
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:SMARCA2-BEND2 (FusionGDB2 ID:HG6595TG139105) |
Fusion Gene Summary for SMARCA2-BEND2 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: SMARCA2-BEND2 | Fusion gene ID: hg6595tg139105 | Hgene | Tgene | Gene symbol | SMARCA2 | BEND2 | Gene ID | 6595 | 139105 |
| Gene name | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 | BEN domain containing 2 | |
| Synonyms | BAF190|BRM|NCBRS|SNF2|SNF2L2|SNF2LA|SWI2|Sth1p|hBRM|hSNF2a | CXorf20 | |
| Cytomap | ('SMARCA2')('BEND2') 9p24.3 | Xp22.13 | |
| Type of gene | protein-coding | protein-coding | |
| Description | probable global transcription activator SNF2L2ATP-dependent helicase SMARCA2BAF190BBRG1-associated factor 190BSNF2-alphaSNF2/SWI2-like protein 2SWI/SNF-related matrix-associated actin-dependent regulator of chromatin a2brahma homologglobal transcr | BEN domain-containing protein 2 | |
| Modification date | 20200315 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000349721, ENST00000357248, ENST00000382194, ENST00000382203, ENST00000302401, ENST00000324954, ENST00000382185, ENST00000382186, ENST00000491574, | ||
| Fusion gene scores | * DoF score | 14 X 9 X 10=1260 | 2 X 2 X 2=8 |
| # samples | 15 | 2 | |
| ** MAII score | log2(15/1260*10)=-3.0703893278914 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(2/8*10)=1.32192809488736 | |
| Context | PubMed: SMARCA2 [Title/Abstract] AND BEND2 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | SMARCA2(2039900)-BEND2(18213562), # samples:4 | ||
| Anticipated loss of major functional domain due to fusion event. | SMARCA2-BEND2 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. SMARCA2-BEND2 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. SMARCA2-BEND2 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. SMARCA2-BEND2 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | SMARCA2 | GO:0008285 | negative regulation of cell proliferation | 14660596 |
| Hgene | SMARCA2 | GO:0045892 | negative regulation of transcription, DNA-templated | 12065415 |
| Hgene | SMARCA2 | GO:0045893 | positive regulation of transcription, DNA-templated | 17984088 |
| Hgene | SMARCA2 | GO:0045944 | positive regulation of transcription by RNA polymerase II | 15774904 |
 Fusion gene breakpoints across SMARCA2 (5'-gene) Fusion gene breakpoints across SMARCA2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
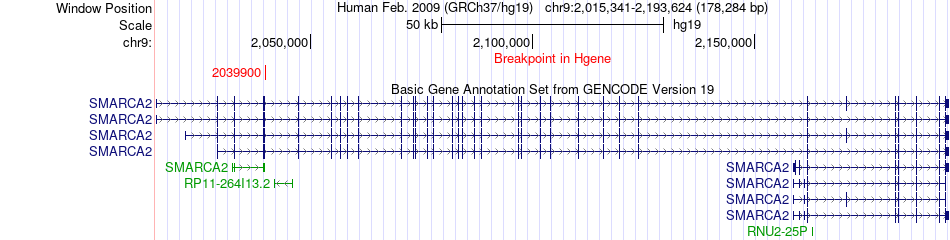 |
 Fusion gene breakpoints across BEND2 (3'-gene) Fusion gene breakpoints across BEND2 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
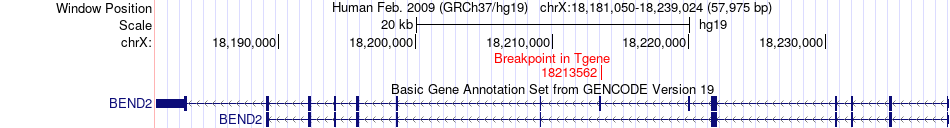 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | PAAD | TCGA-3A-A9IR-01A | SMARCA2 | chr9 | 2039900 | - | BEND2 | chrX | 18213562 | - |
| ChimerDB4 | PAAD | TCGA-3A-A9IR-01A | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
| ChimerDB4 | PAAD | TCGA-3A-A9IR | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
Top |
Fusion Gene ORF analysis for SMARCA2-BEND2 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000349721 | ENST00000380030 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
| 5CDS-intron | ENST00000357248 | ENST00000380030 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
| 5CDS-intron | ENST00000382194 | ENST00000380030 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
| 5CDS-intron | ENST00000382203 | ENST00000380030 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
| In-frame | ENST00000349721 | ENST00000380033 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
| In-frame | ENST00000357248 | ENST00000380033 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
| In-frame | ENST00000382194 | ENST00000380033 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
| In-frame | ENST00000382203 | ENST00000380033 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
| intron-3CDS | ENST00000302401 | ENST00000380033 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
| intron-3CDS | ENST00000324954 | ENST00000380033 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
