
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:SMAD6-NPAS2 (FusionGDB2 ID:HG4091TG4862) |
Fusion Gene Summary for SMAD6-NPAS2 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: SMAD6-NPAS2 | Fusion gene ID: hg4091tg4862 | Hgene | Tgene | Gene symbol | SMAD6 | NPAS2 | Gene ID | 4091 | 4862 |
| Gene name | SMAD family member 6 | neuronal PAS domain protein 2 | |
| Synonyms | AOVD2|HsT17432|MADH6|MADH7 | MOP4|PASD4|bHLHe9 | |
| Cytomap | ('SMAD6')('NPAS2') 15q22.31 | 2q11.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | mothers against decapentaplegic homolog 6MAD homolog 6SMAD, mothers against DPP homolog 6mothers against decapentaplegic, drosophila, homolog of, 6 | neuronal PAS domain-containing protein 2PAS domain-containing protein 4basic-helix-loop-helix-PAS protein MOP4class E basic helix-loop-helix protein 9member of PAS protein 4member of PAS superfamily 4neuronal PAS2 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000288840, ENST00000338426, ENST00000457357, | ||
| Fusion gene scores | * DoF score | 4 X 3 X 4=48 | 9 X 9 X 5=405 |
| # samples | 5 | 9 | |
| ** MAII score | log2(5/48*10)=0.0588936890535686 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | log2(9/405*10)=-2.16992500144231 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: SMAD6 [Title/Abstract] AND NPAS2 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | SMAD6(67004062)-NPAS2(101580519), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | SMAD6-NPAS2 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. SMAD6-NPAS2 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. SMAD6-NPAS2 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. SMAD6-NPAS2 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | SMAD6 | GO:0010991 | negative regulation of SMAD protein complex assembly | 9436979 |
| Hgene | SMAD6 | GO:0030509 | BMP signaling pathway | 23455153 |
| Hgene | SMAD6 | GO:0030514 | negative regulation of BMP signaling pathway | 9436979 |
| Hgene | SMAD6 | GO:0032496 | response to lipopolysaccharide | 19193853 |
| Hgene | SMAD6 | GO:0045444 | fat cell differentiation | 23455153 |
| Hgene | SMAD6 | GO:0060394 | negative regulation of pathway-restricted SMAD protein phosphorylation | 9436979 |
| Tgene | NPAS2 | GO:0045893 | positive regulation of transcription, DNA-templated | 11441146 |
| Tgene | NPAS2 | GO:0051775 | response to redox state | 11441146 |
 Fusion gene breakpoints across SMAD6 (5'-gene) Fusion gene breakpoints across SMAD6 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
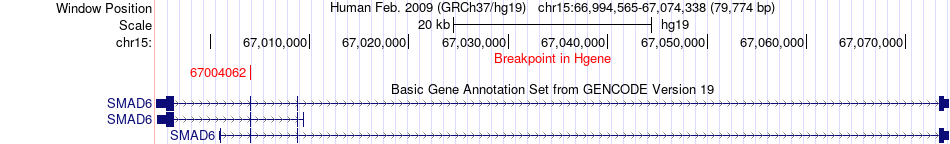 |
 Fusion gene breakpoints across NPAS2 (3'-gene) Fusion gene breakpoints across NPAS2 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
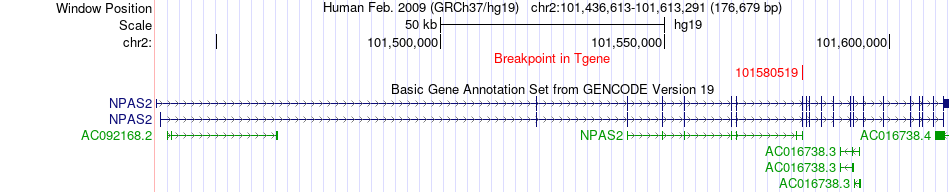 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | ESCA | TCGA-L5-A891 | SMAD6 | chr15 | 67004062 | + | NPAS2 | chr2 | 101580519 | + |
Top |
Fusion Gene ORF analysis for SMAD6-NPAS2 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-3UTR | ENST00000288840 | ENST00000486017 | SMAD6 | chr15 | 67004062 | + | NPAS2 | chr2 | 101580519 | + |
| 5CDS-3UTR | ENST00000338426 | ENST00000486017 | SMAD6 | chr15 | 67004062 | + | NPAS2 | chr2 | 101580519 | + |
| 5CDS-3UTR | ENST00000457357 | ENST00000486017 | SMAD6 | chr15 | 67004062 | + | NPAS2 | chr2 | 101580519 | + |
| In-frame | ENST00000288840 | ENST00000335681 | SMAD6 | chr15 | 67004062 | + | NPAS2 | chr2 | 101580519 | + |
| In-frame | ENST00000288840 | ENST00000542504 | SMAD6 | chr15 | 67004062 | + | NPAS2 | chr2 | 101580519 | + |
| In-frame | ENST00000338426 | ENST00000335681 | SMAD6 | chr15 | 67004062 | + | NPAS2 | chr2 | 101580519 | + |
| In-frame | ENST00000338426 | ENST00000542504 | SMAD6 | chr15 | 67004062 | + | NPAS2 | chr2 | 101580519 | + |
| In-frame | ENST00000457357 | ENST00000335681 | SMAD6 | chr15 | 67004062 | + | NPAS2 | chr2 | 101580519 | + |
| In-frame | ENST00000457357 | ENST00000542504 | SMAD6 | chr15 | 67004062 | + | NPAS2 | chr2 | 101580519 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000288840 | SMAD6 | chr15 | 67004062 | + | ENST00000335681 | NPAS2 | chr2 | 101580519 | + | 5029 | 1905 | 1031 | 3781 | 916 |
| ENST00000288840 | SMAD6 | chr15 | 67004062 | + | ENST00000542504 | NPAS2 | chr2 | 101580519 | + | 3835 | 1905 | 1031 | 3781 | 916 |
| ENST00000457357 | SMAD6 | chr15 | 67004062 | + | ENST00000335681 | NPAS2 | chr2 | 101580519 | + | 4921 | 1797 | 923 | 3673 | 916 |
| ENST00000457357 | SMAD6 | chr15 | 67004062 | + | ENST00000542504 | NPAS2 | chr2 | 101580519 | + | 3727 | 1797 | 923 | 3673 | 916 |
| ENST00000338426 | SMAD6 | chr15 | 67004062 | + | ENST00000335681 | NPAS2 | chr2 | 101580519 | + | 3328 | 204 | 113 | 2080 | 655 |
| ENST00000338426 | SMAD6 | chr15 | 67004062 | + | ENST00000542504 | NPAS2 | chr2 | 101580519 | + | 2134 | 204 | 113 | 2080 | 655 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000288840 | ENST00000335681 | SMAD6 | chr15 | 67004062 | + | NPAS2 | chr2 | 101580519 | + | 0.011618266 | 0.9883817 |
| ENST00000288840 | ENST00000542504 | SMAD6 | chr15 | 67004062 | + | NPAS2 | chr2 | 101580519 | + | 0.026532397 | 0.97346765 |
