
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:ZMYND8-SEP15 (FusionGDB2 ID:HG23613TG9403) |
Fusion Gene Summary for ZMYND8-SEP15 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: ZMYND8-SEP15 | Fusion gene ID: hg23613tg9403 | Hgene | Tgene | Gene symbol | ZMYND8 | SEP15 | Gene ID | 23613 | 9403 |
| Gene name | zinc finger MYND-type containing 8 | selenoprotein F | |
| Synonyms | PRKCBP1|PRO2893|RACK7 | SEP15 | |
| Cytomap | ('ZMYND8')('SEP15') 20q13.12 | 1p22.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | protein kinase C-binding protein 1CTCL tumor antigen se14-3cutaneous T-cell lymphoma-associated antigen se14-3predicted protein of HQ2893zinc finger MYND domain-containing protein 8 | selenoprotein F15 kDa selenoprotein | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000262975, ENST00000311275, ENST00000352431, ENST00000355972, ENST00000360911, ENST00000372023, ENST00000396281, ENST00000446994, ENST00000458360, ENST00000461685, ENST00000471951, ENST00000536340, ENST00000540497, ENST00000468376, | ||
| Fusion gene scores | * DoF score | 25 X 23 X 10=5750 | 2 X 2 X 2=8 |
| # samples | 34 | 2 | |
| ** MAII score | log2(34/5750*10)=-4.0799553045814 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(2/8*10)=1.32192809488736 | |
| Context | PubMed: ZMYND8 [Title/Abstract] AND SEP15 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | ZMYND8(45918930)-SEP15(87333785), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | ZMYND8-SEP15 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. ZMYND8-SEP15 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. ZMYND8-SEP15 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. ZMYND8-SEP15 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. ZMYND8-SEP15 seems lost the major protein functional domain in Hgene partner, which is a epigenetic factor due to the frame-shifted ORF. ZMYND8-SEP15 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. ZMYND8-SEP15 seems lost the major protein functional domain in Hgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
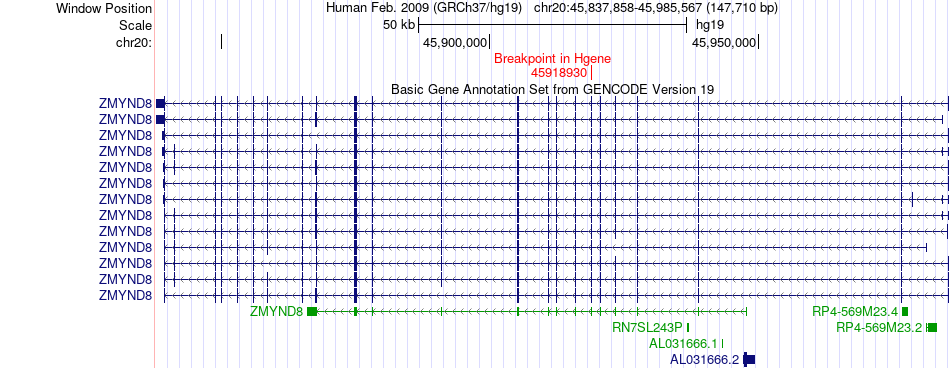
 Fusion gene breakpoints across ZMYND8 (5'-gene) Fusion gene breakpoints across ZMYND8 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
 Fusion gene breakpoints across SEP15 (3'-gene) Fusion gene breakpoints across SEP15 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
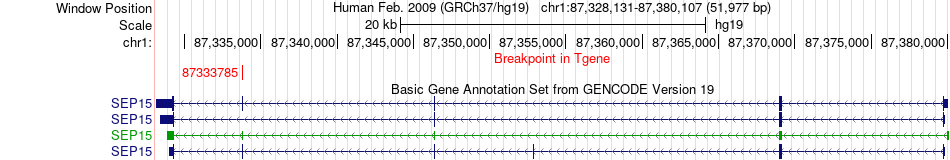 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | Non-Cancer | 269N | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
Top |
Fusion Gene ORF analysis for ZMYND8-SEP15 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000262975 | ENST00000401030 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000262975 | ENST00000469566 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000311275 | ENST00000401030 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000311275 | ENST00000469566 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000352431 | ENST00000401030 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000352431 | ENST00000469566 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000355972 | ENST00000401030 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000355972 | ENST00000469566 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000360911 | ENST00000401030 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000360911 | ENST00000469566 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000372023 | ENST00000401030 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000372023 | ENST00000469566 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000396281 | ENST00000401030 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000396281 | ENST00000469566 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000446994 | ENST00000401030 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000446994 | ENST00000469566 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000458360 | ENST00000401030 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000458360 | ENST00000469566 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000461685 | ENST00000401030 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000461685 | ENST00000469566 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000471951 | ENST00000401030 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000471951 | ENST00000469566 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000536340 | ENST00000401030 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000536340 | ENST00000469566 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000540497 | ENST00000401030 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-5UTR | ENST00000540497 | ENST00000469566 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-intron | ENST00000262975 | ENST00000370554 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-intron | ENST00000311275 | ENST00000370554 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-intron | ENST00000352431 | ENST00000370554 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-intron | ENST00000355972 | ENST00000370554 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-intron | ENST00000360911 | ENST00000370554 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-intron | ENST00000372023 | ENST00000370554 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-intron | ENST00000396281 | ENST00000370554 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-intron | ENST00000446994 | ENST00000370554 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-intron | ENST00000458360 | ENST00000370554 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-intron | ENST00000461685 | ENST00000370554 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-intron | ENST00000471951 | ENST00000370554 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-intron | ENST00000536340 | ENST00000370554 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5CDS-intron | ENST00000540497 | ENST00000370554 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5UTR-3CDS | ENST00000468376 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5UTR-5UTR | ENST00000468376 | ENST00000401030 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5UTR-5UTR | ENST00000468376 | ENST00000469566 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| 5UTR-intron | ENST00000468376 | ENST00000370554 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| Frame-shift | ENST00000262975 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| Frame-shift | ENST00000311275 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| Frame-shift | ENST00000355972 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| Frame-shift | ENST00000446994 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| In-frame | ENST00000352431 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| In-frame | ENST00000360911 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| In-frame | ENST00000372023 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| In-frame | ENST00000396281 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| In-frame | ENST00000458360 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| In-frame | ENST00000461685 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| In-frame | ENST00000471951 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| In-frame | ENST00000536340 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
| In-frame | ENST00000540497 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000360911 | ZMYND8 | chr20 | 45918930 | - | ENST00000331835 | SEP15 | chr1 | 87333785 | - | 1916 | 709 | 36 | 890 | 284 |
| ENST00000458360 | ZMYND8 | chr20 | 45918930 | - | ENST00000331835 | SEP15 | chr1 | 87333785 | - | 1930 | 723 | 50 | 904 | 284 |
| ENST00000471951 | ZMYND8 | chr20 | 45918930 | - | ENST00000331835 | SEP15 | chr1 | 87333785 | - | 1982 | 775 | 27 | 956 | 309 |
| ENST00000352431 | ZMYND8 | chr20 | 45918930 | - | ENST00000331835 | SEP15 | chr1 | 87333785 | - | 1984 | 777 | 29 | 958 | 309 |
| ENST00000396281 | ZMYND8 | chr20 | 45918930 | - | ENST00000331835 | SEP15 | chr1 | 87333785 | - | 2108 | 901 | 213 | 1082 | 289 |
| ENST00000536340 | ZMYND8 | chr20 | 45918930 | - | ENST00000331835 | SEP15 | chr1 | 87333785 | - | 2058 | 851 | 31 | 1032 | 333 |
| ENST00000540497 | ZMYND8 | chr20 | 45918930 | - | ENST00000331835 | SEP15 | chr1 | 87333785 | - | 1880 | 673 | 0 | 854 | 284 |
| ENST00000461685 | ZMYND8 | chr20 | 45918930 | - | ENST00000331835 | SEP15 | chr1 | 87333785 | - | 1955 | 748 | 0 | 929 | 309 |
| ENST00000372023 | ZMYND8 | chr20 | 45918930 | - | ENST00000331835 | SEP15 | chr1 | 87333785 | - | 1880 | 673 | 0 | 854 | 284 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000360911 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - | 0.000280707 | 0.9997193 |
| ENST00000458360 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - | 0.000271205 | 0.99972874 |
| ENST00000471951 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - | 0.000320227 | 0.9996798 |
| ENST00000352431 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - | 0.000307591 | 0.99969244 |
| ENST00000396281 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - | 0.001272315 | 0.9987276 |
| ENST00000536340 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - | 0.000183697 | 0.99981636 |
| ENST00000540497 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - | 0.000300054 | 0.99969995 |
| ENST00000461685 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - | 0.000304498 | 0.99969554 |
| ENST00000372023 | ENST00000331835 | ZMYND8 | chr20 | 45918930 | - | SEP15 | chr1 | 87333785 | - | 0.000300054 | 0.99969995 |
Top |
Fusion Genomic Features for ZMYND8-SEP15 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
 |
Top |
Fusion Protein Features for ZMYND8-SEP15 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr20:45918930/chr1:87333785) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000262975 | - | 8 | 24 | 43_47 | 229 | 1169.0 | Compositional bias | Note=Poly-Lys |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000311275 | - | 7 | 22 | 43_47 | 229 | 1187.0 | Compositional bias | Note=Poly-Lys |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000352431 | - | 7 | 22 | 43_47 | 249 | 1161.0 | Compositional bias | Note=Poly-Lys |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000355972 | - | 8 | 24 | 43_47 | 229 | 1215.0 | Compositional bias | Note=Poly-Lys |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000360911 | - | 7 | 22 | 43_47 | 224 | 1136.0 | Compositional bias | Note=Poly-Lys |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000461685 | - | 7 | 23 | 43_47 | 249 | 1189.0 | Compositional bias | Note=Poly-Lys |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000471951 | - | 7 | 23 | 43_47 | 249 | 1235.0 | Compositional bias | Note=Poly-Lys |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000352431 | - | 7 | 22 | 165_235 | 249 | 1161.0 | Domain | Bromo |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000461685 | - | 7 | 23 | 165_235 | 249 | 1189.0 | Domain | Bromo |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000471951 | - | 7 | 23 | 165_235 | 249 | 1235.0 | Domain | Bromo |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000262975 | - | 8 | 24 | 88_133 | 229 | 1169.0 | Zinc finger | PHD-type |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000311275 | - | 7 | 22 | 88_133 | 229 | 1187.0 | Zinc finger | PHD-type |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000352431 | - | 7 | 22 | 88_133 | 249 | 1161.0 | Zinc finger | PHD-type |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000355972 | - | 8 | 24 | 88_133 | 229 | 1215.0 | Zinc finger | PHD-type |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000360911 | - | 7 | 22 | 88_133 | 224 | 1136.0 | Zinc finger | PHD-type |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000461685 | - | 7 | 23 | 88_133 | 249 | 1189.0 | Zinc finger | PHD-type |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000471951 | - | 7 | 23 | 88_133 | 249 | 1235.0 | Zinc finger | PHD-type |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000262975 | - | 8 | 24 | 1089_1092 | 229 | 1169.0 | Compositional bias | Note=Poly-Ser |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000262975 | - | 8 | 24 | 838_854 | 229 | 1169.0 | Compositional bias | Note=Poly-Gln |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000311275 | - | 7 | 22 | 1089_1092 | 229 | 1187.0 | Compositional bias | Note=Poly-Ser |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000311275 | - | 7 | 22 | 838_854 | 229 | 1187.0 | Compositional bias | Note=Poly-Gln |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000352431 | - | 7 | 22 | 1089_1092 | 249 | 1161.0 | Compositional bias | Note=Poly-Ser |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000352431 | - | 7 | 22 | 838_854 | 249 | 1161.0 | Compositional bias | Note=Poly-Gln |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000355972 | - | 8 | 24 | 1089_1092 | 229 | 1215.0 | Compositional bias | Note=Poly-Ser |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000355972 | - | 8 | 24 | 838_854 | 229 | 1215.0 | Compositional bias | Note=Poly-Gln |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000360911 | - | 7 | 22 | 1089_1092 | 224 | 1136.0 | Compositional bias | Note=Poly-Ser |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000360911 | - | 7 | 22 | 838_854 | 224 | 1136.0 | Compositional bias | Note=Poly-Gln |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000461685 | - | 7 | 23 | 1089_1092 | 249 | 1189.0 | Compositional bias | Note=Poly-Ser |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000461685 | - | 7 | 23 | 838_854 | 249 | 1189.0 | Compositional bias | Note=Poly-Gln |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000471951 | - | 7 | 23 | 1089_1092 | 249 | 1235.0 | Compositional bias | Note=Poly-Ser |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000471951 | - | 7 | 23 | 838_854 | 249 | 1235.0 | Compositional bias | Note=Poly-Gln |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000262975 | - | 8 | 24 | 165_235 | 229 | 1169.0 | Domain | Bromo |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000262975 | - | 8 | 24 | 277_327 | 229 | 1169.0 | Domain | PWWP |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000311275 | - | 7 | 22 | 165_235 | 229 | 1187.0 | Domain | Bromo |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000311275 | - | 7 | 22 | 277_327 | 229 | 1187.0 | Domain | PWWP |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000352431 | - | 7 | 22 | 277_327 | 249 | 1161.0 | Domain | PWWP |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000355972 | - | 8 | 24 | 165_235 | 229 | 1215.0 | Domain | Bromo |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000355972 | - | 8 | 24 | 277_327 | 229 | 1215.0 | Domain | PWWP |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000360911 | - | 7 | 22 | 165_235 | 224 | 1136.0 | Domain | Bromo |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000360911 | - | 7 | 22 | 277_327 | 224 | 1136.0 | Domain | PWWP |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000461685 | - | 7 | 23 | 277_327 | 249 | 1189.0 | Domain | PWWP |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000471951 | - | 7 | 23 | 277_327 | 249 | 1235.0 | Domain | PWWP |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000262975 | - | 8 | 24 | 1028_1062 | 229 | 1169.0 | Zinc finger | MYND-type |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000311275 | - | 7 | 22 | 1028_1062 | 229 | 1187.0 | Zinc finger | MYND-type |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000352431 | - | 7 | 22 | 1028_1062 | 249 | 1161.0 | Zinc finger | MYND-type |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000355972 | - | 8 | 24 | 1028_1062 | 229 | 1215.0 | Zinc finger | MYND-type |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000360911 | - | 7 | 22 | 1028_1062 | 224 | 1136.0 | Zinc finger | MYND-type |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000461685 | - | 7 | 23 | 1028_1062 | 249 | 1189.0 | Zinc finger | MYND-type |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000471951 | - | 7 | 23 | 1028_1062 | 249 | 1235.0 | Zinc finger | MYND-type |
