
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:DNMT3A-KIF3C (FusionGDB2 ID:HG1788TG3797) |
Fusion Gene Summary for DNMT3A-KIF3C |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: DNMT3A-KIF3C | Fusion gene ID: hg1788tg3797 | Hgene | Tgene | Gene symbol | DNMT3A | KIF3C | Gene ID | 1788 | 3797 |
| Gene name | DNA methyltransferase 3 alpha | kinesin family member 3C | |
| Synonyms | DNMT3A2|HESJAS|M.HsaIIIA|TBRS | - | |
| Cytomap | ('DNMT3A')('KIF3C') 2p23.3 | 2p23.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | DNA (cytosine-5)-methyltransferase 3ADNA (cytosine-5-)-methyltransferase 3 alphaDNA MTase HsaIIIADNA cytosine methyltransferase 3A2 | kinesin-like protein KIF3CKIF3C variant protein | |
| Modification date | 20200322 | 20200313 | |
| UniProtAcc | Q9Y6K1 | O14782 | |
| Ensembl transtripts involved in fusion gene | ENST00000264709, ENST00000321117, ENST00000406659, ENST00000380746, ENST00000402667, ENST00000474887, | ||
| Fusion gene scores | * DoF score | 8 X 10 X 7=560 | 5 X 5 X 3=75 |
| # samples | 9 | 5 | |
| ** MAII score | log2(9/560*10)=-2.63742992061529 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(5/75*10)=-0.584962500721156 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: DNMT3A [Title/Abstract] AND KIF3C [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | DNMT3A(25536781)-KIF3C(26152932), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | DNMT3A-KIF3C seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. DNMT3A-KIF3C seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. DNMT3A-KIF3C seems lost the major protein functional domain in Hgene partner, which is a cell metabolism gene due to the frame-shifted ORF. DNMT3A-KIF3C seems lost the major protein functional domain in Hgene partner, which is a CGC due to the frame-shifted ORF. DNMT3A-KIF3C seems lost the major protein functional domain in Hgene partner, which is a epigenetic factor due to the frame-shifted ORF. DNMT3A-KIF3C seems lost the major protein functional domain in Hgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. DNMT3A-KIF3C seems lost the major protein functional domain in Hgene partner, which is a tumor suppressor due to the frame-shifted ORF. DNMT3A-KIF3C seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | DNMT3A | GO:0006306 | DNA methylation | 12138111|19786833|23042785 |
 Fusion gene breakpoints across DNMT3A (5'-gene) Fusion gene breakpoints across DNMT3A (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
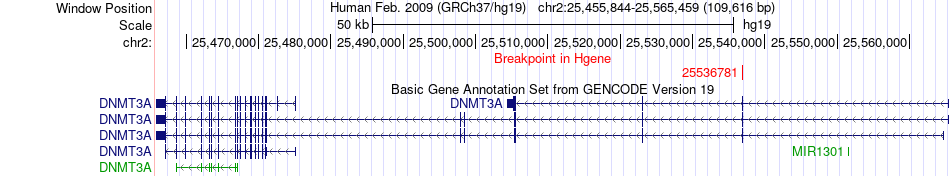 |
 Fusion gene breakpoints across KIF3C (3'-gene) Fusion gene breakpoints across KIF3C (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
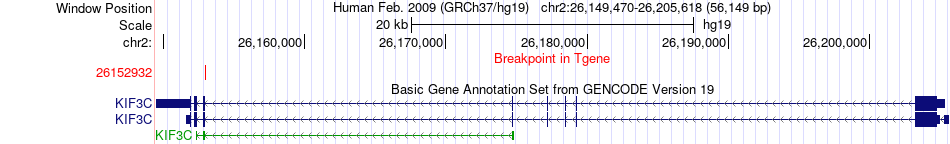 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | STAD | TCGA-VQ-A925 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - |
Top |