| ENST00000457357 | ENST00000335681 | SMAD6 | chr15 | 67004062 | + | NPAS2 | chr2 | 101580519 | + | 0.010349995 | 0.98965 |
| ENST00000457357 | ENST00000542504 | SMAD6 | chr15 | 67004062 | + | NPAS2 | chr2 | 101580519 | + | 0.024498941 | 0.9755011 |
| ENST00000338426 | ENST00000335681 | SMAD6 | chr15 | 67004062 | + | NPAS2 | chr2 | 101580519 | + | 0.062372908 | 0.9376271 |
| ENST00000338426 | ENST00000542504 | SMAD6 | chr15 | 67004062 | + | NPAS2 | chr2 | 101580519 | + | 0.03334287 | 0.9666571 |
Top |
Fusion Genomic Features for SMAD6-NPAS2 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| SMAD6 | chr15 | 67004062 | + | NPAS2 | chr2 | 101580519 | + | 0.10263346 | 0.89736646 |
| SMAD6 | chr15 | 67004062 | + | NPAS2 | chr2 | 101580519 | + | 0.10263346 | 0.89736646 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
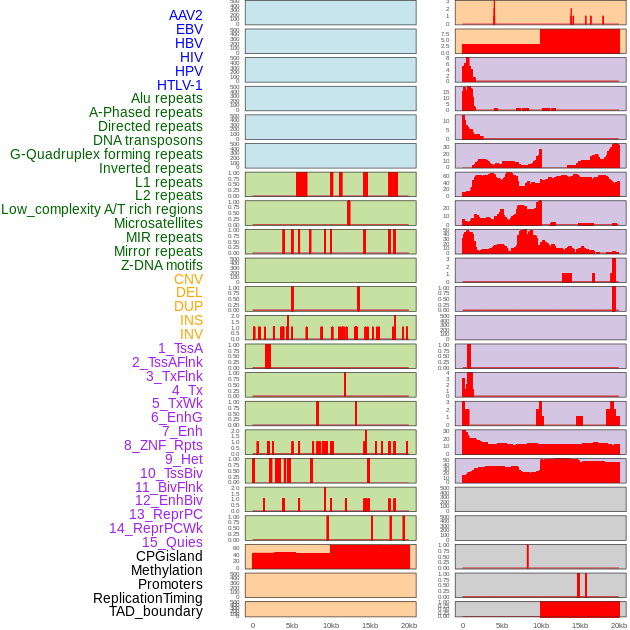 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
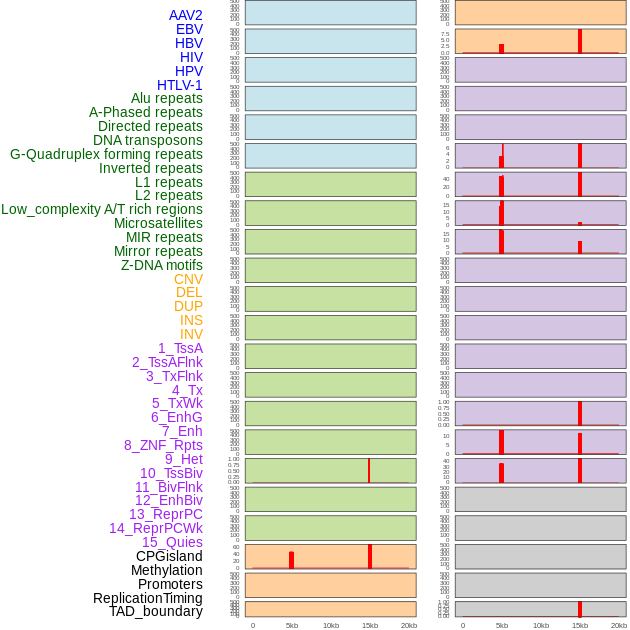 |
Top |
Fusion Protein Features for SMAD6-NPAS2 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr15:67004062/chr2:101580519) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000288840 | + | 2 | 4 | 165_168 | 291 | 497.0 | Compositional bias | Note=Poly-Leu |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000288840 | + | 2 | 4 | 251_254 | 291 | 497.0 | Compositional bias | Note=Poly-Ala |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000288840 | + | 2 | 4 | 25_33 | 291 | 497.0 | Compositional bias | Note=Poly-Gly |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000288840 | + | 2 | 4 | 275_278 | 291 | 497.0 | Compositional bias | Note=Poly-Pro |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000288840 | + | 2 | 4 | 82_85 | 291 | 497.0 | Compositional bias | Note=Poly-Arg |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000338426 | + | 2 | 4 | 25_33 | 30 | 236.0 | Compositional bias | Note=Poly-Gly |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000457357 | + | 2 | 4 | 165_168 | 291 | 339.0 | Compositional bias | Note=Poly-Leu |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000457357 | + | 2 | 4 | 251_254 | 291 | 339.0 | Compositional bias | Note=Poly-Ala |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000457357 | + | 2 | 4 | 25_33 | 291 | 339.0 | Compositional bias | Note=Poly-Gly |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000457357 | + | 2 | 4 | 275_278 | 291 | 339.0 | Compositional bias | Note=Poly-Pro |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000457357 | + | 2 | 4 | 82_85 | 291 | 339.0 | Compositional bias | Note=Poly-Arg |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000288840 | + | 2 | 4 | 148_275 | 291 | 497.0 | Domain | MH1 |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000457357 | + | 2 | 4 | 148_275 | 291 | 339.0 | Domain | MH1 |
| Tgene | NPAS2 | chr15:67004062 | chr2:101580519 | ENST00000335681 | 6 | 21 | 237_307 | 199 | 825.0 | Domain | PAS 2 | |
| Tgene | NPAS2 | chr15:67004062 | chr2:101580519 | ENST00000335681 | 6 | 21 | 311_354 | 199 | 825.0 | Domain | Note=PAC |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000338426 | + | 2 | 4 | 165_168 | 30 | 236.0 | Compositional bias | Note=Poly-Leu |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000338426 | + | 2 | 4 | 251_254 | 30 | 236.0 | Compositional bias | Note=Poly-Ala |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000338426 | + | 2 | 4 | 275_278 | 30 | 236.0 | Compositional bias | Note=Poly-Pro |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000338426 | + | 2 | 4 | 82_85 | 30 | 236.0 | Compositional bias | Note=Poly-Arg |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000288840 | + | 2 | 4 | 331_496 | 291 | 497.0 | Domain | MH2 |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000338426 | + | 2 | 4 | 148_275 | 30 | 236.0 | Domain | MH1 |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000338426 | + | 2 | 4 | 331_496 | 30 | 236.0 | Domain | MH2 |
| Hgene | SMAD6 | chr15:67004062 | chr2:101580519 | ENST00000457357 | + | 2 | 4 | 331_496 | 291 | 339.0 | Domain | MH2 |
| Tgene | NPAS2 | chr15:67004062 | chr2:101580519 | ENST00000335681 | 6 | 21 | 82_152 | 199 | 825.0 | Domain | PAS 1 | |
| Tgene | NPAS2 | chr15:67004062 | chr2:101580519 | ENST00000335681 | 6 | 21 | 9_59 | 199 | 825.0 | Domain | bHLH | |
| Tgene | NPAS2 | chr15:67004062 | chr2:101580519 | ENST00000335681 | 6 | 21 | 1_61 | 199 | 825.0 | Region | Sufficient for heterodimer formation with ARNTL/BMAL1%2C E-box binding and for the effect of NADPH |
Top |