| intron-3CDS | ENST00000382185 | ENST00000380033 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
| intron-3CDS | ENST00000382186 | ENST00000380033 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
| intron-3CDS | ENST00000491574 | ENST00000380033 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
| intron-intron | ENST00000302401 | ENST00000380030 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
| intron-intron | ENST00000324954 | ENST00000380030 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
| intron-intron | ENST00000382185 | ENST00000380030 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
| intron-intron | ENST00000382186 | ENST00000380030 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
| intron-intron | ENST00000491574 | ENST00000380030 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000357248 | SMARCA2 | chr9 | 2039900 | + | ENST00000380033 | BEND2 | chrX | 18213562 | - | 4334 | 889 | 99 | 2255 | 718 |
| ENST00000349721 | SMARCA2 | chr9 | 2039900 | + | ENST00000380033 | BEND2 | chrX | 18213562 | - | 4334 | 889 | 99 | 2255 | 718 |
| ENST00000382203 | SMARCA2 | chr9 | 2039900 | + | ENST00000380033 | BEND2 | chrX | 18213562 | - | 4444 | 999 | 209 | 2365 | 718 |
| ENST00000382194 | SMARCA2 | chr9 | 2039900 | + | ENST00000380033 | BEND2 | chrX | 18213562 | - | 4239 | 794 | 4 | 2160 | 718 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000357248 | ENST00000380033 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - | 0.000453432 | 0.9995466 |
| ENST00000349721 | ENST00000380033 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - | 0.000453432 | 0.9995466 |
| ENST00000382203 | ENST00000380033 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - | 0.000447153 | 0.9995528 |
| ENST00000382194 | ENST00000380033 | SMARCA2 | chr9 | 2039900 | + | BEND2 | chrX | 18213562 | - | 0.000430669 | 0.99956936 |
Top |
Fusion Genomic Features for SMARCA2-BEND2 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
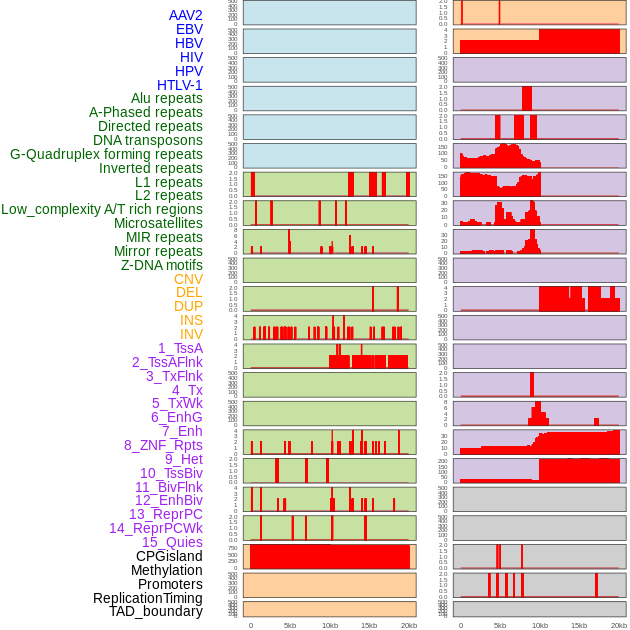 |
Top |
Fusion Protein Features for SMARCA2-BEND2 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr9:2039900/chrX:18213562) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000349721 | + | 4 | 34 | 216_238 | 263 | 1591.0 | Compositional bias | Note=Poly-Gln |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000349721 | + | 4 | 34 | 245_253 | 263 | 1591.0 | Compositional bias | Note=Poly-Gln |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000357248 | + | 4 | 33 | 216_238 | 263 | 1573.0 | Compositional bias | Note=Poly-Gln |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000357248 | + | 4 | 33 | 245_253 | 263 | 1573.0 | Compositional bias | Note=Poly-Gln |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382194 | + | 3 | 32 | 216_238 | 263 | 1573.0 | Compositional bias | Note=Poly-Gln |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382194 | + | 3 | 32 | 245_253 | 263 | 1573.0 | Compositional bias | Note=Poly-Gln |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382203 | + | 4 | 34 | 216_238 | 263 | 1591.0 | Compositional bias | Note=Poly-Gln |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382203 | + | 4 | 34 | 245_253 | 263 | 1591.0 | Compositional bias | Note=Poly-Gln |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000349721 | + | 4 | 34 | 173_208 | 263 | 1591.0 | Domain | QLQ |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000357248 | + | 4 | 33 | 173_208 | 263 | 1573.0 | Domain | QLQ |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382194 | + | 3 | 32 | 173_208 | 263 | 1573.0 | Domain | QLQ |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382203 | + | 4 | 34 | 173_208 | 263 | 1591.0 | Domain | QLQ |
| Tgene | BEND2 | chr9:2039900 | chrX:18213562 | ENST00000380030 | 0 | 11 | 483_582 | 0 | 646.0 | Domain | BEN 1 | |
| Tgene | BEND2 | chr9:2039900 | chrX:18213562 | ENST00000380030 | 0 | 11 | 667_765 | 0 | 646.0 | Domain | BEN 2 | |