Fusion Gene ORF analysis for DNMT3A-KIF3C |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000264709 | ENST00000496378 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - |
| 5CDS-5UTR | ENST00000321117 | ENST00000496378 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - |
| 5CDS-5UTR | ENST00000406659 | ENST00000496378 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - |
| Frame-shift | ENST00000264709 | ENST00000264712 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - |
| Frame-shift | ENST00000264709 | ENST00000405914 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - |
| Frame-shift | ENST00000406659 | ENST00000264712 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - |
| Frame-shift | ENST00000406659 | ENST00000405914 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - |
| In-frame | ENST00000321117 | ENST00000264712 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - |
| In-frame | ENST00000321117 | ENST00000405914 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - |
| intron-3CDS | ENST00000380746 | ENST00000264712 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - |
| intron-3CDS | ENST00000380746 | ENST00000405914 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - |
| intron-3CDS | ENST00000402667 | ENST00000264712 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - |
| intron-3CDS | ENST00000402667 | ENST00000405914 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - |
| intron-3CDS | ENST00000474887 | ENST00000264712 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - |
| intron-3CDS | ENST00000474887 | ENST00000405914 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - |
| intron-5UTR | ENST00000380746 | ENST00000496378 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - |
| intron-5UTR | ENST00000402667 | ENST00000496378 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - |
| intron-5UTR | ENST00000474887 | ENST00000496378 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000321117 | DNMT3A | chr2 | 25536781 | - | ENST00000264712 | KIF3C | chr2 | 26152932 | - | 3061 | 309 | 280 | 684 | 134 |
| ENST00000321117 | DNMT3A | chr2 | 25536781 | - | ENST00000405914 | KIF3C | chr2 | 26152932 | - | 910 | 309 | 280 | 684 | 134 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000321117 | ENST00000264712 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - | 0.69533455 | 0.30466548 |
| ENST00000321117 | ENST00000405914 | DNMT3A | chr2 | 25536781 | - | KIF3C | chr2 | 26152932 | - | 0.22329125 | 0.7767087 |
Top |
Fusion Genomic Features for DNMT3A-KIF3C |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
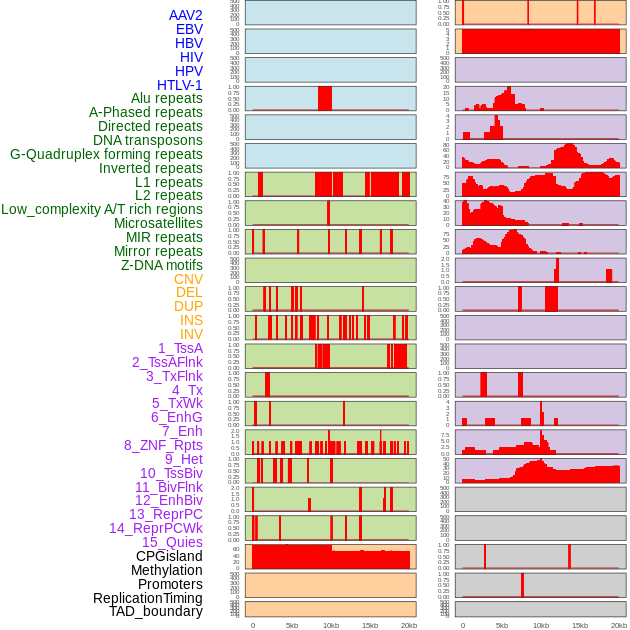 |
Top |
Fusion Protein Features for DNMT3A-KIF3C |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr2:25536781/chr2:26152932) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| DNMT3A | KIF3C |