Fusion Gene Sequence for SMAD6-NPAS2 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >83827_83827_1_SMAD6-NPAS2_SMAD6_chr15_67004062_ENST00000288840_NPAS2_chr2_101580519_ENST00000335681_length(transcript)=5029nt_BP=1905nt AGACTGGCATATGATGGGAGGCAGCCAATGACTCCGCGGCGCTCCTCCGGGGGCCCTCAGTGTGCGTTTGAGGAGAACAAAAAAGAGAGA GAGAGCCGAGCGGGGGAGCGATCGAGGGAGCTGAGCCGAGAGAAAGAGCCGCCGGGCGCTGCCTCGCCAGACCTCGCTGGGACCCCGGGG CCACCGGGAGGCACTTTTGTGGAGGGGGGAGGGGGGGCGACCTCGGCAGCCTCGGCGCACGAAGCGTCCGAGGGCAGCGTGGGGCGGGCT GCGACCTCTGCATCGGTGGACTGCATTTTTAATTAAGGATTCCCAGCAGCTCTTTGGGATTTTTACAGCTTCCACTCATGTGTTGACACC CGCGTCCAGGAGAAACTCGCTCCAAGTGCATCTAGCGCCTGGGACCTGAGACGGCGTTGGCCTTTCGTGCATGCAAATCCAGGGATTTAG GTTTTGTTTGGGATTTCCTTTTCTTTCTTTCCTTTTTTTTTTCTTTTTGCAGGGAGTAAGAAGGGAGCTGGGGGTATCAACAAGCCTGCC TTTCGGATCCTGCGGGAAAAGCCCATGTAGTTAAGCGCTTTGGTTTAAAAAAAAGGCAAGGTAAAGGCAGGGCTTTCCAGACACATTTAG GGGTTCGCGCGAGCGCTTTGTGCTCATGGACCAGCCGCACAACTTTTGAAGGCTCGCCGGCCCATGTGGGGTCTTTCTGGCGGCGCGCCG CCTGCAGCCCCCCTAAAGCGCGGGGGCTGGAGTTGTTGAGCAGCCCCGCCGCTGTGGTCCATGTAGCCGCTGGCCGCGCGCGGACTGCGG CTCGGCGTGCGCGTGTTCCCGGCCGTCCCGCCTCGGCGAGCTCCCTCATGTTGTCGCCCTGCGGCGCCCCTTCGACGACAGGCTGTGCGC GGTCTGCACGGCGCTCCGCGGCGGAGCTTCATGTGGGGCTGCGACCCGCGCAGCCGGCGCCTCGCTGAGGGAACGGACCCCCGGTAACCG GAGACCGCCTCCCCCCCACCCCTGGCGCCAAAGGATATCGTATGTTCAGGTCCAAACGCTCGGGGCTGGTGCGGCGACTTTGGCGAAGTC GTGTGGTCCCCGACCGGGAGGAAGGCGGCAGCGGCGGCGGCGGTGGCGGCGACGAGGATGGGAGCTTGGGCAGCCGAGCTGAGCCGGCCC CGCGGGCAAGAGAGGGCGGAGGCTGCGGCCGCTCCGAAGTCCGCCCGGTAGCCCCGCGGCGGCCCCGGGACGCAGTGGGACAGCGAGGCG CCCAGGGCGCGGGGAGGCGCCGGCGCGCAGGGGGCCCCCCGAGGCCCATGTCGGAGCCAGGGGCCGGCGCTGGGAGCTCCCTGCTGGACG TGGCGGAGCCGGGAGGCCCGGGCTGGCTGCCCGAGAGTGACTGCGAGACGGTGACCTGCTGTCTCTTTTCGGAGCGGGACGCCGCCGGCG CGCCCCGGGACGCCAGCGACCCCCTGGCCGGGGCGGCCCTGGAGCCGGCGGGCGGCGGGCGGAGTCGCGAAGCGCGCTCGCGGCTGCTGC TGCTGGAGCAGGAACTCAAAACCGTCACGTACTCGCTGCTGAAGCGGCTCAAGGAGCGCTCGCTGGACACGCTGCTGGAGGCGGTGGAGT CCCGCGGCGGCGTGCCGGGCGGCTGCGTGCTGGTGCCGCGCGCCGACCTCCGCCTGGGCGGCCAGCCCGCGCCGCCGCAGCTGCTGCTCG GCCGCCTCTTTCGCTGGCCCGACCTGCAGCACGCCGTGGAGCTGAAGCCCCTGTGCGGCTGCCACAGCTTCGCCGCCGCCGCCGACGGCC CTACCGTGTGCTGCAACCCCTACCACTTCAGCCGGCTCTGCGGGCCCGAATCTCCGCCACCTCCCTACTCTCGGCTGTCTCCTCGCGACG AGTACAAGCCACTGGTGCCTAGCCCCTCCTGTAATGGTTTTGACAACACCCTTTCAAGACCTTGCCGGGTGCCACTAGGAAAGGAGGTTT GCTTCATTGCCACCGTTCGTCTGGCAACACCACAATTCTTAAAGGAAATGTGCATAGTTGACGAACCTTTAGAGGAATTCACTTCAAGGC ATAGCTTGGAATGGAAATTTTTATTTCTGGATCACAGAGCACCTCCAATCATAGGATACCTGCCTTTTGAAGTGCTGGGAACCTCAGGCT ATGACTACTACCACATTGATGACCTGGAGCTCCTGGCCAGGTGTCACCAGCACCTGATGCAGTTTGGCAAAGGGAAGTCGTGTTGCTACC GGTTTCTGACCAAAGGTCAGCAGTGGATCTGGCTGCAGACTCACTACTACATCACCTACCATCAGTGGAACTCCAAGCCCGAGTTCATCG TGTGCACACACTCGGTGGTCAGTTACGCAGATGTCCGGGTGGAAAGGAGGCAGGAGCTGGCTCTGGAAGACCCGCCATCCGAGGCCCTCC ACTCCTCAGCACTAAAGGACAAGGGCTCAAGCCTGGAACCTCGGCAGCACTTTAACACACTCGACGTGGGTGCCTCGGGCCTTAATACCA GTCATTCGCCATCGGCGTCCTCAAGAAGTTCCCACAAATCCTCGCACACAGCCATGTCAGAACCCACCTCCACTCCCACCAAGCTGATGG CAGAGGCCAGCACCCCGGCTTTGCCAAGATCAGCCACCCTGCCCCAAGAGTTACCTGTCCCCGGGCTCAGCCAGGCAGCCACCATGCCGG CCCCTCTGCCTTCCCCATCGTCCTGCGACCTCACACAGCAGCTCCTGCCTCAGACCGTTCTGCAGAGCACGCCCGCTCCCATGGCACAGT TTTCGGCACAGTTCAGCATGTTCCAGACCATCAAAGACCAGCTAGAGCAGCGGACGCGGATCCTGCAGGCCAATATCCGGTGGCAACAGG AAGAGCTCCACAAGATCCAGGAGCAGCTCTGCCTGGTCCAGGACTCCAACGTCCAGATGTTCCTGCAGCAGCCAGCTGTATCCCTGAGCT TCAGCAGCACCCAGCGACCTGAGGCTCAGCAGCAGCTACAGCAAAGGTCAGCTGCAGTGACTCAGCCCCAGCTCGGGGCGGGCCCCCAAC TTCCAGGGCAGATCTCCTCTGCCCAGGTCACAAGCCAGCACCTGCTCAGAGAATCAAGTGTGATATCAACCCAGGGTCCAAAGCCAATGA GAAGCTCACAGCTAATGCAGAGCAGCGGCCGCTCTGGAAGCAGCCTAGTGTCCCCGTTCAGCAGCGCCACAGCTGCGCTCCCGCCAAGTC TGAATCTGACCACACCTGCTTCCACCTCCCAGGATGCCAGCCAGTGCCAGCCCAGCCCAGACTTCAGCCATGATCGGCAGCTCAGGCTGT TGCTGAGCCAGCCCATCCAGCCCATGATGCCCGGGTCCTGTGACGCAAGGCAGCCCTCGGAAGTCAGCAGGACGGGACGGCAAGTCAAGT ACGCCCAGAGCCAGACCGTGTTTCAAAATCCAGACGCACACCCCGCCAACAGCAGCAGCGCCCCGATGCCCGTCCTGCTGATGGGGCAGG CGGTGCTCCACCCCAGCTTCCCTGCCTCCCAACCATCGCCCCTGCAGCCTGCACAGGCCCGGCAGCAGCCACCGCAGCACTACCTGCAGG TACAGGCACCAACCTCTTTGCACAGTGAGCAGCAGGACTCGCTACTTCTCTCCACCTACTCACAACAGCCAGGGACCCTGGGCTACCCCC AACCACCCCCAGCACAGCCCCAGCCCCTACGTCCTCCCCGAAGGGTCAGCAGTCTGTCTGAGTCGTCAGGCCTCCAGCAGCCGCCCCGAT AATGCCCCGGCACTGAAGTCGGGACACAATCAGCTTTAACCAATGGATGAGGGGGGTGGCCACAGGAGATGGGGAGAGGAGTCTGAACTA AACCCCTGGCTTTTGTGCACACTGCATACGTTTCAGAACTCCTGGATGGTAACCATCTCTGGAGTGCAGCGCTTGCTGCAGTGGAAATGA TCAGGAATACTGACCGTGTTTCTCTTGCCTCCGAGGTTCTTGGGCACACTCTATAGCCATACTGGACAGGAACCAGGTGCCCCGTGTAGG CATCGTCGGTCGGTTTGCCGTCAGAGATGGCGCATCTCGCTGCATCCCCCGAGAGTACACCGGTTGCTCTAGCCACCTGCGGCCCGCCCA TCTGCGCTAGCTGGCCTTCACGCTCTTGATCGTCTTTCCTTTGTATTGGAGAAGGACTGGGTCAGAGATCTGTTGGAGAGAGAGAATAAA GAGATTATTTTTCATTATTTTTAAATGGTTGTTTTTGTTTTAATTTGCACAGCTACACAGAGGAAATAACTTAGGCACTTTCTGTTTTTT TTAAAAAAATAATAAGGTCTCATGGCTTCATTTAGAGACCACAGTAACAACAGCAGCCCACCAATCAGAGAAGCTGGTTGTTATTAACCA AGCTACAGATTCACACTTTCTGGCCTAAACCCTAATGGGATGAGGCTTTTCACCCCAGGCCATGCTGGTGGTGATTTTTTAGCCCCTAAA TAAAACACTGGACTATTTCCTGTTTACTTCATTGATTGCAACTACAAAGGTGGACTCAAAGCAAAGCACAATCATGCCAGCCAACATTCC AGAATTCTGCTGAGAACTCCAAGTCTGTGAGGGGAGAGGTTTTACAAGCCAGACAGGCCTGGGGGACTGCAGTCCCCAAGGAGACCCTGC CACATGCTGGCCCTTTGAGTGAGAATGCTGCATCTTTCTACATATCTTCATGAGAATACTGAGAATTGGATTTTCCTTTTCAAAATGCAC TTTGCTTTTTTTGTATGTTTTGTTATGTTGAGATGTTTCTAAAGAAAAGATTTTATGTAATTATAAGATGAAGCGTAGTGAATTGTACAG CTGTTGTAATAATGACCTATTTCTATATAAAATAAAATTGTATGGCTTATGTGTAAATTATTTTGTATCTGAGATACCAGTTCCTTTTCC >83827_83827_1_SMAD6-NPAS2_SMAD6_chr15_67004062_ENST00000288840_NPAS2_chr2_101580519_ENST00000335681_length(amino acids)=916AA_BP=289 MFRSKRSGLVRRLWRSRVVPDREEGGSGGGGGGDEDGSLGSRAEPAPRAREGGGCGRSEVRPVAPRRPRDAVGQRGAQGAGRRRRAGGPP RPMSEPGAGAGSSLLDVAEPGGPGWLPESDCETVTCCLFSERDAAGAPRDASDPLAGAALEPAGGGRSREARSRLLLLEQELKTVTYSLL KRLKERSLDTLLEAVESRGGVPGGCVLVPRADLRLGGQPAPPQLLLGRLFRWPDLQHAVELKPLCGCHSFAAAADGPTVCCNPYHFSRLC