
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:PTCH1-FLNA (FusionGDB2 ID:HG5727TG2316) |
Fusion Gene Summary for PTCH1-FLNA |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: PTCH1-FLNA | Fusion gene ID: hg5727tg2316 | Hgene | Tgene | Gene symbol | PTCH1 | FLNA | Gene ID | 5727 | 2316 |
| Gene name | patched 1 | filamin A | |
| Synonyms | BCNS|NBCCS|PTC|PTC1|PTCH | ABP-280|ABPX|CSBS|CVD1|FGS2|FLN|FLN-A|FLN1|FMD|MNS|NHBP|OPD|OPD1|OPD2|XLVD|XMVD | |
| Cytomap | ('PTCH1')('FLNA') 9q22.32 | Xq28 | |
| Type of gene | protein-coding | protein-coding | |
| Description | protein patched homolog 1 | filamin-Aactin binding protein 280alpha-filaminendothelial actin-binding proteinepididymis secretory sperm binding proteinfilamin A, alphafilamin-1non-muscle filamin | |
| Modification date | 20200314 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000331920, ENST00000375274, ENST00000418258, ENST00000421141, ENST00000429896, ENST00000430669, ENST00000437951, ENST00000468211, ENST00000548379, | ||
| Fusion gene scores | * DoF score | 6 X 7 X 4=168 | 25 X 31 X 6=4650 |
| # samples | 7 | 31 | |
| ** MAII score | log2(7/168*10)=-1.26303440583379 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(31/4650*10)=-3.90689059560852 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: PTCH1 [Title/Abstract] AND FLNA [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | PTCH1(98232094)-FLNA(153578575), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | PTCH1-FLNA seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. PTCH1-FLNA seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. PTCH1-FLNA seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. PTCH1-FLNA seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. PTCH1-FLNA seems lost the major protein functional domain in Hgene partner, which is a CGC due to the frame-shifted ORF. PTCH1-FLNA seems lost the major protein functional domain in Hgene partner, which is a tumor suppressor due to the frame-shifted ORF. PTCH1-FLNA seems lost the major protein functional domain in Tgene partner, which is a CGC due to the frame-shifted ORF. PTCH1-FLNA seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. PTCH1-FLNA seems lost the major protein functional domain in Tgene partner, which is a tumor suppressor due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | PTCH1 | GO:0010875 | positive regulation of cholesterol efflux | 21931618 |
| Hgene | PTCH1 | GO:0072659 | protein localization to plasma membrane | 11278759 |
| Tgene | FLNA | GO:0016479 | negative regulation of transcription by RNA polymerase I | 22307607 |
| Tgene | FLNA | GO:0030334 | regulation of cell migration | 16291724 |
| Tgene | FLNA | GO:0031532 | actin cytoskeleton reorganization | 10051605 |
| Tgene | FLNA | GO:0034394 | protein localization to cell surface | 18322202 |
| Tgene | FLNA | GO:0043113 | receptor clustering | 10692483 |
| Tgene | FLNA | GO:0043433 | negative regulation of DNA-binding transcription factor activity | 15684392 |
| Tgene | FLNA | GO:0044319 | wound healing, spreading of cells | 16291724 |
| Tgene | FLNA | GO:0045184 | establishment of protein localization | 18322202 |
| Tgene | FLNA | GO:0051764 | actin crosslink formation | 10051605 |
| Tgene | FLNA | GO:0072659 | protein localization to plasma membrane | 24951510 |
| Tgene | FLNA | GO:0090307 | mitotic spindle assembly | 18548008 |
| Tgene | FLNA | GO:1901381 | positive regulation of potassium ion transmembrane transport | 24951510 |
 Fusion gene breakpoints across PTCH1 (5'-gene) Fusion gene breakpoints across PTCH1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
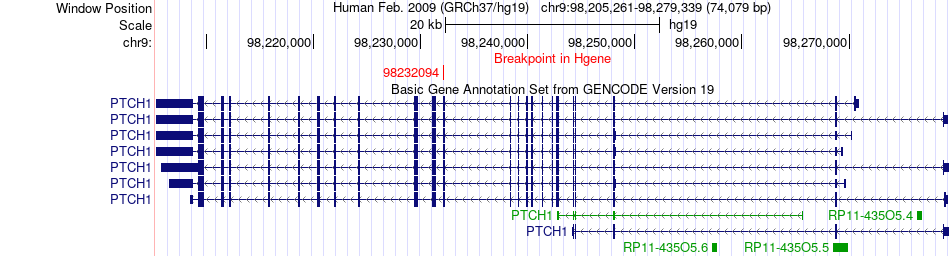 |
 Fusion gene breakpoints across FLNA (3'-gene) Fusion gene breakpoints across FLNA (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
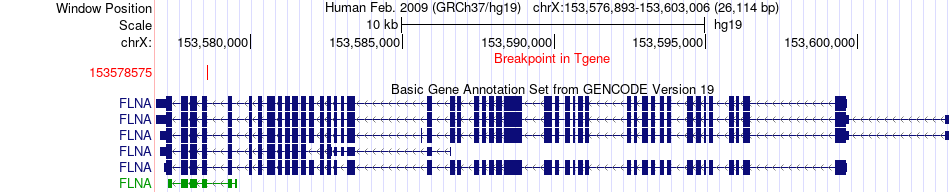 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | Non-Cancer | 173N | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
Top |
Fusion Gene ORF analysis for PTCH1-FLNA |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000331920 | ENST00000498491 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| 5CDS-5UTR | ENST00000375274 | ENST00000498491 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| 5CDS-5UTR | ENST00000418258 | ENST00000498491 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| 5CDS-5UTR | ENST00000421141 | ENST00000498491 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| 5CDS-5UTR | ENST00000429896 | ENST00000498491 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| 5CDS-5UTR | ENST00000430669 | ENST00000498491 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| 5CDS-5UTR | ENST00000437951 | ENST00000498491 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000331920 | ENST00000344736 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000331920 | ENST00000360319 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000331920 | ENST00000369850 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000331920 | ENST00000369856 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000331920 | ENST00000422373 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000375274 | ENST00000344736 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000375274 | ENST00000360319 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000375274 | ENST00000369850 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000375274 | ENST00000369856 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000375274 | ENST00000422373 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000418258 | ENST00000344736 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000418258 | ENST00000360319 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000418258 | ENST00000369850 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000418258 | ENST00000369856 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000418258 | ENST00000422373 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000430669 | ENST00000344736 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000430669 | ENST00000360319 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000430669 | ENST00000369850 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000430669 | ENST00000369856 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000430669 | ENST00000422373 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000437951 | ENST00000344736 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000437951 | ENST00000360319 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000437951 | ENST00000369850 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000437951 | ENST00000369856 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| Frame-shift | ENST00000437951 | ENST00000422373 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| In-frame | ENST00000421141 | ENST00000344736 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| In-frame | ENST00000421141 | ENST00000360319 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| In-frame | ENST00000421141 | ENST00000369850 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| In-frame | ENST00000421141 | ENST00000369856 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| In-frame | ENST00000421141 | ENST00000422373 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| In-frame | ENST00000429896 | ENST00000344736 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| In-frame | ENST00000429896 | ENST00000360319 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| In-frame | ENST00000429896 | ENST00000369850 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| In-frame | ENST00000429896 | ENST00000369856 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| In-frame | ENST00000429896 | ENST00000422373 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| intron-3CDS | ENST00000468211 | ENST00000344736 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| intron-3CDS | ENST00000468211 | ENST00000360319 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| intron-3CDS | ENST00000468211 | ENST00000369850 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| intron-3CDS | ENST00000468211 | ENST00000369856 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| intron-3CDS | ENST00000468211 | ENST00000422373 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| intron-3CDS | ENST00000548379 | ENST00000344736 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| intron-3CDS | ENST00000548379 | ENST00000360319 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| intron-3CDS | ENST00000548379 | ENST00000369850 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| intron-3CDS | ENST00000548379 | ENST00000369856 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| intron-3CDS | ENST00000548379 | ENST00000422373 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| intron-5UTR | ENST00000468211 | ENST00000498491 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
| intron-5UTR | ENST00000548379 | ENST00000498491 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000421141 | PTCH1 | chr9 | 98232094 | - | ENST00000360319 | FLNA | chrX | 153578575 | - | 2864 | 1753 | 134 | 1756 | 540 |
| ENST00000421141 | PTCH1 | chr9 | 98232094 | - | ENST00000422373 | FLNA | chrX | 153578575 | - | 2858 | 1753 | 134 | 1756 | 540 |
| ENST00000421141 | PTCH1 | chr9 | 98232094 | - | ENST00000369850 | FLNA | chrX | 153578575 | - | 2742 | 1753 | 134 | 1756 | 540 |
| ENST00000421141 | PTCH1 | chr9 | 98232094 | - | ENST00000369856 | FLNA | chrX | 153578575 | - | 2734 | 1753 | 134 | 1756 | 540 |
| ENST00000421141 | PTCH1 | chr9 | 98232094 | - | ENST00000344736 | FLNA | chrX | 153578575 | - | 2606 | 1753 | 134 | 1756 | 540 |
| ENST00000429896 | PTCH1 | chr9 | 98232094 | - | ENST00000360319 | FLNA | chrX | 153578575 | - | 2946 | 1835 | 216 | 1838 | 540 |
| ENST00000429896 | PTCH1 | chr9 | 98232094 | - | ENST00000422373 | FLNA | chrX | 153578575 | - | 2940 | 1835 | 216 | 1838 | 540 |
| ENST00000429896 | PTCH1 | chr9 | 98232094 | - | ENST00000369850 | FLNA | chrX | 153578575 | - | 2824 | 1835 | 216 | 1838 | 540 |
| ENST00000429896 | PTCH1 | chr9 | 98232094 | - | ENST00000369856 | FLNA | chrX | 153578575 | - | 2816 | 1835 | 216 | 1838 | 540 |
