
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:CRLS1-NFX1 (FusionGDB2 ID:HG54675TG4799) |
Fusion Gene Summary for CRLS1-NFX1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: CRLS1-NFX1 | Fusion gene ID: hg54675tg4799 | Hgene | Tgene | Gene symbol | CRLS1 | NFX1 | Gene ID | 54675 | 4799 |
| Gene name | cardiolipin synthase 1 | nuclear transcription factor, X-box binding 1 | |
| Synonyms | C20orf155|CLS|CLS1|GCD10|dJ967N21.6 | NFX2|TEG-42|Tex42 | |
| Cytomap | ('CRLS1')('NFX1') 20p12.3 | 9p13.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | cardiolipin synthase (CMP-forming) | transcriptional repressor NF-X1nuclear transcription factor, X box-binding protein 1 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000378863, ENST00000378868, ENST00000452938, ENST00000464921, | ||
| Fusion gene scores | * DoF score | 8 X 8 X 5=320 | 14 X 12 X 9=1512 |
| # samples | 8 | 15 | |
| ** MAII score | log2(8/320*10)=-2 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(15/1512*10)=-3.33342373372519 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: CRLS1 [Title/Abstract] AND NFX1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | CRLS1(5996136)-NFX1(33328579), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | CRLS1-NFX1 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. CRLS1-NFX1 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. CRLS1-NFX1 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. CRLS1-NFX1 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | NFX1 | GO:0000122 | negative regulation of transcription by RNA polymerase II | 7964459 |
| Tgene | NFX1 | GO:0051865 | protein autoubiquitination | 10500182 |
 Fusion gene breakpoints across CRLS1 (5'-gene) Fusion gene breakpoints across CRLS1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
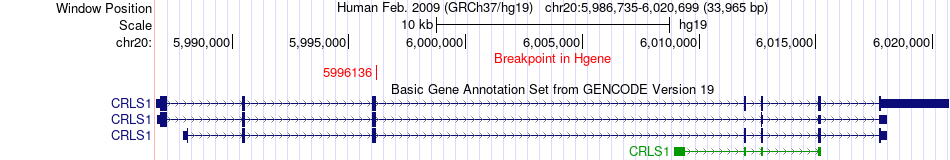 |
 Fusion gene breakpoints across NFX1 (3'-gene) Fusion gene breakpoints across NFX1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
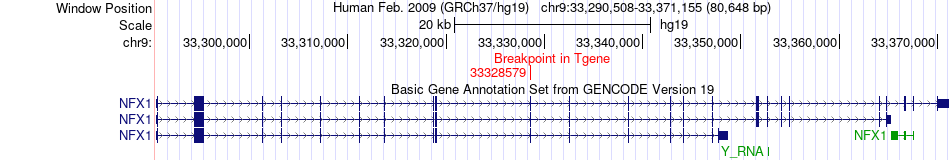 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | KIRC | TCGA-BP-5199-01A | CRLS1 | chr20 | 5996136 | - | NFX1 | chr9 | 33328579 | + |
| ChimerDB4 | KIRC | TCGA-BP-5199-01A | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + |
Top |
Fusion Gene ORF analysis for CRLS1-NFX1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000378863 | ENST00000463421 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + |
| 5CDS-intron | ENST00000378868 | ENST00000463421 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + |
| 5CDS-intron | ENST00000452938 | ENST00000463421 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + |
| In-frame | ENST00000378863 | ENST00000318524 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + |
| In-frame | ENST00000378863 | ENST00000379521 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + |
| In-frame | ENST00000378863 | ENST00000379540 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + |
| In-frame | ENST00000378868 | ENST00000318524 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + |
| In-frame | ENST00000378868 | ENST00000379521 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + |
| In-frame | ENST00000378868 | ENST00000379540 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + |
| In-frame | ENST00000452938 | ENST00000318524 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + |
| In-frame | ENST00000452938 | ENST00000379521 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + |
| In-frame | ENST00000452938 | ENST00000379540 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + |
| intron-3CDS | ENST00000464921 | ENST00000318524 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + |
| intron-3CDS | ENST00000464921 | ENST00000379521 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + |
| intron-3CDS | ENST00000464921 | ENST00000379540 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + |
| intron-intron | ENST00000464921 | ENST00000463421 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000378863 | CRLS1 | chr20 | 5996136 | + | ENST00000379540 | NFX1 | chr9 | 33328579 | + | 3367 | 731 | 157 | 2187 | 676 |
| ENST00000378863 | CRLS1 | chr20 | 5996136 | + | ENST00000379521 | NFX1 | chr9 | 33328579 | + | 2373 | 731 | 157 | 1899 | 580 |
| ENST00000378863 | CRLS1 | chr20 | 5996136 | + | ENST00000318524 | NFX1 | chr9 | 33328579 | + | 2310 | 731 | 157 | 1326 | 389 |
| ENST00000452938 | CRLS1 | chr20 | 5996136 | + | ENST00000379540 | NFX1 | chr9 | 33328579 | + | 3307 | 671 | 97 | 2127 | 676 |
| ENST00000452938 | CRLS1 | chr20 | 5996136 | + | ENST00000379521 | NFX1 | chr9 | 33328579 | + | 2313 | 671 | 97 | 1839 | 580 |
| ENST00000452938 | CRLS1 | chr20 | 5996136 | + | ENST00000318524 | NFX1 | chr9 | 33328579 | + | 2250 | 671 | 97 | 1266 | 389 |
| ENST00000378868 | CRLS1 | chr20 | 5996136 | + | ENST00000379540 | NFX1 | chr9 | 33328579 | + | 3063 | 427 | 150 | 1883 | 577 |
| ENST00000378868 | CRLS1 | chr20 | 5996136 | + | ENST00000379521 | NFX1 | chr9 | 33328579 | + | 2069 | 427 | 150 | 1595 | 481 |
| ENST00000378868 | CRLS1 | chr20 | 5996136 | + | ENST00000318524 | NFX1 | chr9 | 33328579 | + | 2006 | 427 | 150 | 1022 | 290 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000378863 | ENST00000379540 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + | 0.007719378 | 0.9922806 |
| ENST00000378863 | ENST00000379521 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + | 0.013743109 | 0.9862569 |