GPESPPPPYSRLSPRDEYKPLVPSPSCNGFDNTLSRPCRVPLGKEVCFIATVRLATPQFLKEMCIVDEPLEEFTSRHSLEWKFLFLDHRA PPIIGYLPFEVLGTSGYDYYHIDDLELLARCHQHLMQFGKGKSCCYRFLTKGQQWIWLQTHYYITYHQWNSKPEFIVCTHSVVSYADVRV ERRQELALEDPPSEALHSSALKDKGSSLEPRQHFNTLDVGASGLNTSHSPSASSRSSHKSSHTAMSEPTSTPTKLMAEASTPALPRSATL PQELPVPGLSQAATMPAPLPSPSSCDLTQQLLPQTVLQSTPAPMAQFSAQFSMFQTIKDQLEQRTRILQANIRWQQEELHKIQEQLCLVQ DSNVQMFLQQPAVSLSFSSTQRPEAQQQLQQRSAAVTQPQLGAGPQLPGQISSAQVTSQHLLRESSVISTQGPKPMRSSQLMQSSGRSGS SLVSPFSSATAALPPSLNLTTPASTSQDASQCQPSPDFSHDRQLRLLLSQPIQPMMPGSCDARQPSEVSRTGRQVKYAQSQTVFQNPDAH PANSSSAPMPVLLMGQAVLHPSFPASQPSPLQPAQARQQPPQHYLQVQAPTSLHSEQQDSLLLSTYSQQPGTLGYPQPPPAQPQPLRPPR -------------------------------------------------------------- >83827_83827_2_SMAD6-NPAS2_SMAD6_chr15_67004062_ENST00000288840_NPAS2_chr2_101580519_ENST00000542504_length(transcript)=3835nt_BP=1905nt AGACTGGCATATGATGGGAGGCAGCCAATGACTCCGCGGCGCTCCTCCGGGGGCCCTCAGTGTGCGTTTGAGGAGAACAAAAAAGAGAGA GAGAGCCGAGCGGGGGAGCGATCGAGGGAGCTGAGCCGAGAGAAAGAGCCGCCGGGCGCTGCCTCGCCAGACCTCGCTGGGACCCCGGGG CCACCGGGAGGCACTTTTGTGGAGGGGGGAGGGGGGGCGACCTCGGCAGCCTCGGCGCACGAAGCGTCCGAGGGCAGCGTGGGGCGGGCT GCGACCTCTGCATCGGTGGACTGCATTTTTAATTAAGGATTCCCAGCAGCTCTTTGGGATTTTTACAGCTTCCACTCATGTGTTGACACC CGCGTCCAGGAGAAACTCGCTCCAAGTGCATCTAGCGCCTGGGACCTGAGACGGCGTTGGCCTTTCGTGCATGCAAATCCAGGGATTTAG GTTTTGTTTGGGATTTCCTTTTCTTTCTTTCCTTTTTTTTTTCTTTTTGCAGGGAGTAAGAAGGGAGCTGGGGGTATCAACAAGCCTGCC TTTCGGATCCTGCGGGAAAAGCCCATGTAGTTAAGCGCTTTGGTTTAAAAAAAAGGCAAGGTAAAGGCAGGGCTTTCCAGACACATTTAG GGGTTCGCGCGAGCGCTTTGTGCTCATGGACCAGCCGCACAACTTTTGAAGGCTCGCCGGCCCATGTGGGGTCTTTCTGGCGGCGCGCCG CCTGCAGCCCCCCTAAAGCGCGGGGGCTGGAGTTGTTGAGCAGCCCCGCCGCTGTGGTCCATGTAGCCGCTGGCCGCGCGCGGACTGCGG CTCGGCGTGCGCGTGTTCCCGGCCGTCCCGCCTCGGCGAGCTCCCTCATGTTGTCGCCCTGCGGCGCCCCTTCGACGACAGGCTGTGCGC GGTCTGCACGGCGCTCCGCGGCGGAGCTTCATGTGGGGCTGCGACCCGCGCAGCCGGCGCCTCGCTGAGGGAACGGACCCCCGGTAACCG GAGACCGCCTCCCCCCCACCCCTGGCGCCAAAGGATATCGTATGTTCAGGTCCAAACGCTCGGGGCTGGTGCGGCGACTTTGGCGAAGTC GTGTGGTCCCCGACCGGGAGGAAGGCGGCAGCGGCGGCGGCGGTGGCGGCGACGAGGATGGGAGCTTGGGCAGCCGAGCTGAGCCGGCCC CGCGGGCAAGAGAGGGCGGAGGCTGCGGCCGCTCCGAAGTCCGCCCGGTAGCCCCGCGGCGGCCCCGGGACGCAGTGGGACAGCGAGGCG CCCAGGGCGCGGGGAGGCGCCGGCGCGCAGGGGGCCCCCCGAGGCCCATGTCGGAGCCAGGGGCCGGCGCTGGGAGCTCCCTGCTGGACG TGGCGGAGCCGGGAGGCCCGGGCTGGCTGCCCGAGAGTGACTGCGAGACGGTGACCTGCTGTCTCTTTTCGGAGCGGGACGCCGCCGGCG CGCCCCGGGACGCCAGCGACCCCCTGGCCGGGGCGGCCCTGGAGCCGGCGGGCGGCGGGCGGAGTCGCGAAGCGCGCTCGCGGCTGCTGC TGCTGGAGCAGGAACTCAAAACCGTCACGTACTCGCTGCTGAAGCGGCTCAAGGAGCGCTCGCTGGACACGCTGCTGGAGGCGGTGGAGT CCCGCGGCGGCGTGCCGGGCGGCTGCGTGCTGGTGCCGCGCGCCGACCTCCGCCTGGGCGGCCAGCCCGCGCCGCCGCAGCTGCTGCTCG GCCGCCTCTTTCGCTGGCCCGACCTGCAGCACGCCGTGGAGCTGAAGCCCCTGTGCGGCTGCCACAGCTTCGCCGCCGCCGCCGACGGCC CTACCGTGTGCTGCAACCCCTACCACTTCAGCCGGCTCTGCGGGCCCGAATCTCCGCCACCTCCCTACTCTCGGCTGTCTCCTCGCGACG AGTACAAGCCACTGGTGCCTAGCCCCTCCTGTAATGGTTTTGACAACACCCTTTCAAGACCTTGCCGGGTGCCACTAGGAAAGGAGGTTT GCTTCATTGCCACCGTTCGTCTGGCAACACCACAATTCTTAAAGGAAATGTGCATAGTTGACGAACCTTTAGAGGAATTCACTTCAAGGC ATAGCTTGGAATGGAAATTTTTATTTCTGGATCACAGAGCACCTCCAATCATAGGATACCTGCCTTTTGAAGTGCTGGGAACCTCAGGCT ATGACTACTACCACATTGATGACCTGGAGCTCCTGGCCAGGTGTCACCAGCACCTGATGCAGTTTGGCAAAGGGAAGTCGTGTTGCTACC GGTTTCTGACCAAAGGTCAGCAGTGGATCTGGCTGCAGACTCACTACTACATCACCTACCATCAGTGGAACTCCAAGCCCGAGTTCATCG TGTGCACACACTCGGTGGTCAGTTACGCAGATGTCCGGGTGGAAAGGAGGCAGGAGCTGGCTCTGGAAGACCCGCCATCCGAGGCCCTCC ACTCCTCAGCACTAAAGGACAAGGGCTCAAGCCTGGAACCTCGGCAGCACTTTAACACACTCGACGTGGGTGCCTCGGGCCTTAATACCA GTCATTCGCCATCGGCGTCCTCAAGAAGTTCCCACAAATCCTCGCACACAGCCATGTCAGAACCCACCTCCACTCCCACCAAGCTGATGG CAGAGGCCAGCACCCCGGCTTTGCCAAGATCAGCCACCCTGCCCCAAGAGTTACCTGTCCCCGGGCTCAGCCAGGCAGCCACCATGCCGG CCCCTCTGCCTTCCCCATCGTCCTGCGACCTCACACAGCAGCTCCTGCCTCAGACCGTTCTGCAGAGCACGCCCGCTCCCATGGCACAGT TTTCGGCACAGTTCAGCATGTTCCAGACCATCAAAGACCAGCTAGAGCAGCGGACGCGGATCCTGCAGGCCAATATCCGGTGGCAACAGG AAGAGCTCCACAAGATCCAGGAGCAGCTCTGCCTGGTCCAGGACTCCAACGTCCAGATGTTCCTGCAGCAGCCAGCTGTATCCCTGAGCT TCAGCAGCACCCAGCGACCTGAGGCTCAGCAGCAGCTACAGCAAAGGTCAGCTGCAGTGACTCAGCCCCAGCTCGGGGCGGGCCCCCAAC TTCCAGGGCAGATCTCCTCTGCCCAGGTCACAAGCCAGCACCTGCTCAGAGAATCAAGTGTGATATCAACCCAGGGTCCAAAGCCAATGA GAAGCTCACAGCTAATGCAGAGCAGCGGCCGCTCTGGAAGCAGCCTAGTGTCCCCGTTCAGCAGCGCCACAGCTGCGCTCCCGCCAAGTC TGAATCTGACCACACCTGCTTCCACCTCCCAGGATGCCAGCCAGTGCCAGCCCAGCCCAGACTTCAGCCATGATCGGCAGCTCAGGCTGT TGCTGAGCCAGCCCATCCAGCCCATGATGCCCGGGTCCTGTGACGCAAGGCAGCCCTCGGAAGTCAGCAGGACGGGACGGCAAGTCAAGT ACGCCCAGAGCCAGACCGTGTTTCAAAATCCAGACGCACACCCCGCCAACAGCAGCAGCGCCCCGATGCCCGTCCTGCTGATGGGGCAGG CGGTGCTCCACCCCAGCTTCCCTGCCTCCCAACCATCGCCCCTGCAGCCTGCACAGGCCCGGCAGCAGCCACCGCAGCACTACCTGCAGG TACAGGCACCAACCTCTTTGCACAGTGAGCAGCAGGACTCGCTACTTCTCTCCACCTACTCACAACAGCCAGGGACCCTGGGCTACCCCC AACCACCCCCAGCACAGCCCCAGCCCCTACGTCCTCCCCGAAGGGTCAGCAGTCTGTCTGAGTCGTCAGGCCTCCAGCAGCCGCCCCGAT >83827_83827_2_SMAD6-NPAS2_SMAD6_chr15_67004062_ENST00000288840_NPAS2_chr2_101580519_ENST00000542504_length(amino acids)=916AA_BP=289 MFRSKRSGLVRRLWRSRVVPDREEGGSGGGGGGDEDGSLGSRAEPAPRAREGGGCGRSEVRPVAPRRPRDAVGQRGAQGAGRRRRAGGPP RPMSEPGAGAGSSLLDVAEPGGPGWLPESDCETVTCCLFSERDAAGAPRDASDPLAGAALEPAGGGRSREARSRLLLLEQELKTVTYSLL KRLKERSLDTLLEAVESRGGVPGGCVLVPRADLRLGGQPAPPQLLLGRLFRWPDLQHAVELKPLCGCHSFAAAADGPTVCCNPYHFSRLC GPESPPPPYSRLSPRDEYKPLVPSPSCNGFDNTLSRPCRVPLGKEVCFIATVRLATPQFLKEMCIVDEPLEEFTSRHSLEWKFLFLDHRA