| FUNCTION: Required for genome-wide de novo methylation and is essential for the establishment of DNA methylation patterns during development. DNA methylation is coordinated with methylation of histones. It modifies DNA in a non-processive manner and also methylates non-CpG sites. May preferentially methylate DNA linker between 2 nucleosomal cores and is inhibited by histone H1. Plays a role in paternal and maternal imprinting. Required for methylation of most imprinted loci in germ cells. Acts as a transcriptional corepressor for ZBTB18. Recruited to trimethylated 'Lys-36' of histone H3 (H3K36me3) sites. Can actively repress transcription through the recruitment of HDAC activity. {ECO:0000269|PubMed:16357870, ECO:0000269|PubMed:30478443}. | FUNCTION: Microtubule-based anterograde translocator for membranous organelles. {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000264709 | - | 2 | 23 | 292_350 | 24 | 913.0 | Domain | PWWP |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000264709 | - | 2 | 23 | 482_614 | 24 | 913.0 | Domain | ADD |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000264709 | - | 2 | 23 | 634_912 | 24 | 913.0 | Domain | SAM-dependent MTase C5-type |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000321117 | - | 2 | 23 | 292_350 | 24 | 913.0 | Domain | PWWP |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000321117 | - | 2 | 23 | 482_614 | 24 | 913.0 | Domain | ADD |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000321117 | - | 2 | 23 | 634_912 | 24 | 913.0 | Domain | SAM-dependent MTase C5-type |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000380746 | - | 1 | 19 | 292_350 | 0 | 724.0 | Domain | PWWP |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000380746 | - | 1 | 19 | 482_614 | 0 | 724.0 | Domain | ADD |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000380746 | - | 1 | 19 | 634_912 | 0 | 724.0 | Domain | SAM-dependent MTase C5-type |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000402667 | - | 1 | 18 | 292_350 | 0 | 690.0 | Domain | PWWP |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000402667 | - | 1 | 18 | 482_614 | 0 | 690.0 | Domain | ADD |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000402667 | - | 1 | 18 | 634_912 | 0 | 690.0 | Domain | SAM-dependent MTase C5-type |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000406659 | - | 2 | 4 | 292_350 | 24 | 167.0 | Domain | PWWP |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000406659 | - | 2 | 4 | 482_614 | 24 | 167.0 | Domain | ADD |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000406659 | - | 2 | 4 | 634_912 | 24 | 167.0 | Domain | SAM-dependent MTase C5-type |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000264709 | - | 2 | 23 | 641_645 | 24 | 913.0 | Region | S-adenosyl-L-methionine binding |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000264709 | - | 2 | 23 | 686_688 | 24 | 913.0 | Region | S-adenosyl-L-methionine binding |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000264709 | - | 2 | 23 | 891_893 | 24 | 913.0 | Region | S-adenosyl-L-methionine binding |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000321117 | - | 2 | 23 | 641_645 | 24 | 913.0 | Region | S-adenosyl-L-methionine binding |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000321117 | - | 2 | 23 | 686_688 | 24 | 913.0 | Region | S-adenosyl-L-methionine binding |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000321117 | - | 2 | 23 | 891_893 | 24 | 913.0 | Region | S-adenosyl-L-methionine binding |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000380746 | - | 1 | 19 | 641_645 | 0 | 724.0 | Region | S-adenosyl-L-methionine binding |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000380746 | - | 1 | 19 | 686_688 | 0 | 724.0 | Region | S-adenosyl-L-methionine binding |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000380746 | - | 1 | 19 | 891_893 | 0 | 724.0 | Region | S-adenosyl-L-methionine binding |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000402667 | - | 1 | 18 | 641_645 | 0 | 690.0 | Region | S-adenosyl-L-methionine binding |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000402667 | - | 1 | 18 | 686_688 | 0 | 690.0 | Region | S-adenosyl-L-methionine binding |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000402667 | - | 1 | 18 | 891_893 | 0 | 690.0 | Region | S-adenosyl-L-methionine binding |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000406659 | - | 2 | 4 | 641_645 | 24 | 167.0 | Region | S-adenosyl-L-methionine binding |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000406659 | - | 2 | 4 | 686_688 | 24 | 167.0 | Region | S-adenosyl-L-methionine binding |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000406659 | - | 2 | 4 | 891_893 | 24 | 167.0 | Region | S-adenosyl-L-methionine binding |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000264709 | - | 2 | 23 | 493_523 | 24 | 913.0 | Zinc finger | GATA-type%3B atypical |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000264709 | - | 2 | 23 | 534_590 | 24 | 913.0 | Zinc finger | PHD-type%3B atypical |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000321117 | - | 2 | 23 | 493_523 | 24 | 913.0 | Zinc finger | GATA-type%3B atypical |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000321117 | - | 2 | 23 | 534_590 | 24 | 913.0 | Zinc finger | PHD-type%3B atypical |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000380746 | - | 1 | 19 | 493_523 | 0 | 724.0 | Zinc finger | GATA-type%3B atypical |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000380746 | - | 1 | 19 | 534_590 | 0 | 724.0 | Zinc finger | PHD-type%3B atypical |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000402667 | - | 1 | 18 | 493_523 | 0 | 690.0 | Zinc finger | GATA-type%3B atypical |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000402667 | - | 1 | 18 | 534_590 | 0 | 690.0 | Zinc finger | PHD-type%3B atypical |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000406659 | - | 2 | 4 | 493_523 | 24 | 167.0 | Zinc finger | GATA-type%3B atypical |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000406659 | - | 2 | 4 | 534_590 | 24 | 167.0 | Zinc finger | PHD-type%3B atypical |