| ENST00000378863 | ENST00000318524 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + | 0.07748867 | 0.92251134 |
| ENST00000452938 | ENST00000379540 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + | 0.008885396 | 0.9911146 |
| ENST00000452938 | ENST00000379521 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + | 0.017478807 | 0.98252124 |
| ENST00000452938 | ENST00000318524 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + | 0.08388378 | 0.91611624 |
| ENST00000378868 | ENST00000379540 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + | 0.003239489 | 0.99676055 |
| ENST00000378868 | ENST00000379521 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + | 0.003770057 | 0.99622995 |
| ENST00000378868 | ENST00000318524 | CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328579 | + | 0.006904339 | 0.9930957 |
Top |
Fusion Genomic Features for CRLS1-NFX1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328578 | + | 1.60E-06 | 0.99999845 |
| CRLS1 | chr20 | 5996136 | + | NFX1 | chr9 | 33328578 | + | 1.60E-06 | 0.99999845 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
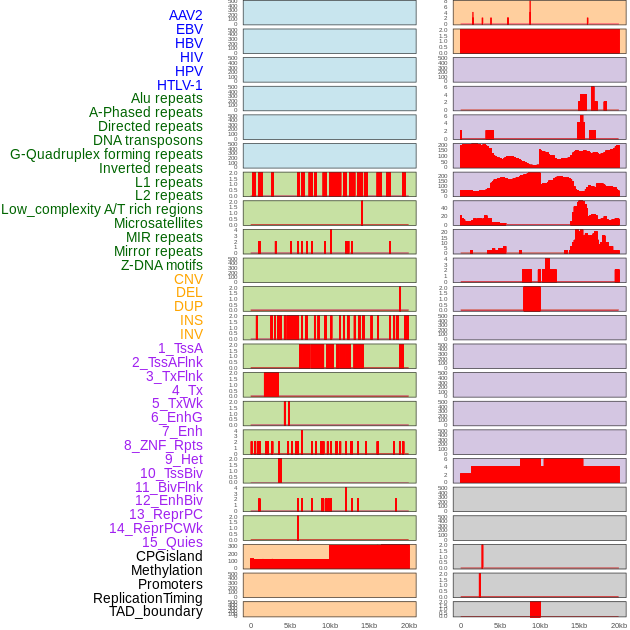 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
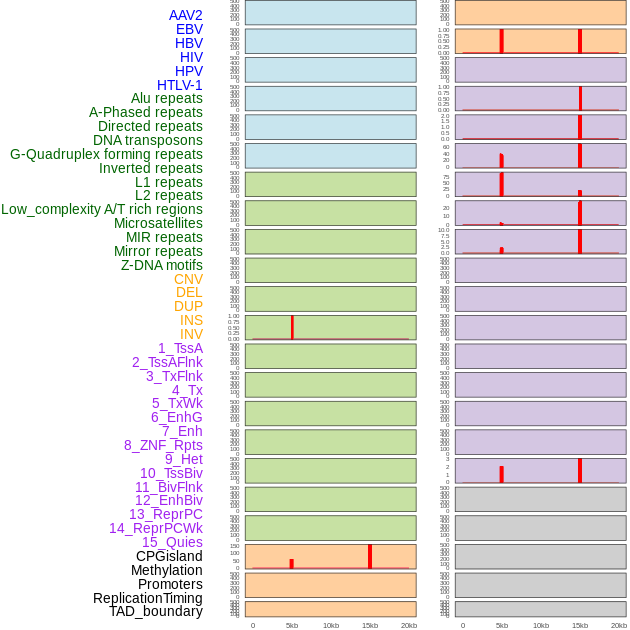 |
Top |
Fusion Protein Features for CRLS1-NFX1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr20:5996136/chr9:33328579) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CRLS1 | chr20:5996136 | chr9:33328579 | ENST00000378863 | + | 3 | 7 | 109_129 | 191 | 302.0 | Transmembrane | Helical |
| Hgene | CRLS1 | chr20:5996136 | chr9:33328579 | ENST00000378863 | + | 3 | 7 | 133_153 | 191 | 302.0 | Transmembrane | Helical |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000318524 | 8 | 16 | 1084_1089 | 635 | 834.0 | Compositional bias | Note=Poly-Pro | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379521 | 8 | 21 | 1084_1089 | 635 | 1025.0 | Compositional bias | Note=Poly-Pro | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379540 | 8 | 24 | 1084_1089 | 635 | 1121.0 | Compositional bias | Note=Poly-Pro | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000318524 | 8 | 16 | 994_1062 | 635 | 834.0 | Domain | R3H | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379521 | 8 | 21 | 994_1062 | 635 | 1025.0 | Domain | R3H | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379540 | 8 | 24 | 994_1062 | 635 | 1121.0 | Domain | R3H | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000318524 | 8 | 16 | 694_713 | 635 | 834.0 | Zinc finger | Note=NF-X1-type 5 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000318524 | 8 | 16 | 721_740 | 635 | 834.0 | Zinc finger | Note=NF-X1-type 6 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000318524 | 8 | 16 | 832_854 | 635 | 834.0 | Zinc finger | Note=NF-X1-type 7 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000318524 | 8 | 16 | 863_884 | 635 | 834.0 | Zinc finger | Note=NF-X1-type 8 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379521 | 8 | 21 | 694_713 | 635 | 1025.0 | Zinc finger | Note=NF-X1-type 5 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379521 | 8 | 21 | 721_740 | 635 | 1025.0 | Zinc finger | Note=NF-X1-type 6 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379521 | 8 | 21 | 832_854 | 635 | 1025.0 | Zinc finger | Note=NF-X1-type 7 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379521 | 8 | 21 | 863_884 | 635 | 1025.0 | Zinc finger | Note=NF-X1-type 8 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379540 | 8 | 24 | 694_713 | 635 | 1121.0 | Zinc finger | Note=NF-X1-type 5 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379540 | 8 | 24 | 721_740 | 635 | 1121.0 | Zinc finger | Note=NF-X1-type 6 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379540 | 8 | 24 | 832_854 | 635 | 1121.0 | Zinc finger | Note=NF-X1-type 7 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379540 | 8 | 24 | 863_884 | 635 | 1121.0 | Zinc finger | Note=NF-X1-type 8 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CRLS1 | chr20:5996136 | chr9:33328579 | ENST00000378863 | + | 3 | 7 | 190_212 | 191 | 302.0 | Transmembrane | Helical |
| Hgene | CRLS1 | chr20:5996136 | chr9:33328579 | ENST00000378863 | + | 3 | 7 | 250_270 | 191 | 302.0 | Transmembrane | Helical |
| Hgene | CRLS1 | chr20:5996136 | chr9:33328579 | ENST00000378863 | + | 3 | 7 | 272_292 | 191 | 302.0 | Transmembrane | Helical |
| Hgene | CRLS1 | chr20:5996136 | chr9:33328579 | ENST00000378868 | + | 3 | 7 | 109_129 | 92 | 203.0 | Transmembrane | Helical |
| Hgene | CRLS1 | chr20:5996136 | chr9:33328579 | ENST00000378868 | + | 3 | 7 | 133_153 | 92 | 203.0 | Transmembrane | Helical |
| Hgene | CRLS1 | chr20:5996136 | chr9:33328579 | ENST00000378868 | + | 3 | 7 | 190_212 | 92 | 203.0 | Transmembrane | Helical |
| Hgene | CRLS1 | chr20:5996136 | chr9:33328579 | ENST00000378868 | + | 3 | 7 | 250_270 | 92 | 203.0 | Transmembrane | Helical |