PPIIGYLPFEVLGTSGYDYYHIDDLELLARCHQHLMQFGKGKSCCYRFLTKGQQWIWLQTHYYITYHQWNSKPEFIVCTHSVVSYADVRV ERRQELALEDPPSEALHSSALKDKGSSLEPRQHFNTLDVGASGLNTSHSPSASSRSSHKSSHTAMSEPTSTPTKLMAEASTPALPRSATL PQELPVPGLSQAATMPAPLPSPSSCDLTQQLLPQTVLQSTPAPMAQFSAQFSMFQTIKDQLEQRTRILQANIRWQQEELHKIQEQLCLVQ DSNVQMFLQQPAVSLSFSSTQRPEAQQQLQQRSAAVTQPQLGAGPQLPGQISSAQVTSQHLLRESSVISTQGPKPMRSSQLMQSSGRSGS SLVSPFSSATAALPPSLNLTTPASTSQDASQCQPSPDFSHDRQLRLLLSQPIQPMMPGSCDARQPSEVSRTGRQVKYAQSQTVFQNPDAH PANSSSAPMPVLLMGQAVLHPSFPASQPSPLQPAQARQQPPQHYLQVQAPTSLHSEQQDSLLLSTYSQQPGTLGYPQPPPAQPQPLRPPR -------------------------------------------------------------- >83827_83827_3_SMAD6-NPAS2_SMAD6_chr15_67004062_ENST00000338426_NPAS2_chr2_101580519_ENST00000335681_length(transcript)=3328nt_BP=204nt AAAAGAACGAAGTCCAGCACCAAAACGTGCTACAACATGGATGAACTTCGATGACTTTGTGCCACATGAAAGAAGAAGCCAGCCACAAAA GGCCATATATTGTATGAAATGAAATGTCCAGAATGGGCAAACCCATAGAGACACAAAAATCTCCGCCACCTCCCTACTCTCGGCTGTCTC CTCGCGACGAGTACAAGCCACTGGTGCCTAGCCCCTCCTGTAATGGTTTTGACAACACCCTTTCAAGACCTTGCCGGGTGCCACTAGGAA AGGAGGTTTGCTTCATTGCCACCGTTCGTCTGGCAACACCACAATTCTTAAAGGAAATGTGCATAGTTGACGAACCTTTAGAGGAATTCA CTTCAAGGCATAGCTTGGAATGGAAATTTTTATTTCTGGATCACAGAGCACCTCCAATCATAGGATACCTGCCTTTTGAAGTGCTGGGAA CCTCAGGCTATGACTACTACCACATTGATGACCTGGAGCTCCTGGCCAGGTGTCACCAGCACCTGATGCAGTTTGGCAAAGGGAAGTCGT GTTGCTACCGGTTTCTGACCAAAGGTCAGCAGTGGATCTGGCTGCAGACTCACTACTACATCACCTACCATCAGTGGAACTCCAAGCCCG AGTTCATCGTGTGCACACACTCGGTGGTCAGTTACGCAGATGTCCGGGTGGAAAGGAGGCAGGAGCTGGCTCTGGAAGACCCGCCATCCG AGGCCCTCCACTCCTCAGCACTAAAGGACAAGGGCTCAAGCCTGGAACCTCGGCAGCACTTTAACACACTCGACGTGGGTGCCTCGGGCC TTAATACCAGTCATTCGCCATCGGCGTCCTCAAGAAGTTCCCACAAATCCTCGCACACAGCCATGTCAGAACCCACCTCCACTCCCACCA AGCTGATGGCAGAGGCCAGCACCCCGGCTTTGCCAAGATCAGCCACCCTGCCCCAAGAGTTACCTGTCCCCGGGCTCAGCCAGGCAGCCA CCATGCCGGCCCCTCTGCCTTCCCCATCGTCCTGCGACCTCACACAGCAGCTCCTGCCTCAGACCGTTCTGCAGAGCACGCCCGCTCCCA TGGCACAGTTTTCGGCACAGTTCAGCATGTTCCAGACCATCAAAGACCAGCTAGAGCAGCGGACGCGGATCCTGCAGGCCAATATCCGGT GGCAACAGGAAGAGCTCCACAAGATCCAGGAGCAGCTCTGCCTGGTCCAGGACTCCAACGTCCAGATGTTCCTGCAGCAGCCAGCTGTAT CCCTGAGCTTCAGCAGCACCCAGCGACCTGAGGCTCAGCAGCAGCTACAGCAAAGGTCAGCTGCAGTGACTCAGCCCCAGCTCGGGGCGG GCCCCCAACTTCCAGGGCAGATCTCCTCTGCCCAGGTCACAAGCCAGCACCTGCTCAGAGAATCAAGTGTGATATCAACCCAGGGTCCAA AGCCAATGAGAAGCTCACAGCTAATGCAGAGCAGCGGCCGCTCTGGAAGCAGCCTAGTGTCCCCGTTCAGCAGCGCCACAGCTGCGCTCC CGCCAAGTCTGAATCTGACCACACCTGCTTCCACCTCCCAGGATGCCAGCCAGTGCCAGCCCAGCCCAGACTTCAGCCATGATCGGCAGC TCAGGCTGTTGCTGAGCCAGCCCATCCAGCCCATGATGCCCGGGTCCTGTGACGCAAGGCAGCCCTCGGAAGTCAGCAGGACGGGACGGC AAGTCAAGTACGCCCAGAGCCAGACCGTGTTTCAAAATCCAGACGCACACCCCGCCAACAGCAGCAGCGCCCCGATGCCCGTCCTGCTGA TGGGGCAGGCGGTGCTCCACCCCAGCTTCCCTGCCTCCCAACCATCGCCCCTGCAGCCTGCACAGGCCCGGCAGCAGCCACCGCAGCACT ACCTGCAGGTACAGGCACCAACCTCTTTGCACAGTGAGCAGCAGGACTCGCTACTTCTCTCCACCTACTCACAACAGCCAGGGACCCTGG GCTACCCCCAACCACCCCCAGCACAGCCCCAGCCCCTACGTCCTCCCCGAAGGGTCAGCAGTCTGTCTGAGTCGTCAGGCCTCCAGCAGC CGCCCCGATAATGCCCCGGCACTGAAGTCGGGACACAATCAGCTTTAACCAATGGATGAGGGGGGTGGCCACAGGAGATGGGGAGAGGAG TCTGAACTAAACCCCTGGCTTTTGTGCACACTGCATACGTTTCAGAACTCCTGGATGGTAACCATCTCTGGAGTGCAGCGCTTGCTGCAG TGGAAATGATCAGGAATACTGACCGTGTTTCTCTTGCCTCCGAGGTTCTTGGGCACACTCTATAGCCATACTGGACAGGAACCAGGTGCC CCGTGTAGGCATCGTCGGTCGGTTTGCCGTCAGAGATGGCGCATCTCGCTGCATCCCCCGAGAGTACACCGGTTGCTCTAGCCACCTGCG GCCCGCCCATCTGCGCTAGCTGGCCTTCACGCTCTTGATCGTCTTTCCTTTGTATTGGAGAAGGACTGGGTCAGAGATCTGTTGGAGAGA GAGAATAAAGAGATTATTTTTCATTATTTTTAAATGGTTGTTTTTGTTTTAATTTGCACAGCTACACAGAGGAAATAACTTAGGCACTTT CTGTTTTTTTTAAAAAAATAATAAGGTCTCATGGCTTCATTTAGAGACCACAGTAACAACAGCAGCCCACCAATCAGAGAAGCTGGTTGT TATTAACCAAGCTACAGATTCACACTTTCTGGCCTAAACCCTAATGGGATGAGGCTTTTCACCCCAGGCCATGCTGGTGGTGATTTTTTA GCCCCTAAATAAAACACTGGACTATTTCCTGTTTACTTCATTGATTGCAACTACAAAGGTGGACTCAAAGCAAAGCACAATCATGCCAGC CAACATTCCAGAATTCTGCTGAGAACTCCAAGTCTGTGAGGGGAGAGGTTTTACAAGCCAGACAGGCCTGGGGGACTGCAGTCCCCAAGG AGACCCTGCCACATGCTGGCCCTTTGAGTGAGAATGCTGCATCTTTCTACATATCTTCATGAGAATACTGAGAATTGGATTTTCCTTTTC AAAATGCACTTTGCTTTTTTTGTATGTTTTGTTATGTTGAGATGTTTCTAAAGAAAAGATTTTATGTAATTATAAGATGAAGCGTAGTGA ATTGTACAGCTGTTGTAATAATGACCTATTTCTATATAAAATAAAATTGTATGGCTTATGTGTAAATTATTTTGTATCTGAGATACCAGT >83827_83827_3_SMAD6-NPAS2_SMAD6_chr15_67004062_ENST00000338426_NPAS2_chr2_101580519_ENST00000335681_length(amino acids)=655AA_BP=28 MSRMGKPIETQKSPPPPYSRLSPRDEYKPLVPSPSCNGFDNTLSRPCRVPLGKEVCFIATVRLATPQFLKEMCIVDEPLEEFTSRHSLEW KFLFLDHRAPPIIGYLPFEVLGTSGYDYYHIDDLELLARCHQHLMQFGKGKSCCYRFLTKGQQWIWLQTHYYITYHQWNSKPEFIVCTHS VVSYADVRVERRQELALEDPPSEALHSSALKDKGSSLEPRQHFNTLDVGASGLNTSHSPSASSRSSHKSSHTAMSEPTSTPTKLMAEAST PALPRSATLPQELPVPGLSQAATMPAPLPSPSSCDLTQQLLPQTVLQSTPAPMAQFSAQFSMFQTIKDQLEQRTRILQANIRWQQEELHK IQEQLCLVQDSNVQMFLQQPAVSLSFSSTQRPEAQQQLQQRSAAVTQPQLGAGPQLPGQISSAQVTSQHLLRESSVISTQGPKPMRSSQL MQSSGRSGSSLVSPFSSATAALPPSLNLTTPASTSQDASQCQPSPDFSHDRQLRLLLSQPIQPMMPGSCDARQPSEVSRTGRQVKYAQSQ TVFQNPDAHPANSSSAPMPVLLMGQAVLHPSFPASQPSPLQPAQARQQPPQHYLQVQAPTSLHSEQQDSLLLSTYSQQPGTLGYPQPPPA -------------------------------------------------------------- >83827_83827_4_SMAD6-NPAS2_SMAD6_chr15_67004062_ENST00000338426_NPAS2_chr2_101580519_ENST00000542504_length(transcript)=2134nt_BP=204nt AAAAGAACGAAGTCCAGCACCAAAACGTGCTACAACATGGATGAACTTCGATGACTTTGTGCCACATGAAAGAAGAAGCCAGCCACAAAA