| Tgene | BEND2 | chr9:2039900 | chrX:18213562 | ENST00000380033 | 5 | 14 | 483_582 | 344 | 800.0 | Domain | BEN 1 | |
| Tgene | BEND2 | chr9:2039900 | chrX:18213562 | ENST00000380033 | 5 | 14 | 667_765 | 344 | 800.0 | Domain | BEN 2 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000349721 | + | 4 | 34 | 1297_1301 | 263 | 1591.0 | Compositional bias | Note=Poly-Glu |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000349721 | + | 4 | 34 | 1518_1529 | 263 | 1591.0 | Compositional bias | Note=Poly-Glu |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000349721 | + | 4 | 34 | 559_562 | 263 | 1591.0 | Compositional bias | Note=Poly-Arg |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000349721 | + | 4 | 34 | 643_650 | 263 | 1591.0 | Compositional bias | Note=Poly-Glu |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000357248 | + | 4 | 33 | 1297_1301 | 263 | 1573.0 | Compositional bias | Note=Poly-Glu |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000357248 | + | 4 | 33 | 1518_1529 | 263 | 1573.0 | Compositional bias | Note=Poly-Glu |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000357248 | + | 4 | 33 | 559_562 | 263 | 1573.0 | Compositional bias | Note=Poly-Arg |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000357248 | + | 4 | 33 | 643_650 | 263 | 1573.0 | Compositional bias | Note=Poly-Glu |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382194 | + | 3 | 32 | 1297_1301 | 263 | 1573.0 | Compositional bias | Note=Poly-Glu |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382194 | + | 3 | 32 | 1518_1529 | 263 | 1573.0 | Compositional bias | Note=Poly-Glu |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382194 | + | 3 | 32 | 559_562 | 263 | 1573.0 | Compositional bias | Note=Poly-Arg |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382194 | + | 3 | 32 | 643_650 | 263 | 1573.0 | Compositional bias | Note=Poly-Glu |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382203 | + | 4 | 34 | 1297_1301 | 263 | 1591.0 | Compositional bias | Note=Poly-Glu |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382203 | + | 4 | 34 | 1518_1529 | 263 | 1591.0 | Compositional bias | Note=Poly-Glu |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382203 | + | 4 | 34 | 559_562 | 263 | 1591.0 | Compositional bias | Note=Poly-Arg |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382203 | + | 4 | 34 | 643_650 | 263 | 1591.0 | Compositional bias | Note=Poly-Glu |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000349721 | + | 4 | 34 | 1054_1216 | 263 | 1591.0 | Domain | Helicase C-terminal |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000349721 | + | 4 | 34 | 1419_1489 | 263 | 1591.0 | Domain | Bromo |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000349721 | + | 4 | 34 | 436_508 | 263 | 1591.0 | Domain | HSA |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000349721 | + | 4 | 34 | 736_901 | 263 | 1591.0 | Domain | Helicase ATP-binding |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000357248 | + | 4 | 33 | 1054_1216 | 263 | 1573.0 | Domain | Helicase C-terminal |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000357248 | + | 4 | 33 | 1419_1489 | 263 | 1573.0 | Domain | Bromo |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000357248 | + | 4 | 33 | 436_508 | 263 | 1573.0 | Domain | HSA |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000357248 | + | 4 | 33 | 736_901 | 263 | 1573.0 | Domain | Helicase ATP-binding |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382194 | + | 3 | 32 | 1054_1216 | 263 | 1573.0 | Domain | Helicase C-terminal |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382194 | + | 3 | 32 | 1419_1489 | 263 | 1573.0 | Domain | Bromo |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382194 | + | 3 | 32 | 436_508 | 263 | 1573.0 | Domain | HSA |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382194 | + | 3 | 32 | 736_901 | 263 | 1573.0 | Domain | Helicase ATP-binding |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382203 | + | 4 | 34 | 1054_1216 | 263 | 1591.0 | Domain | Helicase C-terminal |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382203 | + | 4 | 34 | 1419_1489 | 263 | 1591.0 | Domain | Bromo |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382203 | + | 4 | 34 | 436_508 | 263 | 1591.0 | Domain | HSA |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382203 | + | 4 | 34 | 736_901 | 263 | 1591.0 | Domain | Helicase ATP-binding |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000349721 | + | 4 | 34 | 851_854 | 263 | 1591.0 | Motif | Note=DEGH box |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000357248 | + | 4 | 33 | 851_854 | 263 | 1573.0 | Motif | Note=DEGH box |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382194 | + | 3 | 32 | 851_854 | 263 | 1573.0 | Motif | Note=DEGH box |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382203 | + | 4 | 34 | 851_854 | 263 | 1591.0 | Motif | Note=DEGH box |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000349721 | + | 4 | 34 | 749_756 | 263 | 1591.0 | Nucleotide binding | ATP |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000357248 | + | 4 | 33 | 749_756 | 263 | 1573.0 | Nucleotide binding | ATP |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382194 | + | 3 | 32 | 749_756 | 263 | 1573.0 | Nucleotide binding | ATP |
| Hgene | SMARCA2 | chr9:2039900 | chrX:18213562 | ENST00000382203 | + | 4 | 34 | 749_756 | 263 | 1591.0 | Nucleotide binding | ATP |
Top |
Fusion Gene Sequence for SMARCA2-BEND2 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >83869_83869_1_SMARCA2-BEND2_SMARCA2_chr9_2039900_ENST00000349721_BEND2_chrX_18213562_ENST00000380033_length(transcript)=4334nt_BP=889nt TTTCTGTACTCTGGGTGACTCAGAGAGGGAAGAGATTCAGCCAGCACACTCCTCGCGAGCAAGCATTACTCTACTGACTGGCAGAGACAG GAGAGGTAGATGTCCACGCCCACAGACCCTGGTGCGATGCCCCACCCAGGGCCTTCGCCGGGGCCTGGGCCTTCCCCTGGGCCAATTCTT GGGCCTAGTCCAGGACCAGGACCATCCCCAGGTTCCGTCCACAGCATGATGGGGCCAAGTCCTGGACCTCCAAGTGTCTCCCATCCTATG CCGACGATGGGGTCCACAGACTTCCCACAGGAAGGCATGCATCAAATGCATAAGCCCATCGATGGTATACATGACAAGGGGATTGTAGAA GACATCCATTGTGGATCCATGAAGGGCACTGGTATGCGACCACCTCACCCAGGCATGGGCCCTCCCCAGAGTCCAATGGATCAACACAGC CAAGGTTATATGTCACCACACCCATCTCCATTAGGAGCCCCAGAGCACGTCTCCAGCCCTATGTCTGGAGGAGGCCCAACTCCACCTCAG ATGCCACCAAGCCAGCCGGGGGCCCTCATCCCAGGTGATCCGCAGGCCATGAGCCAGCCCAACAGAGGTCCCTCACCTTTCAGTCCTGTC CAGCTGCATCAGCTTCGAGCTCAGATTTTAGCTTATAAAATGCTGGCCCGAGGCCAGCCCCTCCCCGAAACGCTGCAGCTTGCAGTCCAG GGGAAAAGGACGTTGCCTGGCTTGCAGCAACAACAGCAGCAGCAACAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAACAG CAGCCGCAGCAGCAGCCGCCGCAACCACAGACGCAGCAACAACAGCAGCCGGCCCTTGTTAACTACAACAGACCATCTGCTGAGAAAATT ATACTTACTGAAATGCCAGGAACAACAGAAACCAACGTGGAAAATAACTCTCAGACAGTGTATTACCCAGCTTTATCGGGAAATACGAGT GCCCCATATCCAGCCTCTTCATATCTTCCCATCACTTCTAATTTTGAATCTGGCCCACAAATGAGTTATGGGACAATGAGTTACTCAACT GAAATGAAAAATAACTGTGACCAAGATGATGCTTCAGCATCTGCCTGCCTCACTCCCGATTTTGCACTGTTACCTCTGAATATTTTGGTA AAAGTAGACACCAACACGGAAAACAGCGTCAACACAATGAATCGCTCAACTTTATTGGACAGTGACAGTGGCCAGGATTCTTCCTCATCA TCTGTCTGTATCCCTCCCAAGTATGGCTATCTTGGTGATCCAAAAAGAAATGTCAGAGTACTTAAAATCCATTTGCTGGCCGTACAAAAT ATGGCCAAACCTAAGCAAGCAGCCTGCTACTTGGTTCGTATTTTGTTCTCCAAGGAAATCCTGATTAGCAGCTCAGTGGATATCCATTTG AAAGACAGCCAATCCCTCGACCCGAACAAAATGGCTGCATTGAGAGAATATCTGGCAACAACTTTTCCCACCTGTGATTTGCATGAACAT GGAAAAGACTGGCAGGACTGTATTTCTGGTATCAATTCTATGATCTACTGTTTATGTTCTGAAGGCAAGAGTACTCCAAAAACTGTTCGC AAAAATAAAAAACGTACCAACCGTGTTGCATCGGCATCTGCAGATAGAAATGACCAAAGGGGCAGAGATGGTGGTGAAGGCTGTTCTTGG ATGTTTCAGCCAATGAATAACTCCAAAATGAGAGAAAAGAGGAACTTGCAGCCAAACAGTAATGCTATCCCTGAAGGAATGCGAGAACCT TCCACTGATAATCCAGAGGAACCTGGTGAAGCATGGAGCTATTTTGGAAGACCATGGAGAAATATACGGATGCCATGTTCAGTACTGACT TTGGCAAAAACTAAGTCTTGCGCAAGCCTGTCGGCTAGATACCTTATTCAGAAACTCTTCACAAAAGATGTCCTGGTCCAAAGTAACGTC TATGGCAATCTGAAGCATGGCCTGTGTGCCCTTGACCCCAATAAGATTAGTGCTCTCCGAGAGTTCCTCCAAGAAAACTACCCAATTTGT GATCTCTCGGAAAATGGAAGAGACTGGAAGTCGTGTGTGACCTCCATCAACAGCGGTATCCGTAGCCTTAGACATGACGTCAGAAGGGCT GAAGCCAGGTCTCAGTCGCTTCCAGCAGTGACCCCTCCAGAGCTGGAGCAGGAGTCAAAGCCAGGAGATCCCGACGCCACTGACCCAAGC ACCTGAACGGCAGCTGCCAAACTTTTTTTTTTTTTTTAACTGTTAAGACTATTTTGTAAGTTTCTCAAGTTCTAATTCCAAAATCAGCCT