| Hgene | CRLS1 | chr20:5996136 | chr9:33328579 | ENST00000378868 | + | 3 | 7 | 272_292 | 92 | 203.0 | Transmembrane | Helical |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000318524 | 8 | 16 | 358_409 | 635 | 834.0 | Zinc finger | RING-type%3B atypical | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000318524 | 8 | 16 | 453_471 | 635 | 834.0 | Zinc finger | Note=NF-X1-type 1 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000318524 | 8 | 16 | 506_525 | 635 | 834.0 | Zinc finger | Note=NF-X1-type 2 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000318524 | 8 | 16 | 567_586 | 635 | 834.0 | Zinc finger | Note=NF-X1-type 3 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000318524 | 8 | 16 | 632_655 | 635 | 834.0 | Zinc finger | Note=NF-X1-type 4 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379521 | 8 | 21 | 358_409 | 635 | 1025.0 | Zinc finger | RING-type%3B atypical | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379521 | 8 | 21 | 453_471 | 635 | 1025.0 | Zinc finger | Note=NF-X1-type 1 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379521 | 8 | 21 | 506_525 | 635 | 1025.0 | Zinc finger | Note=NF-X1-type 2 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379521 | 8 | 21 | 567_586 | 635 | 1025.0 | Zinc finger | Note=NF-X1-type 3 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379521 | 8 | 21 | 632_655 | 635 | 1025.0 | Zinc finger | Note=NF-X1-type 4 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379540 | 8 | 24 | 358_409 | 635 | 1121.0 | Zinc finger | RING-type%3B atypical | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379540 | 8 | 24 | 453_471 | 635 | 1121.0 | Zinc finger | Note=NF-X1-type 1 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379540 | 8 | 24 | 506_525 | 635 | 1121.0 | Zinc finger | Note=NF-X1-type 2 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379540 | 8 | 24 | 567_586 | 635 | 1121.0 | Zinc finger | Note=NF-X1-type 3 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379540 | 8 | 24 | 632_655 | 635 | 1121.0 | Zinc finger | Note=NF-X1-type 4 |
Top |
Fusion Gene Sequence for CRLS1-NFX1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >19520_19520_1_CRLS1-NFX1_CRLS1_chr20_5996136_ENST00000378863_NFX1_chr9_33328579_ENST00000318524_length(transcript)=2310nt_BP=731nt TGTATAAGTGGAGTGTGCTGGGGTGTGTAAAGTAGTATGGAGGCAGCGGTAGCCCAGTGTCTGAGTGGTTGCCGGGTCTCCATGGAGAAG CGGCTCGCCAGTGTCCCAGGCTGCTGAGCTCTCGCCGCCCGAGACCCCGCGGCGCGGCCGCAGGGCCATGCTAGCCTTGCGCGTGGCGCG CGGCTCGTGGGGGGCCCTGCGCGGCGCCGCTTGGGCTCCGGGAACGCGGCCGAGTAAGCGACGCGCCTGCTGGGCCCTGCTGCCGCCCGT GCCCTGCTGCTTGGGCTGCCTGGCCGAACGCTGGAGGCTGCGTCCGGCCGCTCTTGGCTTGCGGCTGCCCGGGATCGGCCAGCGGAACCA CTGTTCGGGCGCGGGGAAGGCGGCTCCCAGGCCAGCGGCCGGAGCGGGCGCCGCTGCCGAAGCCCCGGGCGGCCAGTGGGGCCCGGCGAG CACCCCCAGCCTGTATGAAAACCCATGGACAATCCCGAATATGTTGTCAATGACGAGAATTGGCTTGGCCCCAGTTCTGGGCTATTTGAT TATTGAAGAAGATTTTAATATTGCACTAGGAGTTTTTGCTTTAGCTGGACTAACAGATTTGTTGGATGGATTTATTGCTCGAAACTGGGC CAATCAAAGATCAGCTTTGGGAAGTGCTCTTGATCCACTTGCTGATAAAATACTTATCAGTATCTTATATGTTAGCTTGACCTATGCAGA TCTTATTCCAGATTTCATTCATACCTGTGAAAAGCTCTGCCATGAAGGAGACTGTGGACCATGCTCTCGCACATCAGTTATTTCCTGCAG ATGCTCTTTCAGAACAAAGGAGCTTCCATGTACCAGTCTCAAAAGTGAAGATGCTACATTTATGTGTGACAAGCGGTGTAACAAGAAACG GTTGTGTGGACGGCATAAATGTAATGAGATATGCTGTGTGGATAAGGAGCACAAGTGTCCTTTGATTTGTGGGAGGAAACTCCGTTGTGG CCTTCATAGGTGTGAAGAACCTTGTCATCGTGGAAACTGCCAGACATGCTGGCAAGCCAGTTTTGATGAATTAACCTGCCATTGTGGTGC ATCAGTGATTTACCCTCCAGTTCCCTGTGGTACTAGGCCCCCTGAATGTACCCAAACCTGCGCTAGAGTCCATGAGTGTGACCATCCAGT ATATCATTCTTGTCATAGTGAGGAGAAGTGTCCCCCTTGCACTTTCCTAACTCAGAAGTGGTGCATGGGCAAGCATGAGCAGTCCCACTA CTGGGCGTCTACCCAGAAGAAAAGAAGTCATTATATGAAAAAGATACCTGCACACGCATGTTTATAGCAGCACAATTCACAATTGCAAAA ATGTGGAACCAGCCCAACTGCCCATCAGTCAACAAGTGGATAAAGAAATTGTGGTGTATCTATACGTACCATGGAATACTACTCAGCCAG GAACGAAATAATGGCATTCACAGCAACCTGGATGGATTTGGAGACCATTATTCTAAGTGAAGTAACTCAGGAATGGAAAACCAAACATCG TATGTTCTCAATTATAAGAGGGAGCTAAGCTATGAGGACGCAAAGGCATGAGAATGATACAATGGACTTTGGGGACTCTGGGGAAAGGAC GGGAGGCGGGTGAGGGATAAAAAACTACACACTGGGTCCGAGCACGGTGGCTCATGCCTATAATCCCAGCACTTTGGGAGGCCAAGGCAG GTAGATTATGAGGTCGGGAGTTCGAAACCAGCCTGGCCAACAAGGTGAGACCCCCCCCATCTCTACTAAAATAATTAGCCGGGCCTGGTG GCGCATCCCTGTAATCCCAGCTACTTTACTTGGGAGGCTGAGGCAGGAGAATTGCTTGAACTCGGGAGGCGGATGTTGCAGTGAGCCAAT ATCGCACCACTGCACTCCAGCCTGGGCAACAGAGCAAGACTCCGTCTCGAAAAAAAAAATAAATTTTAAAAACTACACATTGGGTACAAT GTACACTCCTCGGGTGATGGGTGCACCAAAATCTCAAAAATCACTACTAAGGAACTTATCCATGTAACCAAACACCACCTGTTCCCCAAG AACTATTGAAATAAGAAAAAAGAAAAAAAAAAAATCACTGATTGCAGGTCACCCTAACAGATATAATAATGAAAAAGTCTGAGATGTTGC GAGAATTACCAAAATGTGATGCAGACACGAAATGAGCATGTGCTGTTGGAAAAGTGGAACAGATAGACTTGCTGGAGACAGGGTTGCCAC >19520_19520_1_CRLS1-NFX1_CRLS1_chr20_5996136_ENST00000378863_NFX1_chr9_33328579_ENST00000318524_length(amino acids)=389AA_BP=189 MLALRVARGSWGALRGAAWAPGTRPSKRRACWALLPPVPCCLGCLAERWRLRPAALGLRLPGIGQRNHCSGAGKAAPRPAAGAGAAAEAP GGQWGPASTPSLYENPWTIPNMLSMTRIGLAPVLGYLIIEEDFNIALGVFALAGLTDLLDGFIARNWANQRSALGSALDPLADKILISIL YVSLTYADLIPDFIHTCEKLCHEGDCGPCSRTSVISCRCSFRTKELPCTSLKSEDATFMCDKRCNKKRLCGRHKCNEICCVDKEHKCPLI CGRKLRCGLHRCEEPCHRGNCQTCWQASFDELTCHCGASVIYPPVPCGTRPPECTQTCARVHECDHPVYHSCHSEEKCPPCTFLTQKWCM -------------------------------------------------------------- >19520_19520_2_CRLS1-NFX1_CRLS1_chr20_5996136_ENST00000378863_NFX1_chr9_33328579_ENST00000379521_length(transcript)=2373nt_BP=731nt TGTATAAGTGGAGTGTGCTGGGGTGTGTAAAGTAGTATGGAGGCAGCGGTAGCCCAGTGTCTGAGTGGTTGCCGGGTCTCCATGGAGAAG CGGCTCGCCAGTGTCCCAGGCTGCTGAGCTCTCGCCGCCCGAGACCCCGCGGCGCGGCCGCAGGGCCATGCTAGCCTTGCGCGTGGCGCG CGGCTCGTGGGGGGCCCTGCGCGGCGCCGCTTGGGCTCCGGGAACGCGGCCGAGTAAGCGACGCGCCTGCTGGGCCCTGCTGCCGCCCGT GCCCTGCTGCTTGGGCTGCCTGGCCGAACGCTGGAGGCTGCGTCCGGCCGCTCTTGGCTTGCGGCTGCCCGGGATCGGCCAGCGGAACCA CTGTTCGGGCGCGGGGAAGGCGGCTCCCAGGCCAGCGGCCGGAGCGGGCGCCGCTGCCGAAGCCCCGGGCGGCCAGTGGGGCCCGGCGAG CACCCCCAGCCTGTATGAAAACCCATGGACAATCCCGAATATGTTGTCAATGACGAGAATTGGCTTGGCCCCAGTTCTGGGCTATTTGAT TATTGAAGAAGATTTTAATATTGCACTAGGAGTTTTTGCTTTAGCTGGACTAACAGATTTGTTGGATGGATTTATTGCTCGAAACTGGGC