| ENST00000429896 | PTCH1 | chr9 | 98232094 | - | ENST00000344736 | FLNA | chrX | 153578575 | - | 2688 | 1835 | 216 | 1838 | 540 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000421141 | ENST00000360319 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - | 0.003723764 | 0.99627626 |
| ENST00000421141 | ENST00000422373 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - | 0.003692471 | 0.9963076 |
| ENST00000421141 | ENST00000369850 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - | 0.005124524 | 0.9948755 |
| ENST00000421141 | ENST00000369856 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - | 0.005194935 | 0.99480504 |
| ENST00000421141 | ENST00000344736 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - | 0.00427159 | 0.99572843 |
| ENST00000429896 | ENST00000360319 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - | 0.003763315 | 0.99623674 |
| ENST00000429896 | ENST00000422373 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - | 0.003731571 | 0.9962684 |
| ENST00000429896 | ENST00000369850 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - | 0.005114782 | 0.99488527 |
| ENST00000429896 | ENST00000369856 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - | 0.005179845 | 0.9948201 |
| ENST00000429896 | ENST00000344736 | PTCH1 | chr9 | 98232094 | - | FLNA | chrX | 153578575 | - | 0.004352502 | 0.9956475 |
Top |
Fusion Genomic Features for PTCH1-FLNA |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
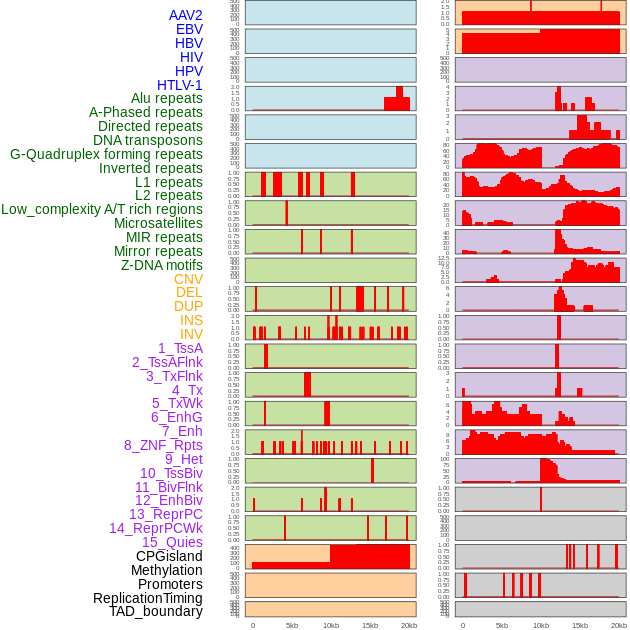 |
Top |
Fusion Protein Features for PTCH1-FLNA |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr9:98232094/chrX:153578575) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 14_31 | 615 | 2534.3333333333335 | Compositional bias | Note=Gly-rich |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 14_31 | 614 | 1526.3333333333333 | Compositional bias | Note=Gly-rich |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 14_31 | 464 | 2461.3333333333335 | Compositional bias | Note=Gly-rich |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 14_31 | 464 | 2513.3333333333335 | Compositional bias | Note=Gly-rich |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 14_31 | 464 | 2093.3333333333335 | Compositional bias | Note=Gly-rich |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 14_31 | 549 | 1382.0 | Compositional bias | Note=Gly-rich |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 14_31 | 549 | 2468.3333333333335 | Compositional bias | Note=Gly-rich |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 438_598 | 615 | 2534.3333333333335 | Domain | SSD |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 438_598 | 614 | 1526.3333333333333 | Domain | SSD |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 122_436 | 615 | 2534.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 1_100 | 615 | 2534.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 458_472 | 615 | 2534.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 494_501 | 615 | 2534.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 523_547 | 615 | 2534.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 569_577 | 615 | 2534.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 122_436 | 614 | 1526.3333333333333 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 1_100 | 614 | 1526.3333333333333 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 458_472 | 614 | 1526.3333333333333 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 494_501 | 614 | 1526.3333333333333 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 523_547 | 614 | 1526.3333333333333 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 569_577 | 614 | 1526.3333333333333 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 122_436 | 464 | 2461.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 1_100 | 464 | 2461.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 122_436 | 464 | 2513.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 1_100 | 464 | 2513.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 122_436 | 464 | 2093.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 1_100 | 464 | 2093.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 122_436 | 549 | 1382.0 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 1_100 | 549 | 1382.0 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 458_472 | 549 | 1382.0 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 494_501 | 549 | 1382.0 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 523_547 | 549 | 1382.0 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 122_436 | 549 | 2468.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 1_100 | 549 | 2468.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 458_472 | 549 | 2468.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 494_501 | 549 | 2468.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 523_547 | 549 | 2468.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 101_121 | 615 | 2534.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 437_457 | 615 | 2534.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 473_493 | 615 | 2534.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 502_522 | 615 | 2534.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 548_568 | 615 | 2534.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 578_598 | 615 | 2534.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 101_121 | 614 | 1526.3333333333333 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 437_457 | 614 | 1526.3333333333333 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 473_493 | 614 | 1526.3333333333333 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 502_522 | 614 | 1526.3333333333333 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 548_568 | 614 | 1526.3333333333333 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 578_598 | 614 | 1526.3333333333333 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 101_121 | 464 | 2461.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 437_457 | 464 | 2461.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 101_121 | 464 | 2513.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 437_457 | 464 | 2513.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 101_121 | 464 | 2093.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 437_457 | 464 | 2093.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 101_121 | 549 | 1382.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 437_457 | 549 | 1382.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 473_493 | 549 | 1382.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 502_522 | 549 | 1382.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 101_121 | 549 | 2468.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 437_457 | 549 | 2468.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 473_493 | 549 | 2468.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 502_522 | 549 | 2468.3333333333335 | Transmembrane | Helical |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 2517_2551 | 2377 | 2640.0 | Region | Note=Hinge 2 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 2517_2647 | 2377 | 2640.0 | Region | Note=Self-association site%2C tail | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 2517_2551 | 2385 | 2648.0 | Region | Note=Hinge 2 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 2517_2647 | 2385 | 2648.0 | Region | Note=Self-association site%2C tail | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 2517_2551 | 2377 | 2640.0 | Region | Note=Hinge 2 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 2517_2647 | 2377 | 2640.0 | Region | Note=Self-association site%2C tail | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 2424_2516 | 2377 | 2640.0 | Repeat | Note=Filamin 23 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 2552_2646 | 2377 | 2640.0 | Repeat | Note=Filamin 24 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 2424_2516 | 2385 | 2648.0 | Repeat | Note=Filamin 23 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 2552_2646 | 2385 | 2648.0 | Repeat | Note=Filamin 24 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 2424_2516 | 2377 | 2640.0 | Repeat | Note=Filamin 23 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 2552_2646 | 2377 | 2640.0 | Repeat | Note=Filamin 24 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 14_31 | 0 | 184.0 | Compositional bias | Note=Gly-rich |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 438_598 | 464 | 2461.3333333333335 | Domain | SSD |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 438_598 | 464 | 2513.3333333333335 | Domain | SSD |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 438_598 | 464 | 2093.3333333333335 | Domain | SSD |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 438_598 | 549 | 1382.0 | Domain | SSD |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 438_598 | 549 | 2468.3333333333335 | Domain | SSD |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 438_598 | 0 | 184.0 | Domain | SSD |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 1049_1055 | 615 | 2534.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 1077_1083 | 615 | 2534.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 1105_1121 | 615 | 2534.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 1142_1154 | 615 | 2534.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 1176_1447 | 615 | 2534.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 599_748 | 615 | 2534.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 770_1027 | 615 | 2534.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 1049_1055 | 614 | 1526.3333333333333 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 1077_1083 | 614 | 1526.3333333333333 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 1105_1121 | 614 | 1526.3333333333333 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 1142_1154 | 614 | 1526.3333333333333 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 1176_1447 | 614 | 1526.3333333333333 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 599_748 | 614 | 1526.3333333333333 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 770_1027 | 614 | 1526.3333333333333 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 1049_1055 | 464 | 2461.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 1077_1083 | 464 | 2461.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 1105_1121 | 464 | 2461.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 1142_1154 | 464 | 2461.