GCTGTACTGGCAGAGTAGTTTGCATTGACACCTGTAGTAATCGCCAAACCTCTTTTTCTGCTCATAAAGGAGGTAGAATCACATTAGGAA GATACGTAATGTTTCATCTGTGGATTTGATGACAACCTCTGATTGAGGGAATGGTAGTAAACTCCTCATGGGCTTGTGACCCATGGCTTT GACTTAGATAATTGACATTAATATACTTTGTTTTTTTGATAAGGTAGAGAAAGAAATTCAATAGAAAGTAAGGAGCCACTCAGTCAAATA ATATAATTCGTATAAAACTTGACAAGTCTTTTTATTTTTGTGACTTGTGAGATAGTTTAAGATATTTTTATGTGACCAATTTGTAGAATG AATACATATTTTCAAAGTTATCTTTTCAAAATTCAACTTTTAGAGTATTTTAGCAAAATATATGTATAGTTTATAATATAGAAGTATAGC TTTATAAAGATATTAAGGATTTATTTTTTAAAATAATTCTCTGAATAGTTTGCTGTAGTAGTTGCCTTCAGCTTAATAAAACCATAGTAC TTCTACATACATCTTTATAAGTTAGGCTATGCCTACAAAGGTCTTAAAAAGTAGATAATTTACCGGGTGTAGTGGCTCACACCTGTAATC CCAGCACTTTGGGAGGCCGAGACGGGTGGATCACCTGAGGTCAGGGGTTCGAGACCAGCCTGGCCAACATGACGAAACCCCATCTCTACT AAAAATACAAAAAAAGTAGCCAGGCATGGTGGCAAGCACCTGTAGTCCCAGTTACTCGGGTGGCTGAGGCAGGAGAATCACTTGAACCTG GGAGGTGGAGACTGCAGTGAGCCGAGATTGTGCCACTGCACTCGAGCCTGGGTGACAGAGTGAGACTCCGTCTCGGGAAAAGAAAAAAAA GAAGTAGATAATTAAAAAAATCTTGCATAAAAAAAAATCTTCCATAAAGGGCAAAAAGGCTTTAACTTCCATTATTCAGATTTAATCCAA CTAAGATAGTCTTTGGCTTCATTCACTGTTCAGTTTCACAAGTTTTTTACAACTACTAGTAACAGAGCTAATACAGAATAGTAATCCACA TTATAAACATAGATCAAAATACCACATCATACCCCATAAATACATGCAGTTATTATTTGTCAATTAAAAATAAAATTTTTTTAAAAGAAA AGTACAGTTCGACCCTAGGCAAGAAGAAATTTCAAATTGTATATGCATATTTTGAATTTGGTTTATATTTTGTGGTATCTGAAATATTCC CTGGACTAGAAGCAAAACCACAGAAACAAAACTTTACATTTAGAAAAAAATTTCAAGACTTCTATTGCTAACTTTATTACATTAGTGGTT TTTAGAAGTTATTTAGAGCAGTCTTGTCTGATAGAACTTTCCACCATGATGGAAATGTAAATAAATATACCACATTAAAAATATAAATTT GCAATGTCCAATAGGCTCGTATGGCTATTGAGCATTTAAGATGTAATTAATTAGACAGAGGAAGTAAATTTAACATTTTCTCTCATTTTA GCTAATTAGAATTTAAAGTGAAATAGCCACAAGTGGATAGTGGCTACTGTGTTGGATGGCACAGTTCAAAATGCCTCTTATTTATAAAAA AAAGTGTTTAAAAGTGGTTCCCATGGCTTGAAGAGGAATAATATACACATAACTTCTTTAAAATCACATTTTGTTTGAGATGGAGATCTT GCTATATTGCCCAGGCTTATCTGGAACTCCTGGGCTCAAGAGGGAGGATCACATTTGACTGGACAAAATACATTTTATAAAGGTGTTTTG TATTTTATGTTAAAGTGGAATTGTGTCTTCAAAATTGTGTCTTCAAAATTTCACAAGTTAATACTATTACTGTATTAGTCCAATGTTAAA CATTTTAAATGCCTGCTCATAAGACTCATTTTTGATAAATAAATGTATCTTTAATGTTTTGTTTCCCAAATAGATACAATAAATAAACAT >83869_83869_1_SMARCA2-BEND2_SMARCA2_chr9_2039900_ENST00000349721_BEND2_chrX_18213562_ENST00000380033_length(amino acids)=718AA_BP=261 MSTPTDPGAMPHPGPSPGPGPSPGPILGPSPGPGPSPGSVHSMMGPSPGPPSVSHPMPTMGSTDFPQEGMHQMHKPIDGIHDKGIVEDIH CGSMKGTGMRPPHPGMGPPQSPMDQHSQGYMSPHPSPLGAPEHVSSPMSGGGPTPPQMPPSQPGALIPGDPQAMSQPNRGPSPFSPVQLH QLRAQILAYKMLARGQPLPETLQLAVQGKRTLPGLQQQQQQQQQQQQQQQQQQQQQQQPQQQPPQPQTQQQQQPALVNYNRPSAEKIILT EMPGTTETNVENNSQTVYYPALSGNTSAPYPASSYLPITSNFESGPQMSYGTMSYSTEMKNNCDQDDASASACLTPDFALLPLNILVKVD TNTENSVNTMNRSTLLDSDSGQDSSSSSVCIPPKYGYLGDPKRNVRVLKIHLLAVQNMAKPKQAACYLVRILFSKEILISSSVDIHLKDS QSLDPNKMAALREYLATTFPTCDLHEHGKDWQDCISGINSMIYCLCSEGKSTPKTVRKNKKRTNRVASASADRNDQRGRDGGEGCSWMFQ PMNNSKMREKRNLQPNSNAIPEGMREPSTDNPEEPGEAWSYFGRPWRNIRMPCSVLTLAKTKSCASLSARYLIQKLFTKDVLVQSNVYGN -------------------------------------------------------------- >83869_83869_2_SMARCA2-BEND2_SMARCA2_chr9_2039900_ENST00000357248_BEND2_chrX_18213562_ENST00000380033_length(transcript)=4334nt_BP=889nt TTTCTGTACTCTGGGTGACTCAGAGAGGGAAGAGATTCAGCCAGCACACTCCTCGCGAGCAAGCATTACTCTACTGACTGGCAGAGACAG GAGAGGTAGATGTCCACGCCCACAGACCCTGGTGCGATGCCCCACCCAGGGCCTTCGCCGGGGCCTGGGCCTTCCCCTGGGCCAATTCTT GGGCCTAGTCCAGGACCAGGACCATCCCCAGGTTCCGTCCACAGCATGATGGGGCCAAGTCCTGGACCTCCAAGTGTCTCCCATCCTATG CCGACGATGGGGTCCACAGACTTCCCACAGGAAGGCATGCATCAAATGCATAAGCCCATCGATGGTATACATGACAAGGGGATTGTAGAA GACATCCATTGTGGATCCATGAAGGGCACTGGTATGCGACCACCTCACCCAGGCATGGGCCCTCCCCAGAGTCCAATGGATCAACACAGC CAAGGTTATATGTCACCACACCCATCTCCATTAGGAGCCCCAGAGCACGTCTCCAGCCCTATGTCTGGAGGAGGCCCAACTCCACCTCAG ATGCCACCAAGCCAGCCGGGGGCCCTCATCCCAGGTGATCCGCAGGCCATGAGCCAGCCCAACAGAGGTCCCTCACCTTTCAGTCCTGTC CAGCTGCATCAGCTTCGAGCTCAGATTTTAGCTTATAAAATGCTGGCCCGAGGCCAGCCCCTCCCCGAAACGCTGCAGCTTGCAGTCCAG GGGAAAAGGACGTTGCCTGGCTTGCAGCAACAACAGCAGCAGCAACAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAACAG CAGCCGCAGCAGCAGCCGCCGCAACCACAGACGCAGCAACAACAGCAGCCGGCCCTTGTTAACTACAACAGACCATCTGCTGAGAAAATT ATACTTACTGAAATGCCAGGAACAACAGAAACCAACGTGGAAAATAACTCTCAGACAGTGTATTACCCAGCTTTATCGGGAAATACGAGT