Top |
Fusion Gene Sequence for ZMYND8-SEP15 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >101415_101415_1_ZMYND8-SEP15_ZMYND8_chr20_45918930_ENST00000352431_SEP15_chr1_87333785_ENST00000331835_length(transcript)=1984nt_BP=777nt ATAAGTATTGACATTACACAGTGTTAACAATGCATCCACAGAGCTTGGCTGAAGAGGAAATAAAAACAGAACAGGAGGTGGTAGAGGGCA TGGATATCTCTACTCGCTCCAAAGATCCTGGCTCTGCAGAGAGAACAGCCCAGAAAAGAAAGTTCCCCAGCCCTCCACATTCTTCCAATG GCCACTCGCCGCAGGACACATCAACAAGCCCCATTAAAAAGAAAAAGAAACCTGGCTTACTGAACAGTAACAATAAGGAGCAGTCAGAAC TAAGACATGGTCCGTTTTACTATATGAAGCAGCCACTCACCACAGACCCTGTTGATGTTGTACCGCAGGATGGACGGAATGATTTCTACT GCTGGGTTTGTCACCGGGAAGGCCAAGTCCTTTGCTGTGAGCTCTGTCCCCGGGTTTATCACGCTAAGTGTCTGAGACTGACATCGGAAC CAGAGGGGGACTGGTTTTGTCCTGAATGTGAGAAAATTACAGTAGCAGAATGCATCGAGACCCAGAGTAAAGCCATGACAATGCTCACCA TTGAACAGTTATCCTACCTGCTCAAGTTTGCCATTCAGAAAATGAAACAGCCAGGGACAGATGCATTCCAGAAGCCCGTTCCATTGGAAC AGCACCCTGACTATGCGGAATACATCTTCCATCCAATGGACCTTTGTACATTGGAAAAGAATGCGAAAAAGAAAATGTATGGCTGCACAG AAGCCTTCCTGGCTGATGCAAAGTGGATTTTGCACAACTGCATCATTTATAATGGGGCTTTTGTTAGGAGTGATAAACCCAAACTGTTCA GAGGACTGCAAATCAAGTATGTCCGTGGTTCAGACCCTGTATTAAAGCTTTTGGACGACAATGGGAACATTGCTGAAGAACTGAGCATTC TCAAATGGAACACAGACAGTGTAGAAGAATTCCTGAGTGAAAAGTTGGAACGCATATAAATCTTGCTTAAATTTTGTCCTATCCTTTTGT TACCTTATCAAATGAAATATTACAGCACCTAGAAAATAATTTAGTTTTGCTTGCTTCCATTGATCAGTCTTTTACTTGAGGCATTAAATA TCTAATTAAATCGTGAAATGGCAGTATAGTCCATGATATCTAAGGAGTTGGCAAGCTTAACAAAACCCATTTTTTATAAATGTCCATCCT CCTGCATTTGTTGATACCACTAACAAAATGCTTTGTAACAGACTTGCGGTTAATTATGCAAATGATAGTTTGTGATAATTGGTCCAGTTT TACGAACAACAGATTTCTAAATTAGAGAGGTTAACAAGACAGATGATTACTATGCCTCATGTGCTGTGTGCTCTTTGAAAGGAATGACAG CAGACTACAAAGCAAATAAGATATACTGAGCCTCAACAGATTGCCTGCTCCTCAGAGTCTCTCCTATTTTTGTATTACCCAGCTTTCTTT TTAATACAAATGTTATTTATAGTTTACAATGAATGCACTGCATAAAAACTTTGTAGCTTCATTATTGTAAAACATATTCAAGATCCTACA GTAAGAGTGAAACATTCACAAAGATTTGCGTTAATGAAGACTACACAGAAAACCTTTCTAGGGATTTGTGTGGATCAGATACATACTTGG CAAATTTTTGAGTTTTACATTCTTACAGAAAAGTCCATTTAAAAGTGATCATTTGTAAGACCAAAATATAAATAAAAAGTTTCAAAAATC TATCTGAATTTGGAATTCTTCTGGTTTGTTCTTTCATGTTTAAAAATGATGTTTTTCAATGCATTTTTTTCATGTAAGCCCTTTTTTTAG CCAAAATGTAAAAATGGCTGTAATATTTAAAACTTATAACATCTTATTGTTGGTAATAGTGCTTTATATTTGTCTGATTTTATTTTTCAA AGTTTTTTCATTTATGAACACATTTTCATTGGTATATTATTTAAGGAATATCTCTTGATATAGAATTTTTATATTAAAAATGATTTTTCT >101415_101415_1_ZMYND8-SEP15_ZMYND8_chr20_45918930_ENST00000352431_SEP15_chr1_87333785_ENST00000331835_length(amino acids)=309AA_BP=249 MHPQSLAEEEIKTEQEVVEGMDISTRSKDPGSAERTAQKRKFPSPPHSSNGHSPQDTSTSPIKKKKKPGLLNSNNKEQSELRHGPFYYMK QPLTTDPVDVVPQDGRNDFYCWVCHREGQVLCCELCPRVYHAKCLRLTSEPEGDWFCPECEKITVAECIETQSKAMTMLTIEQLSYLLKF AIQKMKQPGTDAFQKPVPLEQHPDYAEYIFHPMDLCTLEKNAKKKMYGCTEAFLADAKWILHNCIIYNGAFVRSDKPKLFRGLQIKYVRG -------------------------------------------------------------- >101415_101415_2_ZMYND8-SEP15_ZMYND8_chr20_45918930_ENST00000360911_SEP15_chr1_87333785_ENST00000331835_length(transcript)=1916nt_BP=709nt ATGTTTTATAAGTATTGACATTACACAGTGTTAACAATGCATCCACAGAGCTTGGCTGAAGAGGAAATAAAAACAGAACAGGAGGTGGTA GAGGGCATGGATATCTCTACTCGCTCCAAAGATCCTGGCTCTGCAGAGAGAACAGCCCAGAAAAGAAAGTTCCCCAGCCCTCCACATTCT TCCAATGGCCACTCGCCGCAGGACACATCAACAAGCCCCATTAAAAAGAAAAAGAAACCTGGCTTACTGAACAGTAACAATAAGGAGCAG GATGGACGGAATGATTTCTACTGCTGGGTTTGTCACCGGGAAGGCCAAGTCCTTTGCTGTGAGCTCTGTCCCCGGGTTTATCACGCTAAG TGTCTGAGACTGACATCGGAACCAGAGGGGGACTGGTTTTGTCCTGAATGTGAGAAAATTACAGTAGCAGAATGCATCGAGACCCAGAGT AAAGCCATGACAATGCTCACCATTGAACAGTTATCCTACCTGCTCAAGTTTGCCATTCAGAAAATGAAACAGCCAGGGACAGATGCATTC CAGAAGCCCGTTCCATTGGAACAGCACCCTGACTATGCGGAATACATCTTCCATCCAATGGACCTTTGTACATTGGAAAAGAATGCGAAA AAGAAAATGTATGGCTGCACAGAAGCCTTCCTGGCTGATGCAAAGTGGATTTTGCACAACTGCATCATTTATAATGGGGCTTTTGTTAGG AGTGATAAACCCAAACTGTTCAGAGGACTGCAAATCAAGTATGTCCGTGGTTCAGACCCTGTATTAAAGCTTTTGGACGACAATGGGAAC ATTGCTGAAGAACTGAGCATTCTCAAATGGAACACAGACAGTGTAGAAGAATTCCTGAGTGAAAAGTTGGAACGCATATAAATCTTGCTT AAATTTTGTCCTATCCTTTTGTTACCTTATCAAATGAAATATTACAGCACCTAGAAAATAATTTAGTTTTGCTTGCTTCCATTGATCAGT CTTTTACTTGAGGCATTAAATATCTAATTAAATCGTGAAATGGCAGTATAGTCCATGATATCTAAGGAGTTGGCAAGCTTAACAAAACCC ATTTTTTATAAATGTCCATCCTCCTGCATTTGTTGATACCACTAACAAAATGCTTTGTAACAGACTTGCGGTTAATTATGCAAATGATAG TTTGTGATAATTGGTCCAGTTTTACGAACAACAGATTTCTAAATTAGAGAGGTTAACAAGACAGATGATTACTATGCCTCATGTGCTGTG TGCTCTTTGAAAGGAATGACAGCAGACTACAAAGCAAATAAGATATACTGAGCCTCAACAGATTGCCTGCTCCTCAGAGTCTCTCCTATT TTTGTATTACCCAGCTTTCTTTTTAATACAAATGTTATTTATAGTTTACAATGAATGCACTGCATAAAAACTTTGTAGCTTCATTATTGT AAAACATATTCAAGATCCTACAGTAAGAGTGAAACATTCACAAAGATTTGCGTTAATGAAGACTACACAGAAAACCTTTCTAGGGATTTG TGTGGATCAGATACATACTTGGCAAATTTTTGAGTTTTACATTCTTACAGAAAAGTCCATTTAAAAGTGATCATTTGTAAGACCAAAATA TAAATAAAAAGTTTCAAAAATCTATCTGAATTTGGAATTCTTCTGGTTTGTTCTTTCATGTTTAAAAATGATGTTTTTCAATGCATTTTT TTCATGTAAGCCCTTTTTTTAGCCAAAATGTAAAAATGGCTGTAATATTTAAAACTTATAACATCTTATTGTTGGTAATAGTGCTTTATA TTTGTCTGATTTTATTTTTCAAAGTTTTTTCATTTATGAACACATTTTCATTGGTATATTATTTAAGGAATATCTCTTGATATAGAATTT >101415_101415_2_ZMYND8-SEP15_ZMYND8_chr20_45918930_ENST00000360911_SEP15_chr1_87333785_ENST00000331835_length(amino acids)=284AA_BP=224 