
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:EFR3B-NCOA1 (FusionGDB2 ID:HG22979TG8648) |
Fusion Gene Summary for EFR3B-NCOA1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: EFR3B-NCOA1 | Fusion gene ID: hg22979tg8648 | Hgene | Tgene | Gene symbol | EFR3B | NCOA1 | Gene ID | 22979 | 8648 |
| Gene name | EFR3 homolog B | nuclear receptor coactivator 1 | |
| Synonyms | KIAA0953 | F-SRC-1|KAT13A|RIP160|SRC1|bHLHe42|bHLHe74 | |
| Cytomap | ('EFR3B')('NCOA1') 2p23.3 | 2p23.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | protein EFR3 homolog B | nuclear receptor coactivator 1Hin-2 proteinclass E basic helix-loop-helix protein 74renal carcinoma antigen NY-REN-52steroid receptor coactivator-1 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | Q15788 | |
| Ensembl transtripts involved in fusion gene | ENST00000401432, ENST00000402191, ENST00000403714, ENST00000405108, | ||
| Fusion gene scores | * DoF score | 3 X 3 X 3=27 | 11 X 13 X 7=1001 |
| # samples | 3 | 13 | |
| ** MAII score | log2(3/27*10)=0.15200309344505 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | log2(13/1001*10)=-2.94485844580754 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: EFR3B [Title/Abstract] AND NCOA1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | EFR3B(25367920)-NCOA1(24962301), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | EFR3B-NCOA1 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. EFR3B-NCOA1 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. EFR3B-NCOA1 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. EFR3B-NCOA1 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. EFR3B-NCOA1 seems lost the major protein functional domain in Tgene partner, which is a cell metabolism gene due to the frame-shifted ORF. EFR3B-NCOA1 seems lost the major protein functional domain in Tgene partner, which is a CGC due to the frame-shifted ORF. EFR3B-NCOA1 seems lost the major protein functional domain in Tgene partner, which is a epigenetic factor due to the frame-shifted ORF. EFR3B-NCOA1 seems lost the major protein functional domain in Tgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. EFR3B-NCOA1 seems lost the major protein functional domain in Tgene partner, which is a transcription factor due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | EFR3B | GO:0046854 | phosphatidylinositol phosphorylation | 23229899 |
| Hgene | EFR3B | GO:0072659 | protein localization to plasma membrane | 23229899 |
| Tgene | NCOA1 | GO:0000435 | positive regulation of transcription from RNA polymerase II promoter by galactose | 10207113 |
| Tgene | NCOA1 | GO:0006351 | transcription, DNA-templated | 9223431 |
| Tgene | NCOA1 | GO:0045893 | positive regulation of transcription, DNA-templated | 11891224|15367689 |
| Tgene | NCOA1 | GO:0045944 | positive regulation of transcription by RNA polymerase II | 15919756 |
 Fusion gene breakpoints across EFR3B (5'-gene) Fusion gene breakpoints across EFR3B (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
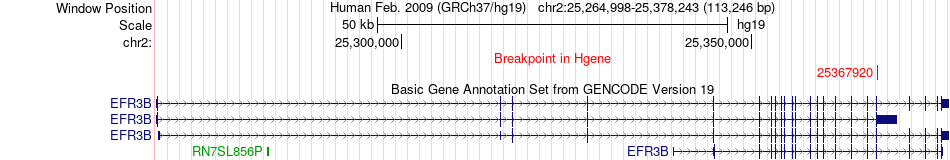 |
 Fusion gene breakpoints across NCOA1 (3'-gene) Fusion gene breakpoints across NCOA1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
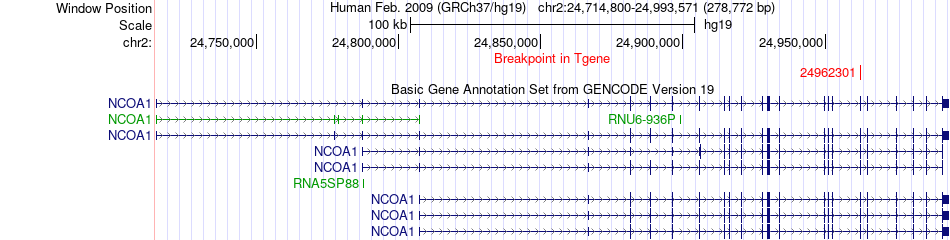 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-AC-A8OQ-01A | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
Top |
Fusion Gene ORF analysis for EFR3B-NCOA1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000401432 | ENST00000288599 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000401432 | ENST00000348332 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000401432 | ENST00000395856 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000401432 | ENST00000405141 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000401432 | ENST00000407230 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000401432 | ENST00000469850 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000401432 | ENST00000538539 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000402191 | ENST00000288599 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000402191 | ENST00000348332 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000402191 | ENST00000395856 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000402191 | ENST00000405141 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000402191 | ENST00000407230 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000402191 | ENST00000469850 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000402191 | ENST00000538539 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000403714 | ENST00000288599 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000403714 | ENST00000348332 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000403714 | ENST00000395856 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000403714 | ENST00000405141 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000403714 | ENST00000407230 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000403714 | ENST00000469850 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000403714 | ENST00000538539 