GCCCCATATCCAGCCTCTTCATATCTTCCCATCACTTCTAATTTTGAATCTGGCCCACAAATGAGTTATGGGACAATGAGTTACTCAACT GAAATGAAAAATAACTGTGACCAAGATGATGCTTCAGCATCTGCCTGCCTCACTCCCGATTTTGCACTGTTACCTCTGAATATTTTGGTA AAAGTAGACACCAACACGGAAAACAGCGTCAACACAATGAATCGCTCAACTTTATTGGACAGTGACAGTGGCCAGGATTCTTCCTCATCA TCTGTCTGTATCCCTCCCAAGTATGGCTATCTTGGTGATCCAAAAAGAAATGTCAGAGTACTTAAAATCCATTTGCTGGCCGTACAAAAT ATGGCCAAACCTAAGCAAGCAGCCTGCTACTTGGTTCGTATTTTGTTCTCCAAGGAAATCCTGATTAGCAGCTCAGTGGATATCCATTTG AAAGACAGCCAATCCCTCGACCCGAACAAAATGGCTGCATTGAGAGAATATCTGGCAACAACTTTTCCCACCTGTGATTTGCATGAACAT GGAAAAGACTGGCAGGACTGTATTTCTGGTATCAATTCTATGATCTACTGTTTATGTTCTGAAGGCAAGAGTACTCCAAAAACTGTTCGC AAAAATAAAAAACGTACCAACCGTGTTGCATCGGCATCTGCAGATAGAAATGACCAAAGGGGCAGAGATGGTGGTGAAGGCTGTTCTTGG ATGTTTCAGCCAATGAATAACTCCAAAATGAGAGAAAAGAGGAACTTGCAGCCAAACAGTAATGCTATCCCTGAAGGAATGCGAGAACCT TCCACTGATAATCCAGAGGAACCTGGTGAAGCATGGAGCTATTTTGGAAGACCATGGAGAAATATACGGATGCCATGTTCAGTACTGACT TTGGCAAAAACTAAGTCTTGCGCAAGCCTGTCGGCTAGATACCTTATTCAGAAACTCTTCACAAAAGATGTCCTGGTCCAAAGTAACGTC TATGGCAATCTGAAGCATGGCCTGTGTGCCCTTGACCCCAATAAGATTAGTGCTCTCCGAGAGTTCCTCCAAGAAAACTACCCAATTTGT GATCTCTCGGAAAATGGAAGAGACTGGAAGTCGTGTGTGACCTCCATCAACAGCGGTATCCGTAGCCTTAGACATGACGTCAGAAGGGCT GAAGCCAGGTCTCAGTCGCTTCCAGCAGTGACCCCTCCAGAGCTGGAGCAGGAGTCAAAGCCAGGAGATCCCGACGCCACTGACCCAAGC ACCTGAACGGCAGCTGCCAAACTTTTTTTTTTTTTTTAACTGTTAAGACTATTTTGTAAGTTTCTCAAGTTCTAATTCCAAAATCAGCCT GCTGTACTGGCAGAGTAGTTTGCATTGACACCTGTAGTAATCGCCAAACCTCTTTTTCTGCTCATAAAGGAGGTAGAATCACATTAGGAA GATACGTAATGTTTCATCTGTGGATTTGATGACAACCTCTGATTGAGGGAATGGTAGTAAACTCCTCATGGGCTTGTGACCCATGGCTTT GACTTAGATAATTGACATTAATATACTTTGTTTTTTTGATAAGGTAGAGAAAGAAATTCAATAGAAAGTAAGGAGCCACTCAGTCAAATA ATATAATTCGTATAAAACTTGACAAGTCTTTTTATTTTTGTGACTTGTGAGATAGTTTAAGATATTTTTATGTGACCAATTTGTAGAATG AATACATATTTTCAAAGTTATCTTTTCAAAATTCAACTTTTAGAGTATTTTAGCAAAATATATGTATAGTTTATAATATAGAAGTATAGC TTTATAAAGATATTAAGGATTTATTTTTTAAAATAATTCTCTGAATAGTTTGCTGTAGTAGTTGCCTTCAGCTTAATAAAACCATAGTAC TTCTACATACATCTTTATAAGTTAGGCTATGCCTACAAAGGTCTTAAAAAGTAGATAATTTACCGGGTGTAGTGGCTCACACCTGTAATC CCAGCACTTTGGGAGGCCGAGACGGGTGGATCACCTGAGGTCAGGGGTTCGAGACCAGCCTGGCCAACATGACGAAACCCCATCTCTACT AAAAATACAAAAAAAGTAGCCAGGCATGGTGGCAAGCACCTGTAGTCCCAGTTACTCGGGTGGCTGAGGCAGGAGAATCACTTGAACCTG GGAGGTGGAGACTGCAGTGAGCCGAGATTGTGCCACTGCACTCGAGCCTGGGTGACAGAGTGAGACTCCGTCTCGGGAAAAGAAAAAAAA GAAGTAGATAATTAAAAAAATCTTGCATAAAAAAAAATCTTCCATAAAGGGCAAAAAGGCTTTAACTTCCATTATTCAGATTTAATCCAA CTAAGATAGTCTTTGGCTTCATTCACTGTTCAGTTTCACAAGTTTTTTACAACTACTAGTAACAGAGCTAATACAGAATAGTAATCCACA TTATAAACATAGATCAAAATACCACATCATACCCCATAAATACATGCAGTTATTATTTGTCAATTAAAAATAAAATTTTTTTAAAAGAAA AGTACAGTTCGACCCTAGGCAAGAAGAAATTTCAAATTGTATATGCATATTTTGAATTTGGTTTATATTTTGTGGTATCTGAAATATTCC CTGGACTAGAAGCAAAACCACAGAAACAAAACTTTACATTTAGAAAAAAATTTCAAGACTTCTATTGCTAACTTTATTACATTAGTGGTT TTTAGAAGTTATTTAGAGCAGTCTTGTCTGATAGAACTTTCCACCATGATGGAAATGTAAATAAATATACCACATTAAAAATATAAATTT GCAATGTCCAATAGGCTCGTATGGCTATTGAGCATTTAAGATGTAATTAATTAGACAGAGGAAGTAAATTTAACATTTTCTCTCATTTTA GCTAATTAGAATTTAAAGTGAAATAGCCACAAGTGGATAGTGGCTACTGTGTTGGATGGCACAGTTCAAAATGCCTCTTATTTATAAAAA AAAGTGTTTAAAAGTGGTTCCCATGGCTTGAAGAGGAATAATATACACATAACTTCTTTAAAATCACATTTTGTTTGAGATGGAGATCTT GCTATATTGCCCAGGCTTATCTGGAACTCCTGGGCTCAAGAGGGAGGATCACATTTGACTGGACAAAATACATTTTATAAAGGTGTTTTG TATTTTATGTTAAAGTGGAATTGTGTCTTCAAAATTGTGTCTTCAAAATTTCACAAGTTAATACTATTACTGTATTAGTCCAATGTTAAA CATTTTAAATGCCTGCTCATAAGACTCATTTTTGATAAATAAATGTATCTTTAATGTTTTGTTTCCCAAATAGATACAATAAATAAACAT >83869_83869_2_SMARCA2-BEND2_SMARCA2_chr9_2039900_ENST00000357248_BEND2_chrX_18213562_ENST00000380033_length(amino acids)=718AA_BP=261 MSTPTDPGAMPHPGPSPGPGPSPGPILGPSPGPGPSPGSVHSMMGPSPGPPSVSHPMPTMGSTDFPQEGMHQMHKPIDGIHDKGIVEDIH CGSMKGTGMRPPHPGMGPPQSPMDQHSQGYMSPHPSPLGAPEHVSSPMSGGGPTPPQMPPSQPGALIPGDPQAMSQPNRGPSPFSPVQLH QLRAQILAYKMLARGQPLPETLQLAVQGKRTLPGLQQQQQQQQQQQQQQQQQQQQQQQPQQQPPQPQTQQQQQPALVNYNRPSAEKIILT EMPGTTETNVENNSQTVYYPALSGNTSAPYPASSYLPITSNFESGPQMSYGTMSYSTEMKNNCDQDDASASACLTPDFALLPLNILVKVD TNTENSVNTMNRSTLLDSDSGQDSSSSSVCIPPKYGYLGDPKRNVRVLKIHLLAVQNMAKPKQAACYLVRILFSKEILISSSVDIHLKDS QSLDPNKMAALREYLATTFPTCDLHEHGKDWQDCISGINSMIYCLCSEGKSTPKTVRKNKKRTNRVASASADRNDQRGRDGGEGCSWMFQ PMNNSKMREKRNLQPNSNAIPEGMREPSTDNPEEPGEAWSYFGRPWRNIRMPCSVLTLAKTKSCASLSARYLIQKLFTKDVLVQSNVYGN -------------------------------------------------------------- >83869_83869_3_SMARCA2-BEND2_SMARCA2_chr9_2039900_ENST00000382194_BEND2_chrX_18213562_ENST00000380033_length(transcript)=4239nt_BP=794nt GTAGATGTCCACGCCCACAGACCCTGGTGCGATGCCCCACCCAGGGCCTTCGCCGGGGCCTGGGCCTTCCCCTGGGCCAATTCTTGGGCC TAGTCCAGGACCAGGACCATCCCCAGGTTCCGTCCACAGCATGATGGGGCCAAGTCCTGGACCTCCAAGTGTCTCCCATCCTATGCCGAC GATGGGGTCCACAGACTTCCCACAGGAAGGCATGCATCAAATGCATAAGCCCATCGATGGTATACATGACAAGGGGATTGTAGAAGACAT CCATTGTGGATCCATGAAGGGCACTGGTATGCGACCACCTCACCCAGGCATGGGCCCTCCCCAGAGTCCAATGGATCAACACAGCCAAGG TTATATGTCACCACACCCATCTCCATTAGGAGCCCCAGAGCACGTCTCCAGCCCTATGTCTGGAGGAGGCCCAACTCCACCTCAGATGCC ACCAAGCCAGCCGGGGGCCCTCATCCCAGGTGATCCGCAGGCCATGAGCCAGCCCAACAGAGGTCCCTCACCTTTCAGTCCTGTCCAGCT GCATCAGCTTCGAGCTCAGATTTTAGCTTATAAAATGCTGGCCCGAGGCCAGCCCCTCCCCGAAACGCTGCAGCTTGCAGTCCAGGGGAA AAGGACGTTGCCTGGCTTGCAGCAACAACAGCAGCAGCAACAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAACAGCAGCC GCAGCAGCAGCCGCCGCAACCACAGACGCAGCAACAACAGCAGCCGGCCCTTGTTAACTACAACAGACCATCTGCTGAGAAAATTATACT TACTGAAATGCCAGGAACAACAGAAACCAACGTGGAAAATAACTCTCAGACAGTGTATTACCCAGCTTTATCGGGAAATACGAGTGCCCC ATATCCAGCCTCTTCATATCTTCCCATCACTTCTAATTTTGAATCTGGCCCACAAATGAGTTATGGGACAATGAGTTACTCAACTGAAAT GAAAAATAACTGTGACCAAGATGATGCTTCAGCATCTGCCTGCCTCACTCCCGATTTTGCACTGTTACCTCTGAATATTTTGGTAAAAGT AGACACCAACACGGAAAACAGCGTCAACACAATGAATCGCTCAACTTTATTGGACAGTGACAGTGGCCAGGATTCTTCCTCATCATCTGT CTGTATCCCTCCCAAGTATGGCTATCTTGGTGATCCAAAAAGAAATGTCAGAGTACTTAAAATCCATTTGCTGGCCGTACAAAATATGGC CAAACCTAAGCAAGCAGCCTGCTACTTGGTTCGTATTTTGTTCTCCAAGGAAATCCTGATTAGCAGCTCAGTGGATATCCATTTGAAAGA CAGCCAATCCCTCGACCCGAACAAAATGGCTGCATTGAGAGAATATCTGGCAACAACTTTTCCCACCTGTGATTTGCATGAACATGGAAA AGACTGGCAGGACTGTATTTCTGGTATCAATTCTATGATCTACTGTTTATGTTCTGAAGGCAAGAGTACTCCAAAAACTGTTCGCAAAAA TAAAAAACGTACCAACCGTGTTGCATCGGCATCTGCAGATAGAAATGACCAAAGGGGCAGAGATGGTGGTGAAGGCTGTTCTTGGATGTT TCAGCCAATGAATAACTCCAAAATGAGAGAAAAGAGGAACTTGCAGCCAAACAGTAATGCTATCCCTGAAGGAATGCGAGAACCTTCCAC TGATAATCCAGAGGAACCTGGTGAAGCATGGAGCTATTTTGGAAGACCATGGAGAAATATACGGATGCCATGTTCAGTACTGACTTTGGC AAAAACTAAGTCTTGCGCAAGCCTGTCGGCTAGATACCTTATTCAGAAACTCTTCACAAAAGATGTCCTGGTCCAAAGTAACGTCTATGG CAATCTGAAGCATGGCCTGTGTGCCCTTGACCCCAATAAGATTAGTGCTCTCCGAGAGTTCCTCCAAGAAAACTACCCAATTTGTGATCT CTCGGAAAATGGAAGAGACTGGAAGTCGTGTGTGACCTCCATCAACAGCGGTATCCGTAGCCTTAGACATGACGTCAGAAGGGCTGAAGC CAGGTCTCAGTCGCTTCCAGCAGTGACCCCTCCAGAGCTGGAGCAGGAGTCAAAGCCAGGAGATCCCGACGCCACTGACCCAAGCACCTG AACGGCAGCTGCCAAACTTTTTTTTTTTTTTTAACTGTTAAGACTATTTTGTAAGTTTCTCAAGTTCTAATTCCAAAATCAGCCTGCTGT ACTGGCAGAGTAGTTTGCATTGACACCTGTAGTAATCGCCAAACCTCTTTTTCTGCTCATAAAGGAGGTAGAATCACATTAGGAAGATAC GTAATGTTTCATCTGTGGATTTGATGACAACCTCTGATTGAGGGAATGGTAGTAAACTCCTCATGGGCTTGTGACCCATGGCTTTGACTT AGATAATTGACATTAATATACTTTGTTTTTTTGATAAGGTAGAGAAAGAAATTCAATAGAAAGTAAGGAGCCACTCAGTCAAATAATATA ATTCGTATAAAACTTGACAAGTCTTTTTATTTTTGTGACTTGTGAGATAGTTTAAGATATTTTTATGTGACCAATTTGTAGAATGAATAC ATATTTTCAAAGTTATCTTTTCAAAATTCAACTTTTAGAGTATTTTAGCAAAATATATGTATAGTTTATAATATAGAAGTATAGCTTTAT AAAGATATTAAGGATTTATTTTTTAAAATAATTCTCTGAATAGTTTGCTGTAGTAGTTGCCTTCAGCTTAATAAAACCATAGTACTTCTA CATACATCTTTATAAGTTAGGCTATGCCTACAAAGGTCTTAAAAAGTAGATAATTTACCGGGTGTAGTGGCTCACACCTGTAATCCCAGC ACTTTGGGAGGCCGAGACGGGTGGATCACCTGAGGTCAGGGGTTCGAGACCAGCCTGGCCAACATGACGAAACCCCATCTCTACTAAAAA TACAAAAAAAGTAGCCAGGCATGGTGGCAAGCACCTGTAGTCCCAGTTACTCGGGTGGCTGAGGCAGGAGAATCACTTGAACCTGGGAGG TGGAGACTGCAGTGAGCCGAGATTGTGCCACTGCACTCGAGCCTGGGTGACAGAGTGAGACTCCGTCTCGGGAAAAGAAAAAAAAGAAGT AGATAATTAAAAAAATCTTGCATAAAAAAAAATCTTCCATAAAGGGCAAAAAGGCTTTAACTTCCATTATTCAGATTTAATCCAACTAAG ATAGTCTTTGGCTTCATTCACTGTTCAGTTTCACAAGTTTTTTACAACTACTAGTAACAGAGCTAATACAGAATAGTAATCCACATTATA AACATAGATCAAAATACCACATCATACCCCATAAATACATGCAGTTATTATTTGTCAATTAAAAATAAAATTTTTTTAAAAGAAAAGTAC AGTTCGACCCTAGGCAAGAAGAAATTTCAAATTGTATATGCATATTTTGAATTTGGTTTATATTTTGTGGTATCTGAAATATTCCCTGGA CTAGAAGCAAAACCACAGAAACAAAACTTTACATTTAGAAAAAAATTTCAAGACTTCTATTGCTAACTTTATTACATTAGTGGTTTTTAG AAGTTATTTAGAGCAGTCTTGTCTGATAGAACTTTCCACCATGATGGAAATGTAAATAAATATACCACATTAAAAATATAAATTTGCAAT