| Tgene | KIF3C | chr2:25536781 | chr2:26152932 | ENST00000264712 | 4 | 8 | 376_629 | 668 | 794.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | KIF3C | chr2:25536781 | chr2:26152932 | ENST00000405914 | 5 | 9 | 376_629 | 668 | 794.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | KIF3C | chr2:25536781 | chr2:26152932 | ENST00000264712 | 4 | 8 | 270_284 | 668 | 794.0 | Compositional bias | Note=Poly-Gly | |
| Tgene | KIF3C | chr2:25536781 | chr2:26152932 | ENST00000264712 | 4 | 8 | 438_441 | 668 | 794.0 | Compositional bias | Note=Poly-Asn | |
| Tgene | KIF3C | chr2:25536781 | chr2:26152932 | ENST00000405914 | 5 | 9 | 270_284 | 668 | 794.0 | Compositional bias | Note=Poly-Gly | |
| Tgene | KIF3C | chr2:25536781 | chr2:26152932 | ENST00000405914 | 5 | 9 | 438_441 | 668 | 794.0 | Compositional bias | Note=Poly-Asn | |
| Tgene | KIF3C | chr2:25536781 | chr2:26152932 | ENST00000264712 | 4 | 8 | 10_365 | 668 | 794.0 | Domain | Kinesin motor | |
| Tgene | KIF3C | chr2:25536781 | chr2:26152932 | ENST00000405914 | 5 | 9 | 10_365 | 668 | 794.0 | Domain | Kinesin motor | |
| Tgene | KIF3C | chr2:25536781 | chr2:26152932 | ENST00000264712 | 4 | 8 | 97_104 | 668 | 794.0 | Nucleotide binding | Note=ATP | |
| Tgene | KIF3C | chr2:25536781 | chr2:26152932 | ENST00000405914 | 5 | 9 | 97_104 | 668 | 794.0 | Nucleotide binding | Note=ATP | |
| Tgene | KIF3C | chr2:25536781 | chr2:26152932 | ENST00000264712 | 4 | 8 | 630_793 | 668 | 794.0 | Region | Globular | |
| Tgene | KIF3C | chr2:25536781 | chr2:26152932 | ENST00000405914 | 5 | 9 | 630_793 | 668 | 794.0 | Region | Globular |
Top |
Fusion Gene Sequence for DNMT3A-KIF3C |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >23711_23711_1_DNMT3A-KIF3C_DNMT3A_chr2_25536781_ENST00000321117_KIF3C_chr2_26152932_ENST00000264712_length(transcript)=3061nt_BP=309nt GACGCGGCGCCGCGGCACCAGGGCGCGCAGCCGGGCCGGCCCGACCCCACCGGCCATACGGTGGAGCCATCGAAGCCCCCACCCACAGGC TGACAGAGGCACCGTTCACCAGAGGGCTCAACACCGGGATCTATGTTTAAGTTTTAACTCTCGCCTCCAAAGACCACGATAATTCCTTCC CCAAAGCCCAGCAGCCCCCCAGCCCCGCGCAGCCCCAGCCTGCCTCCCGGCGCCCAGATGCCCGCCATGCCCTCCAGCGGCCCCGGGGAC ACCAGCAGCTCTGCTGCGGAGCGGGAGGAGGACCGAAAGCAGTAGCAGCCAGATGAAGAAGCGGCCAACATCTGCAGTGGGCTACAAGAG GCCTATCAGCCAGTATGCTCGGGTTGCCATGGCAATGGGGTCCCACCCCAGGTACAGGGCTGAAAACATAATGTTTCTGGAGTTGGATGT GTCCCCTCCAGCTGTCTTTGAGATGGAATTCTCTCACGACCAAGAACAAGACCCTCGTGCGCTACACATGGAGAGGCTCATGCGATTGGA CAGCTTTCTGGAAAGACCTTCCACGTCTAAAGTCCGAAAGTCCAGATCCTGGTGCCAGAGTCCTCAGCGGCCTCCACCTTCCACCACACA TGCCTCCCTGGCCTCTGCTTCTCTGCGCCCTGCAACAGTGGCGGACCATGAGTGACAACCATCACGTCAGGCTGCCCATCCAATAGACTC CTGGGATGGGGCAGCCAACCCTGGCTCATCTCATCTGCCGCTTGGTGCGTGTGCGTGTGCGTGCATGTGCGTGTGCGTGTGTGCAGGGGT