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000405108 | ENST00000288599 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000405108 | ENST00000348332 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000405108 | ENST00000395856 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000405108 | ENST00000405141 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000405108 | ENST00000407230 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000405108 | ENST00000469850 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| 5CDS-intron | ENST00000405108 | ENST00000538539 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| Frame-shift | ENST00000401432 | ENST00000406961 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| In-frame | ENST00000402191 | ENST00000406961 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| In-frame | ENST00000403714 | ENST00000406961 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
| In-frame | ENST00000405108 | ENST00000406961 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000403714 | EFR3B | chr2 | 25367920 | + | ENST00000406961 | NCOA1 | chr2 | 24962301 | + | 5761 | 2325 | 45 | 3449 | 1134 |
| ENST00000402191 | EFR3B | chr2 | 25367920 | + | ENST00000406961 | NCOA1 | chr2 | 24962301 | + | 5775 | 2339 | 17 | 3463 | 1148 |
| ENST00000405108 | EFR3B | chr2 | 25367920 | + | ENST00000406961 | NCOA1 | chr2 | 24962301 | + | 5331 | 1895 | 197 | 3019 | 940 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000403714 | ENST00000406961 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + | 0.001448583 | 0.9985514 |
| ENST00000402191 | ENST00000406961 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + | 0.001565698 | 0.99843425 |
| ENST00000405108 | ENST00000406961 | EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962301 | + | 0.001898578 | 0.9981014 |
Top |
Fusion Genomic Features for EFR3B-NCOA1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962300 | + | 1.97E-05 | 0.99998033 |
| EFR3B | chr2 | 25367920 | + | NCOA1 | chr2 | 24962300 | + | 1.97E-05 | 0.99998033 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
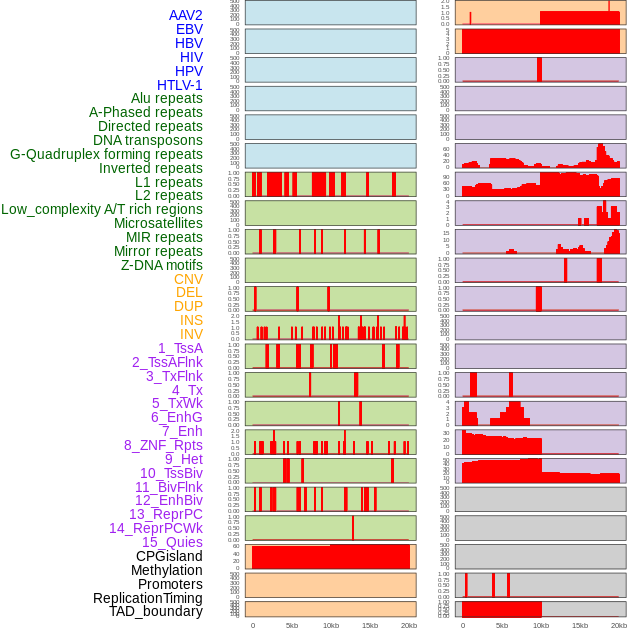 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
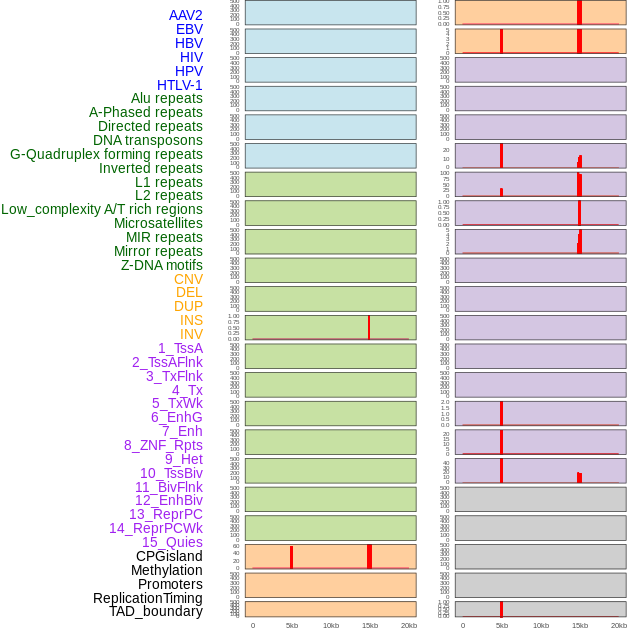 |
Top |
Fusion Protein Features for EFR3B-NCOA1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr2:25367920/chr2:24962301) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | NCOA1 |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Nuclear receptor coactivator that directly binds nuclear receptors and stimulates the transcriptional activities in a hormone-dependent fashion. Involved in the coactivation of different nuclear receptors, such as for steroids (PGR, GR and ER), retinoids (RXRs), thyroid hormone (TRs) and prostanoids (PPARs). Also involved in coactivation mediated by STAT3, STAT5A, STAT5B and STAT6 transcription factors. Displays histone acetyltransferase activity toward H3 and H4; the relevance of such activity remains however unclear. Plays a central role in creating multisubunit coactivator complexes that act via remodeling of chromatin, and possibly acts by participating in both chromatin remodeling and recruitment of general transcription factors. Required with NCOA2 to control energy balance between white and brown adipose tissues. Required for mediating steroid hormone response. Isoform 2 has a higher thyroid hormone-dependent transactivation activity than isoform 1 and isoform 3. {ECO:0000269|PubMed:10449719, ECO:0000269|PubMed:12954634, ECO:0000269|PubMed:7481822, ECO:0000269|PubMed:9223281, ECO:0000269|PubMed:9223431, ECO:0000269|PubMed:9296499, ECO:0000269|PubMed:9427757}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000288599 | 14 | 22 | 1435_1439 | 1067 | 2199.3333333333335 | Motif | Note=LXXLL motif 7 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000348332 | 14 | 21 | 1435_1439 | 1067 | 1442.0 | Motif | Note=LXXLL motif 7 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000395856 | 14 | 21 | 1435_1439 | 1067 | 1441.0 | Motif | Note=LXXLL motif 7 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000405141 | 17 | 25 | 1435_1439 | 1067 | 2183.6666666666665 | Motif | Note=LXXLL motif 7 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000406961 | 16 | 23 | 1435_1439 | 1067 | 1442.0 | Motif | Note=LXXLL motif 7 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000538539 | 15 | 23 | 1435_1439 | 1067 | 1422.6666666666667 | Motif | Note=LXXLL motif 7 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000288599 | 14 | 22 | 1053_1138 | 1067 | 2199.3333333333335 | Compositional bias | Note=Gln-rich | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000288599 | 14 | 22 | 389_682 | 1067 | 2199.3333333333335 | Compositional bias | Note=Ser-rich | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000348332 | 14 | 21 | 1053_1138 | 1067 | 1442.0 | Compositional bias | Note=Gln-rich | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000348332 | 14 | 21 | 389_682 | 1067 | 1442.0 | Compositional bias | Note=Ser-rich | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000395856 | 14 | 21 | 1053_1138 | 1067 | 1441.0 | Compositional bias | Note=Gln-rich | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000395856 | 14 | 21 | 389_682 | 1067 | 1441.0 | Compositional bias | Note=Ser-rich | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000405141 | 17 | 25 | 1053_1138 | 1067 | 2183.6666666666665 | Compositional bias | Note=Gln-rich | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000405141 | 17 | 25 | 389_682 | 1067 | 2183.6666666666665 | Compositional bias | Note=Ser-rich | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000406961 | 16 | 23 | 1053_1138 | 1067 | 1442.0 | Compositional bias | Note=Gln-rich | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000406961 | 16 | 23 | 389_682 | 1067 | 1442.0 | Compositional bias | Note=Ser-rich | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000538539 | 15 | 23 | 1053_1138 | 1067 | 1422.6666666666667 | Compositional bias | Note=Gln-rich | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000538539 | 15 | 23 | 389_682 | 1067 | 1422.6666666666667 | Compositional bias | Note=Ser-rich | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000288599 | 14 | 22 | 109_180 | 1067 | 2199.3333333333335 | Domain | PAS | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000288599 | 14 | 22 | 23_80 | 1067 | 2199.3333333333335 | Domain | bHLH | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000348332 | 14 | 21 | 109_180 | 1067 | 1442.0 | Domain | PAS | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000348332 | 14 | 21 | 23_80 | 1067 | 1442.0 | Domain | bHLH | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000395856 | 14 | 21 | 109_180 | 1067 | 1441.0 | Domain | PAS | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000395856 | 14 | 21 | 23_80 | 1067 | 1441.0 | Domain | bHLH | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000405141 | 17 | 25 | 109_180 | 1067 | 2183.6666666666665 | Domain | PAS | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000405141 | 17 | 25 | 23_80 | 1067 | 2183.6666666666665 | Domain | bHLH | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000406961 | 16 | 23 | 109_180 | 1067 | 1442.0 | Domain | PAS | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000406961 | 16 | 23 | 23_80 | 1067 | 1442.0 | Domain | bHLH | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000538539 | 15 | 23 | 109_180 | 1067 | 1422.6666666666667 | Domain | PAS | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000538539 | 15 | 23 | 23_80 | 1067 | 1422.6666666666667 | Domain | bHLH | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000288599 | 14 | 22 | 112_116 | 1067 | 2199.3333333333335 | Motif | Note=LXXLL motif 2 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000288599 | 14 | 22 | 46_50 | 1067 | 2199.3333333333335 | Motif | Note=LXXLL motif 1 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000288599 | 14 | 22 | 633_637 | 1067 | 2199.3333333333335 | Motif | Note=LXXLL motif 3 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000288599 | 14 | 22 | 690_694 | 1067 | 2199.3333333333335 | Motif | Note=LXXLL motif 4 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000288599 | 14 | 22 | 749_753 | 1067 | 2199.3333333333335 | Motif | Note=LXXLL motif 5 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000288599 | 14 | 22 | 913_917 | 1067 | 2199.3333333333335 | Motif | Note=LXXLL motif 6 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000348332 | 14 | 21 | 112_116 | 1067 | 1442.0 | Motif | Note=LXXLL motif 2 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000348332 | 14 | 21 | 46_50 | 1067 | 1442.0 | Motif | Note=LXXLL motif 1 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000348332 | 14 | 21 | 633_637 | 1067 | 1442.0 | Motif | Note=LXXLL motif 3 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000348332 | 14 | 21 | 690_694 | 1067 | 1442.0 | Motif | Note=LXXLL motif 4 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000348332 | 14 | 21 | 749_753 | 1067 | 1442.0 | Motif | Note=LXXLL motif 5 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000348332 | 14 | 21 | 913_917 | 1067 | 1442.0 | Motif | Note=LXXLL motif 6 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000395856 | 14 | 21 | 112_116 | 1067 | 1441.0 | Motif | Note=LXXLL motif 2 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000395856 | 14 | 21 | 46_50 | 1067 | 1441.0 | Motif | Note=LXXLL motif 1 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000395856 | 14 | 21 | 633_637 | 1067 | 1441.0 | Motif | Note=LXXLL motif 3 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000395856 | 14 | 21 | 690_694 | 1067 | 1441.0 | Motif | Note=LXXLL motif 4 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000395856 | 14 | 21 | 749_753 | 1067 | 1441.0 | Motif | Note=LXXLL motif 5 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000395856 | 14 | 21 | 913_917 | 1067 | 1441.0 | Motif | Note=LXXLL motif 6 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000405141 | 17 | 25 | 112_116 | 1067 | 2183.6666666666665 | Motif | Note=LXXLL motif 2 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000405141 | 17 | 25 | 46_50 | 1067 | 2183.6666666666665 | Motif | Note=LXXLL motif 1 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000405141 | 17 | 25 | 633_637 | 1067 | 2183.6666666666665 | Motif | Note=LXXLL motif 3 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000405141 | 17 | 25 | 690_694 | 1067 | 2183.6666666666665 | Motif | Note=LXXLL motif 4 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000405141 | 17 | 25 | 749_753 | 1067 | 2183.6666666666665 | Motif | Note=LXXLL motif 5 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000405141 | 17 | 25 | 913_917 | 1067 | 2183.6666666666665 | Motif | Note=LXXLL motif 6 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000406961 | 16 | 23 | 112_116 | 1067 | 1442.0 | Motif | Note=LXXLL motif 2 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000406961 | 16 | 23 | 46_50 | 1067 | 1442.0 | Motif | Note=LXXLL motif 1 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000406961 | 16 | 23 | 633_637 | 1067 | 1442.0 | Motif | Note=LXXLL motif 3 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000406961 | 16 | 23 | 690_694 | 1067 | 1442.0 | Motif | Note=LXXLL motif 4 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000406961 | 16 | 23 | 749_753 | 1067 | 1442.0 | Motif | Note=LXXLL motif 5 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000406961 | 16 | 23 | 913_917 | 1067 | 1442.0 | Motif | Note=LXXLL motif 6 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000538539 | 15 | 23 | 112_116 | 1067 | 1422.6666666666667 | Motif | Note=LXXLL motif 2 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000538539 | 15 | 23 | 46_50 | 1067 | 1422.6666666666667 | Motif | Note=LXXLL motif 1 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000538539 | 15 | 23 | 633_637 | 1067 | 1422.6666666666667 | Motif | Note=LXXLL motif 3 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000538539 | 15 | 23 | 690_694 | 1067 | 1422.6666666666667 | Motif | Note=LXXLL motif 4 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000538539 | 15 | 23 | 749_753 | 1067 | 1422.6666666666667 | Motif | Note=LXXLL motif 5 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000538539 | 15 | 23 | 913_917 | 1067 | 1422.6666666666667 | Motif | Note=LXXLL motif 6 |
Top |
Fusion Gene Sequence for EFR3B-NCOA1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >25408_25408_1_EFR3B-NCOA1_EFR3B_chr2_25367920_ENST00000402191_NCOA1_chr2_24962301_ENST00000406961_length(transcript)=5775nt_BP=2339nt GTGCGCGCGCGTCTGCGCTGCGAGGACAAAGATGCCTCGGGCCGGGGACCCTGGGGTCCTCTCCAGGCCCGGCGCGTGCCGGTGCTGGGC GGTGGTCCTTCGGGGGGCGGAGGCTCAGGGGAAAGCGGGTCTCCCGGAGCCGAGCAGACCGGGAGTGCTGGAGGAGGTGGTCGTGTGGCA GCATTTGCGGAATTAACAAAAGCGGTGTGTGTGGCTGCTGTGGTGCCCTACGCCCCAGGTACAAAAGGCTGGTTGACAACATCTTCCCTG AGGATCCCGAGGATGGTCTGGTGAAGACCAACATGGAGAAGCTGACCTTCTATGCCCTCTCAGCTCCAGAAAAACTTGATCGTATTGGCG CCTACCTCTCTGAGAGGCTCATCCGTGACGTGGGTCGCCATCGATATGGGTACGTGTGCATTGCTATGGAGGCTTTGGACCAGCTGCTCA TGGCCTGCCACTGCCAGAGCATCAACCTCTTCGTGGAGAGCTTCCTCAAGATGGTGGCCAAGCTGCTGGAGTCAGAGAAACCCAACCTGC AGATCCTCGGCACCAACTCGTTTGTGAAGTTTGCCAACATCGAGGAGGACACCCCGTCCTATCACCGGAGCTATGACTTCTTTGTGTCCC GATTCAGTGAAATGTGCCACTCGAGCCATGATGACTTAGAAATCAAGACCAAAATTCGAATGTCAGGCATCAAAGGCCTGCAAGGGGTGG TGAGGAAGACGGTGAATGATGAACTGCAGGCCAATATCTGGGACCCACAGCACATGGATAAGATCGTTCCATCACTGCTTTTCAATCTAC AGCATGTAGAGGAGGCAGAGAGCCGGTCTCCCTCACCCCTCCAAGCACCTGAGAAGGAGAAAGAGAGCCCCGCGGAGCTGGCTGAGAGGT GTCTTCGGGAGCTGCTGGGCCGGGCTGCCTTTGGCAACATCAAAAACGCCATCAAGCCTGTTCTCATCCATCTGGATAACCATTCTCTTT GGGAACCCAAGGTGTTTGCCATCCGTTGCTTTAAAATCATCATGTACTCAATTCAGCCGCAGCACTCACACCTGGTCATCCAGCAGCTCC TGGGCCACCTGGACGCCAACAGCCGCAGCGCTGCGACGGTGCGCGCGGGCATCGTGGAAGTCTTGTCGGAAGCCGCGGTCATCGCTGCCA CCGGCTCTGTGGGGCCCACAGTACTGGAGATGTTCAACACGCTGCTGAGGCAGCTGCGGCTCAGCATCGACTACGCGCTGACCGGGAGCT ACGACGGGGCGGTCAGCCTCGGCACCAAGATCATCAAGGAGCACGAGGAGCGCATGTTCCAGGAGGCCGTCATCAAGACCGTGGGCTCCT TTGCCAGCACGCTGCCCACCTACCAGCGCTCCGAGGTGATCCTCTTCATCATGAGCAAGGTCCCGCGGCCATCCCTGCACCAGGCGGTGG ACACAGGCAGGACGGGGGAGAATAGGAACCGTCTGACCCAGATTATGCTGCTAAAATCCCTCCTGCAGGTATCCACAGGTTTCCAGTGCA ACAACATGATGTCAGCCCTGCCTAGCAACTTCCTGGACCGCCTTCTCTCCACCGCCCTCATGGAGGATGCAGAAATTCGACTCTTTGTTC TAGAGATTCTCATCAGTTTCATTGATCGTCATGGCAACCGCCACAAGTTCTCTACCATCAGTACCCTCAGTGACATCTCTGTCCTGAAGC TGAAAGTGGACAAGTGCTCTCGACAGGACACCGTCTTCATGAAGAAGCACTCCCAGCAGCTCTACAGACACATCTACCTGAGCTGCAAGG AGGAAACAAACGTGCAGAAACACTACGAGGCGCTCTATGGCTTGCTGGCCCTCATCAGCATCGAGCTGGCTAACGAGGAGGTGGTGGTGG ACCTCATCCGTCTGGTGCTGGCTGTTCAGGACGTGGCCCAAGTCAATGAGGAGAACTTGCCTGTCTACAACCGCTGTGCCCTCTATGCTC TGGGCGCAGCCTACCTGAACCTCATCAGTCAGCTCACAACAGTGCCTGCCTTCTGCCAGCACATCCATGAGGTGATAGAGACCAGGAAGA AAGAGGCTCCATACATGCTCCCCGAGGATGTGTTTGTGGAGAGGCCCAGGCTGTCTCAGAATCTTGATGGGGTGGTCATTGAGCTCCTCT TCCGCCAGAGCAAGATCAGTGAAGTCCTGGGAGGCAGTGGCTACAACTCGGACCGGCTCTGCCTGCCCTACATTCCTCAGCTGACAGATG AGGATCGTTTATCCAAGAGGAGGAGCATTGGAGAGACCATCTCCCTGCAGGTGGAGGTAGAATCGAGGAACAGTCCGGAGAAGGAGGAGT TGATACACCAAAATCGGCAAGCTATCTTAAACCAGTTTGCAGCAACTGCTCCTGTTGGCATCAATATGAGATCAGGCATGCAACAGCAAA TTACACCTCAGCCACCCCTGAATGCTCAAATGTTGGCACAACGTCAGCGGGAACTGTACAGTCAACAGCACCGACAGAGGCAGCTAATAC AGCAGCAAAGAGCCATGCTTATGAGGCAGCAAAGCTTTGGGAACAACCTCCCTCCCTCATCTGGACTACCAGTTCAAATGGGGAACCCCC GTCTTCCTCAGGGTGCTCCACAGCAATTCCCCTATCCACCAAACTATGGTACAAATCCAGGAACCCCACCTGCTTCTACCAGCCCGTTTT