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 1176_1447 | 464 | 2461.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 458_472 | 464 | 2461.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 494_501 | 464 | 2461.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 523_547 | 464 | 2461.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 569_577 | 464 | 2461.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 599_748 | 464 | 2461.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 770_1027 | 464 | 2461.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 1049_1055 | 464 | 2513.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 1077_1083 | 464 | 2513.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 1105_1121 | 464 | 2513.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 1142_1154 | 464 | 2513.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 1176_1447 | 464 | 2513.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 458_472 | 464 | 2513.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 494_501 | 464 | 2513.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 523_547 | 464 | 2513.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 569_577 | 464 | 2513.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 599_748 | 464 | 2513.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 770_1027 | 464 | 2513.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 1049_1055 | 464 | 2093.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 1077_1083 | 464 | 2093.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 1105_1121 | 464 | 2093.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 1142_1154 | 464 | 2093.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 1176_1447 | 464 | 2093.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 458_472 | 464 | 2093.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 494_501 | 464 | 2093.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 523_547 | 464 | 2093.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 569_577 | 464 | 2093.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 599_748 | 464 | 2093.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 770_1027 | 464 | 2093.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 1049_1055 | 549 | 1382.0 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 1077_1083 | 549 | 1382.0 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 1105_1121 | 549 | 1382.0 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 1142_1154 | 549 | 1382.0 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 1176_1447 | 549 | 1382.0 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 569_577 | 549 | 1382.0 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 599_748 | 549 | 1382.0 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 770_1027 | 549 | 1382.0 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 1049_1055 | 549 | 2468.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 1077_1083 | 549 | 2468.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 1105_1121 | 549 | 2468.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 1142_1154 | 549 | 2468.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 1176_1447 | 549 | 2468.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 569_577 | 549 | 2468.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 599_748 | 549 | 2468.3333333333335 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 770_1027 | 549 | 2468.3333333333335 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 1049_1055 | 0 | 184.0 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 1077_1083 | 0 | 184.0 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 1105_1121 | 0 | 184.0 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 1142_1154 | 0 | 184.0 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 1176_1447 | 0 | 184.0 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 122_436 | 0 | 184.0 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 1_100 | 0 | 184.0 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 458_472 | 0 | 184.0 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 494_501 | 0 | 184.0 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 523_547 | 0 | 184.0 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 569_577 | 0 | 184.0 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 599_748 | 0 | 184.0 | Topological domain | Cytoplasmic |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 770_1027 | 0 | 184.0 | Topological domain | Extracellular |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 1028_1048 | 615 | 2534.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 1056_1076 | 615 | 2534.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 1084_1104 | 615 | 2534.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 1122_1141 | 615 | 2534.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 1155_1175 | 615 | 2534.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000331920 | - | 13 | 24 | 749_769 | 615 | 2534.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 1028_1048 | 614 | 1526.3333333333333 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 1056_1076 | 614 | 1526.3333333333333 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 1084_1104 | 614 | 1526.3333333333333 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 1122_1141 | 614 | 1526.3333333333333 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 1155_1175 | 614 | 1526.3333333333333 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000375274 | - | 13 | 24 | 749_769 | 614 | 1526.3333333333333 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 1028_1048 | 464 | 2461.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 1056_1076 | 464 | 2461.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 1084_1104 | 464 | 2461.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 1122_1141 | 464 | 2461.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 1155_1175 | 464 | 2461.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 473_493 | 464 | 2461.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 502_522 | 464 | 2461.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 548_568 | 464 | 2461.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 578_598 | 464 | 2461.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000418258 | - | 13 | 24 | 749_769 | 464 | 2461.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 1028_1048 | 464 | 2513.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 1056_1076 | 464 | 2513.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 1084_1104 | 464 | 2513.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 1122_1141 | 464 | 2513.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 1155_1175 | 464 | 2513.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 473_493 | 464 | 2513.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 502_522 | 464 | 2513.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 548_568 | 464 | 2513.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 578_598 | 464 | 2513.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000421141 | - | 13 | 24 | 749_769 | 464 | 2513.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 1028_1048 | 464 | 2093.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 1056_1076 | 464 | 2093.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 1084_1104 | 464 | 2093.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 1122_1141 | 464 | 2093.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 1155_1175 | 464 | 2093.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 473_493 | 464 | 2093.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 502_522 | 464 | 2093.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 548_568 | 464 | 2093.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 578_598 | 464 | 2093.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000429896 | - | 13 | 24 | 749_769 | 464 | 2093.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 1028_1048 | 549 | 1382.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 1056_1076 | 549 | 1382.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 1084_1104 | 549 | 1382.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 1122_1141 | 549 | 1382.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 1155_1175 | 549 | 1382.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 548_568 | 549 | 1382.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 578_598 | 549 | 1382.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000430669 | - | 13 | 23 | 749_769 | 549 | 1382.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 1028_1048 | 549 | 2468.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 1056_1076 | 549 | 2468.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 1084_1104 | 549 | 2468.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 1122_1141 | 549 | 2468.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 1155_1175 | 549 | 2468.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 548_568 | 549 | 2468.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 578_598 | 549 | 2468.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000437951 | - | 13 | 24 | 749_769 | 549 | 2468.3333333333335 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 101_121 | 0 | 184.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 1028_1048 | 0 | 184.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 1056_1076 | 0 | 184.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 1084_1104 | 0 | 184.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 1122_1141 | 0 | 184.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 1155_1175 | 0 | 184.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 437_457 | 0 | 184.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 473_493 | 0 | 184.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 502_522 | 0 | 184.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 548_568 | 0 | 184.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 578_598 | 0 | 184.0 | Transmembrane | Helical |