GGCCATATATTGTATGAAATGAAATGTCCAGAATGGGCAAACCCATAGAGACACAAAAATCTCCGCCACCTCCCTACTCTCGGCTGTCTC CTCGCGACGAGTACAAGCCACTGGTGCCTAGCCCCTCCTGTAATGGTTTTGACAACACCCTTTCAAGACCTTGCCGGGTGCCACTAGGAA AGGAGGTTTGCTTCATTGCCACCGTTCGTCTGGCAACACCACAATTCTTAAAGGAAATGTGCATAGTTGACGAACCTTTAGAGGAATTCA CTTCAAGGCATAGCTTGGAATGGAAATTTTTATTTCTGGATCACAGAGCACCTCCAATCATAGGATACCTGCCTTTTGAAGTGCTGGGAA CCTCAGGCTATGACTACTACCACATTGATGACCTGGAGCTCCTGGCCAGGTGTCACCAGCACCTGATGCAGTTTGGCAAAGGGAAGTCGT GTTGCTACCGGTTTCTGACCAAAGGTCAGCAGTGGATCTGGCTGCAGACTCACTACTACATCACCTACCATCAGTGGAACTCCAAGCCCG AGTTCATCGTGTGCACACACTCGGTGGTCAGTTACGCAGATGTCCGGGTGGAAAGGAGGCAGGAGCTGGCTCTGGAAGACCCGCCATCCG AGGCCCTCCACTCCTCAGCACTAAAGGACAAGGGCTCAAGCCTGGAACCTCGGCAGCACTTTAACACACTCGACGTGGGTGCCTCGGGCC TTAATACCAGTCATTCGCCATCGGCGTCCTCAAGAAGTTCCCACAAATCCTCGCACACAGCCATGTCAGAACCCACCTCCACTCCCACCA AGCTGATGGCAGAGGCCAGCACCCCGGCTTTGCCAAGATCAGCCACCCTGCCCCAAGAGTTACCTGTCCCCGGGCTCAGCCAGGCAGCCA CCATGCCGGCCCCTCTGCCTTCCCCATCGTCCTGCGACCTCACACAGCAGCTCCTGCCTCAGACCGTTCTGCAGAGCACGCCCGCTCCCA TGGCACAGTTTTCGGCACAGTTCAGCATGTTCCAGACCATCAAAGACCAGCTAGAGCAGCGGACGCGGATCCTGCAGGCCAATATCCGGT GGCAACAGGAAGAGCTCCACAAGATCCAGGAGCAGCTCTGCCTGGTCCAGGACTCCAACGTCCAGATGTTCCTGCAGCAGCCAGCTGTAT CCCTGAGCTTCAGCAGCACCCAGCGACCTGAGGCTCAGCAGCAGCTACAGCAAAGGTCAGCTGCAGTGACTCAGCCCCAGCTCGGGGCGG GCCCCCAACTTCCAGGGCAGATCTCCTCTGCCCAGGTCACAAGCCAGCACCTGCTCAGAGAATCAAGTGTGATATCAACCCAGGGTCCAA AGCCAATGAGAAGCTCACAGCTAATGCAGAGCAGCGGCCGCTCTGGAAGCAGCCTAGTGTCCCCGTTCAGCAGCGCCACAGCTGCGCTCC CGCCAAGTCTGAATCTGACCACACCTGCTTCCACCTCCCAGGATGCCAGCCAGTGCCAGCCCAGCCCAGACTTCAGCCATGATCGGCAGC TCAGGCTGTTGCTGAGCCAGCCCATCCAGCCCATGATGCCCGGGTCCTGTGACGCAAGGCAGCCCTCGGAAGTCAGCAGGACGGGACGGC AAGTCAAGTACGCCCAGAGCCAGACCGTGTTTCAAAATCCAGACGCACACCCCGCCAACAGCAGCAGCGCCCCGATGCCCGTCCTGCTGA TGGGGCAGGCGGTGCTCCACCCCAGCTTCCCTGCCTCCCAACCATCGCCCCTGCAGCCTGCACAGGCCCGGCAGCAGCCACCGCAGCACT ACCTGCAGGTACAGGCACCAACCTCTTTGCACAGTGAGCAGCAGGACTCGCTACTTCTCTCCACCTACTCACAACAGCCAGGGACCCTGG GCTACCCCCAACCACCCCCAGCACAGCCCCAGCCCCTACGTCCTCCCCGAAGGGTCAGCAGTCTGTCTGAGTCGTCAGGCCTCCAGCAGC >83827_83827_4_SMAD6-NPAS2_SMAD6_chr15_67004062_ENST00000338426_NPAS2_chr2_101580519_ENST00000542504_length(amino acids)=655AA_BP=28 MSRMGKPIETQKSPPPPYSRLSPRDEYKPLVPSPSCNGFDNTLSRPCRVPLGKEVCFIATVRLATPQFLKEMCIVDEPLEEFTSRHSLEW KFLFLDHRAPPIIGYLPFEVLGTSGYDYYHIDDLELLARCHQHLMQFGKGKSCCYRFLTKGQQWIWLQTHYYITYHQWNSKPEFIVCTHS VVSYADVRVERRQELALEDPPSEALHSSALKDKGSSLEPRQHFNTLDVGASGLNTSHSPSASSRSSHKSSHTAMSEPTSTPTKLMAEAST PALPRSATLPQELPVPGLSQAATMPAPLPSPSSCDLTQQLLPQTVLQSTPAPMAQFSAQFSMFQTIKDQLEQRTRILQANIRWQQEELHK IQEQLCLVQDSNVQMFLQQPAVSLSFSSTQRPEAQQQLQQRSAAVTQPQLGAGPQLPGQISSAQVTSQHLLRESSVISTQGPKPMRSSQL MQSSGRSGSSLVSPFSSATAALPPSLNLTTPASTSQDASQCQPSPDFSHDRQLRLLLSQPIQPMMPGSCDARQPSEVSRTGRQVKYAQSQ TVFQNPDAHPANSSSAPMPVLLMGQAVLHPSFPASQPSPLQPAQARQQPPQHYLQVQAPTSLHSEQQDSLLLSTYSQQPGTLGYPQPPPA -------------------------------------------------------------- >83827_83827_5_SMAD6-NPAS2_SMAD6_chr15_67004062_ENST00000457357_NPAS2_chr2_101580519_ENST00000335681_length(transcript)=4921nt_BP=1797nt CGATCGAGGGAGCTGAGCCGAGAGAAAGAGCCGCCGGGCGCTGCCTCGCCAGACCTCGCTGGGACCCCGGGGCCACCGGGAGGCACTTTT GTGGAGGGGGGAGGGGGGGCGACCTCGGCAGCCTCGGCGCACGAAGCGTCCGAGGGCAGCGTGGGGCGGGCTGCGACCTCTGCATCGGTG GACTGCATTTTTAATTAAGGATTCCCAGCAGCTCTTTGGGATTTTTACAGCTTCCACTCATGTGTTGACACCCGCGTCCAGGAGAAACTC GCTCCAAGTGCATCTAGCGCCTGGGACCTGAGACGGCGTTGGCCTTTCGTGCATGCAAATCCAGGGATTTAGGTTTTGTTTGGGATTTCC TTTTCTTTCTTTCCTTTTTTTTTTCTTTTTGCAGGGAGTAAGAAGGGAGCTGGGGGTATCAACAAGCCTGCCTTTCGGATCCTGCGGGAA AAGCCCATGTAGTTAAGCGCTTTGGTTTAAAAAAAAGGCAAGGTAAAGGCAGGGCTTTCCAGACACATTTAGGGGTTCGCGCGAGCGCTT TGTGCTCATGGACCAGCCGCACAACTTTTGAAGGCTCGCCGGCCCATGTGGGGTCTTTCTGGCGGCGCGCCGCCTGCAGCCCCCCTAAAG CGCGGGGGCTGGAGTTGTTGAGCAGCCCCGCCGCTGTGGTCCATGTAGCCGCTGGCCGCGCGCGGACTGCGGCTCGGCGTGCGCGTGTTC CCGGCCGTCCCGCCTCGGCGAGCTCCCTCATGTTGTCGCCCTGCGGCGCCCCTTCGACGACAGGCTGTGCGCGGTCTGCACGGCGCTCCG CGGCGGAGCTTCATGTGGGGCTGCGACCCGCGCAGCCGGCGCCTCGCTGAGGGAACGGACCCCCGGTAACCGGAGACCGCCTCCCCCCCA CCCCTGGCGCCAAAGGATATCGTATGTTCAGGTCCAAACGCTCGGGGCTGGTGCGGCGACTTTGGCGAAGTCGTGTGGTCCCCGACCGGG AGGAAGGCGGCAGCGGCGGCGGCGGTGGCGGCGACGAGGATGGGAGCTTGGGCAGCCGAGCTGAGCCGGCCCCGCGGGCAAGAGAGGGCG GAGGCTGCGGCCGCTCCGAAGTCCGCCCGGTAGCCCCGCGGCGGCCCCGGGACGCAGTGGGACAGCGAGGCGCCCAGGGCGCGGGGAGGC GCCGGCGCGCAGGGGGCCCCCCGAGGCCCATGTCGGAGCCAGGGGCCGGCGCTGGGAGCTCCCTGCTGGACGTGGCGGAGCCGGGAGGCC CGGGCTGGCTGCCCGAGAGTGACTGCGAGACGGTGACCTGCTGTCTCTTTTCGGAGCGGGACGCCGCCGGCGCGCCCCGGGACGCCAGCG ACCCCCTGGCCGGGGCGGCCCTGGAGCCGGCGGGCGGCGGGCGGAGTCGCGAAGCGCGCTCGCGGCTGCTGCTGCTGGAGCAGGAACTCA AAACCGTCACGTACTCGCTGCTGAAGCGGCTCAAGGAGCGCTCGCTGGACACGCTGCTGGAGGCGGTGGAGTCCCGCGGCGGCGTGCCGG GCGGCTGCGTGCTGGTGCCGCGCGCCGACCTCCGCCTGGGCGGCCAGCCCGCGCCGCCGCAGCTGCTGCTCGGCCGCCTCTTTCGCTGGC CCGACCTGCAGCACGCCGTGGAGCTGAAGCCCCTGTGCGGCTGCCACAGCTTCGCCGCCGCCGCCGACGGCCCTACCGTGTGCTGCAACC CCTACCACTTCAGCCGGCTCTGCGGGCCCGAATCTCCGCCACCTCCCTACTCTCGGCTGTCTCCTCGCGACGAGTACAAGCCACTGGTGC CTAGCCCCTCCTGTAATGGTTTTGACAACACCCTTTCAAGACCTTGCCGGGTGCCACTAGGAAAGGAGGTTTGCTTCATTGCCACCGTTC GTCTGGCAACACCACAATTCTTAAAGGAAATGTGCATAGTTGACGAACCTTTAGAGGAATTCACTTCAAGGCATAGCTTGGAATGGAAAT TTTTATTTCTGGATCACAGAGCACCTCCAATCATAGGATACCTGCCTTTTGAAGTGCTGGGAACCTCAGGCTATGACTACTACCACATTG ATGACCTGGAGCTCCTGGCCAGGTGTCACCAGCACCTGATGCAGTTTGGCAAAGGGAAGTCGTGTTGCTACCGGTTTCTGACCAAAGGTC