CACAACTAGCAGCAAATCCTGAAGCATCCTTGGCCAACCGCAACAGCATGGTGAGCAGAGGCATGACAGGAAACATAGGAGGACAGTTTG GCACTGGAATCAATCCTCAGATGCAGCAGAATGTCTTCCAGTATCCAGGAGCAGGAATGGTTCCCCAAGGTGAGGCCAACTTTGCTCCAT CTCTAAGCCCTGGGAGCTCCATGGTGCCGATGCCAATCCCTCCTCCTCAGAGTTCTCTTCTCCAGCAAACTCCACCTGCCTCCGGGTATC AGTCACCAGACATGAAGGCCTGGCAGCAAGGAGCGATAGGAAACAACAATGTGTTCAGTCAAGCTGTCCAGAACCAGCCCACGCCTGCAC AGCCAGGAGTATACAACAACATGAGCATCACCGTTTCCATGGCAGGTGGAAATACGAATGTTCAGAACATGAACCCAATGATGGCCCAGA TGCAGATGAGCTCTTTGCAGATGCCAGGAATGAACACTGTGTGCCCTGAGCAGATAAATGATCCCGCACTGAGACACACAGGCCTCTACT GCAACCAGCTCTCATCCACTGACCTTCTCAAAACAGAAGCAGATGGAACCCAGCAGGTGCAACAGGTTCAGGTGTTTGCTGACGTCCAGT GTACAGTGAATCTGGTAGGCGGGGACCCTTACCTGAACCAGCCTGGTCCACTGGGAACTCAAAAGCCCACGTCAGGACCACAGACCCCCC AGGCCCAGCAGAAGAGCCTCCTTCAGCAGCTACTGACTGAATAACCACTTTTAAAGGAATGTGAAATTTAAATAATAGACATACAGAGAT ATACAAATATATTATATATTTTTCTGAGATTTTTGATATCTCAATCTGCAGCCATTCTTCAGGTCGTAGCATTTGGAGCAAAAAAAAAAA AAAAAAAAAAAAAAGGAGTTTGCTTTTGTCGGGAGATTGAAAGATGTTTTTGTTTCTTTCTTTGTAAAGGCCTTGGATATTGAAAAAATA CCAAGGCAGAACAGTTGGACAATCTATTTCTTGAGCCAAATTTAATTATTCTTATTTTTGTAATCAGTCATTGGCTTCTTATCTGGATGA AGGCTTTTGGAGGAGAACCAAAACGACAAGTTCCAAGAAGAAGATGAAGCTCCGCCTCCGCCGCTTAGTCCCAACCCTGCCCAGGAAGAA GGGCCCGTGGGGCTTTGCCTGTGCCCGTCCACCAAAGGCTGTCATGTGTCTCGAAATCAGCAGCCCTCCCCATCCCAATCCCAGGCAGCT TGTGTGTACAATCAGCTTCTCTAGCAACTCTGTATCTGTTGGCTTCAAGAGAATATTTTGCCTCCACATATGTACCCCTTCTCCTTTTTT TAAAGATGGATTTAAACCAAGATGCCTCCAGGAAAGAGGACGAAATGAGTATATTCACAGAGGAATCCAAAAAATACAGTTTGGGGGAAA ATGCAATAATTTTTGATGAGATGGGTGAAGGACAAGAAGTGAGTTGTGTCAATTATTGTAGATACAATTTTCTGATTAAATCTGGAAAAA TAAAAGGCAGCCTGTTTTTTCTGCTTTTATTGTATTAACAGCTGAGGTAGCTAAAGTTATTTAAAATAAAATTAAATTTATGATCCAAGT AGCTTATTTTTCCCTTTAAATCTCATTGTAAATATATTTGATTTCTTGTAGAAATTGATTTCCTTCTGTTTAATTTTATGCTTTTATTAT ACTCTTGATTTTTCTAAATTTGTGTGTGAAATATAACATTGATTGAATTGCAGTTACATTTGGTTAGTAATATTTCATTATTTTAATAAC TGTGATGTCATGTATGGATTTACTTTGGGGTTCAAATCAAAATGTCACTGCCAGAAAGAGCTGTTCCAGCTGATCTAGAGCATACTGCCC TAGAGTGTCCCTGGGATCATCTGAACAGAAGTGCACAGGCTACTTGTACAGAGAAAAAATTAATACTCAAAGGAAATCTTCATTTTTTAG ATTGACTTTGGGAATTTGAATTTTCATCAGTGCAAATATAAATTTCTCTATCCTGCTCTGAGGCTAATTGGTACCATATTTTCCCTTTGT GTCTTGTGACTCTGCCACATCCCATCTCATCCTGGCCTCTGAGTCAAGAACCCAGTGAACTGACTTTCTAGTTCTAGAAGTTCCGCTGCA AGGCCAGGAAAGCTTGAGAAAGGTATTGTGGAAGAAGCAAAGGTAGACCCCCATCACTCACCTTTGTCTGCATCCCTGGGCCTGTGAATG ATGACAGCACCTGACATTCTGCACCAGCTACCTCTGCCTCCATGGCAGAGAAAAGGCCATAAGAACAGTGGAAGAGGAGCATGGACTCAG ACTTCAAGGAAGAAGCCATTTCCCCAGGTCCTTCCTTCTGCATCTCACCACCCCTAGTTACAAATAACTCCATTGAACAGCATCTATTCA GAAACTATGCCGAATAAAAAGATTGGTGGAAGGGCTCATGTGGTTAGCAACTATGAAACAGAAATAGGACACTCAGTTACAAACATTATC TCCTTTAGTTTTTCAGAAAATGCATCCCTGATTTCATTCATTTCCAGCTTGAAAGCCAGCCATATTACTCTAGTCCCTACCAAACTGCTC TAGAAGGTCATTTCCATTTTGTTTGTGATATTTAGACGCGCAGACTTCAGGAAGTTCACCTTTAACTTCAGCATTCCAGATGAAGTTTCC TGACTCAGTGCTTTTGCATAAGGAACTAGAAAAAAAAAGTAAGGAAAATTGGAGATGCTAACATCCTCCCCCATCCCAACTGCACCTTAA AATATGCATGTCACCTTCAAGGTTTTATAATTGCACTGTTTGTTTTATGTATGTACAGATTAAAATTATTGCTACATTTGAGGAAAATAA AATTGCTTGCTTCTATGTAATTCCTGTCATTCCACAGGAAATTCACTTTTCCAGCTACTGAATAGAATTTGTTTAACAACCTCATGGGTT GCCTCTTCATTTACAGGGAAGAAATGAAATGTACATCTGCAGAAATTGCCAAAGCCCAAATTAACGAGTATTATAAAAGGATAATAAAGC >25408_25408_1_EFR3B-NCOA1_EFR3B_chr2_25367920_ENST00000402191_NCOA1_chr2_24962301_ENST00000406961_length(amino acids)=1148AA_BP=774 MRGQRCLGPGTLGSSPGPARAGAGRWSFGGRRLRGKRVSRSRADRECWRRWSCGSICGINKSGVCGCCGALRPRYKRLVDNIFPEDPEDG LVKTNMEKLTFYALSAPEKLDRIGAYLSERLIRDVGRHRYGYVCIAMEALDQLLMACHCQSINLFVESFLKMVAKLLESEKPNLQILGTN SFVKFANIEEDTPSYHRSYDFFVSRFSEMCHSSHDDLEIKTKIRMSGIKGLQGVVRKTVNDELQANIWDPQHMDKIVPSLLFNLQHVEEA ESRSPSPLQAPEKEKESPAELAERCLRELLGRAAFGNIKNAIKPVLIHLDNHSLWEPKVFAIRCFKIIMYSIQPQHSHLVIQQLLGHLDA NSRSAATVRAGIVEVLSEAAVIAATGSVGPTVLEMFNTLLRQLRLSIDYALTGSYDGAVSLGTKIIKEHEERMFQEAVIKTVGSFASTLP TYQRSEVILFIMSKVPRPSLHQAVDTGRTGENRNRLTQIMLLKSLLQVSTGFQCNNMMSALPSNFLDRLLSTALMEDAEIRLFVLEILIS FIDRHGNRHKFSTISTLSDISVLKLKVDKCSRQDTVFMKKHSQQLYRHIYLSCKEETNVQKHYEALYGLLALISIELANEEVVVDLIRLV LAVQDVAQVNEENLPVYNRCALYALGAAYLNLISQLTTVPAFCQHIHEVIETRKKEAPYMLPEDVFVERPRLSQNLDGVVIELLFRQSKI SEVLGGSGYNSDRLCLPYIPQLTDEDRLSKRRSIGETISLQVEVESRNSPEKEELIHQNRQAILNQFAATAPVGINMRSGMQQQITPQPP LNAQMLAQRQRELYSQQHRQRQLIQQQRAMLMRQQSFGNNLPPSSGLPVQMGNPRLPQGAPQQFPYPPNYGTNPGTPPASTSPFSQLAAN PEASLANRNSMVSRGMTGNIGGQFGTGINPQMQQNVFQYPGAGMVPQGEANFAPSLSPGSSMVPMPIPPPQSSLLQQTPPASGYQSPDMK AWQQGAIGNNNVFSQAVQNQPTPAQPGVYNNMSITVSMAGGNTNVQNMNPMMAQMQMSSLQMPGMNTVCPEQINDPALRHTGLYCNQLSS -------------------------------------------------------------- >25408_25408_2_EFR3B-NCOA1_EFR3B_chr2_25367920_ENST00000403714_NCOA1_chr2_24962301_ENST00000406961_length(transcript)=5761nt_BP=2325nt GCTGGCGGCGCCCGGCTCCGTCCTGCCCGCGGCCGGCCCCCGCGTCTGCTCCCTCCCCGCCCGGGCCCCTGTCGGCCGCCGCCAGTCCCC GCCCCGACTGTGAATGAAAGGCGGGCGCCGCCGAGGGCTGGCTGGGAACGCCGCAGCGACGCCGGCCTCTCGAGAGGCGCGCGCCCCGCC GAGATGTACGGTGTGTGTGGCTGCTGTGGTGCCCTACGCCCCAGGTACAAAAGGCTGGTTGACAACATCTTCCCTGAGGATCCCGAGGAT GGTCTGGTGAAGACCAACATGGAGAAGCTGACCTTCTATGCCCTCTCAGCTCCAGAAAAACTTGATCGTATTGGCGCCTACCTCTCTGAG AGGCTCATCCGTGACGTGGGTCGCCATCGATATGGGTACGTGTGCATTGCTATGGAGGCTTTGGACCAGCTGCTCATGGCCTGCCACTGC CAGAGCATCAACCTCTTCGTGGAGAGCTTCCTCAAGATGGTGGCCAAGCTGCTGGAGTCAGAGAAACCCAACCTGCAGATCCTCGGCACC AACTCGTTTGTGAAGTTTGCCAACATCGAGGAGGACACCCCGTCCTATCACCGGAGCTATGACTTCTTTGTGTCCCGATTCAGTGAAATG