CAATCAAAGATCAGCTTTGGGAAGTGCTCTTGATCCACTTGCTGATAAAATACTTATCAGTATCTTATATGTTAGCTTGACCTATGCAGA TCTTATTCCAGATTTCATTCATACCTGTGAAAAGCTCTGCCATGAAGGAGACTGTGGACCATGCTCTCGCACATCAGTTATTTCCTGCAG ATGCTCTTTCAGAACAAAGGAGCTTCCATGTACCAGTCTCAAAAGTGAAGATGCTACATTTATGTGTGACAAGCGGTGTAACAAGAAACG GTTGTGTGGACGGCATAAATGTAATGAGATATGCTGTGTGGATAAGGAGCACAAGTGTCCTTTGATTTGTGGGAGGAAACTCCGTTGTGG CCTTCATAGGTGTGAAGAACCTTGTCATCGTGGAAACTGCCAGACATGCTGGCAAGCCAGTTTTGATGAATTAACCTGCCATTGTGGTGC ATCAGTGATTTACCCTCCAGTTCCCTGTGGTACTAGGCCCCCTGAATGTACCCAAACCTGCGCTAGAGTCCATGAGTGTGACCATCCAGT ATATCATTCTTGTCATAGTGAGGAGAAGTGTCCCCCTTGCACTTTCCTAACTCAGAAGTGGTGCATGGGCAAGCATGAGTTTCGGAGCAA CATCCCCTGTCACCTGGTTGATATCTCTTGCGGATTACCCTGCAGTGCCACGCTACCATGTGGGATGCACAAATGTCAGAGACTCTGTCA CAAAGGGGAGTGTCTTGTGGATGAGCCCTGCAAGCAGCCCTGCACCACCCCCAGAGCTGACTGTGGTCACCCGTGTATGGCACCCTGCCA TACCAGCTCACCCTGCCCTGTGACTGCTTGTAAAGCTAAGGTAGAGCTACAGTGTGAATGTGGACGAAGAAAAGAGATGGTGATTTGCTC TGAAGCATCTAGTACTTATCAAAGAATAGCTGCAATCTCCATGGCCTCTAAGATAACAGACATGCAGCTTGGAGGTTCAGTGGAGATCAG CAAGTTAATTACCAAAAAGGAAGTTCATCAAGCCAGGCTGGAGTGTGATGAGGAGTGTTCAGCCTTGGAAAGGAAAAAGAGATTAGCAGA GGCATTTCATATCAGTGAGGATTCTGATCCTTTCAATATACGTTCTTCAGGGTCAAAATTCAGTGATAGTTTGAAAGAAGATGCCAGGAA GGACTTAAAGTTTGTCAGTGACGTTGAGAAGGAAATGGAAACCCTCGTGGAGGCCGTGAATAAGGTTGAAGTCGAAACATCCCACTGGAC ATTTCTCTAAGGCCAGCTTGATGAAAAAAAAAAAGCAATCAAGTCCTGGAAACACTTTCAACCTAGAGTAGTTGTAGGAAAAAGTCACAA CTTCTTTGAGGATCCTTATCTTTCTCAAAACACAGCCATTATAACATCTGTGGAAGCCTGTCCTGGTTCAGGACAGTTGACTGCAGAAAA GATCTACAGTCGGCTGGGCACAGTGTCTCATGCCTGTAATGCCAGCACTTTGGGAGGCCAAGGCAGGCAGATCACTTGAGGTCAGGAGTT CTAGATCAGCTTGGCCAACATGACGAAACCCTGTCTCTACTAAAAATATAAAAATTAGCTGGGTGTGGTGGCGGGCACCTGTTATTCCAG CTACTTGCAAGGCTTAGGCAGGAGAATTCCTTGAACCCGGGAGGCAGAGGTTGCTGTGAGCTGAGATTCTGCCACTGCACTCCAGCCTGG >19520_19520_2_CRLS1-NFX1_CRLS1_chr20_5996136_ENST00000378863_NFX1_chr9_33328579_ENST00000379521_length(amino acids)=580AA_BP=189 MLALRVARGSWGALRGAAWAPGTRPSKRRACWALLPPVPCCLGCLAERWRLRPAALGLRLPGIGQRNHCSGAGKAAPRPAAGAGAAAEAP GGQWGPASTPSLYENPWTIPNMLSMTRIGLAPVLGYLIIEEDFNIALGVFALAGLTDLLDGFIARNWANQRSALGSALDPLADKILISIL YVSLTYADLIPDFIHTCEKLCHEGDCGPCSRTSVISCRCSFRTKELPCTSLKSEDATFMCDKRCNKKRLCGRHKCNEICCVDKEHKCPLI CGRKLRCGLHRCEEPCHRGNCQTCWQASFDELTCHCGASVIYPPVPCGTRPPECTQTCARVHECDHPVYHSCHSEEKCPPCTFLTQKWCM GKHEFRSNIPCHLVDISCGLPCSATLPCGMHKCQRLCHKGECLVDEPCKQPCTTPRADCGHPCMAPCHTSSPCPVTACKAKVELQCECGR RKEMVICSEASSTYQRIAAISMASKITDMQLGGSVEISKLITKKEVHQARLECDEECSALERKKRLAEAFHISEDSDPFNIRSSGSKFSD -------------------------------------------------------------- >19520_19520_3_CRLS1-NFX1_CRLS1_chr20_5996136_ENST00000378863_NFX1_chr9_33328579_ENST00000379540_length(transcript)=3367nt_BP=731nt TGTATAAGTGGAGTGTGCTGGGGTGTGTAAAGTAGTATGGAGGCAGCGGTAGCCCAGTGTCTGAGTGGTTGCCGGGTCTCCATGGAGAAG CGGCTCGCCAGTGTCCCAGGCTGCTGAGCTCTCGCCGCCCGAGACCCCGCGGCGCGGCCGCAGGGCCATGCTAGCCTTGCGCGTGGCGCG CGGCTCGTGGGGGGCCCTGCGCGGCGCCGCTTGGGCTCCGGGAACGCGGCCGAGTAAGCGACGCGCCTGCTGGGCCCTGCTGCCGCCCGT GCCCTGCTGCTTGGGCTGCCTGGCCGAACGCTGGAGGCTGCGTCCGGCCGCTCTTGGCTTGCGGCTGCCCGGGATCGGCCAGCGGAACCA CTGTTCGGGCGCGGGGAAGGCGGCTCCCAGGCCAGCGGCCGGAGCGGGCGCCGCTGCCGAAGCCCCGGGCGGCCAGTGGGGCCCGGCGAG CACCCCCAGCCTGTATGAAAACCCATGGACAATCCCGAATATGTTGTCAATGACGAGAATTGGCTTGGCCCCAGTTCTGGGCTATTTGAT TATTGAAGAAGATTTTAATATTGCACTAGGAGTTTTTGCTTTAGCTGGACTAACAGATTTGTTGGATGGATTTATTGCTCGAAACTGGGC CAATCAAAGATCAGCTTTGGGAAGTGCTCTTGATCCACTTGCTGATAAAATACTTATCAGTATCTTATATGTTAGCTTGACCTATGCAGA TCTTATTCCAGATTTCATTCATACCTGTGAAAAGCTCTGCCATGAAGGAGACTGTGGACCATGCTCTCGCACATCAGTTATTTCCTGCAG ATGCTCTTTCAGAACAAAGGAGCTTCCATGTACCAGTCTCAAAAGTGAAGATGCTACATTTATGTGTGACAAGCGGTGTAACAAGAAACG GTTGTGTGGACGGCATAAATGTAATGAGATATGCTGTGTGGATAAGGAGCACAAGTGTCCTTTGATTTGTGGGAGGAAACTCCGTTGTGG CCTTCATAGGTGTGAAGAACCTTGTCATCGTGGAAACTGCCAGACATGCTGGCAAGCCAGTTTTGATGAATTAACCTGCCATTGTGGTGC ATCAGTGATTTACCCTCCAGTTCCCTGTGGTACTAGGCCCCCTGAATGTACCCAAACCTGCGCTAGAGTCCATGAGTGTGACCATCCAGT ATATCATTCTTGTCATAGTGAGGAGAAGTGTCCCCCTTGCACTTTCCTAACTCAGAAGTGGTGCATGGGCAAGCATGAGTTTCGGAGCAA CATCCCCTGTCACCTGGTTGATATCTCTTGCGGATTACCCTGCAGTGCCACGCTACCATGTGGGATGCACAAATGTCAGAGACTCTGTCA CAAAGGGGAGTGTCTTGTGGATGAGCCCTGCAAGCAGCCCTGCACCACCCCCAGAGCTGACTGTGGTCACCCGTGTATGGCACCCTGCCA TACCAGCTCACCCTGCCCTGTGACTGCTTGTAAAGCTAAGGTAGAGCTACAGTGTGAATGTGGACGAAGAAAAGAGATGGTGATTTGCTC TGAAGCATCTAGTACTTATCAAAGAATAGCTGCAATCTCCATGGCCTCTAAGATAACAGACATGCAGCTTGGAGGTTCAGTGGAGATCAG CAAGTTAATTACCAAAAAGGAAGTTCATCAAGCCAGGCTGGAGTGTGATGAGGAGTGTTCAGCCTTGGAAAGGAAAAAGAGATTAGCAGA GGCATTTCATATCAGTGAGGATTCTGATCCTTTCAATATACGTTCTTCAGGGTCAAAATTCAGTGATAGTTTGAAAGAAGATGCCAGGAA GGACTTAAAGTTTGTCAGTGACGTTGAGAAGGAAATGGAAACCCTCGTGGAGGCCGTGAATAAGGGAAAGAATAGTAAGAAAAGCCACAG CTTCCCTCCCATGAACAGAGACCACCGCCGGATCATCCATGACTTGGCCCAAGTTTATGGCCTGGAGAGCGTGAGCTATGACAGTGAACC GAAGCGCAATGTGGTGGTCACTGCCATCAGGGGGAAGTCCGTTTGTCCTCCTACCACGCTGACAGGTGTGCTTGAAAGGGAAATGCAGGC ACGGCCTCCACCACCGATTCCTCATCACAGACATCAGTCAGACAAGAATCCTGGGAGCAGTAATTTACAGAAAATAACCAAGGAGCCAAT AATTGACTATTTTGACGTCCAGGACTAAGAAGATCATGATGCACTTAGATAAAAGAATGATTAGGTATAGTGGAGACTTATTTGCCAGCA GATAAATCATGCCCGTTCCCCTCTGCCTGGCAGAATCACAGTCTCACATACTGTCTTGTACTGACACATCCAAAGCATGAGTGTGTCAGA AATCCCTTGTCTATTCCTGTCTGTATAAAGTGTTTCATTATGACCAGATCTCTGATTGTATGGTCACTAGGTATGCAATCACGCATTCAA AGAGGCTCTTTACACCATCACTGTGATTGCTCTGAGAGTTGAGGGACTATTGGGCTTTATTTGGACAAACCAAACTTTTAGCCTGAAACC AACTTTATGCCACTAAGTCATAGCCTCAGTTGTCCCAGTTATTTGTCCTCCTGAAAATGCCTGAAACATCAGACAGACATTGCTTGCTTT ACCCAAACTGATCAAAATCTTTAGGAGCACAAATGAATTTTTTAGTCTGAAATACCAAATAATGAATTGGTATACCATATCCGGAATCAC ACATGTTATCTTAAACCCAGCCATCATACCTAAGTCTTTTGCCAAAACCTCTCATAGGTATATCTAGCTGAACTTATTTTGGCATTTTCA