| Hgene | PTCH1 | chr9:98232094 | chrX:153578575 | ENST00000468211 | - | 1 | 5 | 749_769 | 0 | 184.0 | Transmembrane | Helical |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 166_269 | 2377 | 2640.0 | Domain | Calponin-homology (CH) 2 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 43_149 | 2377 | 2640.0 | Domain | Calponin-homology (CH) 1 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 166_269 | 2385 | 2648.0 | Domain | Calponin-homology (CH) 2 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 43_149 | 2385 | 2648.0 | Domain | Calponin-homology (CH) 1 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 166_269 | 2377 | 2640.0 | Domain | Calponin-homology (CH) 2 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 43_149 | 2377 | 2640.0 | Domain | Calponin-homology (CH) 1 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 1741_1778 | 2377 | 2640.0 | Region | Note=Hinge 1 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 2_274 | 2377 | 2640.0 | Region | Note=Actin-binding | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 1741_1778 | 2385 | 2648.0 | Region | Note=Hinge 1 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 2_274 | 2385 | 2648.0 | Region | Note=Actin-binding | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 1741_1778 | 2377 | 2640.0 | Region | Note=Hinge 1 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 2_274 | 2377 | 2640.0 | Region | Note=Actin-binding | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 1062_1154 | 2377 | 2640.0 | Repeat | Note=Filamin 9 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 1155_1249 | 2377 | 2640.0 | Repeat | Note=Filamin 10 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 1250_1349 | 2377 | 2640.0 | Repeat | Note=Filamin 11 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 1350_1442 | 2377 | 2640.0 | Repeat | Note=Filamin 12 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 1443_1539 | 2377 | 2640.0 | Repeat | Note=Filamin 13 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 1540_1636 | 2377 | 2640.0 | Repeat | Note=Filamin 14 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 1649_1740 | 2377 | 2640.0 | Repeat | Note=Filamin 15 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 1779_1860 | 2377 | 2640.0 | Repeat | Note=Filamin 16 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 1861_1950 | 2377 | 2640.0 | Repeat | Note=Filamin 17 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 1951_2039 | 2377 | 2640.0 | Repeat | Note=Filamin 18 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 2042_2131 | 2377 | 2640.0 | Repeat | Note=Filamin 19 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 2132_2230 | 2377 | 2640.0 | Repeat | Note=Filamin 20 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 2233_2325 | 2377 | 2640.0 | Repeat | Note=Filamin 21 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 2327_2420 | 2377 | 2640.0 | Repeat | Note=Filamin 22 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 276_374 | 2377 | 2640.0 | Repeat | Note=Filamin 1 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 376_474 | 2377 | 2640.0 | Repeat | Note=Filamin 2 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 475_570 | 2377 | 2640.0 | Repeat | Note=Filamin 3 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 571_663 | 2377 | 2640.0 | Repeat | Note=Filamin 4 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 667_763 | 2377 | 2640.0 | Repeat | Note=Filamin 5 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 764_866 | 2377 | 2640.0 | Repeat | Note=Filamin 6 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 867_965 | 2377 | 2640.0 | Repeat | Note=Filamin 7 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 966_1061 | 2377 | 2640.0 | Repeat | Note=Filamin 8 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 1062_1154 | 2385 | 2648.0 | Repeat | Note=Filamin 9 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 1155_1249 | 2385 | 2648.0 | Repeat | Note=Filamin 10 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 1250_1349 | 2385 | 2648.0 | Repeat | Note=Filamin 11 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 1350_1442 | 2385 | 2648.0 | Repeat | Note=Filamin 12 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 1443_1539 | 2385 | 2648.0 | Repeat | Note=Filamin 13 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 1540_1636 | 2385 | 2648.0 | Repeat | Note=Filamin 14 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 1649_1740 | 2385 | 2648.0 | Repeat | Note=Filamin 15 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 1779_1860 | 2385 | 2648.0 | Repeat | Note=Filamin 16 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 1861_1950 | 2385 | 2648.0 | Repeat | Note=Filamin 17 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 1951_2039 | 2385 | 2648.0 | Repeat | Note=Filamin 18 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 2042_2131 | 2385 | 2648.0 | Repeat | Note=Filamin 19 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 2132_2230 | 2385 | 2648.0 | Repeat | Note=Filamin 20 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 2233_2325 | 2385 | 2648.0 | Repeat | Note=Filamin 21 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 2327_2420 | 2385 | 2648.0 | Repeat | Note=Filamin 22 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 276_374 | 2385 | 2648.0 | Repeat | Note=Filamin 1 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 376_474 | 2385 | 2648.0 | Repeat | Note=Filamin 2 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 475_570 | 2385 | 2648.0 | Repeat | Note=Filamin 3 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 571_663 | 2385 | 2648.0 | Repeat | Note=Filamin 4 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 667_763 | 2385 | 2648.0 | Repeat | Note=Filamin 5 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 764_866 | 2385 | 2648.0 | Repeat | Note=Filamin 6 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 867_965 | 2385 | 2648.0 | Repeat | Note=Filamin 7 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 966_1061 | 2385 | 2648.0 | Repeat | Note=Filamin 8 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 1062_1154 | 2377 | 2640.0 | Repeat | Note=Filamin 9 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 1155_1249 | 2377 | 2640.0 | Repeat | Note=Filamin 10 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 1250_1349 | 2377 | 2640.0 | Repeat | Note=Filamin 11 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 1350_1442 | 2377 | 2640.0 | Repeat | Note=Filamin 12 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 1443_1539 | 2377 | 2640.0 | Repeat | Note=Filamin 13 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 1540_1636 | 2377 | 2640.0 | Repeat | Note=Filamin 14 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 1649_1740 | 2377 | 2640.0 | Repeat | Note=Filamin 15 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 1779_1860 | 2377 | 2640.0 | Repeat | Note=Filamin 16 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 1861_1950 | 2377 | 2640.0 | Repeat | Note=Filamin 17 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 1951_2039 | 2377 | 2640.0 | Repeat | Note=Filamin 18 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 2042_2131 | 2377 | 2640.0 | Repeat | Note=Filamin 19 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 2132_2230 | 2377 | 2640.0 | Repeat | Note=Filamin 20 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 2233_2325 | 2377 | 2640.0 | Repeat | Note=Filamin 21 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 2327_2420 | 2377 | 2640.0 | Repeat | Note=Filamin 22 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 276_374 | 2377 | 2640.0 | Repeat | Note=Filamin 1 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 376_474 | 2377 | 2640.0 | Repeat | Note=Filamin 2 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 475_570 | 2377 | 2640.0 | Repeat | Note=Filamin 3 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 571_663 | 2377 | 2640.0 | Repeat | Note=Filamin 4 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 667_763 | 2377 | 2640.0 | Repeat | Note=Filamin 5 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 764_866 | 2377 | 2640.0 | Repeat | Note=Filamin 6 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 867_965 | 2377 | 2640.0 | Repeat | Note=Filamin 7 | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 966_1061 | 2377 | 2640.0 | Repeat | Note=Filamin 8 |
Top |
Fusion Gene Sequence for PTCH1-FLNA |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >69864_69864_1_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000421141_FLNA_chrX_153578575_ENST00000344736_length(transcript)=2606nt_BP=1753nt ACTCACACCTCTCCAGGAAAAGCAGCAGACAAATGGGGATTGAAAAATTCAAACCCTCCCTCTGGTCCTGGGAGGAAAGGGCTGTCTGAG GTCCGCAGGGGGTGGAGGGGAAGGCTACTGGCCGGAAAGCGCCGCTGTGGCTGAGAGCGAAGTTTCAGAGACTCTTATTTAAACTGGGTT GTTACATTCAAAAAAACTGCGGCAAGTTCTTGGTTGTGGGCCTCCTCATATTTGGGGCCTTCGCGGTGGGATTAAAAGCAGCGAACCTCG AGACCAACGTGGAGGAGCTGTGGGTGGAAGTTGGAGGACGAGTAAGTCGTGAATTAAATTATACTCGCCAGAAGATTGGAGAAGAGGCTA TGTTTAATCCTCAACTCATGATACAGACCCCTAAAGAAGAAGGTGCTAATGTCCTGACCACAGAAGCGCTCCTACAACACCTGGACTCGG CACTCCAGGCCAGCCGTGTCCATGTATACATGTACAACAGGCAGTGGAAATTGGAACATTTGTGTTACAAATCAGGAGAGCTTATCACAG AAACAGGTTACATGGATCAGATAATAGAATATCTTTACCCTTGTTTGATTATTACACCTTTGGACTGCTTCTGGGAAGGGGCGAAATTAC AGTCTGGGACAGCATACCTCCTAGGTAAACCTCCTTTGCGGTGGACAAACTTCGACCCTTTGGAATTCCTGGAAGAGTTAAAGAAAATAA ACTATCAAGTGGACAGCTGGGAGGAAATGCTGAATAAGGCTGAGGTTGGTCATGGTTACATGGACCGCCCCTGCCTCAATCCGGCCGATC CAGACTGCCCCGCCACAGCCCCCAACAAAAATTCAACCAAACCTCTTGATATGGCCCTTGTTTTGAATGGTGGATGTCATGGCTTATCCA GAAAGTATATGCACTGGCAGGAGGAGTTGATTGTGGGTGGCACAGTCAAGAACAGCACTGGAAAACTCGTCAGCGCCCATGCCCTGCAGA CCATGTTCCAGTTAATGACTCCCAAGCAAATGTACGAGCACTTCAAGGGGTACGAGTATGTCTCACACATCAACTGGAACGAGGACAAAG CGGCAGCCATCCTGGAGGCCTGGCAGAGGACATATGTGGAGGTGGTTCATCAGAGTGTCGCACAGAACTCCACTCAAAAGGTGCTTTCCT TCACCACCACGACCCTGGACGACATCCTGAAATCCTTCTCTGACGTCAGTGTCATCCGCGTGGCCAGCGGCTACTTACTCATGCTCGCCT ATGCCTGTCTAACCATGCTGCGCTGGGACTGCTCCAAGTCCCAGGGTGCCGTGGGGCTGGCTGGCGTCCTGCTGGTTGCACTGTCAGTGG CTGCAGGACTGGGCCTGTGCTCATTGATCGGAATTTCCTTTAACGCTGCAACAACTCAGGTTTTGCCATTTCTCGCTCTTGGTGTTGGTG TGGATGATGTTTTTCTTCTGGCCCACGCCTTCAGTGAAACAGGACAGAATAAAAGAATCCCTTTTGAGGACAGGACCGGGGAGTGCCTGA AGCGCACAGGAGCCAGCGTGGCCCTCACGTCCATCAGCAATGTCACAGCCTTCTTCATGGCCGCGTTAATCCCAATTCCCGCTCTGCGGG CGTTCTCCCTCCAGGCAGCGGTAGTAGTGGTGTTCAATTTTGCCATGGTTCTGCTCATTTTTCCTGCAATTCTCAGCATGGATTTATATC GACGCGAGGACAGGAGACTGGATATTTTCTGCTGTTTTACAAGATAAGTATGCTGTGCGCTTCATCCCTCGGGAGAATGGCGTTTACCTG ATTGACGTCAAGTTCAACGGCACCCACATCCCTGGAAGCCCCTTCAAGATCCGAGTTGGGGAGCCTGGGCATGGAGGGGACCCAGGCTTG