AGCAGTGGATCTGGCTGCAGACTCACTACTACATCACCTACCATCAGTGGAACTCCAAGCCCGAGTTCATCGTGTGCACACACTCGGTGG TCAGTTACGCAGATGTCCGGGTGGAAAGGAGGCAGGAGCTGGCTCTGGAAGACCCGCCATCCGAGGCCCTCCACTCCTCAGCACTAAAGG ACAAGGGCTCAAGCCTGGAACCTCGGCAGCACTTTAACACACTCGACGTGGGTGCCTCGGGCCTTAATACCAGTCATTCGCCATCGGCGT CCTCAAGAAGTTCCCACAAATCCTCGCACACAGCCATGTCAGAACCCACCTCCACTCCCACCAAGCTGATGGCAGAGGCCAGCACCCCGG CTTTGCCAAGATCAGCCACCCTGCCCCAAGAGTTACCTGTCCCCGGGCTCAGCCAGGCAGCCACCATGCCGGCCCCTCTGCCTTCCCCAT CGTCCTGCGACCTCACACAGCAGCTCCTGCCTCAGACCGTTCTGCAGAGCACGCCCGCTCCCATGGCACAGTTTTCGGCACAGTTCAGCA TGTTCCAGACCATCAAAGACCAGCTAGAGCAGCGGACGCGGATCCTGCAGGCCAATATCCGGTGGCAACAGGAAGAGCTCCACAAGATCC AGGAGCAGCTCTGCCTGGTCCAGGACTCCAACGTCCAGATGTTCCTGCAGCAGCCAGCTGTATCCCTGAGCTTCAGCAGCACCCAGCGAC CTGAGGCTCAGCAGCAGCTACAGCAAAGGTCAGCTGCAGTGACTCAGCCCCAGCTCGGGGCGGGCCCCCAACTTCCAGGGCAGATCTCCT CTGCCCAGGTCACAAGCCAGCACCTGCTCAGAGAATCAAGTGTGATATCAACCCAGGGTCCAAAGCCAATGAGAAGCTCACAGCTAATGC AGAGCAGCGGCCGCTCTGGAAGCAGCCTAGTGTCCCCGTTCAGCAGCGCCACAGCTGCGCTCCCGCCAAGTCTGAATCTGACCACACCTG CTTCCACCTCCCAGGATGCCAGCCAGTGCCAGCCCAGCCCAGACTTCAGCCATGATCGGCAGCTCAGGCTGTTGCTGAGCCAGCCCATCC AGCCCATGATGCCCGGGTCCTGTGACGCAAGGCAGCCCTCGGAAGTCAGCAGGACGGGACGGCAAGTCAAGTACGCCCAGAGCCAGACCG TGTTTCAAAATCCAGACGCACACCCCGCCAACAGCAGCAGCGCCCCGATGCCCGTCCTGCTGATGGGGCAGGCGGTGCTCCACCCCAGCT TCCCTGCCTCCCAACCATCGCCCCTGCAGCCTGCACAGGCCCGGCAGCAGCCACCGCAGCACTACCTGCAGGTACAGGCACCAACCTCTT TGCACAGTGAGCAGCAGGACTCGCTACTTCTCTCCACCTACTCACAACAGCCAGGGACCCTGGGCTACCCCCAACCACCCCCAGCACAGC CCCAGCCCCTACGTCCTCCCCGAAGGGTCAGCAGTCTGTCTGAGTCGTCAGGCCTCCAGCAGCCGCCCCGATAATGCCCCGGCACTGAAG TCGGGACACAATCAGCTTTAACCAATGGATGAGGGGGGTGGCCACAGGAGATGGGGAGAGGAGTCTGAACTAAACCCCTGGCTTTTGTGC ACACTGCATACGTTTCAGAACTCCTGGATGGTAACCATCTCTGGAGTGCAGCGCTTGCTGCAGTGGAAATGATCAGGAATACTGACCGTG TTTCTCTTGCCTCCGAGGTTCTTGGGCACACTCTATAGCCATACTGGACAGGAACCAGGTGCCCCGTGTAGGCATCGTCGGTCGGTTTGC CGTCAGAGATGGCGCATCTCGCTGCATCCCCCGAGAGTACACCGGTTGCTCTAGCCACCTGCGGCCCGCCCATCTGCGCTAGCTGGCCTT CACGCTCTTGATCGTCTTTCCTTTGTATTGGAGAAGGACTGGGTCAGAGATCTGTTGGAGAGAGAGAATAAAGAGATTATTTTTCATTAT TTTTAAATGGTTGTTTTTGTTTTAATTTGCACAGCTACACAGAGGAAATAACTTAGGCACTTTCTGTTTTTTTTAAAAAAATAATAAGGT CTCATGGCTTCATTTAGAGACCACAGTAACAACAGCAGCCCACCAATCAGAGAAGCTGGTTGTTATTAACCAAGCTACAGATTCACACTT TCTGGCCTAAACCCTAATGGGATGAGGCTTTTCACCCCAGGCCATGCTGGTGGTGATTTTTTAGCCCCTAAATAAAACACTGGACTATTT CCTGTTTACTTCATTGATTGCAACTACAAAGGTGGACTCAAAGCAAAGCACAATCATGCCAGCCAACATTCCAGAATTCTGCTGAGAACT CCAAGTCTGTGAGGGGAGAGGTTTTACAAGCCAGACAGGCCTGGGGGACTGCAGTCCCCAAGGAGACCCTGCCACATGCTGGCCCTTTGA GTGAGAATGCTGCATCTTTCTACATATCTTCATGAGAATACTGAGAATTGGATTTTCCTTTTCAAAATGCACTTTGCTTTTTTTGTATGT TTTGTTATGTTGAGATGTTTCTAAAGAAAAGATTTTATGTAATTATAAGATGAAGCGTAGTGAATTGTACAGCTGTTGTAATAATGACCT ATTTCTATATAAAATAAAATTGTATGGCTTATGTGTAAATTATTTTGTATCTGAGATACCAGTTCCTTTTCCCAAATATAAAAGTATAAA >83827_83827_5_SMAD6-NPAS2_SMAD6_chr15_67004062_ENST00000457357_NPAS2_chr2_101580519_ENST00000335681_length(amino acids)=916AA_BP=289 MFRSKRSGLVRRLWRSRVVPDREEGGSGGGGGGDEDGSLGSRAEPAPRAREGGGCGRSEVRPVAPRRPRDAVGQRGAQGAGRRRRAGGPP RPMSEPGAGAGSSLLDVAEPGGPGWLPESDCETVTCCLFSERDAAGAPRDASDPLAGAALEPAGGGRSREARSRLLLLEQELKTVTYSLL KRLKERSLDTLLEAVESRGGVPGGCVLVPRADLRLGGQPAPPQLLLGRLFRWPDLQHAVELKPLCGCHSFAAAADGPTVCCNPYHFSRLC GPESPPPPYSRLSPRDEYKPLVPSPSCNGFDNTLSRPCRVPLGKEVCFIATVRLATPQFLKEMCIVDEPLEEFTSRHSLEWKFLFLDHRA PPIIGYLPFEVLGTSGYDYYHIDDLELLARCHQHLMQFGKGKSCCYRFLTKGQQWIWLQTHYYITYHQWNSKPEFIVCTHSVVSYADVRV ERRQELALEDPPSEALHSSALKDKGSSLEPRQHFNTLDVGASGLNTSHSPSASSRSSHKSSHTAMSEPTSTPTKLMAEASTPALPRSATL PQELPVPGLSQAATMPAPLPSPSSCDLTQQLLPQTVLQSTPAPMAQFSAQFSMFQTIKDQLEQRTRILQANIRWQQEELHKIQEQLCLVQ DSNVQMFLQQPAVSLSFSSTQRPEAQQQLQQRSAAVTQPQLGAGPQLPGQISSAQVTSQHLLRESSVISTQGPKPMRSSQLMQSSGRSGS SLVSPFSSATAALPPSLNLTTPASTSQDASQCQPSPDFSHDRQLRLLLSQPIQPMMPGSCDARQPSEVSRTGRQVKYAQSQTVFQNPDAH PANSSSAPMPVLLMGQAVLHPSFPASQPSPLQPAQARQQPPQHYLQVQAPTSLHSEQQDSLLLSTYSQQPGTLGYPQPPPAQPQPLRPPR -------------------------------------------------------------- >83827_83827_6_SMAD6-NPAS2_SMAD6_chr15_67004062_ENST00000457357_NPAS2_chr2_101580519_ENST00000542504_length(transcript)=3727nt_BP=1797nt CGATCGAGGGAGCTGAGCCGAGAGAAAGAGCCGCCGGGCGCTGCCTCGCCAGACCTCGCTGGGACCCCGGGGCCACCGGGAGGCACTTTT GTGGAGGGGGGAGGGGGGGCGACCTCGGCAGCCTCGGCGCACGAAGCGTCCGAGGGCAGCGTGGGGCGGGCTGCGACCTCTGCATCGGTG GACTGCATTTTTAATTAAGGATTCCCAGCAGCTCTTTGGGATTTTTACAGCTTCCACTCATGTGTTGACACCCGCGTCCAGGAGAAACTC GCTCCAAGTGCATCTAGCGCCTGGGACCTGAGACGGCGTTGGCCTTTCGTGCATGCAAATCCAGGGATTTAGGTTTTGTTTGGGATTTCC TTTTCTTTCTTTCCTTTTTTTTTTCTTTTTGCAGGGAGTAAGAAGGGAGCTGGGGGTATCAACAAGCCTGCCTTTCGGATCCTGCGGGAA AAGCCCATGTAGTTAAGCGCTTTGGTTTAAAAAAAAGGCAAGGTAAAGGCAGGGCTTTCCAGACACATTTAGGGGTTCGCGCGAGCGCTT TGTGCTCATGGACCAGCCGCACAACTTTTGAAGGCTCGCCGGCCCATGTGGGGTCTTTCTGGCGGCGCGCCGCCTGCAGCCCCCCTAAAG CGCGGGGGCTGGAGTTGTTGAGCAGCCCCGCCGCTGTGGTCCATGTAGCCGCTGGCCGCGCGCGGACTGCGGCTCGGCGTGCGCGTGTTC CCGGCCGTCCCGCCTCGGCGAGCTCCCTCATGTTGTCGCCCTGCGGCGCCCCTTCGACGACAGGCTGTGCGCGGTCTGCACGGCGCTCCG CGGCGGAGCTTCATGTGGGGCTGCGACCCGCGCAGCCGGCGCCTCGCTGAGGGAACGGACCCCCGGTAACCGGAGACCGCCTCCCCCCCA CCCCTGGCGCCAAAGGATATCGTATGTTCAGGTCCAAACGCTCGGGGCTGGTGCGGCGACTTTGGCGAAGTCGTGTGGTCCCCGACCGGG AGGAAGGCGGCAGCGGCGGCGGCGGTGGCGGCGACGAGGATGGGAGCTTGGGCAGCCGAGCTGAGCCGGCCCCGCGGGCAAGAGAGGGCG