ATGTGATCAGTTCTAGACCTAGAAGGGGGTCAGGCTGCTTTACAGAATTCTATTTCCTTAAGTCCCTGGCACTTCTCATACCACATCACT GAACCTGTTCAGTAACAATCAGTTTGGCCGTCCCCCATGATGGTAGGAAATATAGAGAGCAAGTTCTTCTGCCAGGGTCACACTGTGGTC TCTGAACTGACCAGTATATCCCTAACTCCTCTTTGATAGAGAAAGAGTCTCAAATGGACAACTGTCCTGTGTTGCTTTCCCTAGGCCTTC AGCAGCCTATTGGCTCTCCCTGCCTCTGAGCTCTGGACTCTGTTTGAATATTCCAAGTAGTATATGGACAGTCCAGGGCTTATGCCCAGC AGCCCACTGGAGGCATTCTTCAGGCTCCTTTAAGGCAGGTGCATTGATAGTTCCATTAGTGTGACCCTTGCATTGGCACCCCTCCAGCCT GGAGGCCAGGCTTCCAGCAACTTCCTTCTGCCCTAGAGCAAGCCATGAGCCCCAGAGCAGTAGCAGGAGACTTGAGAAGTAGAGTGACAA >19520_19520_3_CRLS1-NFX1_CRLS1_chr20_5996136_ENST00000378863_NFX1_chr9_33328579_ENST00000379540_length(amino acids)=676AA_BP=189 MLALRVARGSWGALRGAAWAPGTRPSKRRACWALLPPVPCCLGCLAERWRLRPAALGLRLPGIGQRNHCSGAGKAAPRPAAGAGAAAEAP GGQWGPASTPSLYENPWTIPNMLSMTRIGLAPVLGYLIIEEDFNIALGVFALAGLTDLLDGFIARNWANQRSALGSALDPLADKILISIL YVSLTYADLIPDFIHTCEKLCHEGDCGPCSRTSVISCRCSFRTKELPCTSLKSEDATFMCDKRCNKKRLCGRHKCNEICCVDKEHKCPLI CGRKLRCGLHRCEEPCHRGNCQTCWQASFDELTCHCGASVIYPPVPCGTRPPECTQTCARVHECDHPVYHSCHSEEKCPPCTFLTQKWCM GKHEFRSNIPCHLVDISCGLPCSATLPCGMHKCQRLCHKGECLVDEPCKQPCTTPRADCGHPCMAPCHTSSPCPVTACKAKVELQCECGR RKEMVICSEASSTYQRIAAISMASKITDMQLGGSVEISKLITKKEVHQARLECDEECSALERKKRLAEAFHISEDSDPFNIRSSGSKFSD SLKEDARKDLKFVSDVEKEMETLVEAVNKGKNSKKSHSFPPMNRDHRRIIHDLAQVYGLESVSYDSEPKRNVVVTAIRGKSVCPPTTLTG -------------------------------------------------------------- >19520_19520_4_CRLS1-NFX1_CRLS1_chr20_5996136_ENST00000378868_NFX1_chr9_33328579_ENST00000318524_length(transcript)=2006nt_BP=427nt AGTTGTACAGCATGGGTTGAGGTGGCCAGATCCTGGCAGGGGTCTCAACTCCACTGATAGCTAAAGCATGTTGGGGTTTGAATGCTGGAT ACTTTGAAGTTGCCATATCCTGACTGAAGTCCTTCCCAGAACGGCAGTAGTTGGTCGAGTATGCCACAGTATGAAAACCCATGGACAATC CCGAATATGTTGTCAATGACGAGAATTGGCTTGGCCCCAGTTCTGGGCTATTTGATTATTGAAGAAGATTTTAATATTGCACTAGGAGTT TTTGCTTTAGCTGGACTAACAGATTTGTTGGATGGATTTATTGCTCGAAACTGGGCCAATCAAAGATCAGCTTTGGGAAGTGCTCTTGAT CCACTTGCTGATAAAATACTTATCAGTATCTTATATGTTAGCTTGACCTATGCAGATCTTATTCCAGATTTCATTCATACCTGTGAAAAG CTCTGCCATGAAGGAGACTGTGGACCATGCTCTCGCACATCAGTTATTTCCTGCAGATGCTCTTTCAGAACAAAGGAGCTTCCATGTACC AGTCTCAAAAGTGAAGATGCTACATTTATGTGTGACAAGCGGTGTAACAAGAAACGGTTGTGTGGACGGCATAAATGTAATGAGATATGC TGTGTGGATAAGGAGCACAAGTGTCCTTTGATTTGTGGGAGGAAACTCCGTTGTGGCCTTCATAGGTGTGAAGAACCTTGTCATCGTGGA AACTGCCAGACATGCTGGCAAGCCAGTTTTGATGAATTAACCTGCCATTGTGGTGCATCAGTGATTTACCCTCCAGTTCCCTGTGGTACT AGGCCCCCTGAATGTACCCAAACCTGCGCTAGAGTCCATGAGTGTGACCATCCAGTATATCATTCTTGTCATAGTGAGGAGAAGTGTCCC CCTTGCACTTTCCTAACTCAGAAGTGGTGCATGGGCAAGCATGAGCAGTCCCACTACTGGGCGTCTACCCAGAAGAAAAGAAGTCATTAT ATGAAAAAGATACCTGCACACGCATGTTTATAGCAGCACAATTCACAATTGCAAAAATGTGGAACCAGCCCAACTGCCCATCAGTCAACA AGTGGATAAAGAAATTGTGGTGTATCTATACGTACCATGGAATACTACTCAGCCAGGAACGAAATAATGGCATTCACAGCAACCTGGATG GATTTGGAGACCATTATTCTAAGTGAAGTAACTCAGGAATGGAAAACCAAACATCGTATGTTCTCAATTATAAGAGGGAGCTAAGCTATG AGGACGCAAAGGCATGAGAATGATACAATGGACTTTGGGGACTCTGGGGAAAGGACGGGAGGCGGGTGAGGGATAAAAAACTACACACTG GGTCCGAGCACGGTGGCTCATGCCTATAATCCCAGCACTTTGGGAGGCCAAGGCAGGTAGATTATGAGGTCGGGAGTTCGAAACCAGCCT GGCCAACAAGGTGAGACCCCCCCCATCTCTACTAAAATAATTAGCCGGGCCTGGTGGCGCATCCCTGTAATCCCAGCTACTTTACTTGGG AGGCTGAGGCAGGAGAATTGCTTGAACTCGGGAGGCGGATGTTGCAGTGAGCCAATATCGCACCACTGCACTCCAGCCTGGGCAACAGAG CAAGACTCCGTCTCGAAAAAAAAAATAAATTTTAAAAACTACACATTGGGTACAATGTACACTCCTCGGGTGATGGGTGCACCAAAATCT CAAAAATCACTACTAAGGAACTTATCCATGTAACCAAACACCACCTGTTCCCCAAGAACTATTGAAATAAGAAAAAAGAAAAAAAAAAAA TCACTGATTGCAGGTCACCCTAACAGATATAATAATGAAAAAGTCTGAGATGTTGCGAGAATTACCAAAATGTGATGCAGACACGAAATG AGCATGTGCTGTTGGAAAAGTGGAACAGATAGACTTGCTGGAGACAGGGTTGCCACAAAGTTTCAATTTGTAAAAAATGTGATATCTATG >19520_19520_4_CRLS1-NFX1_CRLS1_chr20_5996136_ENST00000378868_NFX1_chr9_33328579_ENST00000318524_length(amino acids)=290AA_BP=90 MPQYENPWTIPNMLSMTRIGLAPVLGYLIIEEDFNIALGVFALAGLTDLLDGFIARNWANQRSALGSALDPLADKILISILYVSLTYADL IPDFIHTCEKLCHEGDCGPCSRTSVISCRCSFRTKELPCTSLKSEDATFMCDKRCNKKRLCGRHKCNEICCVDKEHKCPLICGRKLRCGL HRCEEPCHRGNCQTCWQASFDELTCHCGASVIYPPVPCGTRPPECTQTCARVHECDHPVYHSCHSEEKCPPCTFLTQKWCMGKHEQSHYW -------------------------------------------------------------- >19520_19520_5_CRLS1-NFX1_CRLS1_chr20_5996136_ENST00000378868_NFX1_chr9_33328579_ENST00000379521_length(transcript)=2069nt_BP=427nt AGTTGTACAGCATGGGTTGAGGTGGCCAGATCCTGGCAGGGGTCTCAACTCCACTGATAGCTAAAGCATGTTGGGGTTTGAATGCTGGAT ACTTTGAAGTTGCCATATCCTGACTGAAGTCCTTCCCAGAACGGCAGTAGTTGGTCGAGTATGCCACAGTATGAAAACCCATGGACAATC CCGAATATGTTGTCAATGACGAGAATTGGCTTGGCCCCAGTTCTGGGCTATTTGATTATTGAAGAAGATTTTAATATTGCACTAGGAGTT TTTGCTTTAGCTGGACTAACAGATTTGTTGGATGGATTTATTGCTCGAAACTGGGCCAATCAAAGATCAGCTTTGGGAAGTGCTCTTGAT CCACTTGCTGATAAAATACTTATCAGTATCTTATATGTTAGCTTGACCTATGCAGATCTTATTCCAGATTTCATTCATACCTGTGAAAAG CTCTGCCATGAAGGAGACTGTGGACCATGCTCTCGCACATCAGTTATTTCCTGCAGATGCTCTTTCAGAACAAAGGAGCTTCCATGTACC AGTCTCAAAAGTGAAGATGCTACATTTATGTGTGACAAGCGGTGTAACAAGAAACGGTTGTGTGGACGGCATAAATGTAATGAGATATGC TGTGTGGATAAGGAGCACAAGTGTCCTTTGATTTGTGGGAGGAAACTCCGTTGTGGCCTTCATAGGTGTGAAGAACCTTGTCATCGTGGA AACTGCCAGACATGCTGGCAAGCCAGTTTTGATGAATTAACCTGCCATTGTGGTGCATCAGTGATTTACCCTCCAGTTCCCTGTGGTACT AGGCCCCCTGAATGTACCCAAACCTGCGCTAGAGTCCATGAGTGTGACCATCCAGTATATCATTCTTGTCATAGTGAGGAGAAGTGTCCC CCTTGCACTTTCCTAACTCAGAAGTGGTGCATGGGCAAGCATGAGTTTCGGAGCAACATCCCCTGTCACCTGGTTGATATCTCTTGCGGA TTACCCTGCAGTGCCACGCTACCATGTGGGATGCACAAATGTCAGAGACTCTGTCACAAAGGGGAGTGTCTTGTGGATGAGCCCTGCAAG CAGCCCTGCACCACCCCCAGAGCTGACTGTGGTCACCCGTGTATGGCACCCTGCCATACCAGCTCACCCTGCCCTGTGACTGCTTGTAAA GCTAAGGTAGAGCTACAGTGTGAATGTGGACGAAGAAAAGAGATGGTGATTTGCTCTGAAGCATCTAGTACTTATCAAAGAATAGCTGCA