GAGAATCTGGCAGATGGTGCCTCTGCCTGCTCTTCTTCGCCTCCTTTATTTAATTCATGTTATTTATTCGCGGAGCTCTGTTCGTGTTGG GGAGATGCCCTCGCCTGAGCCGTCTGGGCCTACCGTGGTCACTGCGTAGCCTCTTTTTCTTCTGACTTGAGAGCTCCCCCAGTCAGATCT CAGGCTTGTCCCCCTGTCAGCTGCCTCCAGAAGGGAAGGTAGCCAGTGCCTGAGAAGACAGTCCCTTTTCTACCCACCGCACTCCATAAC CTCCATCTTCTCCCACACTGATGGCGAGCAGCCCCTGAGCACTTTCTGGGACTGGGAGACTGCTTGGTGTTCCCTGAGGACAAGAGACAT CCTGACAGTGTTGGGCATCTGCTCCCCGTGGACACAGCCCCACTCTCCACTTTCTGAGCCTCAGACAACCTCATTCAGCCTCTTGGGCTC CTTTTCAAGGACATTAATAACCTCACCAACATAGCTCATGCCCTTCAGCTTTGACAAGAACTCACAGCTTCCCAAACTCTGCTTTCTGCC CACCTTGGATGGGAACTGTGGACCAAGCAATTACCATCGCCTTGGAACCTGCAGGAAATGGAACAGCAATTGAGACAACTTGAACAGTCA TCAACGGAAGTCCCTCCACTGGATTCCTTTGTTTCTGTCCCCTCCGAGGAGTCATTTTGGTCGACAGGCTCTCAAGGCAACTCCCCATTT TCAAGAGGCTGCTCCTGCCTGCTTCGATCATTTCTCCCTGCAGCTGCCTAGACCCCGTTCACAGTGGGAGGAGTCAATGTCATTCTACCC CTCGCTAAACGAAGATATTAACATCTATTGCTTTTTCCCTTCATCTGTCACAGGAAACAGAAGCCCAGGCACAATCTTTTCCAGCTTTGC CTGTTACCCCTGTTTCTGAATTGCATCTTTAAGGTATTATTTTGTTGAAAATAGATCCTTTATTCACTAGTTACGCAAATTGGTTCCTAG GGGGATACTCCTTACCTTCCTTTGTGATGGCCCAAAATGTCTCTAGGTATCTCAAGTGATAAGTAAATTTCTACAAAAAAAAATGGTTAA TGTTCATTGACTGGCTTTTTAAGTGTATATTTTGGAGGACGGGTGAAGAGGTCATAACGAAAGCAAGCGAGTGAATTAGGATTTCAAAGT GCCCTAATAGTGTGAGTCTCCAGTTCCTAGAATATGAAGAGTGCTGTCGTTGGGGTGAAACCATGAGACTGACAGATCTGCCTGAAATGG GGGGTGTGGGAGGTGGTGGCGGGGGTTATTCTCTTTCCTTCAGGAAATGAACCCTTCTTACATCATTCAAGTTCTGCTCTGAGGATCAAG CTTGGGTCTGATTTAACTCAGCGACACTGTCATTTCTGCTTCATTACTGGACTAGAGGGTTGAGCCACCCACTTGCCATTTGCTCCTGTC CTTCCAGGAAATCACAATTTTCATCAGAGCCCAAGAGATTATTTGAGACTCAGGATTCAGATCAGAGGTTCGACTGTGGCTGGGACAGGA GTTGTGTGTAGAAATTCACCAGGTGGCCTGAGCGCAGGGGGACCTCCAGGGCTGCGTTGAGCAGCCTCTCCCACTGACCTCTTTCTCGTT TGTGGACAAAGCAGCACGTATCACCTCATTCATCACTTGGACACATCGCCTTTGCATTGTCTTGTCACACCTCCCTCACAGTCTTATAGC ACAATATACCCAAATCAGCCCCCCCAGTCCGAGGGCTGGGCCCGAGGTATGGTCGGAGGAGGAGCTCCTGCCTGCGGTTTTGTGTATGTG TGTATGTGTGTGCGTGTTTGTGTGCGTGTTTACCTCCACAGGGGACACTCTACACTCAGTGTAAGATCTGCTGGGAACAGGGCCACCAGG AGTGGCTGGATCTCAGTCTCTCTGTCTCTCTTTCTCTCCTTTTCCTTTTGGTGTATCAAATATTTGATTGACAAAGTAAGGGCCTTGATT AGGACCAAATTCTCGTGTGTTGCTATGGTCTTTATTTAGGACAACAATTAACAATGCAGTGGCCCATTCTTGTCACTCTACACATATGAC TATACGGGACATATGTAATATATAAATATATATATAAAACATTCCCCTCTGTCCCCTTGGCTTCGGATGGAGGCCTTTCTGTTGAGCTGA AATGCACCTGCAGCTGGGTGCTGCCAGCAGCTTGCAGGCCCCAGCCCTGTTCCAATCAATGCAGTTGACAATAAAGGAATGAGTATCGTC >23711_23711_1_DNMT3A-KIF3C_DNMT3A_chr2_25536781_ENST00000321117_KIF3C_chr2_26152932_ENST00000264712_length(amino acids)=134AA_BP=9 MLRSGRRTESSSSQMKKRPTSAVGYKRPISQYARVAMAMGSHPRYRAENIMFLELDVSPPAVFEMEFSHDQEQDPRALHMERLMRLDSFL -------------------------------------------------------------- >23711_23711_2_DNMT3A-KIF3C_DNMT3A_chr2_25536781_ENST00000321117_KIF3C_chr2_26152932_ENST00000405914_length(transcript)=910nt_BP=309nt GACGCGGCGCCGCGGCACCAGGGCGCGCAGCCGGGCCGGCCCGACCCCACCGGCCATACGGTGGAGCCATCGAAGCCCCCACCCACAGGC TGACAGAGGCACCGTTCACCAGAGGGCTCAACACCGGGATCTATGTTTAAGTTTTAACTCTCGCCTCCAAAGACCACGATAATTCCTTCC CCAAAGCCCAGCAGCCCCCCAGCCCCGCGCAGCCCCAGCCTGCCTCCCGGCGCCCAGATGCCCGCCATGCCCTCCAGCGGCCCCGGGGAC