GTGTCTGCTTACGGAGCAGGTCTGGAAGGCGGTGTCACAGGGAACCCAGCTGAGTTCGTCGTGAACACGAGCAATGCGGGAGCTGGTGCC CTGTCGGTGACCATTGACGGCCCCTCCAAGGTGAAGATGGATTGCCAGGAGTGCCCTGAGGGCTACCGCGTCACCTATACCCCCATGGCA CCTGGCAGCTACCTCATCTCCATCAAGTACGGCGGCCCCTACCACATTGGGGGCAGCCCCTTCAAGGCCAAAGTCACAGGCCCCCGTCTC GTCAGCAACCACAGCCTCCACGAGACATCATCAGTGTTTGTAGACTCTCTGACCAAGGCCACCTGTGCCCCCCAGCATGGGGCCCCGGGT CCTGGGCCTGCTGACGCCAGCAAGGTGGTGGCCAAGGGCCTGGGGCTGAGCAAGGCCTACGTAGGCCAGAAGAGCAGCTTCACAGTAGAC TGCAGCAAAGCAGGCAACAACATGCTGCTGGTGGGGGTTCATGGCCCAAGGACCCCCTGCGAGGAGATCCTGGTGAAGCACGTGGGCAGC CGGCTCTACAGCGTGTCCTACCTGCTCAAGGACAAGGGGGAGTACACACTGGTGGTCAAATGGGGGGACGAGCACATCCCAGGCAGCCCC >69864_69864_1_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000421141_FLNA_chrX_153578575_ENST00000344736_length(amino acids)=540AA_BP= MWLRAKFQRLLFKLGCYIQKNCGKFLVVGLLIFGAFAVGLKAANLETNVEELWVEVGGRVSRELNYTRQKIGEEAMFNPQLMIQTPKEEG ANVLTTEALLQHLDSALQASRVHVYMYNRQWKLEHLCYKSGELITETGYMDQIIEYLYPCLIITPLDCFWEGAKLQSGTAYLLGKPPLRW TNFDPLEFLEELKKINYQVDSWEEMLNKAEVGHGYMDRPCLNPADPDCPATAPNKNSTKPLDMALVLNGGCHGLSRKYMHWQEELIVGGT VKNSTGKLVSAHALQTMFQLMTPKQMYEHFKGYEYVSHINWNEDKAAAILEAWQRTYVEVVHQSVAQNSTQKVLSFTTTTLDDILKSFSD VSVIRVASGYLLMLAYACLTMLRWDCSKSQGAVGLAGVLLVALSVAAGLGLCSLIGISFNAATTQVLPFLALGVGVDDVFLLAHAFSETG QNKRIPFEDRTGECLKRTGASVALTSISNVTAFFMAALIPIPALRAFSLQAAVVVVFNFAMVLLIFPAILSMDLYRREDRRLDIFCCFTR -------------------------------------------------------------- >69864_69864_2_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000421141_FLNA_chrX_153578575_ENST00000360319_length(transcript)=2864nt_BP=1753nt ACTCACACCTCTCCAGGAAAAGCAGCAGACAAATGGGGATTGAAAAATTCAAACCCTCCCTCTGGTCCTGGGAGGAAAGGGCTGTCTGAG GTCCGCAGGGGGTGGAGGGGAAGGCTACTGGCCGGAAAGCGCCGCTGTGGCTGAGAGCGAAGTTTCAGAGACTCTTATTTAAACTGGGTT GTTACATTCAAAAAAACTGCGGCAAGTTCTTGGTTGTGGGCCTCCTCATATTTGGGGCCTTCGCGGTGGGATTAAAAGCAGCGAACCTCG AGACCAACGTGGAGGAGCTGTGGGTGGAAGTTGGAGGACGAGTAAGTCGTGAATTAAATTATACTCGCCAGAAGATTGGAGAAGAGGCTA TGTTTAATCCTCAACTCATGATACAGACCCCTAAAGAAGAAGGTGCTAATGTCCTGACCACAGAAGCGCTCCTACAACACCTGGACTCGG CACTCCAGGCCAGCCGTGTCCATGTATACATGTACAACAGGCAGTGGAAATTGGAACATTTGTGTTACAAATCAGGAGAGCTTATCACAG AAACAGGTTACATGGATCAGATAATAGAATATCTTTACCCTTGTTTGATTATTACACCTTTGGACTGCTTCTGGGAAGGGGCGAAATTAC AGTCTGGGACAGCATACCTCCTAGGTAAACCTCCTTTGCGGTGGACAAACTTCGACCCTTTGGAATTCCTGGAAGAGTTAAAGAAAATAA ACTATCAAGTGGACAGCTGGGAGGAAATGCTGAATAAGGCTGAGGTTGGTCATGGTTACATGGACCGCCCCTGCCTCAATCCGGCCGATC CAGACTGCCCCGCCACAGCCCCCAACAAAAATTCAACCAAACCTCTTGATATGGCCCTTGTTTTGAATGGTGGATGTCATGGCTTATCCA GAAAGTATATGCACTGGCAGGAGGAGTTGATTGTGGGTGGCACAGTCAAGAACAGCACTGGAAAACTCGTCAGCGCCCATGCCCTGCAGA CCATGTTCCAGTTAATGACTCCCAAGCAAATGTACGAGCACTTCAAGGGGTACGAGTATGTCTCACACATCAACTGGAACGAGGACAAAG CGGCAGCCATCCTGGAGGCCTGGCAGAGGACATATGTGGAGGTGGTTCATCAGAGTGTCGCACAGAACTCCACTCAAAAGGTGCTTTCCT TCACCACCACGACCCTGGACGACATCCTGAAATCCTTCTCTGACGTCAGTGTCATCCGCGTGGCCAGCGGCTACTTACTCATGCTCGCCT ATGCCTGTCTAACCATGCTGCGCTGGGACTGCTCCAAGTCCCAGGGTGCCGTGGGGCTGGCTGGCGTCCTGCTGGTTGCACTGTCAGTGG CTGCAGGACTGGGCCTGTGCTCATTGATCGGAATTTCCTTTAACGCTGCAACAACTCAGGTTTTGCCATTTCTCGCTCTTGGTGTTGGTG TGGATGATGTTTTTCTTCTGGCCCACGCCTTCAGTGAAACAGGACAGAATAAAAGAATCCCTTTTGAGGACAGGACCGGGGAGTGCCTGA AGCGCACAGGAGCCAGCGTGGCCCTCACGTCCATCAGCAATGTCACAGCCTTCTTCATGGCCGCGTTAATCCCAATTCCCGCTCTGCGGG CGTTCTCCCTCCAGGCAGCGGTAGTAGTGGTGTTCAATTTTGCCATGGTTCTGCTCATTTTTCCTGCAATTCTCAGCATGGATTTATATC GACGCGAGGACAGGAGACTGGATATTTTCTGCTGTTTTACAAGATAAGTATGCTGTGCGCTTCATCCCTCGGGAGAATGGCGTTTACCTG ATTGACGTCAAGTTCAACGGCACCCACATCCCTGGAAGCCCCTTCAAGATCCGAGTTGGGGAGCCTGGGCATGGAGGGGACCCAGGCTTG GTGTCTGCTTACGGAGCAGGTCTGGAAGGCGGTGTCACAGGGAACCCAGCTGAGTTCGTCGTGAACACGAGCAATGCGGGAGCTGGTGCC CTGTCGGTGACCATTGACGGCCCCTCCAAGGTGAAGATGGATTGCCAGGAGTGCCCTGAGGGCTACCGCGTCACCTATACCCCCATGGCA CCTGGCAGCTACCTCATCTCCATCAAGTACGGCGGCCCCTACCACATTGGGGGCAGCCCCTTCAAGGCCAAAGTCACAGGCCCCCGTCTC GTCAGCAACCACAGCCTCCACGAGACATCATCAGTGTTTGTAGACTCTCTGACCAAGGCCACCTGTGCCCCCCAGCATGGGGCCCCGGGT CCTGGGCCTGCTGACGCCAGCAAGGTGGTGGCCAAGGGCCTGGGGCTGAGCAAGGCCTACGTAGGCCAGAAGAGCAGCTTCACAGTAGAC TGCAGCAAAGCAGGCAACAACATGCTGCTGGTGGGGGTTCATGGCCCAAGGACCCCCTGCGAGGAGATCCTGGTGAAGCACGTGGGCAGC CGGCTCTACAGCGTGTCCTACCTGCTCAAGGACAAGGGGGAGTACACACTGGTGGTCAAATGGGGGGACGAGCACATCCCAGGCAGCCCC TACCGCGTTGTGGTGCCCTGAGTCTGGGGCCCGTGCCAGCCGGCAGCCCCCAAGCCTGCCCCGCTACCCAAGCAGCCCCGCCCTCTTCCC CTCAACCCCGGCCCAGGCCGCCCTGGCCGCCCGCCTGTCACTGCAGCCGCCCCTGCCCTGTGCCGTGCTGCGCTCACCTGCCTCCCCAGC CAGCCGCTGACCTCTCGGCTTTCACTTGGGCAGAGGGAGCCATTTGGTGGCGCTGCTTGTCTTCTTTGGTTCTGGGAGGGGTGAGGGATG >69864_69864_2_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000421141_FLNA_chrX_153578575_ENST00000360319_length(amino acids)=540AA_BP= MWLRAKFQRLLFKLGCYIQKNCGKFLVVGLLIFGAFAVGLKAANLETNVEELWVEVGGRVSRELNYTRQKIGEEAMFNPQLMIQTPKEEG ANVLTTEALLQHLDSALQASRVHVYMYNRQWKLEHLCYKSGELITETGYMDQIIEYLYPCLIITPLDCFWEGAKLQSGTAYLLGKPPLRW TNFDPLEFLEELKKINYQVDSWEEMLNKAEVGHGYMDRPCLNPADPDCPATAPNKNSTKPLDMALVLNGGCHGLSRKYMHWQEELIVGGT VKNSTGKLVSAHALQTMFQLMTPKQMYEHFKGYEYVSHINWNEDKAAAILEAWQRTYVEVVHQSVAQNSTQKVLSFTTTTLDDILKSFSD VSVIRVASGYLLMLAYACLTMLRWDCSKSQGAVGLAGVLLVALSVAAGLGLCSLIGISFNAATTQVLPFLALGVGVDDVFLLAHAFSETG QNKRIPFEDRTGECLKRTGASVALTSISNVTAFFMAALIPIPALRAFSLQAAVVVVFNFAMVLLIFPAILSMDLYRREDRRLDIFCCFTR -------------------------------------------------------------- >69864_69864_3_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000421141_FLNA_chrX_153578575_ENST00000369850_length(transcript)=2742nt_BP=1753nt ACTCACACCTCTCCAGGAAAAGCAGCAGACAAATGGGGATTGAAAAATTCAAACCCTCCCTCTGGTCCTGGGAGGAAAGGGCTGTCTGAG GTCCGCAGGGGGTGGAGGGGAAGGCTACTGGCCGGAAAGCGCCGCTGTGGCTGAGAGCGAAGTTTCAGAGACTCTTATTTAAACTGGGTT GTTACATTCAAAAAAACTGCGGCAAGTTCTTGGTTGTGGGCCTCCTCATATTTGGGGCCTTCGCGGTGGGATTAAAAGCAGCGAACCTCG AGACCAACGTGGAGGAGCTGTGGGTGGAAGTTGGAGGACGAGTAAGTCGTGAATTAAATTATACTCGCCAGAAGATTGGAGAAGAGGCTA TGTTTAATCCTCAACTCATGATACAGACCCCTAAAGAAGAAGGTGCTAATGTCCTGACCACAGAAGCGCTCCTACAACACCTGGACTCGG CACTCCAGGCCAGCCGTGTCCATGTATACATGTACAACAGGCAGTGGAAATTGGAACATTTGTGTTACAAATCAGGAGAGCTTATCACAG AAACAGGTTACATGGATCAGATAATAGAATATCTTTACCCTTGTTTGATTATTACACCTTTGGACTGCTTCTGGGAAGGGGCGAAATTAC AGTCTGGGACAGCATACCTCCTAGGTAAACCTCCTTTGCGGTGGACAAACTTCGACCCTTTGGAATTCCTGGAAGAGTTAAAGAAAATAA ACTATCAAGTGGACAGCTGGGAGGAAATGCTGAATAAGGCTGAGGTTGGTCATGGTTACATGGACCGCCCCTGCCTCAATCCGGCCGATC CAGACTGCCCCGCCACAGCCCCCAACAAAAATTCAACCAAACCTCTTGATATGGCCCTTGTTTTGAATGGTGGATGTCATGGCTTATCCA GAAAGTATATGCACTGGCAGGAGGAGTTGATTGTGGGTGGCACAGTCAAGAACAGCACTGGAAAACTCGTCAGCGCCCATGCCCTGCAGA CCATGTTCCAGTTAATGACTCCCAAGCAAATGTACGAGCACTTCAAGGGGTACGAGTATGTCTCACACATCAACTGGAACGAGGACAAAG CGGCAGCCATCCTGGAGGCCTGGCAGAGGACATATGTGGAGGTGGTTCATCAGAGTGTCGCACAGAACTCCACTCAAAAGGTGCTTTCCT TCACCACCACGACCCTGGACGACATCCTGAAATCCTTCTCTGACGTCAGTGTCATCCGCGTGGCCAGCGGCTACTTACTCATGCTCGCCT ATGCCTGTCTAACCATGCTGCGCTGGGACTGCTCCAAGTCCCAGGGTGCCGTGGGGCTGGCTGGCGTCCTGCTGGTTGCACTGTCAGTGG CTGCAGGACTGGGCCTGTGCTCATTGATCGGAATTTCCTTTAACGCTGCAACAACTCAGGTTTTGCCATTTCTCGCTCTTGGTGTTGGTG TGGATGATGTTTTTCTTCTGGCCCACGCCTTCAGTGAAACAGGACAGAATAAAAGAATCCCTTTTGAGGACAGGACCGGGGAGTGCCTGA AGCGCACAGGAGCCAGCGTGGCCCTCACGTCCATCAGCAATGTCACAGCCTTCTTCATGGCCGCGTTAATCCCAATTCCCGCTCTGCGGG CGTTCTCCCTCCAGGCAGCGGTAGTAGTGGTGTTCAATTTTGCCATGGTTCTGCTCATTTTTCCTGCAATTCTCAGCATGGATTTATATC GACGCGAGGACAGGAGACTGGATATTTTCTGCTGTTTTACAAGATAAGTATGCTGTGCGCTTCATCCCTCGGGAGAATGGCGTTTACCTG ATTGACGTCAAGTTCAACGGCACCCACATCCCTGGAAGCCCCTTCAAGATCCGAGTTGGGGAGCCTGGGCATGGAGGGGACCCAGGCTTG GTGTCTGCTTACGGAGCAGGTCTGGAAGGCGGTGTCACAGGGAACCCAGCTGAGTTCGTCGTGAACACGAGCAATGCGGGAGCTGGTGCC CTGTCGGTGACCATTGACGGCCCCTCCAAGGTGAAGATGGATTGCCAGGAGTGCCCTGAGGGCTACCGCGTCACCTATACCCCCATGGCA CCTGGCAGCTACCTCATCTCCATCAAGTACGGCGGCCCCTACCACATTGGGGGCAGCCCCTTCAAGGCCAAAGTCACAGGCCCCCGTCTC GTCAGCAACCACAGCCTCCACGAGACATCATCAGTGTTTGTAGACTCTCTGACCAAGGCCACCTGTGCCCCCCAGCATGGGGCCCCGGGT CCTGGGCCTGCTGACGCCAGCAAGGTGGTGGCCAAGGGCCTGGGGCTGAGCAAGGCCTACGTAGGCCAGAAGAGCAGCTTCACAGTAGAC TGCAGCAAAGCAGGCAACAACATGCTGCTGGTGGGGGTTCATGGCCCAAGGACCCCCTGCGAGGAGATCCTGGTGAAGCACGTGGGCAGC CGGCTCTACAGCGTGTCCTACCTGCTCAAGGACAAGGGGGAGTACACACTGGTGGTCAAATGGGGGGACGAGCACATCCCAGGCAGCCCC TACCGCGTTGTGGTGCCCTGAGTCTGGGGCCCGTGCCAGCCGGCAGCCCCCAAGCCTGCCCCGCTACCCAAGCAGCCCCGCCCTCTTCCC CTCAACCCCGGCCCAGGCCGCCCTGGCCGCCCGCCTGTCACTGCAGCCGCCCCTGCCCTGTGCCGTGCTGCGCTCACCTGCCTCCCCAGC >69864_69864_3_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000421141_FLNA_chrX_153578575_ENST00000369850_length(amino acids)=540AA_BP= MWLRAKFQRLLFKLGCYIQKNCGKFLVVGLLIFGAFAVGLKAANLETNVEELWVEVGGRVSRELNYTRQKIGEEAMFNPQLMIQTPKEEG ANVLTTEALLQHLDSALQASRVHVYMYNRQWKLEHLCYKSGELITETGYMDQIIEYLYPCLIITPLDCFWEGAKLQSGTAYLLGKPPLRW TNFDPLEFLEELKKINYQVDSWEEMLNKAEVGHGYMDRPCLNPADPDCPATAPNKNSTKPLDMALVLNGGCHGLSRKYMHWQEELIVGGT VKNSTGKLVSAHALQTMFQLMTPKQMYEHFKGYEYVSHINWNEDKAAAILEAWQRTYVEVVHQSVAQNSTQKVLSFTTTTLDDILKSFSD VSVIRVASGYLLMLAYACLTMLRWDCSKSQGAVGLAGVLLVALSVAAGLGLCSLIGISFNAATTQVLPFLALGVGVDDVFLLAHAFSETG QNKRIPFEDRTGECLKRTGASVALTSISNVTAFFMAALIPIPALRAFSLQAAVVVVFNFAMVLLIFPAILSMDLYRREDRRLDIFCCFTR -------------------------------------------------------------- >69864_69864_4_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000421141_FLNA_chrX_153578575_ENST00000369856_length(transcript)=2734nt_BP=1753nt ACTCACACCTCTCCAGGAAAAGCAGCAGACAAATGGGGATTGAAAAATTCAAACCCTCCCTCTGGTCCTGGGAGGAAAGGGCTGTCTGAG GTCCGCAGGGGGTGGAGGGGAAGGCTACTGGCCGGAAAGCGCCGCTGTGGCTGAGAGCGAAGTTTCAGAGACTCTTATTTAAACTGGGTT GTTACATTCAAAAAAACTGCGGCAAGTTCTTGGTTGTGGGCCTCCTCATATTTGGGGCCTTCGCGGTGGGATTAAAAGCAGCGAACCTCG AGACCAACGTGGAGGAGCTGTGGGTGGAAGTTGGAGGACGAGTAAGTCGTGAATTAAATTATACTCGCCAGAAGATTGGAGAAGAGGCTA TGTTTAATCCTCAACTCATGATACAGACCCCTAAAGAAGAAGGTGCTAATGTCCTGACCACAGAAGCGCTCCTACAACACCTGGACTCGG CACTCCAGGCCAGCCGTGTCCATGTATACATGTACAACAGGCAGTGGAAATTGGAACATTTGTGTTACAAATCAGGAGAGCTTATCACAG AAACAGGTTACATGGATCAGATAATAGAATATCTTTACCCTTGTTTGATTATTACACCTTTGGACTGCTTCTGGGAAGGGGCGAAATTAC AGTCTGGGACAGCATACCTCCTAGGTAAACCTCCTTTGCGGTGGACAAACTTCGACCCTTTGGAATTCCTGGAAGAGTTAAAGAAAATAA ACTATCAAGTGGACAGCTGGGAGGAAATGCTGAATAAGGCTGAGGTTGGTCATGGTTACATGGACCGCCCCTGCCTCAATCCGGCCGATC CAGACTGCCCCGCCACAGCCCCCAACAAAAATTCAACCAAACCTCTTGATATGGCCCTTGTTTTGAATGGTGGATGTCATGGCTTATCCA GAAAGTATATGCACTGGCAGGAGGAGTTGATTGTGGGTGGCACAGTCAAGAACAGCACTGGAAAACTCGTCAGCGCCCATGCCCTGCAGA CCATGTTCCAGTTAATGACTCCCAAGCAAATGTACGAGCACTTCAAGGGGTACGAGTATGTCTCACACATCAACTGGAACGAGGACAAAG CGGCAGCCATCCTGGAGGCCTGGCAGAGGACATATGTGGAGGTGGTTCATCAGAGTGTCGCACAGAACTCCACTCAAAAGGTGCTTTCCT TCACCACCACGACCCTGGACGACATCCTGAAATCCTTCTCTGACGTCAGTGTCATCCGCGTGGCCAGCGGCTACTTACTCATGCTCGCCT ATGCCTGTCTAACCATGCTGCGCTGGGACTGCTCCAAGTCCCAGGGTGCCGTGGGGCTGGCTGGCGTCCTGCTGGTTGCACTGTCAGTGG CTGCAGGACTGGGCCTGTGCTCATTGATCGGAATTTCCTTTAACGCTGCAACAACTCAGGTTTTGCCATTTCTCGCTCTTGGTGTTGGTG