ATCTCCATGGCCTCTAAGATAACAGACATGCAGCTTGGAGGTTCAGTGGAGATCAGCAAGTTAATTACCAAAAAGGAAGTTCATCAAGCC AGGCTGGAGTGTGATGAGGAGTGTTCAGCCTTGGAAAGGAAAAAGAGATTAGCAGAGGCATTTCATATCAGTGAGGATTCTGATCCTTTC AATATACGTTCTTCAGGGTCAAAATTCAGTGATAGTTTGAAAGAAGATGCCAGGAAGGACTTAAAGTTTGTCAGTGACGTTGAGAAGGAA ATGGAAACCCTCGTGGAGGCCGTGAATAAGGTTGAAGTCGAAACATCCCACTGGACATTTCTCTAAGGCCAGCTTGATGAAAAAAAAAAA GCAATCAAGTCCTGGAAACACTTTCAACCTAGAGTAGTTGTAGGAAAAAGTCACAACTTCTTTGAGGATCCTTATCTTTCTCAAAACACA GCCATTATAACATCTGTGGAAGCCTGTCCTGGTTCAGGACAGTTGACTGCAGAAAAGATCTACAGTCGGCTGGGCACAGTGTCTCATGCC TGTAATGCCAGCACTTTGGGAGGCCAAGGCAGGCAGATCACTTGAGGTCAGGAGTTCTAGATCAGCTTGGCCAACATGACGAAACCCTGT CTCTACTAAAAATATAAAAATTAGCTGGGTGTGGTGGCGGGCACCTGTTATTCCAGCTACTTGCAAGGCTTAGGCAGGAGAATTCCTTGA >19520_19520_5_CRLS1-NFX1_CRLS1_chr20_5996136_ENST00000378868_NFX1_chr9_33328579_ENST00000379521_length(amino acids)=481AA_BP=90 MPQYENPWTIPNMLSMTRIGLAPVLGYLIIEEDFNIALGVFALAGLTDLLDGFIARNWANQRSALGSALDPLADKILISILYVSLTYADL IPDFIHTCEKLCHEGDCGPCSRTSVISCRCSFRTKELPCTSLKSEDATFMCDKRCNKKRLCGRHKCNEICCVDKEHKCPLICGRKLRCGL HRCEEPCHRGNCQTCWQASFDELTCHCGASVIYPPVPCGTRPPECTQTCARVHECDHPVYHSCHSEEKCPPCTFLTQKWCMGKHEFRSNI PCHLVDISCGLPCSATLPCGMHKCQRLCHKGECLVDEPCKQPCTTPRADCGHPCMAPCHTSSPCPVTACKAKVELQCECGRRKEMVICSE ASSTYQRIAAISMASKITDMQLGGSVEISKLITKKEVHQARLECDEECSALERKKRLAEAFHISEDSDPFNIRSSGSKFSDSLKEDARKD -------------------------------------------------------------- >19520_19520_6_CRLS1-NFX1_CRLS1_chr20_5996136_ENST00000378868_NFX1_chr9_33328579_ENST00000379540_length(transcript)=3063nt_BP=427nt AGTTGTACAGCATGGGTTGAGGTGGCCAGATCCTGGCAGGGGTCTCAACTCCACTGATAGCTAAAGCATGTTGGGGTTTGAATGCTGGAT ACTTTGAAGTTGCCATATCCTGACTGAAGTCCTTCCCAGAACGGCAGTAGTTGGTCGAGTATGCCACAGTATGAAAACCCATGGACAATC CCGAATATGTTGTCAATGACGAGAATTGGCTTGGCCCCAGTTCTGGGCTATTTGATTATTGAAGAAGATTTTAATATTGCACTAGGAGTT TTTGCTTTAGCTGGACTAACAGATTTGTTGGATGGATTTATTGCTCGAAACTGGGCCAATCAAAGATCAGCTTTGGGAAGTGCTCTTGAT CCACTTGCTGATAAAATACTTATCAGTATCTTATATGTTAGCTTGACCTATGCAGATCTTATTCCAGATTTCATTCATACCTGTGAAAAG CTCTGCCATGAAGGAGACTGTGGACCATGCTCTCGCACATCAGTTATTTCCTGCAGATGCTCTTTCAGAACAAAGGAGCTTCCATGTACC AGTCTCAAAAGTGAAGATGCTACATTTATGTGTGACAAGCGGTGTAACAAGAAACGGTTGTGTGGACGGCATAAATGTAATGAGATATGC TGTGTGGATAAGGAGCACAAGTGTCCTTTGATTTGTGGGAGGAAACTCCGTTGTGGCCTTCATAGGTGTGAAGAACCTTGTCATCGTGGA AACTGCCAGACATGCTGGCAAGCCAGTTTTGATGAATTAACCTGCCATTGTGGTGCATCAGTGATTTACCCTCCAGTTCCCTGTGGTACT AGGCCCCCTGAATGTACCCAAACCTGCGCTAGAGTCCATGAGTGTGACCATCCAGTATATCATTCTTGTCATAGTGAGGAGAAGTGTCCC CCTTGCACTTTCCTAACTCAGAAGTGGTGCATGGGCAAGCATGAGTTTCGGAGCAACATCCCCTGTCACCTGGTTGATATCTCTTGCGGA TTACCCTGCAGTGCCACGCTACCATGTGGGATGCACAAATGTCAGAGACTCTGTCACAAAGGGGAGTGTCTTGTGGATGAGCCCTGCAAG CAGCCCTGCACCACCCCCAGAGCTGACTGTGGTCACCCGTGTATGGCACCCTGCCATACCAGCTCACCCTGCCCTGTGACTGCTTGTAAA GCTAAGGTAGAGCTACAGTGTGAATGTGGACGAAGAAAAGAGATGGTGATTTGCTCTGAAGCATCTAGTACTTATCAAAGAATAGCTGCA ATCTCCATGGCCTCTAAGATAACAGACATGCAGCTTGGAGGTTCAGTGGAGATCAGCAAGTTAATTACCAAAAAGGAAGTTCATCAAGCC AGGCTGGAGTGTGATGAGGAGTGTTCAGCCTTGGAAAGGAAAAAGAGATTAGCAGAGGCATTTCATATCAGTGAGGATTCTGATCCTTTC AATATACGTTCTTCAGGGTCAAAATTCAGTGATAGTTTGAAAGAAGATGCCAGGAAGGACTTAAAGTTTGTCAGTGACGTTGAGAAGGAA ATGGAAACCCTCGTGGAGGCCGTGAATAAGGGAAAGAATAGTAAGAAAAGCCACAGCTTCCCTCCCATGAACAGAGACCACCGCCGGATC ATCCATGACTTGGCCCAAGTTTATGGCCTGGAGAGCGTGAGCTATGACAGTGAACCGAAGCGCAATGTGGTGGTCACTGCCATCAGGGGG AAGTCCGTTTGTCCTCCTACCACGCTGACAGGTGTGCTTGAAAGGGAAATGCAGGCACGGCCTCCACCACCGATTCCTCATCACAGACAT CAGTCAGACAAGAATCCTGGGAGCAGTAATTTACAGAAAATAACCAAGGAGCCAATAATTGACTATTTTGACGTCCAGGACTAAGAAGAT CATGATGCACTTAGATAAAAGAATGATTAGGTATAGTGGAGACTTATTTGCCAGCAGATAAATCATGCCCGTTCCCCTCTGCCTGGCAGA ATCACAGTCTCACATACTGTCTTGTACTGACACATCCAAAGCATGAGTGTGTCAGAAATCCCTTGTCTATTCCTGTCTGTATAAAGTGTT TCATTATGACCAGATCTCTGATTGTATGGTCACTAGGTATGCAATCACGCATTCAAAGAGGCTCTTTACACCATCACTGTGATTGCTCTG AGAGTTGAGGGACTATTGGGCTTTATTTGGACAAACCAAACTTTTAGCCTGAAACCAACTTTATGCCACTAAGTCATAGCCTCAGTTGTC CCAGTTATTTGTCCTCCTGAAAATGCCTGAAACATCAGACAGACATTGCTTGCTTTACCCAAACTGATCAAAATCTTTAGGAGCACAAAT GAATTTTTTAGTCTGAAATACCAAATAATGAATTGGTATACCATATCCGGAATCACACATGTTATCTTAAACCCAGCCATCATACCTAAG TCTTTTGCCAAAACCTCTCATAGGTATATCTAGCTGAACTTATTTTGGCATTTTCAATGTGATCAGTTCTAGACCTAGAAGGGGGTCAGG CTGCTTTACAGAATTCTATTTCCTTAAGTCCCTGGCACTTCTCATACCACATCACTGAACCTGTTCAGTAACAATCAGTTTGGCCGTCCC CCATGATGGTAGGAAATATAGAGAGCAAGTTCTTCTGCCAGGGTCACACTGTGGTCTCTGAACTGACCAGTATATCCCTAACTCCTCTTT GATAGAGAAAGAGTCTCAAATGGACAACTGTCCTGTGTTGCTTTCCCTAGGCCTTCAGCAGCCTATTGGCTCTCCCTGCCTCTGAGCTCT GGACTCTGTTTGAATATTCCAAGTAGTATATGGACAGTCCAGGGCTTATGCCCAGCAGCCCACTGGAGGCATTCTTCAGGCTCCTTTAAG GCAGGTGCATTGATAGTTCCATTAGTGTGACCCTTGCATTGGCACCCCTCCAGCCTGGAGGCCAGGCTTCCAGCAACTTCCTTCTGCCCT AGAGCAAGCCATGAGCCCCAGAGCAGTAGCAGGAGACTTGAGAAGTAGAGTGACAAAAACAAGCACTTAATTAAATTATAAAATTTAACT >19520_19520_6_CRLS1-NFX1_CRLS1_chr20_5996136_ENST00000378868_NFX1_chr9_33328579_ENST00000379540_length(amino acids)=577AA_BP=90 MPQYENPWTIPNMLSMTRIGLAPVLGYLIIEEDFNIALGVFALAGLTDLLDGFIARNWANQRSALGSALDPLADKILISILYVSLTYADL IPDFIHTCEKLCHEGDCGPCSRTSVISCRCSFRTKELPCTSLKSEDATFMCDKRCNKKRLCGRHKCNEICCVDKEHKCPLICGRKLRCGL HRCEEPCHRGNCQTCWQASFDELTCHCGASVIYPPVPCGTRPPECTQTCARVHECDHPVYHSCHSEEKCPPCTFLTQKWCMGKHEFRSNI PCHLVDISCGLPCSATLPCGMHKCQRLCHKGECLVDEPCKQPCTTPRADCGHPCMAPCHTSSPCPVTACKAKVELQCECGRRKEMVICSE ASSTYQRIAAISMASKITDMQLGGSVEISKLITKKEVHQARLECDEECSALERKKRLAEAFHISEDSDPFNIRSSGSKFSDSLKEDARKD LKFVSDVEKEMETLVEAVNKGKNSKKSHSFPPMNRDHRRIIHDLAQVYGLESVSYDSEPKRNVVVTAIRGKSVCPPTTLTGVLEREMQAR -------------------------------------------------------------- >19520_19520_7_CRLS1-NFX1_CRLS1_chr20_5996136_ENST00000452938_NFX1_chr9_33328579_ENST00000318524_length(transcript)=2250nt_BP=671nt