MHPQSLAEEEIKTEQEVVEGMDISTRSKDPGSAERTAQKRKFPSPPHSSNGHSPQDTSTSPIKKKKKPGLLNSNNKEQDGRNDFYCWVCH REGQVLCCELCPRVYHAKCLRLTSEPEGDWFCPECEKITVAECIETQSKAMTMLTIEQLSYLLKFAIQKMKQPGTDAFQKPVPLEQHPDY AEYIFHPMDLCTLEKNAKKKMYGCTEAFLADAKWILHNCIIYNGAFVRSDKPKLFRGLQIKYVRGSDPVLKLLDDNGNIAEELSILKWNT -------------------------------------------------------------- >101415_101415_3_ZMYND8-SEP15_ZMYND8_chr20_45918930_ENST00000372023_SEP15_chr1_87333785_ENST00000331835_length(transcript)=1880nt_BP=673nt ATGCATCCACAGAGCTTGGCTGAAGAGGAAATAAAAACAGAACAGGAGGTGGTAGAGGGCATGGATATCTCTACTCGCTCCAAAGATCCT GGCTCTGCAGAGAGAACAGCCCAGAAAAGAAAGTTCCCCAGCCCTCCACATTCTTCCAATGGCCACTCGCCGCAGGACACATCAACAAGC CCCATTAAAAAGAAAAAGAAACCTGGCTTACTGAACAGTAACAATAAGGAGCAGGATGGACGGAATGATTTCTACTGCTGGGTTTGTCAC CGGGAAGGCCAAGTCCTTTGCTGTGAGCTCTGTCCCCGGGTTTATCACGCTAAGTGTCTGAGACTGACATCGGAACCAGAGGGGGACTGG TTTTGTCCTGAATGTGAGAAAATTACAGTAGCAGAATGCATCGAGACCCAGAGTAAAGCCATGACAATGCTCACCATTGAACAGTTATCC TACCTGCTCAAGTTTGCCATTCAGAAAATGAAACAGCCAGGGACAGATGCATTCCAGAAGCCCGTTCCATTGGAACAGCACCCTGACTAT GCGGAATACATCTTCCATCCAATGGACCTTTGTACATTGGAAAAGAATGCGAAAAAGAAAATGTATGGCTGCACAGAAGCCTTCCTGGCT GATGCAAAGTGGATTTTGCACAACTGCATCATTTATAATGGGGCTTTTGTTAGGAGTGATAAACCCAAACTGTTCAGAGGACTGCAAATC AAGTATGTCCGTGGTTCAGACCCTGTATTAAAGCTTTTGGACGACAATGGGAACATTGCTGAAGAACTGAGCATTCTCAAATGGAACACA GACAGTGTAGAAGAATTCCTGAGTGAAAAGTTGGAACGCATATAAATCTTGCTTAAATTTTGTCCTATCCTTTTGTTACCTTATCAAATG AAATATTACAGCACCTAGAAAATAATTTAGTTTTGCTTGCTTCCATTGATCAGTCTTTTACTTGAGGCATTAAATATCTAATTAAATCGT GAAATGGCAGTATAGTCCATGATATCTAAGGAGTTGGCAAGCTTAACAAAACCCATTTTTTATAAATGTCCATCCTCCTGCATTTGTTGA TACCACTAACAAAATGCTTTGTAACAGACTTGCGGTTAATTATGCAAATGATAGTTTGTGATAATTGGTCCAGTTTTACGAACAACAGAT TTCTAAATTAGAGAGGTTAACAAGACAGATGATTACTATGCCTCATGTGCTGTGTGCTCTTTGAAAGGAATGACAGCAGACTACAAAGCA AATAAGATATACTGAGCCTCAACAGATTGCCTGCTCCTCAGAGTCTCTCCTATTTTTGTATTACCCAGCTTTCTTTTTAATACAAATGTT ATTTATAGTTTACAATGAATGCACTGCATAAAAACTTTGTAGCTTCATTATTGTAAAACATATTCAAGATCCTACAGTAAGAGTGAAACA TTCACAAAGATTTGCGTTAATGAAGACTACACAGAAAACCTTTCTAGGGATTTGTGTGGATCAGATACATACTTGGCAAATTTTTGAGTT TTACATTCTTACAGAAAAGTCCATTTAAAAGTGATCATTTGTAAGACCAAAATATAAATAAAAAGTTTCAAAAATCTATCTGAATTTGGA ATTCTTCTGGTTTGTTCTTTCATGTTTAAAAATGATGTTTTTCAATGCATTTTTTTCATGTAAGCCCTTTTTTTAGCCAAAATGTAAAAA TGGCTGTAATATTTAAAACTTATAACATCTTATTGTTGGTAATAGTGCTTTATATTTGTCTGATTTTATTTTTCAAAGTTTTTTCATTTA >101415_101415_3_ZMYND8-SEP15_ZMYND8_chr20_45918930_ENST00000372023_SEP15_chr1_87333785_ENST00000331835_length(amino acids)=284AA_BP=224 MHPQSLAEEEIKTEQEVVEGMDISTRSKDPGSAERTAQKRKFPSPPHSSNGHSPQDTSTSPIKKKKKPGLLNSNNKEQDGRNDFYCWVCH REGQVLCCELCPRVYHAKCLRLTSEPEGDWFCPECEKITVAECIETQSKAMTMLTIEQLSYLLKFAIQKMKQPGTDAFQKPVPLEQHPDY AEYIFHPMDLCTLEKNAKKKMYGCTEAFLADAKWILHNCIIYNGAFVRSDKPKLFRGLQIKYVRGSDPVLKLLDDNGNIAEELSILKWNT -------------------------------------------------------------- >101415_101415_4_ZMYND8-SEP15_ZMYND8_chr20_45918930_ENST00000396281_SEP15_chr1_87333785_ENST00000331835_length(transcript)=2108nt_BP=901nt ATAAGTATTGACATTACACAGTGTTAACAATGCATCCACAGAGTCCCAAATCTGCCAGCCTGTTCTCTCTTCCTAACTGTGGAAAAATGA GACGGTTGCTTGAACATTTGTTGGCAGCATTCCTGCCTGAAGTATTCAGCTCCTCTAGGCTGTTACAAGCCAAATAATGGTTCCCTTGAA TGCAGCCTTCCTCCATGATCAATACACCTTTGAATGTTTTCATCGCTGCAGAAATCCTTCAACTTGGCTGAAGAGGAAATAAAAACAGAA CAGGAGGTGGTAGAGGGCATGGATATCTCTACTCGCTCCAAAGATCCTGGCTCTGCAGAGAGAACAGCCCAGAAAAGAAAGTTCCCCAGC CCTCCACATTCTTCCAATGGCCACTCGCCGCAGGACACATCAACAAGCCCCATTAAAAAGAAAAAGAAACCTGGCTTACTGAACAGTAAC AATAAGGAGCAGGATGGACGGAATGATTTCTACTGCTGGGTTTGTCACCGGGAAGGCCAAGTCCTTTGCTGTGAGCTCTGTCCCCGGGTT TATCACGCTAAGTGTCTGAGACTGACATCGGAACCAGAGGGGGACTGGTTTTGTCCTGAATGTGAGAAAATTACAGTAGCAGAATGCATC GAGACCCAGAGTAAAGCCATGACAATGCTCACCATTGAACAGTTATCCTACCTGCTCAAGTTTGCCATTCAGAAAATGAAACAGCCAGGG ACAGATGCATTCCAGAAGCCCGTTCCATTGGAACAGCACCCTGACTATGCGGAATACATCTTCCATCCAATGGACCTTTGTACATTGGAA AAGAATGCGAAAAAGAAAATGTATGGCTGCACAGAAGCCTTCCTGGCTGATGCAAAGTGGATTTTGCACAACTGCATCATTTATAATGGG GCTTTTGTTAGGAGTGATAAACCCAAACTGTTCAGAGGACTGCAAATCAAGTATGTCCGTGGTTCAGACCCTGTATTAAAGCTTTTGGAC GACAATGGGAACATTGCTGAAGAACTGAGCATTCTCAAATGGAACACAGACAGTGTAGAAGAATTCCTGAGTGAAAAGTTGGAACGCATA TAAATCTTGCTTAAATTTTGTCCTATCCTTTTGTTACCTTATCAAATGAAATATTACAGCACCTAGAAAATAATTTAGTTTTGCTTGCTT CCATTGATCAGTCTTTTACTTGAGGCATTAAATATCTAATTAAATCGTGAAATGGCAGTATAGTCCATGATATCTAAGGAGTTGGCAAGC TTAACAAAACCCATTTTTTATAAATGTCCATCCTCCTGCATTTGTTGATACCACTAACAAAATGCTTTGTAACAGACTTGCGGTTAATTA TGCAAATGATAGTTTGTGATAATTGGTCCAGTTTTACGAACAACAGATTTCTAAATTAGAGAGGTTAACAAGACAGATGATTACTATGCC TCATGTGCTGTGTGCTCTTTGAAAGGAATGACAGCAGACTACAAAGCAAATAAGATATACTGAGCCTCAACAGATTGCCTGCTCCTCAGA GTCTCTCCTATTTTTGTATTACCCAGCTTTCTTTTTAATACAAATGTTATTTATAGTTTACAATGAATGCACTGCATAAAAACTTTGTAG CTTCATTATTGTAAAACATATTCAAGATCCTACAGTAAGAGTGAAACATTCACAAAGATTTGCGTTAATGAAGACTACACAGAAAACCTT TCTAGGGATTTGTGTGGATCAGATACATACTTGGCAAATTTTTGAGTTTTACATTCTTACAGAAAAGTCCATTTAAAAGTGATCATTTGT AAGACCAAAATATAAATAAAAAGTTTCAAAAATCTATCTGAATTTGGAATTCTTCTGGTTTGTTCTTTCATGTTTAAAAATGATGTTTTT CAATGCATTTTTTTCATGTAAGCCCTTTTTTTAGCCAAAATGTAAAAATGGCTGTAATATTTAAAACTTATAACATCTTATTGTTGGTAA TAGTGCTTTATATTTGTCTGATTTTATTTTTCAAAGTTTTTTCATTTATGAACACATTTTCATTGGTATATTATTTAAGGAATATCTCTT >101415_101415_4_ZMYND8-SEP15_ZMYND8_chr20_45918930_ENST00000396281_SEP15_chr1_87333785_ENST00000331835_length(amino acids)=289AA_BP=229 MFSSLQKSFNLAEEEIKTEQEVVEGMDISTRSKDPGSAERTAQKRKFPSPPHSSNGHSPQDTSTSPIKKKKKPGLLNSNNKEQDGRNDFY CWVCHREGQVLCCELCPRVYHAKCLRLTSEPEGDWFCPECEKITVAECIETQSKAMTMLTIEQLSYLLKFAIQKMKQPGTDAFQKPVPLE