TGGATGATGTTTTTCTTCTGGCCCACGCCTTCAGTGAAACAGGACAGAATAAAAGAATCCCTTTTGAGGACAGGACCGGGGAGTGCCTGA AGCGCACAGGAGCCAGCGTGGCCCTCACGTCCATCAGCAATGTCACAGCCTTCTTCATGGCCGCGTTAATCCCAATTCCCGCTCTGCGGG CGTTCTCCCTCCAGGCAGCGGTAGTAGTGGTGTTCAATTTTGCCATGGTTCTGCTCATTTTTCCTGCAATTCTCAGCATGGATTTATATC GACGCGAGGACAGGAGACTGGATATTTTCTGCTGTTTTACAAGATAAGTATGCTGTGCGCTTCATCCCTCGGGAGAATGGCGTTTACCTG ATTGACGTCAAGTTCAACGGCACCCACATCCCTGGAAGCCCCTTCAAGATCCGAGTTGGGGAGCCTGGGCATGGAGGGGACCCAGGCTTG GTGTCTGCTTACGGAGCAGGTCTGGAAGGCGGTGTCACAGGGAACCCAGCTGAGTTCGTCGTGAACACGAGCAATGCGGGAGCTGGTGCC CTGTCGGTGACCATTGACGGCCCCTCCAAGGTGAAGATGGATTGCCAGGAGTGCCCTGAGGGCTACCGCGTCACCTATACCCCCATGGCA CCTGGCAGCTACCTCATCTCCATCAAGTACGGCGGCCCCTACCACATTGGGGGCAGCCCCTTCAAGGCCAAAGTCACAGGCCCCCGTCTC GTCAGCAACCACAGCCTCCACGAGACATCATCAGTGTTTGTAGACTCTCTGACCAAGGCCACCTGTGCCCCCCAGCATGGGGCCCCGGGT CCTGGGCCTGCTGACGCCAGCAAGGTGGTGGCCAAGGGCCTGGGGCTGAGCAAGGCCTACGTAGGCCAGAAGAGCAGCTTCACAGTAGAC TGCAGCAAAGCAGGCAACAACATGCTGCTGGTGGGGGTTCATGGCCCAAGGACCCCCTGCGAGGAGATCCTGGTGAAGCACGTGGGCAGC CGGCTCTACAGCGTGTCCTACCTGCTCAAGGACAAGGGGGAGTACACACTGGTGGTCAAATGGGGGGACGAGCACATCCCAGGCAGCCCC TACCGCGTTGTGGTGCCCTGAGTCTGGGGCCCGTGCCAGCCGGCAGCCCCCAAGCCTGCCCCGCTACCCAAGCAGCCCCGCCCTCTTCCC CTCAACCCCGGCCCAGGCCGCCCTGGCCGCCCGCCTGTCACTGCAGCCGCCCCTGCCCTGTGCCGTGCTGCGCTCACCTGCCTCCCCAGC >69864_69864_4_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000421141_FLNA_chrX_153578575_ENST00000369856_length(amino acids)=540AA_BP= MWLRAKFQRLLFKLGCYIQKNCGKFLVVGLLIFGAFAVGLKAANLETNVEELWVEVGGRVSRELNYTRQKIGEEAMFNPQLMIQTPKEEG ANVLTTEALLQHLDSALQASRVHVYMYNRQWKLEHLCYKSGELITETGYMDQIIEYLYPCLIITPLDCFWEGAKLQSGTAYLLGKPPLRW TNFDPLEFLEELKKINYQVDSWEEMLNKAEVGHGYMDRPCLNPADPDCPATAPNKNSTKPLDMALVLNGGCHGLSRKYMHWQEELIVGGT VKNSTGKLVSAHALQTMFQLMTPKQMYEHFKGYEYVSHINWNEDKAAAILEAWQRTYVEVVHQSVAQNSTQKVLSFTTTTLDDILKSFSD VSVIRVASGYLLMLAYACLTMLRWDCSKSQGAVGLAGVLLVALSVAAGLGLCSLIGISFNAATTQVLPFLALGVGVDDVFLLAHAFSETG QNKRIPFEDRTGECLKRTGASVALTSISNVTAFFMAALIPIPALRAFSLQAAVVVVFNFAMVLLIFPAILSMDLYRREDRRLDIFCCFTR -------------------------------------------------------------- >69864_69864_5_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000421141_FLNA_chrX_153578575_ENST00000422373_length(transcript)=2858nt_BP=1753nt ACTCACACCTCTCCAGGAAAAGCAGCAGACAAATGGGGATTGAAAAATTCAAACCCTCCCTCTGGTCCTGGGAGGAAAGGGCTGTCTGAG GTCCGCAGGGGGTGGAGGGGAAGGCTACTGGCCGGAAAGCGCCGCTGTGGCTGAGAGCGAAGTTTCAGAGACTCTTATTTAAACTGGGTT GTTACATTCAAAAAAACTGCGGCAAGTTCTTGGTTGTGGGCCTCCTCATATTTGGGGCCTTCGCGGTGGGATTAAAAGCAGCGAACCTCG AGACCAACGTGGAGGAGCTGTGGGTGGAAGTTGGAGGACGAGTAAGTCGTGAATTAAATTATACTCGCCAGAAGATTGGAGAAGAGGCTA TGTTTAATCCTCAACTCATGATACAGACCCCTAAAGAAGAAGGTGCTAATGTCCTGACCACAGAAGCGCTCCTACAACACCTGGACTCGG CACTCCAGGCCAGCCGTGTCCATGTATACATGTACAACAGGCAGTGGAAATTGGAACATTTGTGTTACAAATCAGGAGAGCTTATCACAG AAACAGGTTACATGGATCAGATAATAGAATATCTTTACCCTTGTTTGATTATTACACCTTTGGACTGCTTCTGGGAAGGGGCGAAATTAC AGTCTGGGACAGCATACCTCCTAGGTAAACCTCCTTTGCGGTGGACAAACTTCGACCCTTTGGAATTCCTGGAAGAGTTAAAGAAAATAA ACTATCAAGTGGACAGCTGGGAGGAAATGCTGAATAAGGCTGAGGTTGGTCATGGTTACATGGACCGCCCCTGCCTCAATCCGGCCGATC CAGACTGCCCCGCCACAGCCCCCAACAAAAATTCAACCAAACCTCTTGATATGGCCCTTGTTTTGAATGGTGGATGTCATGGCTTATCCA GAAAGTATATGCACTGGCAGGAGGAGTTGATTGTGGGTGGCACAGTCAAGAACAGCACTGGAAAACTCGTCAGCGCCCATGCCCTGCAGA CCATGTTCCAGTTAATGACTCCCAAGCAAATGTACGAGCACTTCAAGGGGTACGAGTATGTCTCACACATCAACTGGAACGAGGACAAAG CGGCAGCCATCCTGGAGGCCTGGCAGAGGACATATGTGGAGGTGGTTCATCAGAGTGTCGCACAGAACTCCACTCAAAAGGTGCTTTCCT TCACCACCACGACCCTGGACGACATCCTGAAATCCTTCTCTGACGTCAGTGTCATCCGCGTGGCCAGCGGCTACTTACTCATGCTCGCCT ATGCCTGTCTAACCATGCTGCGCTGGGACTGCTCCAAGTCCCAGGGTGCCGTGGGGCTGGCTGGCGTCCTGCTGGTTGCACTGTCAGTGG CTGCAGGACTGGGCCTGTGCTCATTGATCGGAATTTCCTTTAACGCTGCAACAACTCAGGTTTTGCCATTTCTCGCTCTTGGTGTTGGTG TGGATGATGTTTTTCTTCTGGCCCACGCCTTCAGTGAAACAGGACAGAATAAAAGAATCCCTTTTGAGGACAGGACCGGGGAGTGCCTGA AGCGCACAGGAGCCAGCGTGGCCCTCACGTCCATCAGCAATGTCACAGCCTTCTTCATGGCCGCGTTAATCCCAATTCCCGCTCTGCGGG CGTTCTCCCTCCAGGCAGCGGTAGTAGTGGTGTTCAATTTTGCCATGGTTCTGCTCATTTTTCCTGCAATTCTCAGCATGGATTTATATC GACGCGAGGACAGGAGACTGGATATTTTCTGCTGTTTTACAAGATAAGTATGCTGTGCGCTTCATCCCTCGGGAGAATGGCGTTTACCTG ATTGACGTCAAGTTCAACGGCACCCACATCCCTGGAAGCCCCTTCAAGATCCGAGTTGGGGAGCCTGGGCATGGAGGGGACCCAGGCTTG GTGTCTGCTTACGGAGCAGGTCTGGAAGGCGGTGTCACAGGGAACCCAGCTGAGTTCGTCGTGAACACGAGCAATGCGGGAGCTGGTGCC CTGTCGGTGACCATTGACGGCCCCTCCAAGGTGAAGATGGATTGCCAGGAGTGCCCTGAGGGCTACCGCGTCACCTATACCCCCATGGCA CCTGGCAGCTACCTCATCTCCATCAAGTACGGCGGCCCCTACCACATTGGGGGCAGCCCCTTCAAGGCCAAAGTCACAGGCCCCCGTCTC GTCAGCAACCACAGCCTCCACGAGACATCATCAGTGTTTGTAGACTCTCTGACCAAGGCCACCTGTGCCCCCCAGCATGGGGCCCCGGGT CCTGGGCCTGCTGACGCCAGCAAGGTGGTGGCCAAGGGCCTGGGGCTGAGCAAGGCCTACGTAGGCCAGAAGAGCAGCTTCACAGTAGAC TGCAGCAAAGCAGGCAACAACATGCTGCTGGTGGGGGTTCATGGCCCAAGGACCCCCTGCGAGGAGATCCTGGTGAAGCACGTGGGCAGC CGGCTCTACAGCGTGTCCTACCTGCTCAAGGACAAGGGGGAGTACACACTGGTGGTCAAATGGGGGGACGAGCACATCCCAGGCAGCCCC TACCGCGTTGTGGTGCCCTGAGTCTGGGGCCCGTGCCAGCCGGCAGCCCCCAAGCCTGCCCCGCTACCCAAGCAGCCCCGCCCTCTTCCC CTCAACCCCGGCCCAGGCCGCCCTGGCCGCCCGCCTGTCACTGCAGCCGCCCCTGCCCTGTGCCGTGCTGCGCTCACCTGCCTCCCCAGC CAGCCGCTGACCTCTCGGCTTTCACTTGGGCAGAGGGAGCCATTTGGTGGCGCTGCTTGTCTTCTTTGGTTCTGGGAGGGGTGAGGGATG >69864_69864_5_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000421141_FLNA_chrX_153578575_ENST00000422373_length(amino acids)=540AA_BP= MWLRAKFQRLLFKLGCYIQKNCGKFLVVGLLIFGAFAVGLKAANLETNVEELWVEVGGRVSRELNYTRQKIGEEAMFNPQLMIQTPKEEG ANVLTTEALLQHLDSALQASRVHVYMYNRQWKLEHLCYKSGELITETGYMDQIIEYLYPCLIITPLDCFWEGAKLQSGTAYLLGKPPLRW TNFDPLEFLEELKKINYQVDSWEEMLNKAEVGHGYMDRPCLNPADPDCPATAPNKNSTKPLDMALVLNGGCHGLSRKYMHWQEELIVGGT VKNSTGKLVSAHALQTMFQLMTPKQMYEHFKGYEYVSHINWNEDKAAAILEAWQRTYVEVVHQSVAQNSTQKVLSFTTTTLDDILKSFSD VSVIRVASGYLLMLAYACLTMLRWDCSKSQGAVGLAGVLLVALSVAAGLGLCSLIGISFNAATTQVLPFLALGVGVDDVFLLAHAFSETG QNKRIPFEDRTGECLKRTGASVALTSISNVTAFFMAALIPIPALRAFSLQAAVVVVFNFAMVLLIFPAILSMDLYRREDRRLDIFCCFTR -------------------------------------------------------------- >69864_69864_6_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000429896_FLNA_chrX_153578575_ENST00000344736_length(transcript)=2688nt_BP=1835nt GAAGGCGAGCACCCAGACGGGGGCCCGCCGGGGCCGCGGCCAGCGCCGGGGAAATGCCGCGCCGGGGAGCAGCCTGCGCCGGCCTGAGCC CTTCCCTTTGCACTCGGCTGTTTTTTACGTTTAACCAGAAAGGAAGGGAGAGGAGGGAAAGATCCATGTGGCTGCCCTCTTCCGATCACA AATATTGTCGGGAAGGCTACTGGCCGGAAAGCGCCGCTGTGGCTGAGAGCGAAGTTTCAGAGACTCTTATTTAAACTGGGTTGTTACATT CAAAAAAACTGCGGCAAGTTCTTGGTTGTGGGCCTCCTCATATTTGGGGCCTTCGCGGTGGGATTAAAAGCAGCGAACCTCGAGACCAAC GTGGAGGAGCTGTGGGTGGAAGTTGGAGGACGAGTAAGTCGTGAATTAAATTATACTCGCCAGAAGATTGGAGAAGAGGCTATGTTTAAT CCTCAACTCATGATACAGACCCCTAAAGAAGAAGGTGCTAATGTCCTGACCACAGAAGCGCTCCTACAACACCTGGACTCGGCACTCCAG GCCAGCCGTGTCCATGTATACATGTACAACAGGCAGTGGAAATTGGAACATTTGTGTTACAAATCAGGAGAGCTTATCACAGAAACAGGT TACATGGATCAGATAATAGAATATCTTTACCCTTGTTTGATTATTACACCTTTGGACTGCTTCTGGGAAGGGGCGAAATTACAGTCTGGG ACAGCATACCTCCTAGGTAAACCTCCTTTGCGGTGGACAAACTTCGACCCTTTGGAATTCCTGGAAGAGTTAAAGAAAATAAACTATCAA GTGGACAGCTGGGAGGAAATGCTGAATAAGGCTGAGGTTGGTCATGGTTACATGGACCGCCCCTGCCTCAATCCGGCCGATCCAGACTGC CCCGCCACAGCCCCCAACAAAAATTCAACCAAACCTCTTGATATGGCCCTTGTTTTGAATGGTGGATGTCATGGCTTATCCAGAAAGTAT ATGCACTGGCAGGAGGAGTTGATTGTGGGTGGCACAGTCAAGAACAGCACTGGAAAACTCGTCAGCGCCCATGCCCTGCAGACCATGTTC CAGTTAATGACTCCCAAGCAAATGTACGAGCACTTCAAGGGGTACGAGTATGTCTCACACATCAACTGGAACGAGGACAAAGCGGCAGCC ATCCTGGAGGCCTGGCAGAGGACATATGTGGAGGTGGTTCATCAGAGTGTCGCACAGAACTCCACTCAAAAGGTGCTTTCCTTCACCACC ACGACCCTGGACGACATCCTGAAATCCTTCTCTGACGTCAGTGTCATCCGCGTGGCCAGCGGCTACTTACTCATGCTCGCCTATGCCTGT CTAACCATGCTGCGCTGGGACTGCTCCAAGTCCCAGGGTGCCGTGGGGCTGGCTGGCGTCCTGCTGGTTGCACTGTCAGTGGCTGCAGGA CTGGGCCTGTGCTCATTGATCGGAATTTCCTTTAACGCTGCAACAACTCAGGTTTTGCCATTTCTCGCTCTTGGTGTTGGTGTGGATGAT GTTTTTCTTCTGGCCCACGCCTTCAGTGAAACAGGACAGAATAAAAGAATCCCTTTTGAGGACAGGACCGGGGAGTGCCTGAAGCGCACA GGAGCCAGCGTGGCCCTCACGTCCATCAGCAATGTCACAGCCTTCTTCATGGCCGCGTTAATCCCAATTCCCGCTCTGCGGGCGTTCTCC CTCCAGGCAGCGGTAGTAGTGGTGTTCAATTTTGCCATGGTTCTGCTCATTTTTCCTGCAATTCTCAGCATGGATTTATATCGACGCGAG GACAGGAGACTGGATATTTTCTGCTGTTTTACAAGATAAGTATGCTGTGCGCTTCATCCCTCGGGAGAATGGCGTTTACCTGATTGACGT CAAGTTCAACGGCACCCACATCCCTGGAAGCCCCTTCAAGATCCGAGTTGGGGAGCCTGGGCATGGAGGGGACCCAGGCTTGGTGTCTGC TTACGGAGCAGGTCTGGAAGGCGGTGTCACAGGGAACCCAGCTGAGTTCGTCGTGAACACGAGCAATGCGGGAGCTGGTGCCCTGTCGGT GACCATTGACGGCCCCTCCAAGGTGAAGATGGATTGCCAGGAGTGCCCTGAGGGCTACCGCGTCACCTATACCCCCATGGCACCTGGCAG CTACCTCATCTCCATCAAGTACGGCGGCCCCTACCACATTGGGGGCAGCCCCTTCAAGGCCAAAGTCACAGGCCCCCGTCTCGTCAGCAA CCACAGCCTCCACGAGACATCATCAGTGTTTGTAGACTCTCTGACCAAGGCCACCTGTGCCCCCCAGCATGGGGCCCCGGGTCCTGGGCC TGCTGACGCCAGCAAGGTGGTGGCCAAGGGCCTGGGGCTGAGCAAGGCCTACGTAGGCCAGAAGAGCAGCTTCACAGTAGACTGCAGCAA AGCAGGCAACAACATGCTGCTGGTGGGGGTTCATGGCCCAAGGACCCCCTGCGAGGAGATCCTGGTGAAGCACGTGGGCAGCCGGCTCTA CAGCGTGTCCTACCTGCTCAAGGACAAGGGGGAGTACACACTGGTGGTCAAATGGGGGGACGAGCACATCCCAGGCAGCCCCTACCGCGT >69864_69864_6_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000429896_FLNA_chrX_153578575_ENST00000344736_length(amino acids)=540AA_BP= MWLRAKFQRLLFKLGCYIQKNCGKFLVVGLLIFGAFAVGLKAANLETNVEELWVEVGGRVSRELNYTRQKIGEEAMFNPQLMIQTPKEEG ANVLTTEALLQHLDSALQASRVHVYMYNRQWKLEHLCYKSGELITETGYMDQIIEYLYPCLIITPLDCFWEGAKLQSGTAYLLGKPPLRW TNFDPLEFLEELKKINYQVDSWEEMLNKAEVGHGYMDRPCLNPADPDCPATAPNKNSTKPLDMALVLNGGCHGLSRKYMHWQEELIVGGT VKNSTGKLVSAHALQTMFQLMTPKQMYEHFKGYEYVSHINWNEDKAAAILEAWQRTYVEVVHQSVAQNSTQKVLSFTTTTLDDILKSFSD VSVIRVASGYLLMLAYACLTMLRWDCSKSQGAVGLAGVLLVALSVAAGLGLCSLIGISFNAATTQVLPFLALGVGVDDVFLLAHAFSETG QNKRIPFEDRTGECLKRTGASVALTSISNVTAFFMAALIPIPALRAFSLQAAVVVVFNFAMVLLIFPAILSMDLYRREDRRLDIFCCFTR -------------------------------------------------------------- >69864_69864_7_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000429896_FLNA_chrX_153578575_ENST00000360319_length(transcript)=2946nt_BP=1835nt GAAGGCGAGCACCCAGACGGGGGCCCGCCGGGGCCGCGGCCAGCGCCGGGGAAATGCCGCGCCGGGGAGCAGCCTGCGCCGGCCTGAGCC CTTCCCTTTGCACTCGGCTGTTTTTTACGTTTAACCAGAAAGGAAGGGAGAGGAGGGAAAGATCCATGTGGCTGCCCTCTTCCGATCACA AATATTGTCGGGAAGGCTACTGGCCGGAAAGCGCCGCTGTGGCTGAGAGCGAAGTTTCAGAGACTCTTATTTAAACTGGGTTGTTACATT CAAAAAAACTGCGGCAAGTTCTTGGTTGTGGGCCTCCTCATATTTGGGGCCTTCGCGGTGGGATTAAAAGCAGCGAACCTCGAGACCAAC GTGGAGGAGCTGTGGGTGGAAGTTGGAGGACGAGTAAGTCGTGAATTAAATTATACTCGCCAGAAGATTGGAGAAGAGGCTATGTTTAAT CCTCAACTCATGATACAGACCCCTAAAGAAGAAGGTGCTAATGTCCTGACCACAGAAGCGCTCCTACAACACCTGGACTCGGCACTCCAG GCCAGCCGTGTCCATGTATACATGTACAACAGGCAGTGGAAATTGGAACATTTGTGTTACAAATCAGGAGAGCTTATCACAGAAACAGGT TACATGGATCAGATAATAGAATATCTTTACCCTTGTTTGATTATTACACCTTTGGACTGCTTCTGGGAAGGGGCGAAATTACAGTCTGGG ACAGCATACCTCCTAGGTAAACCTCCTTTGCGGTGGACAAACTTCGACCCTTTGGAATTCCTGGAAGAGTTAAAGAAAATAAACTATCAA GTGGACAGCTGGGAGGAAATGCTGAATAAGGCTGAGGTTGGTCATGGTTACATGGACCGCCCCTGCCTCAATCCGGCCGATCCAGACTGC