CTGAGTGGTTGCCGGGTCTCCATGGAGAAGCGGCTCGCCAGTGTCCCAGGCTGCTGAGCTCTCGCCGCCCGAGACCCCGCGGCGCGGCCG CAGGGCCATGCTAGCCTTGCGCGTGGCGCGCGGCTCGTGGGGGGCCCTGCGCGGCGCCGCTTGGGCTCCGGGAACGCGGCCGAGTAAGCG ACGCGCCTGCTGGGCCCTGCTGCCGCCCGTGCCCTGCTGCTTGGGCTGCCTGGCCGAACGCTGGAGGCTGCGTCCGGCCGCTCTTGGCTT GCGGCTGCCCGGGATCGGCCAGCGGAACCACTGTTCGGGCGCGGGGAAGGCGGCTCCCAGGCCAGCGGCCGGAGCGGGCGCCGCTGCCGA AGCCCCGGGCGGCCAGTGGGGCCCGGCGAGCACCCCCAGCCTGTATGAAAACCCATGGACAATCCCGAATATGTTGTCAATGACGAGAAT TGGCTTGGCCCCAGTTCTGGGCTATTTGATTATTGAAGAAGATTTTAATATTGCACTAGGAGTTTTTGCTTTAGCTGGACTAACAGATTT GTTGGATGGATTTATTGCTCGAAACTGGGCCAATCAAAGATCAGCTTTGGGAAGTGCTCTTGATCCACTTGCTGATAAAATACTTATCAG TATCTTATATGTTAGCTTGACCTATGCAGATCTTATTCCAGATTTCATTCATACCTGTGAAAAGCTCTGCCATGAAGGAGACTGTGGACC ATGCTCTCGCACATCAGTTATTTCCTGCAGATGCTCTTTCAGAACAAAGGAGCTTCCATGTACCAGTCTCAAAAGTGAAGATGCTACATT TATGTGTGACAAGCGGTGTAACAAGAAACGGTTGTGTGGACGGCATAAATGTAATGAGATATGCTGTGTGGATAAGGAGCACAAGTGTCC TTTGATTTGTGGGAGGAAACTCCGTTGTGGCCTTCATAGGTGTGAAGAACCTTGTCATCGTGGAAACTGCCAGACATGCTGGCAAGCCAG TTTTGATGAATTAACCTGCCATTGTGGTGCATCAGTGATTTACCCTCCAGTTCCCTGTGGTACTAGGCCCCCTGAATGTACCCAAACCTG CGCTAGAGTCCATGAGTGTGACCATCCAGTATATCATTCTTGTCATAGTGAGGAGAAGTGTCCCCCTTGCACTTTCCTAACTCAGAAGTG GTGCATGGGCAAGCATGAGCAGTCCCACTACTGGGCGTCTACCCAGAAGAAAAGAAGTCATTATATGAAAAAGATACCTGCACACGCATG TTTATAGCAGCACAATTCACAATTGCAAAAATGTGGAACCAGCCCAACTGCCCATCAGTCAACAAGTGGATAAAGAAATTGTGGTGTATC TATACGTACCATGGAATACTACTCAGCCAGGAACGAAATAATGGCATTCACAGCAACCTGGATGGATTTGGAGACCATTATTCTAAGTGA AGTAACTCAGGAATGGAAAACCAAACATCGTATGTTCTCAATTATAAGAGGGAGCTAAGCTATGAGGACGCAAAGGCATGAGAATGATAC AATGGACTTTGGGGACTCTGGGGAAAGGACGGGAGGCGGGTGAGGGATAAAAAACTACACACTGGGTCCGAGCACGGTGGCTCATGCCTA TAATCCCAGCACTTTGGGAGGCCAAGGCAGGTAGATTATGAGGTCGGGAGTTCGAAACCAGCCTGGCCAACAAGGTGAGACCCCCCCCAT CTCTACTAAAATAATTAGCCGGGCCTGGTGGCGCATCCCTGTAATCCCAGCTACTTTACTTGGGAGGCTGAGGCAGGAGAATTGCTTGAA CTCGGGAGGCGGATGTTGCAGTGAGCCAATATCGCACCACTGCACTCCAGCCTGGGCAACAGAGCAAGACTCCGTCTCGAAAAAAAAAAT AAATTTTAAAAACTACACATTGGGTACAATGTACACTCCTCGGGTGATGGGTGCACCAAAATCTCAAAAATCACTACTAAGGAACTTATC CATGTAACCAAACACCACCTGTTCCCCAAGAACTATTGAAATAAGAAAAAAGAAAAAAAAAAAATCACTGATTGCAGGTCACCCTAACAG ATATAATAATGAAAAAGTCTGAGATGTTGCGAGAATTACCAAAATGTGATGCAGACACGAAATGAGCATGTGCTGTTGGAAAAGTGGAAC AGATAGACTTGCTGGAGACAGGGTTGCCACAAAGTTTCAATTTGTAAAAAATGTGATATCTATGAAATGCAATAAAGTGAAGTCCAATAA >19520_19520_7_CRLS1-NFX1_CRLS1_chr20_5996136_ENST00000452938_NFX1_chr9_33328579_ENST00000318524_length(amino acids)=389AA_BP=189 MLALRVARGSWGALRGAAWAPGTRPSKRRACWALLPPVPCCLGCLAERWRLRPAALGLRLPGIGQRNHCSGAGKAAPRPAAGAGAAAEAP GGQWGPASTPSLYENPWTIPNMLSMTRIGLAPVLGYLIIEEDFNIALGVFALAGLTDLLDGFIARNWANQRSALGSALDPLADKILISIL YVSLTYADLIPDFIHTCEKLCHEGDCGPCSRTSVISCRCSFRTKELPCTSLKSEDATFMCDKRCNKKRLCGRHKCNEICCVDKEHKCPLI CGRKLRCGLHRCEEPCHRGNCQTCWQASFDELTCHCGASVIYPPVPCGTRPPECTQTCARVHECDHPVYHSCHSEEKCPPCTFLTQKWCM -------------------------------------------------------------- >19520_19520_8_CRLS1-NFX1_CRLS1_chr20_5996136_ENST00000452938_NFX1_chr9_33328579_ENST00000379521_length(transcript)=2313nt_BP=671nt CTGAGTGGTTGCCGGGTCTCCATGGAGAAGCGGCTCGCCAGTGTCCCAGGCTGCTGAGCTCTCGCCGCCCGAGACCCCGCGGCGCGGCCG CAGGGCCATGCTAGCCTTGCGCGTGGCGCGCGGCTCGTGGGGGGCCCTGCGCGGCGCCGCTTGGGCTCCGGGAACGCGGCCGAGTAAGCG ACGCGCCTGCTGGGCCCTGCTGCCGCCCGTGCCCTGCTGCTTGGGCTGCCTGGCCGAACGCTGGAGGCTGCGTCCGGCCGCTCTTGGCTT GCGGCTGCCCGGGATCGGCCAGCGGAACCACTGTTCGGGCGCGGGGAAGGCGGCTCCCAGGCCAGCGGCCGGAGCGGGCGCCGCTGCCGA AGCCCCGGGCGGCCAGTGGGGCCCGGCGAGCACCCCCAGCCTGTATGAAAACCCATGGACAATCCCGAATATGTTGTCAATGACGAGAAT TGGCTTGGCCCCAGTTCTGGGCTATTTGATTATTGAAGAAGATTTTAATATTGCACTAGGAGTTTTTGCTTTAGCTGGACTAACAGATTT GTTGGATGGATTTATTGCTCGAAACTGGGCCAATCAAAGATCAGCTTTGGGAAGTGCTCTTGATCCACTTGCTGATAAAATACTTATCAG TATCTTATATGTTAGCTTGACCTATGCAGATCTTATTCCAGATTTCATTCATACCTGTGAAAAGCTCTGCCATGAAGGAGACTGTGGACC ATGCTCTCGCACATCAGTTATTTCCTGCAGATGCTCTTTCAGAACAAAGGAGCTTCCATGTACCAGTCTCAAAAGTGAAGATGCTACATT TATGTGTGACAAGCGGTGTAACAAGAAACGGTTGTGTGGACGGCATAAATGTAATGAGATATGCTGTGTGGATAAGGAGCACAAGTGTCC TTTGATTTGTGGGAGGAAACTCCGTTGTGGCCTTCATAGGTGTGAAGAACCTTGTCATCGTGGAAACTGCCAGACATGCTGGCAAGCCAG TTTTGATGAATTAACCTGCCATTGTGGTGCATCAGTGATTTACCCTCCAGTTCCCTGTGGTACTAGGCCCCCTGAATGTACCCAAACCTG CGCTAGAGTCCATGAGTGTGACCATCCAGTATATCATTCTTGTCATAGTGAGGAGAAGTGTCCCCCTTGCACTTTCCTAACTCAGAAGTG GTGCATGGGCAAGCATGAGTTTCGGAGCAACATCCCCTGTCACCTGGTTGATATCTCTTGCGGATTACCCTGCAGTGCCACGCTACCATG TGGGATGCACAAATGTCAGAGACTCTGTCACAAAGGGGAGTGTCTTGTGGATGAGCCCTGCAAGCAGCCCTGCACCACCCCCAGAGCTGA CTGTGGTCACCCGTGTATGGCACCCTGCCATACCAGCTCACCCTGCCCTGTGACTGCTTGTAAAGCTAAGGTAGAGCTACAGTGTGAATG TGGACGAAGAAAAGAGATGGTGATTTGCTCTGAAGCATCTAGTACTTATCAAAGAATAGCTGCAATCTCCATGGCCTCTAAGATAACAGA CATGCAGCTTGGAGGTTCAGTGGAGATCAGCAAGTTAATTACCAAAAAGGAAGTTCATCAAGCCAGGCTGGAGTGTGATGAGGAGTGTTC AGCCTTGGAAAGGAAAAAGAGATTAGCAGAGGCATTTCATATCAGTGAGGATTCTGATCCTTTCAATATACGTTCTTCAGGGTCAAAATT CAGTGATAGTTTGAAAGAAGATGCCAGGAAGGACTTAAAGTTTGTCAGTGACGTTGAGAAGGAAATGGAAACCCTCGTGGAGGCCGTGAA TAAGGTTGAAGTCGAAACATCCCACTGGACATTTCTCTAAGGCCAGCTTGATGAAAAAAAAAAAGCAATCAAGTCCTGGAAACACTTTCA ACCTAGAGTAGTTGTAGGAAAAAGTCACAACTTCTTTGAGGATCCTTATCTTTCTCAAAACACAGCCATTATAACATCTGTGGAAGCCTG TCCTGGTTCAGGACAGTTGACTGCAGAAAAGATCTACAGTCGGCTGGGCACAGTGTCTCATGCCTGTAATGCCAGCACTTTGGGAGGCCA AGGCAGGCAGATCACTTGAGGTCAGGAGTTCTAGATCAGCTTGGCCAACATGACGAAACCCTGTCTCTACTAAAAATATAAAAATTAGCT GGGTGTGGTGGCGGGCACCTGTTATTCCAGCTACTTGCAAGGCTTAGGCAGGAGAATTCCTTGAACCCGGGAGGCAGAGGTTGCTGTGAG >19520_19520_8_CRLS1-NFX1_CRLS1_chr20_5996136_ENST00000452938_NFX1_chr9_33328579_ENST00000379521_length(amino acids)=580AA_BP=189 