TGCCACTCGAGCCATGATGACTTAGAAATCAAGACCAAAATTCGAATGTCAGGCATCAAAGGCCTGCAAGGGGTGGTGAGGAAGACGGTG AATGATGAACTGCAGGCCAATATCTGGGACCCACAGCACATGGATAAGATCGTTCCATCACTGCTTTTCAATCTACAGCATGTAGAGGAG GCAGAGAGCCGGTCTCCCTCACCCCTCCAAGCACCTGAGAAGGAGAAAGAGAGCCCCGCGGAGCTGGCTGAGAGGTGTCTTCGGGAGCTG CTGGGCCGGGCTGCCTTTGGCAACATCAAAAACGCCATCAAGCCTGTTCTCATCCATCTGGATAACCATTCTCTTTGGGAACCCAAGGTG TTTGCCATCCGTTGCTTTAAAATCATCATGTACTCAATTCAGCCGCAGCACTCACACCTGGTCATCCAGCAGCTCCTGGGCCACCTGGAC GCCAACAGCCGCAGCGCTGCGACGGTGCGCGCGGGCATCGTGGAAGTCTTGTCGGAAGCCGCGGTCATCGCTGCCACCGGCTCTGTGGGG CCCACAGTACTGGAGATGTTCAACACGCTGCTGAGGCAGCTGCGGCTCAGCATCGACTACGCGCTGACCGGGAGCTACGACGGGGCGGTC AGCCTCGGCACCAAGATCATCAAGGAGCACGAGGAGCGCATGTTCCAGGAGGCCGTCATCAAGACCGTGGGCTCCTTTGCCAGCACGCTG CCCACCTACCAGCGCTCCGAGGTGATCCTCTTCATCATGAGCAAGGTCCCGCGGCCATCCCTGCACCAGGCGGTGGACACAGGCAGGACG GGGGAGAATAGGAACCGTCTGACCCAGATTATGCTGCTAAAATCCCTCCTGCAGGTATCCACAGGTTTCCAGTGCAACAACATGATGTCA GCCCTGCCTAGCAACTTCCTGGACCGCCTTCTCTCCACCGCCCTCATGGAGGATGCAGAAATTCGACTCTTTGTTCTAGAGATTCTCATC AGTTTCATTGATCGTCATGGCAACCGCCACAAGTTCTCTACCATCAGTACCCTCAGTGACATCTCTGTCCTGAAGCTGAAAGTGGACAAG TGCTCTCGACAGGACACCGTCTTCATGAAGAAGCACTCCCAGCAGCTCTACAGACACATCTACCTGAGCTGCAAGGAGGAAACAAACGTG CAGAAACACTACGAGGCGCTCTATGGCTTGCTGGCCCTCATCAGCATCGAGCTGGCTAACGAGGAGGTGGTGGTGGACCTCATCCGTCTG GTGCTGGCTGTTCAGGACGTGGCCCAAGTCAATGAGGAGAACTTGCCTGTCTACAACCGCTGTGCCCTCTATGCTCTGGGCGCAGCCTAC CTGAACCTCATCAGTCAGCTCACAACAGTGCCTGCCTTCTGCCAGCACATCCATGAGGTGATAGAGACCAGGAAGAAAGAGGCTCCATAC ATGCTCCCCGAGGATGTGTTTGTGGAGAGGCCCAGGCTGTCTCAGAATCTTGATGGGGTGGTCATTGAGCTCCTCTTCCGCCAGAGCAAG ATCAGTGAAGTCCTGGGAGGCAGTGGCTACAACTCGGACCGGCTCTGCCTGCCCTACATTCCTCAGCTGACAGATGAGGATCGTTTATCC AAGAGGAGGAGCATTGGAGAGACCATCTCCCTGCAGGTGGAGGTAGAATCGAGGAACAGTCCGGAGAAGGAGGAGTTGATACACCAAAAT CGGCAAGCTATCTTAAACCAGTTTGCAGCAACTGCTCCTGTTGGCATCAATATGAGATCAGGCATGCAACAGCAAATTACACCTCAGCCA CCCCTGAATGCTCAAATGTTGGCACAACGTCAGCGGGAACTGTACAGTCAACAGCACCGACAGAGGCAGCTAATACAGCAGCAAAGAGCC ATGCTTATGAGGCAGCAAAGCTTTGGGAACAACCTCCCTCCCTCATCTGGACTACCAGTTCAAATGGGGAACCCCCGTCTTCCTCAGGGT GCTCCACAGCAATTCCCCTATCCACCAAACTATGGTACAAATCCAGGAACCCCACCTGCTTCTACCAGCCCGTTTTCACAACTAGCAGCA AATCCTGAAGCATCCTTGGCCAACCGCAACAGCATGGTGAGCAGAGGCATGACAGGAAACATAGGAGGACAGTTTGGCACTGGAATCAAT CCTCAGATGCAGCAGAATGTCTTCCAGTATCCAGGAGCAGGAATGGTTCCCCAAGGTGAGGCCAACTTTGCTCCATCTCTAAGCCCTGGG AGCTCCATGGTGCCGATGCCAATCCCTCCTCCTCAGAGTTCTCTTCTCCAGCAAACTCCACCTGCCTCCGGGTATCAGTCACCAGACATG AAGGCCTGGCAGCAAGGAGCGATAGGAAACAACAATGTGTTCAGTCAAGCTGTCCAGAACCAGCCCACGCCTGCACAGCCAGGAGTATAC AACAACATGAGCATCACCGTTTCCATGGCAGGTGGAAATACGAATGTTCAGAACATGAACCCAATGATGGCCCAGATGCAGATGAGCTCT TTGCAGATGCCAGGAATGAACACTGTGTGCCCTGAGCAGATAAATGATCCCGCACTGAGACACACAGGCCTCTACTGCAACCAGCTCTCA TCCACTGACCTTCTCAAAACAGAAGCAGATGGAACCCAGCAGGTGCAACAGGTTCAGGTGTTTGCTGACGTCCAGTGTACAGTGAATCTG GTAGGCGGGGACCCTTACCTGAACCAGCCTGGTCCACTGGGAACTCAAAAGCCCACGTCAGGACCACAGACCCCCCAGGCCCAGCAGAAG AGCCTCCTTCAGCAGCTACTGACTGAATAACCACTTTTAAAGGAATGTGAAATTTAAATAATAGACATACAGAGATATACAAATATATTA TATATTTTTCTGAGATTTTTGATATCTCAATCTGCAGCCATTCTTCAGGTCGTAGCATTTGGAGCAAAAAAAAAAAAAAAAAAAAAAAAA GGAGTTTGCTTTTGTCGGGAGATTGAAAGATGTTTTTGTTTCTTTCTTTGTAAAGGCCTTGGATATTGAAAAAATACCAAGGCAGAACAG TTGGACAATCTATTTCTTGAGCCAAATTTAATTATTCTTATTTTTGTAATCAGTCATTGGCTTCTTATCTGGATGAAGGCTTTTGGAGGA GAACCAAAACGACAAGTTCCAAGAAGAAGATGAAGCTCCGCCTCCGCCGCTTAGTCCCAACCCTGCCCAGGAAGAAGGGCCCGTGGGGCT TTGCCTGTGCCCGTCCACCAAAGGCTGTCATGTGTCTCGAAATCAGCAGCCCTCCCCATCCCAATCCCAGGCAGCTTGTGTGTACAATCA GCTTCTCTAGCAACTCTGTATCTGTTGGCTTCAAGAGAATATTTTGCCTCCACATATGTACCCCTTCTCCTTTTTTTAAAGATGGATTTA AACCAAGATGCCTCCAGGAAAGAGGACGAAATGAGTATATTCACAGAGGAATCCAAAAAATACAGTTTGGGGGAAAATGCAATAATTTTT GATGAGATGGGTGAAGGACAAGAAGTGAGTTGTGTCAATTATTGTAGATACAATTTTCTGATTAAATCTGGAAAAATAAAAGGCAGCCTG TTTTTTCTGCTTTTATTGTATTAACAGCTGAGGTAGCTAAAGTTATTTAAAATAAAATTAAATTTATGATCCAAGTAGCTTATTTTTCCC TTTAAATCTCATTGTAAATATATTTGATTTCTTGTAGAAATTGATTTCCTTCTGTTTAATTTTATGCTTTTATTATACTCTTGATTTTTC TAAATTTGTGTGTGAAATATAACATTGATTGAATTGCAGTTACATTTGGTTAGTAATATTTCATTATTTTAATAACTGTGATGTCATGTA TGGATTTACTTTGGGGTTCAAATCAAAATGTCACTGCCAGAAAGAGCTGTTCCAGCTGATCTAGAGCATACTGCCCTAGAGTGTCCCTGG GATCATCTGAACAGAAGTGCACAGGCTACTTGTACAGAGAAAAAATTAATACTCAAAGGAAATCTTCATTTTTTAGATTGACTTTGGGAA TTTGAATTTTCATCAGTGCAAATATAAATTTCTCTATCCTGCTCTGAGGCTAATTGGTACCATATTTTCCCTTTGTGTCTTGTGACTCTG CCACATCCCATCTCATCCTGGCCTCTGAGTCAAGAACCCAGTGAACTGACTTTCTAGTTCTAGAAGTTCCGCTGCAAGGCCAGGAAAGCT TGAGAAAGGTATTGTGGAAGAAGCAAAGGTAGACCCCCATCACTCACCTTTGTCTGCATCCCTGGGCCTGTGAATGATGACAGCACCTGA CATTCTGCACCAGCTACCTCTGCCTCCATGGCAGAGAAAAGGCCATAAGAACAGTGGAAGAGGAGCATGGACTCAGACTTCAAGGAAGAA GCCATTTCCCCAGGTCCTTCCTTCTGCATCTCACCACCCCTAGTTACAAATAACTCCATTGAACAGCATCTATTCAGAAACTATGCCGAA TAAAAAGATTGGTGGAAGGGCTCATGTGGTTAGCAACTATGAAACAGAAATAGGACACTCAGTTACAAACATTATCTCCTTTAGTTTTTC AGAAAATGCATCCCTGATTTCATTCATTTCCAGCTTGAAAGCCAGCCATATTACTCTAGTCCCTACCAAACTGCTCTAGAAGGTCATTTC CATTTTGTTTGTGATATTTAGACGCGCAGACTTCAGGAAGTTCACCTTTAACTTCAGCATTCCAGATGAAGTTTCCTGACTCAGTGCTTT TGCATAAGGAACTAGAAAAAAAAAGTAAGGAAAATTGGAGATGCTAACATCCTCCCCCATCCCAACTGCACCTTAAAATATGCATGTCAC CTTCAAGGTTTTATAATTGCACTGTTTGTTTTATGTATGTACAGATTAAAATTATTGCTACATTTGAGGAAAATAAAATTGCTTGCTTCT ATGTAATTCCTGTCATTCCACAGGAAATTCACTTTTCCAGCTACTGAATAGAATTTGTTTAACAACCTCATGGGTTGCCTCTTCATTTAC