | 1 | CTD_human | |
| Tgene | C0162871 | Aortic Aneurysm, Abdominal | 1 | CTD_human | |
| Tgene | C0162872 | Aortic Aneurysm, Thoracic | 1 | CTD_human | |
| Tgene | C0205768 | Subependymal Giant Cell Astrocytoma | 1 | CTD_human | |
| Tgene | C0221227 | Centriacinar Emphysema | 1 | CTD_human | |
| Tgene | C0238281 | Middle Cerebral Artery Syndrome | 1 | CTD_human | |
| Tgene | C0243026 | Sepsis | 1 | CTD_human | |
| Tgene | C0264393 | Panacinar Emphysema | 1 | CTD_human | |
| Tgene | C0271673 | Symmetric Diabetic Proximal Motor Neuropathy | 1 | CTD_human | |
| Tgene | C0271674 | Asymmetric Diabetic Proximal Motor Neuropathy | 1 | CTD_human | |
| Tgene | C0271678 | Diabetic Mononeuropathy | 1 | CTD_human | |
| Tgene | C0271680 | Diabetic Polyneuropathies | 1 | CTD_human | |
| Tgene | C0271685 | Diabetic Amyotrophy | 1 | CTD_human | |
| Tgene | C0271686 | Diabetic Autonomic Neuropathy | 1 | CTD_human | |
| Tgene | C0279530 | Malignant Bone Neoplasm | 1 | CTD_human | |
| Tgene | C0280783 | Juvenile Pilocytic Astrocytoma | 1 | CTD_human | |
| Tgene | C0280785 | Diffuse Astrocytoma | 1 | CTD_human | |
| Tgene | C0334579 | Anaplastic astrocytoma | 1 | CTD_human | |
| Tgene | C0334580 | Protoplasmic astrocytoma | 1 | CTD_human | |
| Tgene | C0334581 | Gemistocytic astrocytoma | 1 | CTD_human | |
| Tgene | C0334582 | Fibrillary Astrocytoma | 1 | CTD_human | |
| Tgene | C0334583 | Pilocytic Astrocytoma | 1 | CTD_human | |
| Tgene | C0334588 | Giant Cell Glioblastoma | 1 | CTD_human | |
| Tgene | C0338070 | Childhood Cerebral Astrocytoma | 1 | CTD_human | |
| Tgene | C0340288 | Stable angina | 1 | CTD_human | |
| Tgene | C0340630 | Aortic Aneurysm, Thoracoabdominal | 1 | CTD_human | |
| Tgene | C0346647 | Malignant neoplasm of pancreas | 1 | CTD_human | |
| Tgene | C0376358 | Malignant neoplasm of prostate | 1 | CTD_human | |
| Tgene | C0393835 | Diabetic Asymmetric Polyneuropathy | 1 | CTD_human | |
| Tgene | C0432226 | Metaphyseal anadysplasia | 1 | ORPHANET | |
| Tgene | C0432291 | Mandibuloacral dysostosis | 1 | CTD_human | |
| Tgene | C0472387 | Vasogenic Cerebral Edema | 1 | CTD_human | |
| Tgene | C0472388 | Cytotoxic Cerebral Edema | 1 | CTD_human | |
| Tgene | C0547065 | Mixed oligoastrocytoma | 1 | CTD_human | |
| Tgene | C0600272 | Morphine Abuse | 1 | CTD_human | |
| Tgene | C0678222 | Breast Carcinoma | 1 | CTD_human | |
| Tgene | C0740376 | Middle Cerebral Artery Thrombosis | 1 | CTD_human | |
| Tgene | C0740391 | Middle Cerebral Artery Occlusion | 1 | CTD_human | |
| Tgene | C0740392 | Infarction, Middle Cerebral Artery | 1 | CTD_human | |
| Tgene | C0741160 | Aortic Aneurysm, Ruptured | 1 | CTD_human | |
| Tgene | C0750935 | Cerebral Astrocytoma | 1 | CTD_human | |
| Tgene | C0750936 | Intracranial Astrocytoma | 1 | CTD_human | |
| Tgene | C0750969 | Vasogenic Brain Edema | 1 | CTD_human | |
| Tgene | C0750970 | Cytotoxic Brain Edema | 1 | CTD_human | |
| Tgene | C0751074 | Diabetic Neuralgia | 1 | CTD_human | |
| Tgene | C0751845 | Middle Cerebral Artery Embolus | 1 | CTD_human | |
| Tgene | C0751846 | Left Middle Cerebral Artery Infarction | 1 | CTD_human | |
| Tgene | C0751847 | Embolic Infarction, Middle Cerebral Artery | 1 | CTD_human | |
| Tgene | C0751848 | Thrombotic Infarction, Middle Cerebral Artery | 1 | CTD_human | |
| Tgene | C0751849 | Right Middle Cerebral Artery Infarction | 1 | CTD_human | |
| Tgene | C0860207 | Drug-Induced Liver Disease | 1 | CTD_human | |
| Tgene | C0917798 | Cerebral Ischemia | 1 | CTD_human | |
| Tgene | C0948089 | Acute Coronary Syndrome | 1 | CTD_human | |
| Tgene | C1257931 | Mammary Neoplasms, Human | 1 | CTD_human | |
| Tgene | C1262760 | Hepatitis, Drug-Induced | 1 | CTD_human | |
| Tgene | C1458155 | Mammary Neoplasms | 1 | CTD_human | |
| Tgene | C1527303 | Chronic Airflow Obstruction | 1 | CTD_human | |
| Tgene | C1527311 | Brain Edema | 1 | CTD_human | |
| Tgene | C1621958 | Glioblastoma Multiforme | 1 | CTD_human | |
| Tgene | C1704230 | Grade I Astrocytoma | 1 | CTD_human | |
| Tgene | C1719672 | Severe Sepsis | 1 | CTD_human | |
| Tgene | C2239176 | Liver carcinoma | 1 | CTD_human | |
| Tgene | C2350878 | Focal Emphysema | 1 | CTD_human | |
| Tgene | C2751322 | Metaphyseal Anadysplasia 2 | 1 | CTD_human;GENOMICS_ENGLAND | |
| Tgene | C2936380 | Neointima | 1 | CTD_human | |
| Tgene | C2936381 | Neointima Formation | 1 | CTD_human | |
| Tgene | C3658290 | Drug-Induced Acute Liver Injury | 1 | CTD_human | |
| Tgene | C4277682 | Chemical and Drug Induced Liver Injury | 1 | CTD_human | |
| Tgene | C4279912 | Chemically-Induced Liver Toxicity | 1 | CTD_human | |
| Tgene | C4704874 | Mammary Carcinoma, Human | 1 | CTD_human | |
| Tgene | C4721507 | Alveolitis, Fibrosing | 1 | CTD_human | |
| Tgene | C4721845 | Marfan Syndrome, Type I | 1 | CTD_human |