CCCGCCACAGCCCCCAACAAAAATTCAACCAAACCTCTTGATATGGCCCTTGTTTTGAATGGTGGATGTCATGGCTTATCCAGAAAGTAT ATGCACTGGCAGGAGGAGTTGATTGTGGGTGGCACAGTCAAGAACAGCACTGGAAAACTCGTCAGCGCCCATGCCCTGCAGACCATGTTC CAGTTAATGACTCCCAAGCAAATGTACGAGCACTTCAAGGGGTACGAGTATGTCTCACACATCAACTGGAACGAGGACAAAGCGGCAGCC ATCCTGGAGGCCTGGCAGAGGACATATGTGGAGGTGGTTCATCAGAGTGTCGCACAGAACTCCACTCAAAAGGTGCTTTCCTTCACCACC ACGACCCTGGACGACATCCTGAAATCCTTCTCTGACGTCAGTGTCATCCGCGTGGCCAGCGGCTACTTACTCATGCTCGCCTATGCCTGT CTAACCATGCTGCGCTGGGACTGCTCCAAGTCCCAGGGTGCCGTGGGGCTGGCTGGCGTCCTGCTGGTTGCACTGTCAGTGGCTGCAGGA CTGGGCCTGTGCTCATTGATCGGAATTTCCTTTAACGCTGCAACAACTCAGGTTTTGCCATTTCTCGCTCTTGGTGTTGGTGTGGATGAT GTTTTTCTTCTGGCCCACGCCTTCAGTGAAACAGGACAGAATAAAAGAATCCCTTTTGAGGACAGGACCGGGGAGTGCCTGAAGCGCACA GGAGCCAGCGTGGCCCTCACGTCCATCAGCAATGTCACAGCCTTCTTCATGGCCGCGTTAATCCCAATTCCCGCTCTGCGGGCGTTCTCC CTCCAGGCAGCGGTAGTAGTGGTGTTCAATTTTGCCATGGTTCTGCTCATTTTTCCTGCAATTCTCAGCATGGATTTATATCGACGCGAG GACAGGAGACTGGATATTTTCTGCTGTTTTACAAGATAAGTATGCTGTGCGCTTCATCCCTCGGGAGAATGGCGTTTACCTGATTGACGT CAAGTTCAACGGCACCCACATCCCTGGAAGCCCCTTCAAGATCCGAGTTGGGGAGCCTGGGCATGGAGGGGACCCAGGCTTGGTGTCTGC TTACGGAGCAGGTCTGGAAGGCGGTGTCACAGGGAACCCAGCTGAGTTCGTCGTGAACACGAGCAATGCGGGAGCTGGTGCCCTGTCGGT GACCATTGACGGCCCCTCCAAGGTGAAGATGGATTGCCAGGAGTGCCCTGAGGGCTACCGCGTCACCTATACCCCCATGGCACCTGGCAG CTACCTCATCTCCATCAAGTACGGCGGCCCCTACCACATTGGGGGCAGCCCCTTCAAGGCCAAAGTCACAGGCCCCCGTCTCGTCAGCAA CCACAGCCTCCACGAGACATCATCAGTGTTTGTAGACTCTCTGACCAAGGCCACCTGTGCCCCCCAGCATGGGGCCCCGGGTCCTGGGCC TGCTGACGCCAGCAAGGTGGTGGCCAAGGGCCTGGGGCTGAGCAAGGCCTACGTAGGCCAGAAGAGCAGCTTCACAGTAGACTGCAGCAA AGCAGGCAACAACATGCTGCTGGTGGGGGTTCATGGCCCAAGGACCCCCTGCGAGGAGATCCTGGTGAAGCACGTGGGCAGCCGGCTCTA CAGCGTGTCCTACCTGCTCAAGGACAAGGGGGAGTACACACTGGTGGTCAAATGGGGGGACGAGCACATCCCAGGCAGCCCCTACCGCGT TGTGGTGCCCTGAGTCTGGGGCCCGTGCCAGCCGGCAGCCCCCAAGCCTGCCCCGCTACCCAAGCAGCCCCGCCCTCTTCCCCTCAACCC CGGCCCAGGCCGCCCTGGCCGCCCGCCTGTCACTGCAGCCGCCCCTGCCCTGTGCCGTGCTGCGCTCACCTGCCTCCCCAGCCAGCCGCT GACCTCTCGGCTTTCACTTGGGCAGAGGGAGCCATTTGGTGGCGCTGCTTGTCTTCTTTGGTTCTGGGAGGGGTGAGGGATGGGGGTCCT >69864_69864_7_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000429896_FLNA_chrX_153578575_ENST00000360319_length(amino acids)=540AA_BP= MWLRAKFQRLLFKLGCYIQKNCGKFLVVGLLIFGAFAVGLKAANLETNVEELWVEVGGRVSRELNYTRQKIGEEAMFNPQLMIQTPKEEG ANVLTTEALLQHLDSALQASRVHVYMYNRQWKLEHLCYKSGELITETGYMDQIIEYLYPCLIITPLDCFWEGAKLQSGTAYLLGKPPLRW TNFDPLEFLEELKKINYQVDSWEEMLNKAEVGHGYMDRPCLNPADPDCPATAPNKNSTKPLDMALVLNGGCHGLSRKYMHWQEELIVGGT VKNSTGKLVSAHALQTMFQLMTPKQMYEHFKGYEYVSHINWNEDKAAAILEAWQRTYVEVVHQSVAQNSTQKVLSFTTTTLDDILKSFSD VSVIRVASGYLLMLAYACLTMLRWDCSKSQGAVGLAGVLLVALSVAAGLGLCSLIGISFNAATTQVLPFLALGVGVDDVFLLAHAFSETG QNKRIPFEDRTGECLKRTGASVALTSISNVTAFFMAALIPIPALRAFSLQAAVVVVFNFAMVLLIFPAILSMDLYRREDRRLDIFCCFTR -------------------------------------------------------------- >69864_69864_8_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000429896_FLNA_chrX_153578575_ENST00000369850_length(transcript)=2824nt_BP=1835nt GAAGGCGAGCACCCAGACGGGGGCCCGCCGGGGCCGCGGCCAGCGCCGGGGAAATGCCGCGCCGGGGAGCAGCCTGCGCCGGCCTGAGCC CTTCCCTTTGCACTCGGCTGTTTTTTACGTTTAACCAGAAAGGAAGGGAGAGGAGGGAAAGATCCATGTGGCTGCCCTCTTCCGATCACA AATATTGTCGGGAAGGCTACTGGCCGGAAAGCGCCGCTGTGGCTGAGAGCGAAGTTTCAGAGACTCTTATTTAAACTGGGTTGTTACATT CAAAAAAACTGCGGCAAGTTCTTGGTTGTGGGCCTCCTCATATTTGGGGCCTTCGCGGTGGGATTAAAAGCAGCGAACCTCGAGACCAAC GTGGAGGAGCTGTGGGTGGAAGTTGGAGGACGAGTAAGTCGTGAATTAAATTATACTCGCCAGAAGATTGGAGAAGAGGCTATGTTTAAT CCTCAACTCATGATACAGACCCCTAAAGAAGAAGGTGCTAATGTCCTGACCACAGAAGCGCTCCTACAACACCTGGACTCGGCACTCCAG GCCAGCCGTGTCCATGTATACATGTACAACAGGCAGTGGAAATTGGAACATTTGTGTTACAAATCAGGAGAGCTTATCACAGAAACAGGT TACATGGATCAGATAATAGAATATCTTTACCCTTGTTTGATTATTACACCTTTGGACTGCTTCTGGGAAGGGGCGAAATTACAGTCTGGG ACAGCATACCTCCTAGGTAAACCTCCTTTGCGGTGGACAAACTTCGACCCTTTGGAATTCCTGGAAGAGTTAAAGAAAATAAACTATCAA GTGGACAGCTGGGAGGAAATGCTGAATAAGGCTGAGGTTGGTCATGGTTACATGGACCGCCCCTGCCTCAATCCGGCCGATCCAGACTGC CCCGCCACAGCCCCCAACAAAAATTCAACCAAACCTCTTGATATGGCCCTTGTTTTGAATGGTGGATGTCATGGCTTATCCAGAAAGTAT ATGCACTGGCAGGAGGAGTTGATTGTGGGTGGCACAGTCAAGAACAGCACTGGAAAACTCGTCAGCGCCCATGCCCTGCAGACCATGTTC CAGTTAATGACTCCCAAGCAAATGTACGAGCACTTCAAGGGGTACGAGTATGTCTCACACATCAACTGGAACGAGGACAAAGCGGCAGCC ATCCTGGAGGCCTGGCAGAGGACATATGTGGAGGTGGTTCATCAGAGTGTCGCACAGAACTCCACTCAAAAGGTGCTTTCCTTCACCACC ACGACCCTGGACGACATCCTGAAATCCTTCTCTGACGTCAGTGTCATCCGCGTGGCCAGCGGCTACTTACTCATGCTCGCCTATGCCTGT CTAACCATGCTGCGCTGGGACTGCTCCAAGTCCCAGGGTGCCGTGGGGCTGGCTGGCGTCCTGCTGGTTGCACTGTCAGTGGCTGCAGGA CTGGGCCTGTGCTCATTGATCGGAATTTCCTTTAACGCTGCAACAACTCAGGTTTTGCCATTTCTCGCTCTTGGTGTTGGTGTGGATGAT GTTTTTCTTCTGGCCCACGCCTTCAGTGAAACAGGACAGAATAAAAGAATCCCTTTTGAGGACAGGACCGGGGAGTGCCTGAAGCGCACA GGAGCCAGCGTGGCCCTCACGTCCATCAGCAATGTCACAGCCTTCTTCATGGCCGCGTTAATCCCAATTCCCGCTCTGCGGGCGTTCTCC CTCCAGGCAGCGGTAGTAGTGGTGTTCAATTTTGCCATGGTTCTGCTCATTTTTCCTGCAATTCTCAGCATGGATTTATATCGACGCGAG GACAGGAGACTGGATATTTTCTGCTGTTTTACAAGATAAGTATGCTGTGCGCTTCATCCCTCGGGAGAATGGCGTTTACCTGATTGACGT CAAGTTCAACGGCACCCACATCCCTGGAAGCCCCTTCAAGATCCGAGTTGGGGAGCCTGGGCATGGAGGGGACCCAGGCTTGGTGTCTGC TTACGGAGCAGGTCTGGAAGGCGGTGTCACAGGGAACCCAGCTGAGTTCGTCGTGAACACGAGCAATGCGGGAGCTGGTGCCCTGTCGGT GACCATTGACGGCCCCTCCAAGGTGAAGATGGATTGCCAGGAGTGCCCTGAGGGCTACCGCGTCACCTATACCCCCATGGCACCTGGCAG CTACCTCATCTCCATCAAGTACGGCGGCCCCTACCACATTGGGGGCAGCCCCTTCAAGGCCAAAGTCACAGGCCCCCGTCTCGTCAGCAA CCACAGCCTCCACGAGACATCATCAGTGTTTGTAGACTCTCTGACCAAGGCCACCTGTGCCCCCCAGCATGGGGCCCCGGGTCCTGGGCC TGCTGACGCCAGCAAGGTGGTGGCCAAGGGCCTGGGGCTGAGCAAGGCCTACGTAGGCCAGAAGAGCAGCTTCACAGTAGACTGCAGCAA AGCAGGCAACAACATGCTGCTGGTGGGGGTTCATGGCCCAAGGACCCCCTGCGAGGAGATCCTGGTGAAGCACGTGGGCAGCCGGCTCTA CAGCGTGTCCTACCTGCTCAAGGACAAGGGGGAGTACACACTGGTGGTCAAATGGGGGGACGAGCACATCCCAGGCAGCCCCTACCGCGT TGTGGTGCCCTGAGTCTGGGGCCCGTGCCAGCCGGCAGCCCCCAAGCCTGCCCCGCTACCCAAGCAGCCCCGCCCTCTTCCCCTCAACCC CGGCCCAGGCCGCCCTGGCCGCCCGCCTGTCACTGCAGCCGCCCCTGCCCTGTGCCGTGCTGCGCTCACCTGCCTCCCCAGCCAGCCGCT >69864_69864_8_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000429896_FLNA_chrX_153578575_ENST00000369850_length(amino acids)=540AA_BP= MWLRAKFQRLLFKLGCYIQKNCGKFLVVGLLIFGAFAVGLKAANLETNVEELWVEVGGRVSRELNYTRQKIGEEAMFNPQLMIQTPKEEG ANVLTTEALLQHLDSALQASRVHVYMYNRQWKLEHLCYKSGELITETGYMDQIIEYLYPCLIITPLDCFWEGAKLQSGTAYLLGKPPLRW TNFDPLEFLEELKKINYQVDSWEEMLNKAEVGHGYMDRPCLNPADPDCPATAPNKNSTKPLDMALVLNGGCHGLSRKYMHWQEELIVGGT VKNSTGKLVSAHALQTMFQLMTPKQMYEHFKGYEYVSHINWNEDKAAAILEAWQRTYVEVVHQSVAQNSTQKVLSFTTTTLDDILKSFSD VSVIRVASGYLLMLAYACLTMLRWDCSKSQGAVGLAGVLLVALSVAAGLGLCSLIGISFNAATTQVLPFLALGVGVDDVFLLAHAFSETG QNKRIPFEDRTGECLKRTGASVALTSISNVTAFFMAALIPIPALRAFSLQAAVVVVFNFAMVLLIFPAILSMDLYRREDRRLDIFCCFTR -------------------------------------------------------------- >69864_69864_9_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000429896_FLNA_chrX_153578575_ENST00000369856_length(transcript)=2816nt_BP=1835nt GAAGGCGAGCACCCAGACGGGGGCCCGCCGGGGCCGCGGCCAGCGCCGGGGAAATGCCGCGCCGGGGAGCAGCCTGCGCCGGCCTGAGCC CTTCCCTTTGCACTCGGCTGTTTTTTACGTTTAACCAGAAAGGAAGGGAGAGGAGGGAAAGATCCATGTGGCTGCCCTCTTCCGATCACA AATATTGTCGGGAAGGCTACTGGCCGGAAAGCGCCGCTGTGGCTGAGAGCGAAGTTTCAGAGACTCTTATTTAAACTGGGTTGTTACATT CAAAAAAACTGCGGCAAGTTCTTGGTTGTGGGCCTCCTCATATTTGGGGCCTTCGCGGTGGGATTAAAAGCAGCGAACCTCGAGACCAAC GTGGAGGAGCTGTGGGTGGAAGTTGGAGGACGAGTAAGTCGTGAATTAAATTATACTCGCCAGAAGATTGGAGAAGAGGCTATGTTTAAT CCTCAACTCATGATACAGACCCCTAAAGAAGAAGGTGCTAATGTCCTGACCACAGAAGCGCTCCTACAACACCTGGACTCGGCACTCCAG GCCAGCCGTGTCCATGTATACATGTACAACAGGCAGTGGAAATTGGAACATTTGTGTTACAAATCAGGAGAGCTTATCACAGAAACAGGT TACATGGATCAGATAATAGAATATCTTTACCCTTGTTTGATTATTACACCTTTGGACTGCTTCTGGGAAGGGGCGAAATTACAGTCTGGG ACAGCATACCTCCTAGGTAAACCTCCTTTGCGGTGGACAAACTTCGACCCTTTGGAATTCCTGGAAGAGTTAAAGAAAATAAACTATCAA GTGGACAGCTGGGAGGAAATGCTGAATAAGGCTGAGGTTGGTCATGGTTACATGGACCGCCCCTGCCTCAATCCGGCCGATCCAGACTGC CCCGCCACAGCCCCCAACAAAAATTCAACCAAACCTCTTGATATGGCCCTTGTTTTGAATGGTGGATGTCATGGCTTATCCAGAAAGTAT ATGCACTGGCAGGAGGAGTTGATTGTGGGTGGCACAGTCAAGAACAGCACTGGAAAACTCGTCAGCGCCCATGCCCTGCAGACCATGTTC CAGTTAATGACTCCCAAGCAAATGTACGAGCACTTCAAGGGGTACGAGTATGTCTCACACATCAACTGGAACGAGGACAAAGCGGCAGCC ATCCTGGAGGCCTGGCAGAGGACATATGTGGAGGTGGTTCATCAGAGTGTCGCACAGAACTCCACTCAAAAGGTGCTTTCCTTCACCACC ACGACCCTGGACGACATCCTGAAATCCTTCTCTGACGTCAGTGTCATCCGCGTGGCCAGCGGCTACTTACTCATGCTCGCCTATGCCTGT CTAACCATGCTGCGCTGGGACTGCTCCAAGTCCCAGGGTGCCGTGGGGCTGGCTGGCGTCCTGCTGGTTGCACTGTCAGTGGCTGCAGGA CTGGGCCTGTGCTCATTGATCGGAATTTCCTTTAACGCTGCAACAACTCAGGTTTTGCCATTTCTCGCTCTTGGTGTTGGTGTGGATGAT GTTTTTCTTCTGGCCCACGCCTTCAGTGAAACAGGACAGAATAAAAGAATCCCTTTTGAGGACAGGACCGGGGAGTGCCTGAAGCGCACA GGAGCCAGCGTGGCCCTCACGTCCATCAGCAATGTCACAGCCTTCTTCATGGCCGCGTTAATCCCAATTCCCGCTCTGCGGGCGTTCTCC CTCCAGGCAGCGGTAGTAGTGGTGTTCAATTTTGCCATGGTTCTGCTCATTTTTCCTGCAATTCTCAGCATGGATTTATATCGACGCGAG GACAGGAGACTGGATATTTTCTGCTGTTTTACAAGATAAGTATGCTGTGCGCTTCATCCCTCGGGAGAATGGCGTTTACCTGATTGACGT CAAGTTCAACGGCACCCACATCCCTGGAAGCCCCTTCAAGATCCGAGTTGGGGAGCCTGGGCATGGAGGGGACCCAGGCTTGGTGTCTGC TTACGGAGCAGGTCTGGAAGGCGGTGTCACAGGGAACCCAGCTGAGTTCGTCGTGAACACGAGCAATGCGGGAGCTGGTGCCCTGTCGGT GACCATTGACGGCCCCTCCAAGGTGAAGATGGATTGCCAGGAGTGCCCTGAGGGCTACCGCGTCACCTATACCCCCATGGCACCTGGCAG CTACCTCATCTCCATCAAGTACGGCGGCCCCTACCACATTGGGGGCAGCCCCTTCAAGGCCAAAGTCACAGGCCCCCGTCTCGTCAGCAA CCACAGCCTCCACGAGACATCATCAGTGTTTGTAGACTCTCTGACCAAGGCCACCTGTGCCCCCCAGCATGGGGCCCCGGGTCCTGGGCC TGCTGACGCCAGCAAGGTGGTGGCCAAGGGCCTGGGGCTGAGCAAGGCCTACGTAGGCCAGAAGAGCAGCTTCACAGTAGACTGCAGCAA AGCAGGCAACAACATGCTGCTGGTGGGGGTTCATGGCCCAAGGACCCCCTGCGAGGAGATCCTGGTGAAGCACGTGGGCAGCCGGCTCTA CAGCGTGTCCTACCTGCTCAAGGACAAGGGGGAGTACACACTGGTGGTCAAATGGGGGGACGAGCACATCCCAGGCAGCCCCTACCGCGT TGTGGTGCCCTGAGTCTGGGGCCCGTGCCAGCCGGCAGCCCCCAAGCCTGCCCCGCTACCCAAGCAGCCCCGCCCTCTTCCCCTCAACCC CGGCCCAGGCCGCCCTGGCCGCCCGCCTGTCACTGCAGCCGCCCCTGCCCTGTGCCGTGCTGCGCTCACCTGCCTCCCCAGCCAGCCGCT >69864_69864_9_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000429896_FLNA_chrX_153578575_ENST00000369856_length(amino acids)=540AA_BP= MWLRAKFQRLLFKLGCYIQKNCGKFLVVGLLIFGAFAVGLKAANLETNVEELWVEVGGRVSRELNYTRQKIGEEAMFNPQLMIQTPKEEG ANVLTTEALLQHLDSALQASRVHVYMYNRQWKLEHLCYKSGELITETGYMDQIIEYLYPCLIITPLDCFWEGAKLQSGTAYLLGKPPLRW TNFDPLEFLEELKKINYQVDSWEEMLNKAEVGHGYMDRPCLNPADPDCPATAPNKNSTKPLDMALVLNGGCHGLSRKYMHWQEELIVGGT VKNSTGKLVSAHALQTMFQLMTPKQMYEHFKGYEYVSHINWNEDKAAAILEAWQRTYVEVVHQSVAQNSTQKVLSFTTTTLDDILKSFSD VSVIRVASGYLLMLAYACLTMLRWDCSKSQGAVGLAGVLLVALSVAAGLGLCSLIGISFNAATTQVLPFLALGVGVDDVFLLAHAFSETG QNKRIPFEDRTGECLKRTGASVALTSISNVTAFFMAALIPIPALRAFSLQAAVVVVFNFAMVLLIFPAILSMDLYRREDRRLDIFCCFTR -------------------------------------------------------------- >69864_69864_10_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000429896_FLNA_chrX_153578575_ENST00000422373_length(transcript)=2940nt_BP=1835nt