AGGGAAGAAATGAAATGTACATCTGCAGAAATTGCCAAAGCCCAAATTAACGAGTATTATAAAAGGATAATAAAGCATCCCTGAAGCCAA >25408_25408_2_EFR3B-NCOA1_EFR3B_chr2_25367920_ENST00000403714_NCOA1_chr2_24962301_ENST00000406961_length(amino acids)=1134AA_BP=760 MLPPRPGPCRPPPVPAPTVNERRAPPRAGWERRSDAGLSRGARPAEMYGVCGCCGALRPRYKRLVDNIFPEDPEDGLVKTNMEKLTFYAL SAPEKLDRIGAYLSERLIRDVGRHRYGYVCIAMEALDQLLMACHCQSINLFVESFLKMVAKLLESEKPNLQILGTNSFVKFANIEEDTPS YHRSYDFFVSRFSEMCHSSHDDLEIKTKIRMSGIKGLQGVVRKTVNDELQANIWDPQHMDKIVPSLLFNLQHVEEAESRSPSPLQAPEKE KESPAELAERCLRELLGRAAFGNIKNAIKPVLIHLDNHSLWEPKVFAIRCFKIIMYSIQPQHSHLVIQQLLGHLDANSRSAATVRAGIVE VLSEAAVIAATGSVGPTVLEMFNTLLRQLRLSIDYALTGSYDGAVSLGTKIIKEHEERMFQEAVIKTVGSFASTLPTYQRSEVILFIMSK VPRPSLHQAVDTGRTGENRNRLTQIMLLKSLLQVSTGFQCNNMMSALPSNFLDRLLSTALMEDAEIRLFVLEILISFIDRHGNRHKFSTI STLSDISVLKLKVDKCSRQDTVFMKKHSQQLYRHIYLSCKEETNVQKHYEALYGLLALISIELANEEVVVDLIRLVLAVQDVAQVNEENL PVYNRCALYALGAAYLNLISQLTTVPAFCQHIHEVIETRKKEAPYMLPEDVFVERPRLSQNLDGVVIELLFRQSKISEVLGGSGYNSDRL CLPYIPQLTDEDRLSKRRSIGETISLQVEVESRNSPEKEELIHQNRQAILNQFAATAPVGINMRSGMQQQITPQPPLNAQMLAQRQRELY SQQHRQRQLIQQQRAMLMRQQSFGNNLPPSSGLPVQMGNPRLPQGAPQQFPYPPNYGTNPGTPPASTSPFSQLAANPEASLANRNSMVSR GMTGNIGGQFGTGINPQMQQNVFQYPGAGMVPQGEANFAPSLSPGSSMVPMPIPPPQSSLLQQTPPASGYQSPDMKAWQQGAIGNNNVFS QAVQNQPTPAQPGVYNNMSITVSMAGGNTNVQNMNPMMAQMQMSSLQMPGMNTVCPEQINDPALRHTGLYCNQLSSTDLLKTEADGTQQV -------------------------------------------------------------- >25408_25408_3_EFR3B-NCOA1_EFR3B_chr2_25367920_ENST00000405108_NCOA1_chr2_24962301_ENST00000406961_length(transcript)=5331nt_BP=1895nt GCAAATAAGCTTTACTACAACCATATCTGAAATGTGTTTTCCTGATCAAGTGGGATGGGAAAGAACAGTGAGTGGAGTCCCAGGCATGTT AAGAATGGCGTGCTGTTTTTTAAGTGTTTGTGAAGTTTGCCAACATCGAGGAGGACACCCCGTCCTATCACCGGAGCTATGACTTCTTTG TGTCCCGATTCAGTGAAATGTGCCACTCGAGCCATGATGACTTAGAAATCAAGACCAAAATTCGAATGTCAGGCATCAAAGGCCTGCAAG GGGTGGTGAGGAAGACGGTGAATGATGAACTGCAGGCCAATATCTGGGACCCACAGCACATGGATAAGATCGTTCCATCACTGCTTTTCA ATCTACAGCATGTAGAGGAGGCAGAGAGCCGGTCTCCCTCACCCCTCCAAGCACCTGAGAAGGAGAAAGAGAGCCCCGCGGAGCTGGCTG AGAGGTGTCTTCGGGAGCTGCTGGGCCGGGCTGCCTTTGGCAACATCAAAAACGCCATCAAGCCTGTTCTCATCCATCTGGATAACCATT CTCTTTGGGAACCCAAGGTGTTTGCCATCCGTTGCTTTAAAATCATCATGTACTCAATTCAGCCGCAGCACTCACACCTGGTCATCCAGC AGCTCCTGGGCCACCTGGACGCCAACAGCCGCAGCGCTGCGACGGTGCGCGCGGGCATCGTGGAAGTCTTGTCGGAAGCCGCGGTCATCG CTGCCACCGGCTCTGTGGGGCCCACAGTACTGGAGATGTTCAACACGCTGCTGAGGCAGCTGCGGCTCAGCATCGACTACGCGCTGACCG GGAGCTACGACGGGGCGGTCAGCCTCGGCACCAAGATCATCAAGGAGCACGAGGAGCGCATGTTCCAGGAGGCCGTCATCAAGACCGTGG GCTCCTTTGCCAGCACGCTGCCCACCTACCAGCGCTCCGAGGTGATCCTCTTCATCATGAGCAAGGTCCCGCGGCCATCCCTGCACCAGG CGGTGGACACAGGCAGGACGGGGGAGAATAGGAACCGTCTGACCCAGATTATGCTGCTAAAATCCCTCCTGCAGGTATCCACAGGTTTCC AGTGCAACAACATGATGTCAGCCCTGCCTAGCAACTTCCTGGACCGCCTTCTCTCCACCGCCCTCATGGAGGATGCAGAAATTCGACTCT TTGTTCTAGAGATTCTCATCAGTTTCATTGATCGTCATGGCAACCGCCACAAGTTCTCTACCATCAGTACCCTCAGTGACATCTCTGTCC TGAAGCTGAAAGTGGACAAGTGCTCTCGACAGGACACCGTCTTCATGAAGAAGCACTCCCAGCAGCTCTACAGACACATCTACCTGAGCT GCAAGGAGGAAACAAACGTGCAGAAACACTACGAGGCGCTCTATGGCTTGCTGGCCCTCATCAGCATCGAGCTGGCTAACGAGGAGGTGG TGGTGGACCTCATCCGTCTGGTGCTGGCTGTTCAGGACGTGGCCCAAGTCAATGAGGAGAACTTGCCTGTCTACAACCGCTGTGCCCTCT ATGCTCTGGGCGCAGCCTACCTGAACCTCATCAGTCAGCTCACAACAGTGCCTGCCTTCTGCCAGCACATCCATGAGGTGATAGAGACCA GGAAGAAAGAGGCTCCATACATGCTCCCCGAGGATGTGTTTGTGGAGAGGCCCAGGCTGTCTCAGAATCTTGATGGGGTGGTCATTGAGC TCCTCTTCCGCCAGAGCAAGATCAGTGAAGTCCTGGGAGGCAGTGGCTACAACTCGGACCGGCTCTGCCTGCCCTACATTCCTCAGCTGA CAGATGAGGATCGTTTATCCAAGAGGAGGAGCATTGGAGAGACCATCTCCCTGCAGGTGGAGGTAGAATCGAGGAACAGTCCGGAGAAGG AGGAGTTGATACACCAAAATCGGCAAGCTATCTTAAACCAGTTTGCAGCAACTGCTCCTGTTGGCATCAATATGAGATCAGGCATGCAAC AGCAAATTACACCTCAGCCACCCCTGAATGCTCAAATGTTGGCACAACGTCAGCGGGAACTGTACAGTCAACAGCACCGACAGAGGCAGC TAATACAGCAGCAAAGAGCCATGCTTATGAGGCAGCAAAGCTTTGGGAACAACCTCCCTCCCTCATCTGGACTACCAGTTCAAATGGGGA ACCCCCGTCTTCCTCAGGGTGCTCCACAGCAATTCCCCTATCCACCAAACTATGGTACAAATCCAGGAACCCCACCTGCTTCTACCAGCC CGTTTTCACAACTAGCAGCAAATCCTGAAGCATCCTTGGCCAACCGCAACAGCATGGTGAGCAGAGGCATGACAGGAAACATAGGAGGAC AGTTTGGCACTGGAATCAATCCTCAGATGCAGCAGAATGTCTTCCAGTATCCAGGAGCAGGAATGGTTCCCCAAGGTGAGGCCAACTTTG CTCCATCTCTAAGCCCTGGGAGCTCCATGGTGCCGATGCCAATCCCTCCTCCTCAGAGTTCTCTTCTCCAGCAAACTCCACCTGCCTCCG GGTATCAGTCACCAGACATGAAGGCCTGGCAGCAAGGAGCGATAGGAAACAACAATGTGTTCAGTCAAGCTGTCCAGAACCAGCCCACGC CTGCACAGCCAGGAGTATACAACAACATGAGCATCACCGTTTCCATGGCAGGTGGAAATACGAATGTTCAGAACATGAACCCAATGATGG CCCAGATGCAGATGAGCTCTTTGCAGATGCCAGGAATGAACACTGTGTGCCCTGAGCAGATAAATGATCCCGCACTGAGACACACAGGCC TCTACTGCAACCAGCTCTCATCCACTGACCTTCTCAAAACAGAAGCAGATGGAACCCAGCAGGTGCAACAGGTTCAGGTGTTTGCTGACG TCCAGTGTACAGTGAATCTGGTAGGCGGGGACCCTTACCTGAACCAGCCTGGTCCACTGGGAACTCAAAAGCCCACGTCAGGACCACAGA CCCCCCAGGCCCAGCAGAAGAGCCTCCTTCAGCAGCTACTGACTGAATAACCACTTTTAAAGGAATGTGAAATTTAAATAATAGACATAC AGAGATATACAAATATATTATATATTTTTCTGAGATTTTTGATATCTCAATCTGCAGCCATTCTTCAGGTCGTAGCATTTGGAGCAAAAA AAAAAAAAAAAAAAAAAAAAGGAGTTTGCTTTTGTCGGGAGATTGAAAGATGTTTTTGTTTCTTTCTTTGTAAAGGCCTTGGATATTGAA AAAATACCAAGGCAGAACAGTTGGACAATCTATTTCTTGAGCCAAATTTAATTATTCTTATTTTTGTAATCAGTCATTGGCTTCTTATCT GGATGAAGGCTTTTGGAGGAGAACCAAAACGACAAGTTCCAAGAAGAAGATGAAGCTCCGCCTCCGCCGCTTAGTCCCAACCCTGCCCAG GAAGAAGGGCCCGTGGGGCTTTGCCTGTGCCCGTCCACCAAAGGCTGTCATGTGTCTCGAAATCAGCAGCCCTCCCCATCCCAATCCCAG GCAGCTTGTGTGTACAATCAGCTTCTCTAGCAACTCTGTATCTGTTGGCTTCAAGAGAATATTTTGCCTCCACATATGTACCCCTTCTCC