MLALRVARGSWGALRGAAWAPGTRPSKRRACWALLPPVPCCLGCLAERWRLRPAALGLRLPGIGQRNHCSGAGKAAPRPAAGAGAAAEAP GGQWGPASTPSLYENPWTIPNMLSMTRIGLAPVLGYLIIEEDFNIALGVFALAGLTDLLDGFIARNWANQRSALGSALDPLADKILISIL YVSLTYADLIPDFIHTCEKLCHEGDCGPCSRTSVISCRCSFRTKELPCTSLKSEDATFMCDKRCNKKRLCGRHKCNEICCVDKEHKCPLI CGRKLRCGLHRCEEPCHRGNCQTCWQASFDELTCHCGASVIYPPVPCGTRPPECTQTCARVHECDHPVYHSCHSEEKCPPCTFLTQKWCM GKHEFRSNIPCHLVDISCGLPCSATLPCGMHKCQRLCHKGECLVDEPCKQPCTTPRADCGHPCMAPCHTSSPCPVTACKAKVELQCECGR RKEMVICSEASSTYQRIAAISMASKITDMQLGGSVEISKLITKKEVHQARLECDEECSALERKKRLAEAFHISEDSDPFNIRSSGSKFSD -------------------------------------------------------------- >19520_19520_9_CRLS1-NFX1_CRLS1_chr20_5996136_ENST00000452938_NFX1_chr9_33328579_ENST00000379540_length(transcript)=3307nt_BP=671nt CTGAGTGGTTGCCGGGTCTCCATGGAGAAGCGGCTCGCCAGTGTCCCAGGCTGCTGAGCTCTCGCCGCCCGAGACCCCGCGGCGCGGCCG CAGGGCCATGCTAGCCTTGCGCGTGGCGCGCGGCTCGTGGGGGGCCCTGCGCGGCGCCGCTTGGGCTCCGGGAACGCGGCCGAGTAAGCG ACGCGCCTGCTGGGCCCTGCTGCCGCCCGTGCCCTGCTGCTTGGGCTGCCTGGCCGAACGCTGGAGGCTGCGTCCGGCCGCTCTTGGCTT GCGGCTGCCCGGGATCGGCCAGCGGAACCACTGTTCGGGCGCGGGGAAGGCGGCTCCCAGGCCAGCGGCCGGAGCGGGCGCCGCTGCCGA AGCCCCGGGCGGCCAGTGGGGCCCGGCGAGCACCCCCAGCCTGTATGAAAACCCATGGACAATCCCGAATATGTTGTCAATGACGAGAAT TGGCTTGGCCCCAGTTCTGGGCTATTTGATTATTGAAGAAGATTTTAATATTGCACTAGGAGTTTTTGCTTTAGCTGGACTAACAGATTT GTTGGATGGATTTATTGCTCGAAACTGGGCCAATCAAAGATCAGCTTTGGGAAGTGCTCTTGATCCACTTGCTGATAAAATACTTATCAG TATCTTATATGTTAGCTTGACCTATGCAGATCTTATTCCAGATTTCATTCATACCTGTGAAAAGCTCTGCCATGAAGGAGACTGTGGACC ATGCTCTCGCACATCAGTTATTTCCTGCAGATGCTCTTTCAGAACAAAGGAGCTTCCATGTACCAGTCTCAAAAGTGAAGATGCTACATT TATGTGTGACAAGCGGTGTAACAAGAAACGGTTGTGTGGACGGCATAAATGTAATGAGATATGCTGTGTGGATAAGGAGCACAAGTGTCC TTTGATTTGTGGGAGGAAACTCCGTTGTGGCCTTCATAGGTGTGAAGAACCTTGTCATCGTGGAAACTGCCAGACATGCTGGCAAGCCAG TTTTGATGAATTAACCTGCCATTGTGGTGCATCAGTGATTTACCCTCCAGTTCCCTGTGGTACTAGGCCCCCTGAATGTACCCAAACCTG CGCTAGAGTCCATGAGTGTGACCATCCAGTATATCATTCTTGTCATAGTGAGGAGAAGTGTCCCCCTTGCACTTTCCTAACTCAGAAGTG GTGCATGGGCAAGCATGAGTTTCGGAGCAACATCCCCTGTCACCTGGTTGATATCTCTTGCGGATTACCCTGCAGTGCCACGCTACCATG TGGGATGCACAAATGTCAGAGACTCTGTCACAAAGGGGAGTGTCTTGTGGATGAGCCCTGCAAGCAGCCCTGCACCACCCCCAGAGCTGA CTGTGGTCACCCGTGTATGGCACCCTGCCATACCAGCTCACCCTGCCCTGTGACTGCTTGTAAAGCTAAGGTAGAGCTACAGTGTGAATG TGGACGAAGAAAAGAGATGGTGATTTGCTCTGAAGCATCTAGTACTTATCAAAGAATAGCTGCAATCTCCATGGCCTCTAAGATAACAGA CATGCAGCTTGGAGGTTCAGTGGAGATCAGCAAGTTAATTACCAAAAAGGAAGTTCATCAAGCCAGGCTGGAGTGTGATGAGGAGTGTTC AGCCTTGGAAAGGAAAAAGAGATTAGCAGAGGCATTTCATATCAGTGAGGATTCTGATCCTTTCAATATACGTTCTTCAGGGTCAAAATT CAGTGATAGTTTGAAAGAAGATGCCAGGAAGGACTTAAAGTTTGTCAGTGACGTTGAGAAGGAAATGGAAACCCTCGTGGAGGCCGTGAA TAAGGGAAAGAATAGTAAGAAAAGCCACAGCTTCCCTCCCATGAACAGAGACCACCGCCGGATCATCCATGACTTGGCCCAAGTTTATGG CCTGGAGAGCGTGAGCTATGACAGTGAACCGAAGCGCAATGTGGTGGTCACTGCCATCAGGGGGAAGTCCGTTTGTCCTCCTACCACGCT GACAGGTGTGCTTGAAAGGGAAATGCAGGCACGGCCTCCACCACCGATTCCTCATCACAGACATCAGTCAGACAAGAATCCTGGGAGCAG TAATTTACAGAAAATAACCAAGGAGCCAATAATTGACTATTTTGACGTCCAGGACTAAGAAGATCATGATGCACTTAGATAAAAGAATGA TTAGGTATAGTGGAGACTTATTTGCCAGCAGATAAATCATGCCCGTTCCCCTCTGCCTGGCAGAATCACAGTCTCACATACTGTCTTGTA CTGACACATCCAAAGCATGAGTGTGTCAGAAATCCCTTGTCTATTCCTGTCTGTATAAAGTGTTTCATTATGACCAGATCTCTGATTGTA TGGTCACTAGGTATGCAATCACGCATTCAAAGAGGCTCTTTACACCATCACTGTGATTGCTCTGAGAGTTGAGGGACTATTGGGCTTTAT TTGGACAAACCAAACTTTTAGCCTGAAACCAACTTTATGCCACTAAGTCATAGCCTCAGTTGTCCCAGTTATTTGTCCTCCTGAAAATGC CTGAAACATCAGACAGACATTGCTTGCTTTACCCAAACTGATCAAAATCTTTAGGAGCACAAATGAATTTTTTAGTCTGAAATACCAAAT AATGAATTGGTATACCATATCCGGAATCACACATGTTATCTTAAACCCAGCCATCATACCTAAGTCTTTTGCCAAAACCTCTCATAGGTA TATCTAGCTGAACTTATTTTGGCATTTTCAATGTGATCAGTTCTAGACCTAGAAGGGGGTCAGGCTGCTTTACAGAATTCTATTTCCTTA AGTCCCTGGCACTTCTCATACCACATCACTGAACCTGTTCAGTAACAATCAGTTTGGCCGTCCCCCATGATGGTAGGAAATATAGAGAGC AAGTTCTTCTGCCAGGGTCACACTGTGGTCTCTGAACTGACCAGTATATCCCTAACTCCTCTTTGATAGAGAAAGAGTCTCAAATGGACA ACTGTCCTGTGTTGCTTTCCCTAGGCCTTCAGCAGCCTATTGGCTCTCCCTGCCTCTGAGCTCTGGACTCTGTTTGAATATTCCAAGTAG TATATGGACAGTCCAGGGCTTATGCCCAGCAGCCCACTGGAGGCATTCTTCAGGCTCCTTTAAGGCAGGTGCATTGATAGTTCCATTAGT GTGACCCTTGCATTGGCACCCCTCCAGCCTGGAGGCCAGGCTTCCAGCAACTTCCTTCTGCCCTAGAGCAAGCCATGAGCCCCAGAGCAG >19520_19520_9_CRLS1-NFX1_CRLS1_chr20_5996136_ENST00000452938_NFX1_chr9_33328579_ENST00000379540_length(amino acids)=676AA_BP=189 MLALRVARGSWGALRGAAWAPGTRPSKRRACWALLPPVPCCLGCLAERWRLRPAALGLRLPGIGQRNHCSGAGKAAPRPAAGAGAAAEAP GGQWGPASTPSLYENPWTIPNMLSMTRIGLAPVLGYLIIEEDFNIALGVFALAGLTDLLDGFIARNWANQRSALGSALDPLADKILISIL YVSLTYADLIPDFIHTCEKLCHEGDCGPCSRTSVISCRCSFRTKELPCTSLKSEDATFMCDKRCNKKRLCGRHKCNEICCVDKEHKCPLI CGRKLRCGLHRCEEPCHRGNCQTCWQASFDELTCHCGASVIYPPVPCGTRPPECTQTCARVHECDHPVYHSCHSEEKCPPCTFLTQKWCM GKHEFRSNIPCHLVDISCGLPCSATLPCGMHKCQRLCHKGECLVDEPCKQPCTTPRADCGHPCMAPCHTSSPCPVTACKAKVELQCECGR RKEMVICSEASSTYQRIAAISMASKITDMQLGGSVEISKLITKKEVHQARLECDEECSALERKKRLAEAFHISEDSDPFNIRSSGSKFSD SLKEDARKDLKFVSDVEKEMETLVEAVNKGKNSKKSHSFPPMNRDHRRIIHDLAQVYGLESVSYDSEPKRNVVVTAIRGKSVCPPTTLTG -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for CRLS1-NFX1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000318524 | 8 | 16 | 9_26 | 635.3333333333334 | 834.0 | PABPC1 and PABC4 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379521 | 8 | 21 | 9_26 | 635.3333333333334 | 1025.0 | PABPC1 and PABC4 | |
| Tgene | NFX1 | chr20:5996136 | chr9:33328579 | ENST00000379540 | 8 | 24 | 9_26 | 635.3333333333334 | 1121.0 | PABPC1 and PABC4 |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for CRLS1-NFX1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for CRLS1-NFX1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