QHPDYAEYIFHPMDLCTLEKNAKKKMYGCTEAFLADAKWILHNCIIYNGAFVRSDKPKLFRGLQIKYVRGSDPVLKLLDDNGNIAEELSI -------------------------------------------------------------- >101415_101415_5_ZMYND8-SEP15_ZMYND8_chr20_45918930_ENST00000458360_SEP15_chr1_87333785_ENST00000331835_length(transcript)=1930nt_BP=723nt GGAGCCTGTCCTCCATGTTTTATAAGTATTGACATTACACAGTGTTAACAATGCATCCACAGAGCTTGGCTGAAGAGGAAATAAAAACAG AACAGGAGGTGGTAGAGGGCATGGATATCTCTACTCGCTCCAAAGATCCTGGCTCTGCAGAGAGAACAGCCCAGAAAAGAAAGTTCCCCA GCCCTCCACATTCTTCCAATGGCCACTCGCCGCAGGACACATCAACAAGCCCCATTAAAAAGAAAAAGAAACCTGGCTTACTGAACAGTA ACAATAAGGAGCAGGATGGACGGAATGATTTCTACTGCTGGGTTTGTCACCGGGAAGGCCAAGTCCTTTGCTGTGAGCTCTGTCCCCGGG TTTATCACGCTAAGTGTCTGAGACTGACATCGGAACCAGAGGGGGACTGGTTTTGTCCTGAATGTGAGAAAATTACAGTAGCAGAATGCA TCGAGACCCAGAGTAAAGCCATGACAATGCTCACCATTGAACAGTTATCCTACCTGCTCAAGTTTGCCATTCAGAAAATGAAACAGCCAG GGACAGATGCATTCCAGAAGCCCGTTCCATTGGAACAGCACCCTGACTATGCGGAATACATCTTCCATCCAATGGACCTTTGTACATTGG AAAAGAATGCGAAAAAGAAAATGTATGGCTGCACAGAAGCCTTCCTGGCTGATGCAAAGTGGATTTTGCACAACTGCATCATTTATAATG GGGCTTTTGTTAGGAGTGATAAACCCAAACTGTTCAGAGGACTGCAAATCAAGTATGTCCGTGGTTCAGACCCTGTATTAAAGCTTTTGG ACGACAATGGGAACATTGCTGAAGAACTGAGCATTCTCAAATGGAACACAGACAGTGTAGAAGAATTCCTGAGTGAAAAGTTGGAACGCA TATAAATCTTGCTTAAATTTTGTCCTATCCTTTTGTTACCTTATCAAATGAAATATTACAGCACCTAGAAAATAATTTAGTTTTGCTTGC TTCCATTGATCAGTCTTTTACTTGAGGCATTAAATATCTAATTAAATCGTGAAATGGCAGTATAGTCCATGATATCTAAGGAGTTGGCAA GCTTAACAAAACCCATTTTTTATAAATGTCCATCCTCCTGCATTTGTTGATACCACTAACAAAATGCTTTGTAACAGACTTGCGGTTAAT TATGCAAATGATAGTTTGTGATAATTGGTCCAGTTTTACGAACAACAGATTTCTAAATTAGAGAGGTTAACAAGACAGATGATTACTATG CCTCATGTGCTGTGTGCTCTTTGAAAGGAATGACAGCAGACTACAAAGCAAATAAGATATACTGAGCCTCAACAGATTGCCTGCTCCTCA GAGTCTCTCCTATTTTTGTATTACCCAGCTTTCTTTTTAATACAAATGTTATTTATAGTTTACAATGAATGCACTGCATAAAAACTTTGT AGCTTCATTATTGTAAAACATATTCAAGATCCTACAGTAAGAGTGAAACATTCACAAAGATTTGCGTTAATGAAGACTACACAGAAAACC TTTCTAGGGATTTGTGTGGATCAGATACATACTTGGCAAATTTTTGAGTTTTACATTCTTACAGAAAAGTCCATTTAAAAGTGATCATTT GTAAGACCAAAATATAAATAAAAAGTTTCAAAAATCTATCTGAATTTGGAATTCTTCTGGTTTGTTCTTTCATGTTTAAAAATGATGTTT TTCAATGCATTTTTTTCATGTAAGCCCTTTTTTTAGCCAAAATGTAAAAATGGCTGTAATATTTAAAACTTATAACATCTTATTGTTGGT AATAGTGCTTTATATTTGTCTGATTTTATTTTTCAAAGTTTTTTCATTTATGAACACATTTTCATTGGTATATTATTTAAGGAATATCTC >101415_101415_5_ZMYND8-SEP15_ZMYND8_chr20_45918930_ENST00000458360_SEP15_chr1_87333785_ENST00000331835_length(amino acids)=284AA_BP=224 MHPQSLAEEEIKTEQEVVEGMDISTRSKDPGSAERTAQKRKFPSPPHSSNGHSPQDTSTSPIKKKKKPGLLNSNNKEQDGRNDFYCWVCH REGQVLCCELCPRVYHAKCLRLTSEPEGDWFCPECEKITVAECIETQSKAMTMLTIEQLSYLLKFAIQKMKQPGTDAFQKPVPLEQHPDY AEYIFHPMDLCTLEKNAKKKMYGCTEAFLADAKWILHNCIIYNGAFVRSDKPKLFRGLQIKYVRGSDPVLKLLDDNGNIAEELSILKWNT -------------------------------------------------------------- >101415_101415_6_ZMYND8-SEP15_ZMYND8_chr20_45918930_ENST00000461685_SEP15_chr1_87333785_ENST00000331835_length(transcript)=1955nt_BP=748nt ATGCATCCACAGAGCTTGGCTGAAGAGGAAATAAAAACAGAACAGGAGGTGGTAGAGGGCATGGATATCTCTACTCGCTCCAAAGATCCT GGCTCTGCAGAGAGAACAGCCCAGAAAAGAAAGTTCCCCAGCCCTCCACATTCTTCCAATGGCCACTCGCCGCAGGACACATCAACAAGC CCCATTAAAAAGAAAAAGAAACCTGGCTTACTGAACAGTAACAATAAGGAGCAGTCAGAACTAAGACATGGTCCGTTTTACTATATGAAG CAGCCACTCACCACAGACCCTGTTGATGTTGTACCGCAGGATGGACGGAATGATTTCTACTGCTGGGTTTGTCACCGGGAAGGCCAAGTC CTTTGCTGTGAGCTCTGTCCCCGGGTTTATCACGCTAAGTGTCTGAGACTGACATCGGAACCAGAGGGGGACTGGTTTTGTCCTGAATGT GAGAAAATTACAGTAGCAGAATGCATCGAGACCCAGAGTAAAGCCATGACAATGCTCACCATTGAACAGTTATCCTACCTGCTCAAGTTT GCCATTCAGAAAATGAAACAGCCAGGGACAGATGCATTCCAGAAGCCCGTTCCATTGGAACAGCACCCTGACTATGCGGAATACATCTTC CATCCAATGGACCTTTGTACATTGGAAAAGAATGCGAAAAAGAAAATGTATGGCTGCACAGAAGCCTTCCTGGCTGATGCAAAGTGGATT TTGCACAACTGCATCATTTATAATGGGGCTTTTGTTAGGAGTGATAAACCCAAACTGTTCAGAGGACTGCAAATCAAGTATGTCCGTGGT TCAGACCCTGTATTAAAGCTTTTGGACGACAATGGGAACATTGCTGAAGAACTGAGCATTCTCAAATGGAACACAGACAGTGTAGAAGAA TTCCTGAGTGAAAAGTTGGAACGCATATAAATCTTGCTTAAATTTTGTCCTATCCTTTTGTTACCTTATCAAATGAAATATTACAGCACC TAGAAAATAATTTAGTTTTGCTTGCTTCCATTGATCAGTCTTTTACTTGAGGCATTAAATATCTAATTAAATCGTGAAATGGCAGTATAG TCCATGATATCTAAGGAGTTGGCAAGCTTAACAAAACCCATTTTTTATAAATGTCCATCCTCCTGCATTTGTTGATACCACTAACAAAAT GCTTTGTAACAGACTTGCGGTTAATTATGCAAATGATAGTTTGTGATAATTGGTCCAGTTTTACGAACAACAGATTTCTAAATTAGAGAG GTTAACAAGACAGATGATTACTATGCCTCATGTGCTGTGTGCTCTTTGAAAGGAATGACAGCAGACTACAAAGCAAATAAGATATACTGA GCCTCAACAGATTGCCTGCTCCTCAGAGTCTCTCCTATTTTTGTATTACCCAGCTTTCTTTTTAATACAAATGTTATTTATAGTTTACAA TGAATGCACTGCATAAAAACTTTGTAGCTTCATTATTGTAAAACATATTCAAGATCCTACAGTAAGAGTGAAACATTCACAAAGATTTGC GTTAATGAAGACTACACAGAAAACCTTTCTAGGGATTTGTGTGGATCAGATACATACTTGGCAAATTTTTGAGTTTTACATTCTTACAGA AAAGTCCATTTAAAAGTGATCATTTGTAAGACCAAAATATAAATAAAAAGTTTCAAAAATCTATCTGAATTTGGAATTCTTCTGGTTTGT TCTTTCATGTTTAAAAATGATGTTTTTCAATGCATTTTTTTCATGTAAGCCCTTTTTTTAGCCAAAATGTAAAAATGGCTGTAATATTTA AAACTTATAACATCTTATTGTTGGTAATAGTGCTTTATATTTGTCTGATTTTATTTTTCAAAGTTTTTTCATTTATGAACACATTTTCAT >101415_101415_6_ZMYND8-SEP15_ZMYND8_chr20_45918930_ENST00000461685_SEP15_chr1_87333785_ENST00000331835_length(amino acids)=309AA_BP=249 MHPQSLAEEEIKTEQEVVEGMDISTRSKDPGSAERTAQKRKFPSPPHSSNGHSPQDTSTSPIKKKKKPGLLNSNNKEQSELRHGPFYYMK QPLTTDPVDVVPQDGRNDFYCWVCHREGQVLCCELCPRVYHAKCLRLTSEPEGDWFCPECEKITVAECIETQSKAMTMLTIEQLSYLLKF AIQKMKQPGTDAFQKPVPLEQHPDYAEYIFHPMDLCTLEKNAKKKMYGCTEAFLADAKWILHNCIIYNGAFVRSDKPKLFRGLQIKYVRG -------------------------------------------------------------- >101415_101415_7_ZMYND8-SEP15_ZMYND8_chr20_45918930_ENST00000471951_SEP15_chr1_87333785_ENST00000331835_length(transcript)=1982nt_BP=775nt AAGTATTGACATTACACAGTGTTAACAATGCATCCACAGAGCTTGGCTGAAGAGGAAATAAAAACAGAACAGGAGGTGGTAGAGGGCATG GATATCTCTACTCGCTCCAAAGATCCTGGCTCTGCAGAGAGAACAGCCCAGAAAAGAAAGTTCCCCAGCCCTCCACATTCTTCCAATGGC CACTCGCCGCAGGACACATCAACAAGCCCCATTAAAAAGAAAAAGAAACCTGGCTTACTGAACAGTAACAATAAGGAGCAGTCAGAACTA AGACATGGTCCGTTTTACTATATGAAGCAGCCACTCACCACAGACCCTGTTGATGTTGTACCGCAGGATGGACGGAATGATTTCTACTGC TGGGTTTGTCACCGGGAAGGCCAAGTCCTTTGCTGTGAGCTCTGTCCCCGGGTTTATCACGCTAAGTGTCTGAGACTGACATCGGAACCA GAGGGGGACTGGTTTTGTCCTGAATGTGAGAAAATTACAGTAGCAGAATGCATCGAGACCCAGAGTAAAGCCATGACAATGCTCACCATT GAACAGTTATCCTACCTGCTCAAGTTTGCCATTCAGAAAATGAAACAGCCAGGGACAGATGCATTCCAGAAGCCCGTTCCATTGGAACAG CACCCTGACTATGCGGAATACATCTTCCATCCAATGGACCTTTGTACATTGGAAAAGAATGCGAAAAAGAAAATGTATGGCTGCACAGAA GCCTTCCTGGCTGATGCAAAGTGGATTTTGCACAACTGCATCATTTATAATGGGGCTTTTGTTAGGAGTGATAAACCCAAACTGTTCAGA GGACTGCAAATCAAGTATGTCCGTGGTTCAGACCCTGTATTAAAGCTTTTGGACGACAATGGGAACATTGCTGAAGAACTGAGCATTCTC AAATGGAACACAGACAGTGTAGAAGAATTCCTGAGTGAAAAGTTGGAACGCATATAAATCTTGCTTAAATTTTGTCCTATCCTTTTGTTA CCTTATCAAATGAAATATTACAGCACCTAGAAAATAATTTAGTTTTGCTTGCTTCCATTGATCAGTCTTTTACTTGAGGCATTAAATATC TAATTAAATCGTGAAATGGCAGTATAGTCCATGATATCTAAGGAGTTGGCAAGCTTAACAAAACCCATTTTTTATAAATGTCCATCCTCC TGCATTTGTTGATACCACTAACAAAATGCTTTGTAACAGACTTGCGGTTAATTATGCAAATGATAGTTTGTGATAATTGGTCCAGTTTTA CGAACAACAGATTTCTAAATTAGAGAGGTTAACAAGACAGATGATTACTATGCCTCATGTGCTGTGTGCTCTTTGAAAGGAATGACAGCA GACTACAAAGCAAATAAGATATACTGAGCCTCAACAGATTGCCTGCTCCTCAGAGTCTCTCCTATTTTTGTATTACCCAGCTTTCTTTTT AATACAAATGTTATTTATAGTTTACAATGAATGCACTGCATAAAAACTTTGTAGCTTCATTATTGTAAAACATATTCAAGATCCTACAGT AAGAGTGAAACATTCACAAAGATTTGCGTTAATGAAGACTACACAGAAAACCTTTCTAGGGATTTGTGTGGATCAGATACATACTTGGCA AATTTTTGAGTTTTACATTCTTACAGAAAAGTCCATTTAAAAGTGATCATTTGTAAGACCAAAATATAAATAAAAAGTTTCAAAAATCTA TCTGAATTTGGAATTCTTCTGGTTTGTTCTTTCATGTTTAAAAATGATGTTTTTCAATGCATTTTTTTCATGTAAGCCCTTTTTTTAGCC AAAATGTAAAAATGGCTGTAATATTTAAAACTTATAACATCTTATTGTTGGTAATAGTGCTTTATATTTGTCTGATTTTATTTTTCAAAG TTTTTTCATTTATGAACACATTTTCATTGGTATATTATTTAAGGAATATCTCTTGATATAGAATTTTTATATTAAAAATGATTTTTCTTT >101415_101415_7_ZMYND8-SEP15_ZMYND8_chr20_45918930_ENST00000471951_SEP15_chr1_87333785_ENST00000331835_length(amino acids)=309AA_BP=249 MHPQSLAEEEIKTEQEVVEGMDISTRSKDPGSAERTAQKRKFPSPPHSSNGHSPQDTSTSPIKKKKKPGLLNSNNKEQSELRHGPFYYMK QPLTTDPVDVVPQDGRNDFYCWVCHREGQVLCCELCPRVYHAKCLRLTSEPEGDWFCPECEKITVAECIETQSKAMTMLTIEQLSYLLKF AIQKMKQPGTDAFQKPVPLEQHPDYAEYIFHPMDLCTLEKNAKKKMYGCTEAFLADAKWILHNCIIYNGAFVRSDKPKLFRGLQIKYVRG -------------------------------------------------------------- >101415_101415_8_ZMYND8-SEP15_ZMYND8_chr20_45918930_ENST00000536340_SEP15_chr1_87333785_ENST00000331835_length(transcript)=2058nt_BP=851nt GTCACATGGCTCAGTAGCCATTAGCCATAGCTTGATTTTGCAAACTGGCAAGGGCCTGGCACACCTTTCCTTCCACCAGGGCATGGTGTT TCTGGAAGAGTTTGAAGCTAGATCCTGCTTGGCTGAAGAGGAAATAAAAACAGAACAGGAGGTGGTAGAGGGCATGGATATCTCTACTCG CTCCAAAGATCCTGGCTCTGCAGAGAGAACAGCCCAGAAAAGAAAGTTCCCCAGCCCTCCACATTCTTCCAATGGCCACTCGCCGCAGGA CACATCAACAAGCCCCATTAAAAAGAAAAAGAAACCTGGCTTACTGAACAGTAACAATAAGGAGCAGTCAGAACTAAGACATGGTCCGTT TTACTATATGAAGCAGCCACTCACCACAGACCCTGTTGATGTTGTACCGCAGGATGGACGGAATGATTTCTACTGCTGGGTTTGTCACCG GGAAGGCCAAGTCCTTTGCTGTGAGCTCTGTCCCCGGGTTTATCACGCTAAGTGTCTGAGACTGACATCGGAACCAGAGGGGGACTGGTT TTGTCCTGAATGTGAGAAAATTACAGTAGCAGAATGCATCGAGACCCAGAGTAAAGCCATGACAATGCTCACCATTGAACAGTTATCCTA CCTGCTCAAGTTTGCCATTCAGAAAATGAAACAGCCAGGGACAGATGCATTCCAGAAGCCCGTTCCATTGGAACAGCACCCTGACTATGC GGAATACATCTTCCATCCAATGGACCTTTGTACATTGGAAAAGAATGCGAAAAAGAAAATGTATGGCTGCACAGAAGCCTTCCTGGCTGA TGCAAAGTGGATTTTGCACAACTGCATCATTTATAATGGGGCTTTTGTTAGGAGTGATAAACCCAAACTGTTCAGAGGACTGCAAATCAA GTATGTCCGTGGTTCAGACCCTGTATTAAAGCTTTTGGACGACAATGGGAACATTGCTGAAGAACTGAGCATTCTCAAATGGAACACAGA CAGTGTAGAAGAATTCCTGAGTGAAAAGTTGGAACGCATATAAATCTTGCTTAAATTTTGTCCTATCCTTTTGTTACCTTATCAAATGAA ATATTACAGCACCTAGAAAATAATTTAGTTTTGCTTGCTTCCATTGATCAGTCTTTTACTTGAGGCATTAAATATCTAATTAAATCGTGA AATGGCAGTATAGTCCATGATATCTAAGGAGTTGGCAAGCTTAACAAAACCCATTTTTTATAAATGTCCATCCTCCTGCATTTGTTGATA CCACTAACAAAATGCTTTGTAACAGACTTGCGGTTAATTATGCAAATGATAGTTTGTGATAATTGGTCCAGTTTTACGAACAACAGATTT CTAAATTAGAGAGGTTAACAAGACAGATGATTACTATGCCTCATGTGCTGTGTGCTCTTTGAAAGGAATGACAGCAGACTACAAAGCAAA TAAGATATACTGAGCCTCAACAGATTGCCTGCTCCTCAGAGTCTCTCCTATTTTTGTATTACCCAGCTTTCTTTTTAATACAAATGTTAT TTATAGTTTACAATGAATGCACTGCATAAAAACTTTGTAGCTTCATTATTGTAAAACATATTCAAGATCCTACAGTAAGAGTGAAACATT CACAAAGATTTGCGTTAATGAAGACTACACAGAAAACCTTTCTAGGGATTTGTGTGGATCAGATACATACTTGGCAAATTTTTGAGTTTT ACATTCTTACAGAAAAGTCCATTTAAAAGTGATCATTTGTAAGACCAAAATATAAATAAAAAGTTTCAAAAATCTATCTGAATTTGGAAT TCTTCTGGTTTGTTCTTTCATGTTTAAAAATGATGTTTTTCAATGCATTTTTTTCATGTAAGCCCTTTTTTTAGCCAAAATGTAAAAATG GCTGTAATATTTAAAACTTATAACATCTTATTGTTGGTAATAGTGCTTTATATTTGTCTGATTTTATTTTTCAAAGTTTTTTCATTTATG >101415_101415_8_ZMYND8-SEP15_ZMYND8_chr20_45918930_ENST00000536340_SEP15_chr1_87333785_ENST00000331835_length(amino acids)=333AA_BP=273 MILQTGKGLAHLSFHQGMVFLEEFEARSCLAEEEIKTEQEVVEGMDISTRSKDPGSAERTAQKRKFPSPPHSSNGHSPQDTSTSPIKKKK KPGLLNSNNKEQSELRHGPFYYMKQPLTTDPVDVVPQDGRNDFYCWVCHREGQVLCCELCPRVYHAKCLRLTSEPEGDWFCPECEKITVA ECIETQSKAMTMLTIEQLSYLLKFAIQKMKQPGTDAFQKPVPLEQHPDYAEYIFHPMDLCTLEKNAKKKMYGCTEAFLADAKWILHNCII -------------------------------------------------------------- >101415_101415_9_ZMYND8-SEP15_ZMYND8_chr20_45918930_ENST00000540497_SEP15_chr1_87333785_ENST00000331835_length(transcript)=1880nt_BP=673nt