TTTTTTTAAAGATGGATTTAAACCAAGATGCCTCCAGGAAAGAGGACGAAATGAGTATATTCACAGAGGAATCCAAAAAATACAGTTTGG GGGAAAATGCAATAATTTTTGATGAGATGGGTGAAGGACAAGAAGTGAGTTGTGTCAATTATTGTAGATACAATTTTCTGATTAAATCTG GAAAAATAAAAGGCAGCCTGTTTTTTCTGCTTTTATTGTATTAACAGCTGAGGTAGCTAAAGTTATTTAAAATAAAATTAAATTTATGAT CCAAGTAGCTTATTTTTCCCTTTAAATCTCATTGTAAATATATTTGATTTCTTGTAGAAATTGATTTCCTTCTGTTTAATTTTATGCTTT TATTATACTCTTGATTTTTCTAAATTTGTGTGTGAAATATAACATTGATTGAATTGCAGTTACATTTGGTTAGTAATATTTCATTATTTT AATAACTGTGATGTCATGTATGGATTTACTTTGGGGTTCAAATCAAAATGTCACTGCCAGAAAGAGCTGTTCCAGCTGATCTAGAGCATA CTGCCCTAGAGTGTCCCTGGGATCATCTGAACAGAAGTGCACAGGCTACTTGTACAGAGAAAAAATTAATACTCAAAGGAAATCTTCATT TTTTAGATTGACTTTGGGAATTTGAATTTTCATCAGTGCAAATATAAATTTCTCTATCCTGCTCTGAGGCTAATTGGTACCATATTTTCC CTTTGTGTCTTGTGACTCTGCCACATCCCATCTCATCCTGGCCTCTGAGTCAAGAACCCAGTGAACTGACTTTCTAGTTCTAGAAGTTCC GCTGCAAGGCCAGGAAAGCTTGAGAAAGGTATTGTGGAAGAAGCAAAGGTAGACCCCCATCACTCACCTTTGTCTGCATCCCTGGGCCTG TGAATGATGACAGCACCTGACATTCTGCACCAGCTACCTCTGCCTCCATGGCAGAGAAAAGGCCATAAGAACAGTGGAAGAGGAGCATGG ACTCAGACTTCAAGGAAGAAGCCATTTCCCCAGGTCCTTCCTTCTGCATCTCACCACCCCTAGTTACAAATAACTCCATTGAACAGCATC TATTCAGAAACTATGCCGAATAAAAAGATTGGTGGAAGGGCTCATGTGGTTAGCAACTATGAAACAGAAATAGGACACTCAGTTACAAAC ATTATCTCCTTTAGTTTTTCAGAAAATGCATCCCTGATTTCATTCATTTCCAGCTTGAAAGCCAGCCATATTACTCTAGTCCCTACCAAA CTGCTCTAGAAGGTCATTTCCATTTTGTTTGTGATATTTAGACGCGCAGACTTCAGGAAGTTCACCTTTAACTTCAGCATTCCAGATGAA GTTTCCTGACTCAGTGCTTTTGCATAAGGAACTAGAAAAAAAAAGTAAGGAAAATTGGAGATGCTAACATCCTCCCCCATCCCAACTGCA CCTTAAAATATGCATGTCACCTTCAAGGTTTTATAATTGCACTGTTTGTTTTATGTATGTACAGATTAAAATTATTGCTACATTTGAGGA AAATAAAATTGCTTGCTTCTATGTAATTCCTGTCATTCCACAGGAAATTCACTTTTCCAGCTACTGAATAGAATTTGTTTAACAACCTCA TGGGTTGCCTCTTCATTTACAGGGAAGAAATGAAATGTACATCTGCAGAAATTGCCAAAGCCCAAATTAACGAGTATTATAAAAGGATAA >25408_25408_3_EFR3B-NCOA1_EFR3B_chr2_25367920_ENST00000405108_NCOA1_chr2_24962301_ENST00000406961_length(amino acids)=940AA_BP=566 MCHSSHDDLEIKTKIRMSGIKGLQGVVRKTVNDELQANIWDPQHMDKIVPSLLFNLQHVEEAESRSPSPLQAPEKEKESPAELAERCLRE LLGRAAFGNIKNAIKPVLIHLDNHSLWEPKVFAIRCFKIIMYSIQPQHSHLVIQQLLGHLDANSRSAATVRAGIVEVLSEAAVIAATGSV GPTVLEMFNTLLRQLRLSIDYALTGSYDGAVSLGTKIIKEHEERMFQEAVIKTVGSFASTLPTYQRSEVILFIMSKVPRPSLHQAVDTGR TGENRNRLTQIMLLKSLLQVSTGFQCNNMMSALPSNFLDRLLSTALMEDAEIRLFVLEILISFIDRHGNRHKFSTISTLSDISVLKLKVD KCSRQDTVFMKKHSQQLYRHIYLSCKEETNVQKHYEALYGLLALISIELANEEVVVDLIRLVLAVQDVAQVNEENLPVYNRCALYALGAA YLNLISQLTTVPAFCQHIHEVIETRKKEAPYMLPEDVFVERPRLSQNLDGVVIELLFRQSKISEVLGGSGYNSDRLCLPYIPQLTDEDRL SKRRSIGETISLQVEVESRNSPEKEELIHQNRQAILNQFAATAPVGINMRSGMQQQITPQPPLNAQMLAQRQRELYSQQHRQRQLIQQQR AMLMRQQSFGNNLPPSSGLPVQMGNPRLPQGAPQQFPYPPNYGTNPGTPPASTSPFSQLAANPEASLANRNSMVSRGMTGNIGGQFGTGI NPQMQQNVFQYPGAGMVPQGEANFAPSLSPGSSMVPMPIPPPQSSLLQQTPPASGYQSPDMKAWQQGAIGNNNVFSQAVQNQPTPAQPGV YNNMSITVSMAGGNTNVQNMNPMMAQMQMSSLQMPGMNTVCPEQINDPALRHTGLYCNQLSSTDLLKTEADGTQQVQQVQVFADVQCTVN -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for EFR3B-NCOA1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000288599 | 14 | 22 | 781_988 | 1067.0 | 2199.3333333333335 | CREBBP | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000348332 | 14 | 21 | 781_988 | 1067.0 | 1442.0 | CREBBP | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000395856 | 14 | 21 | 781_988 | 1067.0 | 1441.0 | CREBBP | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000405141 | 17 | 25 | 781_988 | 1067.0 | 2183.6666666666665 | CREBBP | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000406961 | 16 | 23 | 781_988 | 1067.0 | 1442.0 | CREBBP | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000538539 | 15 | 23 | 781_988 | 1067.0 | 1422.6666666666667 | CREBBP | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000288599 | 14 | 22 | 361_567 | 1067.0 | 2199.3333333333335 | STAT3 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000348332 | 14 | 21 | 361_567 | 1067.0 | 1442.0 | STAT3 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000395856 | 14 | 21 | 361_567 | 1067.0 | 1441.0 | STAT3 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000405141 | 17 | 25 | 361_567 | 1067.0 | 2183.6666666666665 | STAT3 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000406961 | 16 | 23 | 361_567 | 1067.0 | 1442.0 | STAT3 | |
| Tgene | NCOA1 | chr2:25367920 | chr2:24962301 | ENST00000538539 | 15 | 23 | 361_567 | 1067.0 | 1422.6666666666667 | STAT3 |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for EFR3B-NCOA1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for EFR3B-NCOA1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
| Tgene | C0006142 | Malignant neoplasm of breast | 1 | CTD_human | |
| Tgene | C0014175 | Endometriosis | 1 | CTD_human | |
| Tgene | C0024668 | Mammary Neoplasms, Experimental | 1 | CTD_human | |
| Tgene | C0027627 | Neoplasm Metastasis | 1 | CTD_human | |
| Tgene | C0033578 | Prostatic Neoplasms | 1 | CTD_human | |
| Tgene | C0269102 | Endometrioma | 1 | CTD_human | |
| Tgene | C0376358 | Malignant neoplasm of prostate | 1 | CTD_human | |
| Tgene | C0678222 | Breast Carcinoma | 1 | CTD_human | |
| Tgene | C1257931 | Mammary Neoplasms, Human | 1 | CTD_human | |
| Tgene | C1458155 | Mammary Neoplasms | 1 | CTD_human | |
| Tgene | C4704874 | Mammary Carcinoma, Human | 1 | CTD_human |