GAAGGCGAGCACCCAGACGGGGGCCCGCCGGGGCCGCGGCCAGCGCCGGGGAAATGCCGCGCCGGGGAGCAGCCTGCGCCGGCCTGAGCC CTTCCCTTTGCACTCGGCTGTTTTTTACGTTTAACCAGAAAGGAAGGGAGAGGAGGGAAAGATCCATGTGGCTGCCCTCTTCCGATCACA AATATTGTCGGGAAGGCTACTGGCCGGAAAGCGCCGCTGTGGCTGAGAGCGAAGTTTCAGAGACTCTTATTTAAACTGGGTTGTTACATT CAAAAAAACTGCGGCAAGTTCTTGGTTGTGGGCCTCCTCATATTTGGGGCCTTCGCGGTGGGATTAAAAGCAGCGAACCTCGAGACCAAC GTGGAGGAGCTGTGGGTGGAAGTTGGAGGACGAGTAAGTCGTGAATTAAATTATACTCGCCAGAAGATTGGAGAAGAGGCTATGTTTAAT CCTCAACTCATGATACAGACCCCTAAAGAAGAAGGTGCTAATGTCCTGACCACAGAAGCGCTCCTACAACACCTGGACTCGGCACTCCAG GCCAGCCGTGTCCATGTATACATGTACAACAGGCAGTGGAAATTGGAACATTTGTGTTACAAATCAGGAGAGCTTATCACAGAAACAGGT TACATGGATCAGATAATAGAATATCTTTACCCTTGTTTGATTATTACACCTTTGGACTGCTTCTGGGAAGGGGCGAAATTACAGTCTGGG ACAGCATACCTCCTAGGTAAACCTCCTTTGCGGTGGACAAACTTCGACCCTTTGGAATTCCTGGAAGAGTTAAAGAAAATAAACTATCAA GTGGACAGCTGGGAGGAAATGCTGAATAAGGCTGAGGTTGGTCATGGTTACATGGACCGCCCCTGCCTCAATCCGGCCGATCCAGACTGC CCCGCCACAGCCCCCAACAAAAATTCAACCAAACCTCTTGATATGGCCCTTGTTTTGAATGGTGGATGTCATGGCTTATCCAGAAAGTAT ATGCACTGGCAGGAGGAGTTGATTGTGGGTGGCACAGTCAAGAACAGCACTGGAAAACTCGTCAGCGCCCATGCCCTGCAGACCATGTTC CAGTTAATGACTCCCAAGCAAATGTACGAGCACTTCAAGGGGTACGAGTATGTCTCACACATCAACTGGAACGAGGACAAAGCGGCAGCC ATCCTGGAGGCCTGGCAGAGGACATATGTGGAGGTGGTTCATCAGAGTGTCGCACAGAACTCCACTCAAAAGGTGCTTTCCTTCACCACC ACGACCCTGGACGACATCCTGAAATCCTTCTCTGACGTCAGTGTCATCCGCGTGGCCAGCGGCTACTTACTCATGCTCGCCTATGCCTGT CTAACCATGCTGCGCTGGGACTGCTCCAAGTCCCAGGGTGCCGTGGGGCTGGCTGGCGTCCTGCTGGTTGCACTGTCAGTGGCTGCAGGA CTGGGCCTGTGCTCATTGATCGGAATTTCCTTTAACGCTGCAACAACTCAGGTTTTGCCATTTCTCGCTCTTGGTGTTGGTGTGGATGAT GTTTTTCTTCTGGCCCACGCCTTCAGTGAAACAGGACAGAATAAAAGAATCCCTTTTGAGGACAGGACCGGGGAGTGCCTGAAGCGCACA GGAGCCAGCGTGGCCCTCACGTCCATCAGCAATGTCACAGCCTTCTTCATGGCCGCGTTAATCCCAATTCCCGCTCTGCGGGCGTTCTCC CTCCAGGCAGCGGTAGTAGTGGTGTTCAATTTTGCCATGGTTCTGCTCATTTTTCCTGCAATTCTCAGCATGGATTTATATCGACGCGAG GACAGGAGACTGGATATTTTCTGCTGTTTTACAAGATAAGTATGCTGTGCGCTTCATCCCTCGGGAGAATGGCGTTTACCTGATTGACGT CAAGTTCAACGGCACCCACATCCCTGGAAGCCCCTTCAAGATCCGAGTTGGGGAGCCTGGGCATGGAGGGGACCCAGGCTTGGTGTCTGC TTACGGAGCAGGTCTGGAAGGCGGTGTCACAGGGAACCCAGCTGAGTTCGTCGTGAACACGAGCAATGCGGGAGCTGGTGCCCTGTCGGT GACCATTGACGGCCCCTCCAAGGTGAAGATGGATTGCCAGGAGTGCCCTGAGGGCTACCGCGTCACCTATACCCCCATGGCACCTGGCAG CTACCTCATCTCCATCAAGTACGGCGGCCCCTACCACATTGGGGGCAGCCCCTTCAAGGCCAAAGTCACAGGCCCCCGTCTCGTCAGCAA CCACAGCCTCCACGAGACATCATCAGTGTTTGTAGACTCTCTGACCAAGGCCACCTGTGCCCCCCAGCATGGGGCCCCGGGTCCTGGGCC TGCTGACGCCAGCAAGGTGGTGGCCAAGGGCCTGGGGCTGAGCAAGGCCTACGTAGGCCAGAAGAGCAGCTTCACAGTAGACTGCAGCAA AGCAGGCAACAACATGCTGCTGGTGGGGGTTCATGGCCCAAGGACCCCCTGCGAGGAGATCCTGGTGAAGCACGTGGGCAGCCGGCTCTA CAGCGTGTCCTACCTGCTCAAGGACAAGGGGGAGTACACACTGGTGGTCAAATGGGGGGACGAGCACATCCCAGGCAGCCCCTACCGCGT TGTGGTGCCCTGAGTCTGGGGCCCGTGCCAGCCGGCAGCCCCCAAGCCTGCCCCGCTACCCAAGCAGCCCCGCCCTCTTCCCCTCAACCC CGGCCCAGGCCGCCCTGGCCGCCCGCCTGTCACTGCAGCCGCCCCTGCCCTGTGCCGTGCTGCGCTCACCTGCCTCCCCAGCCAGCCGCT GACCTCTCGGCTTTCACTTGGGCAGAGGGAGCCATTTGGTGGCGCTGCTTGTCTTCTTTGGTTCTGGGAGGGGTGAGGGATGGGGGTCCT >69864_69864_10_PTCH1-FLNA_PTCH1_chr9_98232094_ENST00000429896_FLNA_chrX_153578575_ENST00000422373_length(amino acids)=540AA_BP= MWLRAKFQRLLFKLGCYIQKNCGKFLVVGLLIFGAFAVGLKAANLETNVEELWVEVGGRVSRELNYTRQKIGEEAMFNPQLMIQTPKEEG ANVLTTEALLQHLDSALQASRVHVYMYNRQWKLEHLCYKSGELITETGYMDQIIEYLYPCLIITPLDCFWEGAKLQSGTAYLLGKPPLRW TNFDPLEFLEELKKINYQVDSWEEMLNKAEVGHGYMDRPCLNPADPDCPATAPNKNSTKPLDMALVLNGGCHGLSRKYMHWQEELIVGGT VKNSTGKLVSAHALQTMFQLMTPKQMYEHFKGYEYVSHINWNEDKAAAILEAWQRTYVEVVHQSVAQNSTQKVLSFTTTTLDDILKSFSD VSVIRVASGYLLMLAYACLTMLRWDCSKSQGAVGLAGVLLVALSVAAGLGLCSLIGISFNAATTQVLPFLALGVGVDDVFLLAHAFSETG QNKRIPFEDRTGECLKRTGASVALTSISNVTAFFMAALIPIPALRAFSLQAAVVVVFNFAMVLLIFPAILSMDLYRREDRRLDIFCCFTR -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for PTCH1-FLNA |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000360319 | 41 | 46 | 1490_1607 | 2377.3333333333335 | 2640.0 | furin | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000369850 | 43 | 48 | 1490_1607 | 2385.3333333333335 | 2648.0 | furin | |
| Tgene | FLNA | chr9:98232094 | chrX:153578575 | ENST00000422373 | 42 | 47 | 1490_1607 | 2377.3333333333335 | 2640.0 | furin |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for PTCH1-FLNA |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for PTCH1-FLNA |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
| Hgene | PTCH1 | C0004779 | Basal Cell Nevus Syndrome | 23 | CLINGEN;CTD_human;GENOMICS_ENGLAND;ORPHANET;UNIPROT |
| Hgene | PTCH1 | C1835820 | HOLOPROSENCEPHALY 7 | 5 | CTD_human;GENOMICS_ENGLAND;UNIPROT |
| Hgene | PTCH1 | C0025149 | Medulloblastoma | 3 | CGI;CTD_human |
| Hgene | PTCH1 | C0205833 | Medullomyoblastoma | 3 | CTD_human |
| Hgene | PTCH1 | C0278510 | Childhood Medulloblastoma | 3 | CTD_human |
| Hgene | PTCH1 | C0278876 | Adult Medulloblastoma | 3 | CTD_human |
| Hgene | PTCH1 | C0751291 | Desmoplastic Medulloblastoma | 3 | CTD_human |
| Hgene | PTCH1 | C1275668 | Melanotic medulloblastoma | 3 | CTD_human |
| Hgene | PTCH1 | C1368275 | Pigmented Basal Cell Carcinoma | 3 | CTD_human |
| Hgene | PTCH1 | C4721806 | Carcinoma, Basal Cell | 3 | CTD_human |
| Hgene | PTCH1 | C0376634 | Craniofacial Abnormalities | 2 | CTD_human |
| Hgene | PTCH1 | C2751544 | BASAL CELL CARCINOMA, SUSCEPTIBILITY TO, 1 | 2 | GENOMICS_ENGLAND;UNIPROT |
| Hgene | PTCH1 | C0006118 | Brain Neoplasms | 1 | CTD_human |
| Hgene | PTCH1 | C0007114 | Malignant neoplasm of skin | 1 | CTD_human |
| Hgene | PTCH1 | C0008924 | Cleft upper lip | 1 | CTD_human |
| Hgene | PTCH1 | C0008925 | Cleft Palate | 1 | CTD_human |
| Hgene | PTCH1 | C0019693 | HIV Infections | 1 | CTD_human |
| Hgene | PTCH1 | C0030297 | Pancreatic Neoplasm | 1 | CTD_human |
| Hgene | PTCH1 | C0035412 | Rhabdomyosarcoma | 1 | CTD_human |
| Hgene | PTCH1 | C0037286 | Skin Neoplasms | 1 | CTD_human |
| Hgene | PTCH1 | C0153633 | Malignant neoplasm of brain | 1 | CTD_human |
| Hgene | PTCH1 | C0206663 | Neuroectodermal Tumor, Primitive | 1 | CTD_human |
| Hgene | PTCH1 | C0238198 | Gastrointestinal Stromal Tumors | 1 | CTD_human |
| Hgene | PTCH1 | C0334584 | Spongioblastoma | 1 | CTD_human |
| Hgene | PTCH1 | C0334596 | Medulloepithelioma | 1 | CTD_human |
| Hgene | PTCH1 | C0346647 | Malignant neoplasm of pancreas | 1 | CTD_human |
| Hgene | PTCH1 | C0496899 | Benign neoplasm of brain, unspecified | 1 | CTD_human |
| Hgene | PTCH1 | C0700367 | Ependymoblastoma | 1 | CTD_human |
| Hgene | PTCH1 | C0750974 | Brain Tumor, Primary | 1 | CTD_human |
| Hgene | PTCH1 | C0750977 | Recurrent Brain Neoplasm | 1 | CTD_human |
| Hgene | PTCH1 | C0750979 | Primary malignant neoplasm of brain | 1 | CTD_human |
| Hgene | PTCH1 | C0751675 | Cerebral Primitive Neuroectodermal Tumor | 1 | CTD_human |
| Hgene | PTCH1 | C0812437 | Oculo-dento-digital syndrome | 1 | ORPHANET |
| Hgene | PTCH1 | C1527390 | Neoplasms, Intracranial | 1 | CTD_human |
| Hgene | PTCH1 | C1834038 | Schilbach-Rott Syndrome | 1 | ORPHANET |
| Hgene | PTCH1 | C1837218 | Cleft palate, isolated | 1 | CTD_human |
| Hgene | PTCH1 | C3151617 | ANTERIOR SEGMENT DYSGENESIS 7 | 1 | GENOMICS_ENGLAND |
| Hgene | PTCH1 | C3179349 | Gastrointestinal Stromal Sarcoma | 1 | CTD_human |
| Hgene | PTCH1 | C3711390 | 9q22.3 Microdeletion | 1 | ORPHANET |
| Hgene | PTCH1 | C4505456 | HIV Coinfection | 1 | CTD_human |
| Tgene | C1848213 | Periventricular Heterotopia, X-Linked | 9 | CTD_human;GENOMICS_ENGLAND;UNIPROT | |
| Tgene | C0025237 | Melnick-Needles Syndrome | 7 | CTD_human;GENOMICS_ENGLAND;ORPHANET;UNIPROT | |
| Tgene | C4707243 | Familial thoracic aortic aneurysm and aortic dissection | 7 | CLINGEN;GENOMICS_ENGLAND | |
| Tgene | C0265251 | Oto-Palato-digital syndrome type 1 | 6 | CTD_human;GENOMICS_ENGLAND;UNIPROT | |
| Tgene | C1844696 | OTOPALATODIGITAL SYNDROME, TYPE II | 6 | CTD_human;GENOMICS_ENGLAND;UNIPROT | |
| Tgene | C0265293 | Frontometaphyseal dysplasia | 5 | CTD_human;GENOMICS_ENGLAND;ORPHANET | |
| Tgene | C1846129 | Terminal Osseous Dysplasia and Pigmentary Defects | 4 | CTD_human;GENOMICS_ENGLAND;ORPHANET | |
| Tgene | C4281559 | FRONTOMETAPHYSEAL DYSPLASIA 1 | 4 | GENOMICS_ENGLAND;UNIPROT | |
| Tgene | C0262436 | Cardiac valvular dysplasia, X-linked | 2 | CTD_human;GENOMICS_ENGLAND;UNIPROT | |
| Tgene | C0543888 | Epileptic encephalopathy | 2 | GENOMICS_ENGLAND | |
| Tgene | C1845902 | FG SYNDROME 2 | 2 | CTD_human;GENOMICS_ENGLAND;UNIPROT | |
| Tgene | C1868720 | Periventricular Nodular Heterotopia | 2 | CTD_human | |
| Tgene | C2746068 | Congenital idiopathic intestinal pseudoobstruction | 2 | CTD_human;GENOMICS_ENGLAND | |
| Tgene | C4551969 | Bilateral Periventricular Nodular Heterotopia | 2 | CTD_human | |
| Tgene | C0006142 | Malignant neoplasm of breast | 1 | CTD_human | |
| Tgene | C0010701 | Phyllodes Tumor | 1 | CTD_human | |
| Tgene | C0013366 | Dyschondroplasias | 1 | CTD_human | |
| Tgene | C0014544 | Epilepsy | 1 | CTD_human | |
| Tgene | C0021847 | Intestinal Pseudo-Obstruction | 1 | ORPHANET | |
| Tgene | C0023893 | Liver Cirrhosis, Experimental | 1 | CTD_human | |
| Tgene | C0026760 | Multiple Epiphyseal Dysplasia | 1 | CTD_human | |
| Tgene | C0029422 | Osteochondrodysplasias | 1 | CTD_human | |
| Tgene | C0036391 | Schwartz-Jampel Syndrome | 1 | CTD_human | |
| Tgene | C0038015 | Spondyloepiphyseal Dysplasia | 1 | CTD_human | |
| Tgene | C0086237 | Epilepsy, Cryptogenic | 1 | CTD_human | |
| Tgene | C0220769 | FG syndrome | 1 | CTD_human | |
| Tgene | C0236018 | Aura | 1 | CTD_human | |
| Tgene | C0268341 | Ehlers-Danlos syndrome type 5 | 1 | ORPHANET | |
| Tgene | C0432272 | Van Buchem disease | 1 | CTD_human | |
| Tgene | C0600066 | Malignant Cystosarcoma Phyllodes | 1 | CTD_human | |
| Tgene | C0678222 | Breast Carcinoma | 1 | CTD_human | |
| Tgene | C0751111 | Awakening Epilepsy | 1 | CTD_human | |
| Tgene | C1257931 | Mammary Neoplasms, Human | 1 | CTD_human | |
| Tgene | C1458155 | Mammary Neoplasms | 1 | CTD_human | |
| Tgene | C1845235 | Heterotopia, Periventricular, Ehlers-Danlos Variant | 1 | CTD_human | |
| Tgene | C1845546 | FG SYNDROME 4 (disorder) | 1 | CTD_human | |
| Tgene | C1845567 | FG SYNDROME 3 | 1 | CTD_human | |
| Tgene | C1855733 | Neuronal intestinal pseudoobstruction | 1 | ORPHANET | |
| Tgene | C2751260 | Macrothrombocytopenia | 1 | GENOMICS_ENGLAND | |
| Tgene | C3541456 | Spondyloepiphyseal Dysplasia Tarda, X-Linked | 1 | CTD_human | |
| Tgene | C4551479 | Schwartz-Jampel Syndrome, Type 1 | 1 | CTD_human | |
| Tgene | C4704874 | Mammary Carcinoma, Human | 1 | CTD_human |