ATGCATCCACAGAGCTTGGCTGAAGAGGAAATAAAAACAGAACAGGAGGTGGTAGAGGGCATGGATATCTCTACTCGCTCCAAAGATCCT GGCTCTGCAGAGAGAACAGCCCAGAAAAGAAAGTTCCCCAGCCCTCCACATTCTTCCAATGGCCACTCGCCGCAGGACACATCAACAAGC CCCATTAAAAAGAAAAAGAAACCTGGCTTACTGAACAGTAACAATAAGGAGCAGGATGGACGGAATGATTTCTACTGCTGGGTTTGTCAC CGGGAAGGCCAAGTCCTTTGCTGTGAGCTCTGTCCCCGGGTTTATCACGCTAAGTGTCTGAGACTGACATCGGAACCAGAGGGGGACTGG TTTTGTCCTGAATGTGAGAAAATTACAGTAGCAGAATGCATCGAGACCCAGAGTAAAGCCATGACAATGCTCACCATTGAACAGTTATCC TACCTGCTCAAGTTTGCCATTCAGAAAATGAAACAGCCAGGGACAGATGCATTCCAGAAGCCCGTTCCATTGGAACAGCACCCTGACTAT GCGGAATACATCTTCCATCCAATGGACCTTTGTACATTGGAAAAGAATGCGAAAAAGAAAATGTATGGCTGCACAGAAGCCTTCCTGGCT GATGCAAAGTGGATTTTGCACAACTGCATCATTTATAATGGGGCTTTTGTTAGGAGTGATAAACCCAAACTGTTCAGAGGACTGCAAATC AAGTATGTCCGTGGTTCAGACCCTGTATTAAAGCTTTTGGACGACAATGGGAACATTGCTGAAGAACTGAGCATTCTCAAATGGAACACA GACAGTGTAGAAGAATTCCTGAGTGAAAAGTTGGAACGCATATAAATCTTGCTTAAATTTTGTCCTATCCTTTTGTTACCTTATCAAATG AAATATTACAGCACCTAGAAAATAATTTAGTTTTGCTTGCTTCCATTGATCAGTCTTTTACTTGAGGCATTAAATATCTAATTAAATCGT GAAATGGCAGTATAGTCCATGATATCTAAGGAGTTGGCAAGCTTAACAAAACCCATTTTTTATAAATGTCCATCCTCCTGCATTTGTTGA TACCACTAACAAAATGCTTTGTAACAGACTTGCGGTTAATTATGCAAATGATAGTTTGTGATAATTGGTCCAGTTTTACGAACAACAGAT TTCTAAATTAGAGAGGTTAACAAGACAGATGATTACTATGCCTCATGTGCTGTGTGCTCTTTGAAAGGAATGACAGCAGACTACAAAGCA AATAAGATATACTGAGCCTCAACAGATTGCCTGCTCCTCAGAGTCTCTCCTATTTTTGTATTACCCAGCTTTCTTTTTAATACAAATGTT ATTTATAGTTTACAATGAATGCACTGCATAAAAACTTTGTAGCTTCATTATTGTAAAACATATTCAAGATCCTACAGTAAGAGTGAAACA TTCACAAAGATTTGCGTTAATGAAGACTACACAGAAAACCTTTCTAGGGATTTGTGTGGATCAGATACATACTTGGCAAATTTTTGAGTT TTACATTCTTACAGAAAAGTCCATTTAAAAGTGATCATTTGTAAGACCAAAATATAAATAAAAAGTTTCAAAAATCTATCTGAATTTGGA ATTCTTCTGGTTTGTTCTTTCATGTTTAAAAATGATGTTTTTCAATGCATTTTTTTCATGTAAGCCCTTTTTTTAGCCAAAATGTAAAAA TGGCTGTAATATTTAAAACTTATAACATCTTATTGTTGGTAATAGTGCTTTATATTTGTCTGATTTTATTTTTCAAAGTTTTTTCATTTA >101415_101415_9_ZMYND8-SEP15_ZMYND8_chr20_45918930_ENST00000540497_SEP15_chr1_87333785_ENST00000331835_length(amino acids)=284AA_BP=224 MHPQSLAEEEIKTEQEVVEGMDISTRSKDPGSAERTAQKRKFPSPPHSSNGHSPQDTSTSPIKKKKKPGLLNSNNKEQDGRNDFYCWVCH REGQVLCCELCPRVYHAKCLRLTSEPEGDWFCPECEKITVAECIETQSKAMTMLTIEQLSYLLKFAIQKMKQPGTDAFQKPVPLEQHPDY AEYIFHPMDLCTLEKNAKKKMYGCTEAFLADAKWILHNCIIYNGAFVRSDKPKLFRGLQIKYVRGSDPVLKLLDDNGNIAEELSILKWNT -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for ZMYND8-SEP15 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000262975 | - | 8 | 24 | 75_406 | 229.33333333333334 | 1169.0 | histone H3K14ac |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000311275 | - | 7 | 22 | 75_406 | 229.33333333333334 | 1187.0 | histone H3K14ac |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000352431 | - | 7 | 22 | 75_406 | 249.33333333333334 | 1161.0 | histone H3K14ac |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000355972 | - | 8 | 24 | 75_406 | 229.33333333333334 | 1215.0 | histone H3K14ac |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000360911 | - | 7 | 22 | 75_406 | 224.33333333333334 | 1136.0 | histone H3K14ac |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000461685 | - | 7 | 23 | 75_406 | 249.33333333333334 | 1189.0 | histone H3K14ac |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000471951 | - | 7 | 23 | 75_406 | 249.33333333333334 | 1235.0 | histone H3K14ac |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000262975 | - | 8 | 24 | 75_268 | 229.33333333333334 | 1169.0 | histone H3K4me0 |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000311275 | - | 7 | 22 | 75_268 | 229.33333333333334 | 1187.0 | histone H3K4me0 |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000352431 | - | 7 | 22 | 75_268 | 249.33333333333334 | 1161.0 | histone H3K4me0 |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000355972 | - | 8 | 24 | 75_268 | 229.33333333333334 | 1215.0 | histone H3K4me0 |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000360911 | - | 7 | 22 | 75_268 | 224.33333333333334 | 1136.0 | histone H3K4me0 |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000461685 | - | 7 | 23 | 75_268 | 249.33333333333334 | 1189.0 | histone H3K4me0 |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000471951 | - | 7 | 23 | 75_268 | 249.33333333333334 | 1235.0 | histone H3K4me0 |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000262975 | - | 8 | 24 | 1147_1186 | 229.33333333333334 | 1169.0 | PRKCB1 |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000311275 | - | 7 | 22 | 1147_1186 | 229.33333333333334 | 1187.0 | PRKCB1 |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000352431 | - | 7 | 22 | 1147_1186 | 249.33333333333334 | 1161.0 | PRKCB1 |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000355972 | - | 8 | 24 | 1147_1186 | 229.33333333333334 | 1215.0 | PRKCB1 |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000360911 | - | 7 | 22 | 1147_1186 | 224.33333333333334 | 1136.0 | PRKCB1 |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000461685 | - | 7 | 23 | 1147_1186 | 249.33333333333334 | 1189.0 | PRKCB1 |
| Hgene | ZMYND8 | chr20:45918930 | chr1:87333785 | ENST00000471951 | - | 7 | 23 | 1147_1186 | 249.33333333333334 | 1235.0 | PRKCB1 |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for ZMYND8-SEP15 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for ZMYND8-SEP15 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