ACCAGCAGCTCTGCTGCGGAGCGGGAGGAGGACCGAAAGCAGTAGCAGCCAGATGAAGAAGCGGCCAACATCTGCAGTGGGCTACAAGAG GCCTATCAGCCAGTATGCTCGGGTTGCCATGGCAATGGGGTCCCACCCCAGGTACAGGGCTGAAAACATAATGTTTCTGGAGTTGGATGT GTCCCCTCCAGCTGTCTTTGAGATGGAATTCTCTCACGACCAAGAACAAGACCCTCGTGCGCTACACATGGAGAGGCTCATGCGATTGGA CAGCTTTCTGGAAAGACCTTCCACGTCTAAAGTCCGAAAGTCCAGATCCTGGTGCCAGAGTCCTCAGCGGCCTCCACCTTCCACCACACA TGCCTCCCTGGCCTCTGCTTCTCTGCGCCCTGCAACAGTGGCGGACCATGAGTGACAACCATCACGTCAGGCTGCCCATCCAATAGACTC CTGGGATGGGGCAGCCAACCCTGGCTCATCTCATCTGCCGCTTGGTGCGTGTGCGTGTGCGTGCATGTGCGTGTGCGTGTGTGCAGGGGT GAGAATCTGGCAGATGGTGCCTCTGCCTGCTCTTCTTCGCCTCCTTTATTTAATTCATGTTATTTATTCGCGGAGCTCTGTTCGTGTTGG >23711_23711_2_DNMT3A-KIF3C_DNMT3A_chr2_25536781_ENST00000321117_KIF3C_chr2_26152932_ENST00000405914_length(amino acids)=134AA_BP=9 MLRSGRRTESSSSQMKKRPTSAVGYKRPISQYARVAMAMGSHPRYRAENIMFLELDVSPPAVFEMEFSHDQEQDPRALHMERLMRLDSFL -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for DNMT3A-KIF3C |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000264709 | - | 2 | 23 | 199_403 | 24.0 | 913.0 | DNMT1 and DNMT3B |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000321117 | - | 2 | 23 | 199_403 | 24.0 | 913.0 | DNMT1 and DNMT3B |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000380746 | - | 1 | 19 | 199_403 | 0 | 724.0 | DNMT1 and DNMT3B |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000402667 | - | 1 | 18 | 199_403 | 0 | 690.0 | DNMT1 and DNMT3B |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000406659 | - | 2 | 4 | 199_403 | 24.0 | 167.0 | DNMT1 and DNMT3B |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000264709 | - | 2 | 23 | 494_586 | 24.0 | 913.0 | the PRC2/EED-EZH2 complex |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000321117 | - | 2 | 23 | 494_586 | 24.0 | 913.0 | the PRC2/EED-EZH2 complex |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000380746 | - | 1 | 19 | 494_586 | 0 | 724.0 | the PRC2/EED-EZH2 complex |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000402667 | - | 1 | 18 | 494_586 | 0 | 690.0 | the PRC2/EED-EZH2 complex |
| Hgene | DNMT3A | chr2:25536781 | chr2:26152932 | ENST00000406659 | - | 2 | 4 | 494_586 | 24.0 | 167.0 | the PRC2/EED-EZH2 complex |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for DNMT3A-KIF3C |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
| Hgene | DNMT3A | Q9Y6K1 | DB01262 | Decitabine | Inhibitor | Small molecule | Approved|Investigational |
| Hgene | DNMT3A | Q9Y6K1 | DB00721 | Procaine | Inhibitor | Small molecule | Approved|Investigational|Vet_approved |
Top |
Related Diseases for DNMT3A-KIF3C |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
| Hgene | DNMT3A | C4014545 | Tatton Brown Rahman syndrome | 4 | CTD_human;GENOMICS_ENGLAND;ORPHANET;UNIPROT |
| Hgene | DNMT3A | C0018273 | Growth Disorders | 2 | CTD_human |
| Hgene | DNMT3A | C0079774 | Peripheral T-Cell Lymphoma | 2 | CTD_human |