GAGGCTGCGGCCGCTCCGAAGTCCGCCCGGTAGCCCCGCGGCGGCCCCGGGACGCAGTGGGACAGCGAGGCGCCCAGGGCGCGGGGAGGC GCCGGCGCGCAGGGGGCCCCCCGAGGCCCATGTCGGAGCCAGGGGCCGGCGCTGGGAGCTCCCTGCTGGACGTGGCGGAGCCGGGAGGCC CGGGCTGGCTGCCCGAGAGTGACTGCGAGACGGTGACCTGCTGTCTCTTTTCGGAGCGGGACGCCGCCGGCGCGCCCCGGGACGCCAGCG ACCCCCTGGCCGGGGCGGCCCTGGAGCCGGCGGGCGGCGGGCGGAGTCGCGAAGCGCGCTCGCGGCTGCTGCTGCTGGAGCAGGAACTCA AAACCGTCACGTACTCGCTGCTGAAGCGGCTCAAGGAGCGCTCGCTGGACACGCTGCTGGAGGCGGTGGAGTCCCGCGGCGGCGTGCCGG GCGGCTGCGTGCTGGTGCCGCGCGCCGACCTCCGCCTGGGCGGCCAGCCCGCGCCGCCGCAGCTGCTGCTCGGCCGCCTCTTTCGCTGGC CCGACCTGCAGCACGCCGTGGAGCTGAAGCCCCTGTGCGGCTGCCACAGCTTCGCCGCCGCCGCCGACGGCCCTACCGTGTGCTGCAACC CCTACCACTTCAGCCGGCTCTGCGGGCCCGAATCTCCGCCACCTCCCTACTCTCGGCTGTCTCCTCGCGACGAGTACAAGCCACTGGTGC CTAGCCCCTCCTGTAATGGTTTTGACAACACCCTTTCAAGACCTTGCCGGGTGCCACTAGGAAAGGAGGTTTGCTTCATTGCCACCGTTC GTCTGGCAACACCACAATTCTTAAAGGAAATGTGCATAGTTGACGAACCTTTAGAGGAATTCACTTCAAGGCATAGCTTGGAATGGAAAT TTTTATTTCTGGATCACAGAGCACCTCCAATCATAGGATACCTGCCTTTTGAAGTGCTGGGAACCTCAGGCTATGACTACTACCACATTG ATGACCTGGAGCTCCTGGCCAGGTGTCACCAGCACCTGATGCAGTTTGGCAAAGGGAAGTCGTGTTGCTACCGGTTTCTGACCAAAGGTC AGCAGTGGATCTGGCTGCAGACTCACTACTACATCACCTACCATCAGTGGAACTCCAAGCCCGAGTTCATCGTGTGCACACACTCGGTGG TCAGTTACGCAGATGTCCGGGTGGAAAGGAGGCAGGAGCTGGCTCTGGAAGACCCGCCATCCGAGGCCCTCCACTCCTCAGCACTAAAGG ACAAGGGCTCAAGCCTGGAACCTCGGCAGCACTTTAACACACTCGACGTGGGTGCCTCGGGCCTTAATACCAGTCATTCGCCATCGGCGT CCTCAAGAAGTTCCCACAAATCCTCGCACACAGCCATGTCAGAACCCACCTCCACTCCCACCAAGCTGATGGCAGAGGCCAGCACCCCGG CTTTGCCAAGATCAGCCACCCTGCCCCAAGAGTTACCTGTCCCCGGGCTCAGCCAGGCAGCCACCATGCCGGCCCCTCTGCCTTCCCCAT CGTCCTGCGACCTCACACAGCAGCTCCTGCCTCAGACCGTTCTGCAGAGCACGCCCGCTCCCATGGCACAGTTTTCGGCACAGTTCAGCA TGTTCCAGACCATCAAAGACCAGCTAGAGCAGCGGACGCGGATCCTGCAGGCCAATATCCGGTGGCAACAGGAAGAGCTCCACAAGATCC AGGAGCAGCTCTGCCTGGTCCAGGACTCCAACGTCCAGATGTTCCTGCAGCAGCCAGCTGTATCCCTGAGCTTCAGCAGCACCCAGCGAC CTGAGGCTCAGCAGCAGCTACAGCAAAGGTCAGCTGCAGTGACTCAGCCCCAGCTCGGGGCGGGCCCCCAACTTCCAGGGCAGATCTCCT CTGCCCAGGTCACAAGCCAGCACCTGCTCAGAGAATCAAGTGTGATATCAACCCAGGGTCCAAAGCCAATGAGAAGCTCACAGCTAATGC AGAGCAGCGGCCGCTCTGGAAGCAGCCTAGTGTCCCCGTTCAGCAGCGCCACAGCTGCGCTCCCGCCAAGTCTGAATCTGACCACACCTG CTTCCACCTCCCAGGATGCCAGCCAGTGCCAGCCCAGCCCAGACTTCAGCCATGATCGGCAGCTCAGGCTGTTGCTGAGCCAGCCCATCC AGCCCATGATGCCCGGGTCCTGTGACGCAAGGCAGCCCTCGGAAGTCAGCAGGACGGGACGGCAAGTCAAGTACGCCCAGAGCCAGACCG TGTTTCAAAATCCAGACGCACACCCCGCCAACAGCAGCAGCGCCCCGATGCCCGTCCTGCTGATGGGGCAGGCGGTGCTCCACCCCAGCT TCCCTGCCTCCCAACCATCGCCCCTGCAGCCTGCACAGGCCCGGCAGCAGCCACCGCAGCACTACCTGCAGGTACAGGCACCAACCTCTT TGCACAGTGAGCAGCAGGACTCGCTACTTCTCTCCACCTACTCACAACAGCCAGGGACCCTGGGCTACCCCCAACCACCCCCAGCACAGC CCCAGCCCCTACGTCCTCCCCGAAGGGTCAGCAGTCTGTCTGAGTCGTCAGGCCTCCAGCAGCCGCCCCGATAATGCCCCGGCACTGAAG >83827_83827_6_SMAD6-NPAS2_SMAD6_chr15_67004062_ENST00000457357_NPAS2_chr2_101580519_ENST00000542504_length(amino acids)=916AA_BP=289 MFRSKRSGLVRRLWRSRVVPDREEGGSGGGGGGDEDGSLGSRAEPAPRAREGGGCGRSEVRPVAPRRPRDAVGQRGAQGAGRRRRAGGPP RPMSEPGAGAGSSLLDVAEPGGPGWLPESDCETVTCCLFSERDAAGAPRDASDPLAGAALEPAGGGRSREARSRLLLLEQELKTVTYSLL KRLKERSLDTLLEAVESRGGVPGGCVLVPRADLRLGGQPAPPQLLLGRLFRWPDLQHAVELKPLCGCHSFAAAADGPTVCCNPYHFSRLC GPESPPPPYSRLSPRDEYKPLVPSPSCNGFDNTLSRPCRVPLGKEVCFIATVRLATPQFLKEMCIVDEPLEEFTSRHSLEWKFLFLDHRA PPIIGYLPFEVLGTSGYDYYHIDDLELLARCHQHLMQFGKGKSCCYRFLTKGQQWIWLQTHYYITYHQWNSKPEFIVCTHSVVSYADVRV ERRQELALEDPPSEALHSSALKDKGSSLEPRQHFNTLDVGASGLNTSHSPSASSRSSHKSSHTAMSEPTSTPTKLMAEASTPALPRSATL PQELPVPGLSQAATMPAPLPSPSSCDLTQQLLPQTVLQSTPAPMAQFSAQFSMFQTIKDQLEQRTRILQANIRWQQEELHKIQEQLCLVQ DSNVQMFLQQPAVSLSFSSTQRPEAQQQLQQRSAAVTQPQLGAGPQLPGQISSAQVTSQHLLRESSVISTQGPKPMRSSQLMQSSGRSGS SLVSPFSSATAALPPSLNLTTPASTSQDASQCQPSPDFSHDRQLRLLLSQPIQPMMPGSCDARQPSEVSRTGRQVKYAQSQTVFQNPDAH PANSSSAPMPVLLMGQAVLHPSFPASQPSPLQPAQARQQPPQHYLQVQAPTSLHSEQQDSLLLSTYSQQPGTLGYPQPPPAQPQPLRPPR -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for SMAD6-NPAS2 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for SMAD6-NPAS2 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for SMAD6-NPAS2 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
| Hgene | SMAD6 | C0149630 | Bicuspid aortic valve | 2 | CTD_human;ORPHANET |
| Hgene | SMAD6 | C3542024 | AORTIC VALVE DISEASE 2 | 2 | CTD_human;GENOMICS_ENGLAND;UNIPROT |
| Hgene | SMAD6 | C4479496 | CRANIOSYNOSTOSIS 7 | 2 | GENOMICS_ENGLAND;UNIPROT |
| Hgene | SMAD6 | C0158761 | Radioulnar Synostosis | 1 | GENOMICS_ENGLAND |
| Hgene | SMAD6 | C0265534 | Scaphycephaly | 1 | CTD_human;GENOMICS_ENGLAND |
| Hgene | SMAD6 | C0428791 | Aortic valve calcification | 1 | ORPHANET |
| Hgene | SMAD6 | C1260873 | Aortic valve disorder | 1 | ORPHANET |
| Hgene | SMAD6 | C1860819 | Metopic synostosis | 1 | CTD_human;GENOMICS_ENGLAND |
| Hgene | SMAD6 | C3887892 | Aortic Valve Disease 1 | 1 | ORPHANET |
| Tgene | C0005586 | Bipolar Disorder | 5 | PSYGENET | |
| Tgene | C0085159 | Seasonal Affective Disorder | 4 | PSYGENET | |
| Tgene | C0011581 | Depressive disorder | 2 | PSYGENET | |
| Tgene | C0004352 | Autistic Disorder | 1 | CTD_human | |
| Tgene | C0011570 | Mental Depression | 1 | PSYGENET | |
| Tgene | C0023893 | Liver Cirrhosis, Experimental | 1 | CTD_human | |
| Tgene | C0036337 | Schizoaffective Disorder | 1 | PSYGENET | |
| Tgene | C0036341 | Schizophrenia | 1 | PSYGENET |