| Hgene | DNMT3A | C0006142 | Malignant neoplasm of breast | 1 | CTD_human |
| Hgene | DNMT3A | C0010346 | Crohn Disease | 1 | CTD_human |
| Hgene | DNMT3A | C0013336 | Dwarfism | 1 | CTD_human |
| Hgene | DNMT3A | C0020796 | Profound Mental Retardation | 1 | CTD_human |
| Hgene | DNMT3A | C0020981 | Angioimmunoblastic Lymphadenopathy | 1 | CTD_human |
| Hgene | DNMT3A | C0023465 | Acute monocytic leukemia | 1 | CTD_human |
| Hgene | DNMT3A | C0023487 | Acute Promyelocytic Leukemia | 1 | CTD_human |
| Hgene | DNMT3A | C0024121 | Lung Neoplasms | 1 | CTD_human |
| Hgene | DNMT3A | C0025363 | Mental Retardation, Psychosocial | 1 | CTD_human |
| Hgene | DNMT3A | C0025958 | Microcephaly | 1 | CTD_human |
| Hgene | DNMT3A | C0027643 | Neoplasm Recurrence, Local | 1 | CTD_human |
| Hgene | DNMT3A | C0036920 | Sezary Syndrome | 1 | CTD_human |
| Hgene | DNMT3A | C0079773 | Lymphoma, T-Cell, Cutaneous | 1 | CTD_human |
| Hgene | DNMT3A | C0156147 | Crohn's disease of large bowel | 1 | CTD_human |
| Hgene | DNMT3A | C0242379 | Malignant neoplasm of lung | 1 | CTD_human |
| Hgene | DNMT3A | C0265202 | Seckel syndrome | 1 | GENOMICS_ENGLAND |
| Hgene | DNMT3A | C0267380 | Crohn's disease of the ileum | 1 | CTD_human |
| Hgene | DNMT3A | C0282631 | Facies | 1 | CTD_human |
| Hgene | DNMT3A | C0349639 | Juvenile Myelomonocytic Leukemia | 1 | CTD_human |
| Hgene | DNMT3A | C0376407 | Granulomatous Slack Skin | 1 | CTD_human |
| Hgene | DNMT3A | C0376634 | Craniofacial Abnormalities | 1 | CTD_human |
| Hgene | DNMT3A | C0678202 | Regional enteritis | 1 | CTD_human |
| Hgene | DNMT3A | C0678222 | Breast Carcinoma | 1 | CTD_human |
| Hgene | DNMT3A | C0917816 | Mental deficiency | 1 | CTD_human |
| Hgene | DNMT3A | C0949272 | IIeocolitis | 1 | CTD_human |
| Hgene | DNMT3A | C1257931 | Mammary Neoplasms, Human | 1 | CTD_human |
| Hgene | DNMT3A | C1458155 | Mammary Neoplasms | 1 | CTD_human |
| Hgene | DNMT3A | C1510586 | Autism Spectrum Disorders | 1 | CTD_human |
| Hgene | DNMT3A | C1956147 | Microlissencephaly | 1 | CTD_human |
| Hgene | DNMT3A | C2931852 | Clear-cell metastatic renal cell carcinoma | 1 | CTD_human |
| Hgene | DNMT3A | C3496069 | cocaine use | 1 | PSYGENET |
| Hgene | DNMT3A | C3714756 | Intellectual Disability | 1 | CTD_human |
| Hgene | DNMT3A | C3853041 | Severe Congenital Microcephaly | 1 | CTD_human |
| Hgene | DNMT3A | C4704874 | Mammary Carcinoma, Human | 1 | CTD_human |
| Tgene | C0004238 | Atrial Fibrillation | 2 | CTD_human | |
| Tgene | C0235480 | Paroxysmal atrial fibrillation | 2 | CTD_human | |
| Tgene | C2585653 | Persistent atrial fibrillation | 2 | CTD_human | |
| Tgene | C3468561 | familial atrial fibrillation | 2 | CTD_human | |
| Tgene | C1862939 | AMYOTROPHIC LATERAL SCLEROSIS 1 | 1 | CTD_human | |
| Tgene | C1862941 | Amyotrophic Lateral Sclerosis, Sporadic | 1 | CTD_human | |
| Tgene | C4551993 | Amyotrophic Lateral Sclerosis, Familial | 1 | CTD_human |