GTCCAATAGGCTCGTATGGCTATTGAGCATTTAAGATGTAATTAATTAGACAGAGGAAGTAAATTTAACATTTTCTCTCATTTTAGCTAA TTAGAATTTAAAGTGAAATAGCCACAAGTGGATAGTGGCTACTGTGTTGGATGGCACAGTTCAAAATGCCTCTTATTTATAAAAAAAAGT GTTTAAAAGTGGTTCCCATGGCTTGAAGAGGAATAATATACACATAACTTCTTTAAAATCACATTTTGTTTGAGATGGAGATCTTGCTAT ATTGCCCAGGCTTATCTGGAACTCCTGGGCTCAAGAGGGAGGATCACATTTGACTGGACAAAATACATTTTATAAAGGTGTTTTGTATTT TATGTTAAAGTGGAATTGTGTCTTCAAAATTGTGTCTTCAAAATTTCACAAGTTAATACTATTACTGTATTAGTCCAATGTTAAACATTT TAAATGCCTGCTCATAAGACTCATTTTTGATAAATAAATGTATCTTTAATGTTTTGTTTCCCAAATAGATACAATAAATAAACATTTAAA >83869_83869_3_SMARCA2-BEND2_SMARCA2_chr9_2039900_ENST00000382194_BEND2_chrX_18213562_ENST00000380033_length(amino acids)=718AA_BP=261 MSTPTDPGAMPHPGPSPGPGPSPGPILGPSPGPGPSPGSVHSMMGPSPGPPSVSHPMPTMGSTDFPQEGMHQMHKPIDGIHDKGIVEDIH CGSMKGTGMRPPHPGMGPPQSPMDQHSQGYMSPHPSPLGAPEHVSSPMSGGGPTPPQMPPSQPGALIPGDPQAMSQPNRGPSPFSPVQLH QLRAQILAYKMLARGQPLPETLQLAVQGKRTLPGLQQQQQQQQQQQQQQQQQQQQQQQPQQQPPQPQTQQQQQPALVNYNRPSAEKIILT EMPGTTETNVENNSQTVYYPALSGNTSAPYPASSYLPITSNFESGPQMSYGTMSYSTEMKNNCDQDDASASACLTPDFALLPLNILVKVD TNTENSVNTMNRSTLLDSDSGQDSSSSSVCIPPKYGYLGDPKRNVRVLKIHLLAVQNMAKPKQAACYLVRILFSKEILISSSVDIHLKDS QSLDPNKMAALREYLATTFPTCDLHEHGKDWQDCISGINSMIYCLCSEGKSTPKTVRKNKKRTNRVASASADRNDQRGRDGGEGCSWMFQ PMNNSKMREKRNLQPNSNAIPEGMREPSTDNPEEPGEAWSYFGRPWRNIRMPCSVLTLAKTKSCASLSARYLIQKLFTKDVLVQSNVYGN -------------------------------------------------------------- >83869_83869_4_SMARCA2-BEND2_SMARCA2_chr9_2039900_ENST00000382203_BEND2_chrX_18213562_ENST00000380033_length(transcript)=4444nt_BP=999nt TCAGAAGAAAGCCCCGAGATCACAGAGACCCGGCGAGATCACAGAGACCCGGCCTGAAGGAACGTGGAAAGACCAATGTACCTGTTTTGA CCGGTTGCCTGGAGCAAGAAGTTCCAGTTGGGGAGAATTTTCAGAAGATAAAGTCGGAGATTGTGGAAAGACTTGACTTGCAGCATTACT CTACTGACTGGCAGAGACAGGAGAGGTAGATGTCCACGCCCACAGACCCTGGTGCGATGCCCCACCCAGGGCCTTCGCCGGGGCCTGGGC CTTCCCCTGGGCCAATTCTTGGGCCTAGTCCAGGACCAGGACCATCCCCAGGTTCCGTCCACAGCATGATGGGGCCAAGTCCTGGACCTC CAAGTGTCTCCCATCCTATGCCGACGATGGGGTCCACAGACTTCCCACAGGAAGGCATGCATCAAATGCATAAGCCCATCGATGGTATAC ATGACAAGGGGATTGTAGAAGACATCCATTGTGGATCCATGAAGGGCACTGGTATGCGACCACCTCACCCAGGCATGGGCCCTCCCCAGA GTCCAATGGATCAACACAGCCAAGGTTATATGTCACCACACCCATCTCCATTAGGAGCCCCAGAGCACGTCTCCAGCCCTATGTCTGGAG GAGGCCCAACTCCACCTCAGATGCCACCAAGCCAGCCGGGGGCCCTCATCCCAGGTGATCCGCAGGCCATGAGCCAGCCCAACAGAGGTC CCTCACCTTTCAGTCCTGTCCAGCTGCATCAGCTTCGAGCTCAGATTTTAGCTTATAAAATGCTGGCCCGAGGCCAGCCCCTCCCCGAAA CGCTGCAGCTTGCAGTCCAGGGGAAAAGGACGTTGCCTGGCTTGCAGCAACAACAGCAGCAGCAACAGCAGCAGCAGCAGCAGCAGCAGC AGCAGCAGCAGCAGCAACAGCAGCCGCAGCAGCAGCCGCCGCAACCACAGACGCAGCAACAACAGCAGCCGGCCCTTGTTAACTACAACA GACCATCTGCTGAGAAAATTATACTTACTGAAATGCCAGGAACAACAGAAACCAACGTGGAAAATAACTCTCAGACAGTGTATTACCCAG CTTTATCGGGAAATACGAGTGCCCCATATCCAGCCTCTTCATATCTTCCCATCACTTCTAATTTTGAATCTGGCCCACAAATGAGTTATG GGACAATGAGTTACTCAACTGAAATGAAAAATAACTGTGACCAAGATGATGCTTCAGCATCTGCCTGCCTCACTCCCGATTTTGCACTGT TACCTCTGAATATTTTGGTAAAAGTAGACACCAACACGGAAAACAGCGTCAACACAATGAATCGCTCAACTTTATTGGACAGTGACAGTG GCCAGGATTCTTCCTCATCATCTGTCTGTATCCCTCCCAAGTATGGCTATCTTGGTGATCCAAAAAGAAATGTCAGAGTACTTAAAATCC ATTTGCTGGCCGTACAAAATATGGCCAAACCTAAGCAAGCAGCCTGCTACTTGGTTCGTATTTTGTTCTCCAAGGAAATCCTGATTAGCA GCTCAGTGGATATCCATTTGAAAGACAGCCAATCCCTCGACCCGAACAAAATGGCTGCATTGAGAGAATATCTGGCAACAACTTTTCCCA CCTGTGATTTGCATGAACATGGAAAAGACTGGCAGGACTGTATTTCTGGTATCAATTCTATGATCTACTGTTTATGTTCTGAAGGCAAGA GTACTCCAAAAACTGTTCGCAAAAATAAAAAACGTACCAACCGTGTTGCATCGGCATCTGCAGATAGAAATGACCAAAGGGGCAGAGATG GTGGTGAAGGCTGTTCTTGGATGTTTCAGCCAATGAATAACTCCAAAATGAGAGAAAAGAGGAACTTGCAGCCAAACAGTAATGCTATCC CTGAAGGAATGCGAGAACCTTCCACTGATAATCCAGAGGAACCTGGTGAAGCATGGAGCTATTTTGGAAGACCATGGAGAAATATACGGA TGCCATGTTCAGTACTGACTTTGGCAAAAACTAAGTCTTGCGCAAGCCTGTCGGCTAGATACCTTATTCAGAAACTCTTCACAAAAGATG TCCTGGTCCAAAGTAACGTCTATGGCAATCTGAAGCATGGCCTGTGTGCCCTTGACCCCAATAAGATTAGTGCTCTCCGAGAGTTCCTCC AAGAAAACTACCCAATTTGTGATCTCTCGGAAAATGGAAGAGACTGGAAGTCGTGTGTGACCTCCATCAACAGCGGTATCCGTAGCCTTA GACATGACGTCAGAAGGGCTGAAGCCAGGTCTCAGTCGCTTCCAGCAGTGACCCCTCCAGAGCTGGAGCAGGAGTCAAAGCCAGGAGATC CCGACGCCACTGACCCAAGCACCTGAACGGCAGCTGCCAAACTTTTTTTTTTTTTTTAACTGTTAAGACTATTTTGTAAGTTTCTCAAGT TCTAATTCCAAAATCAGCCTGCTGTACTGGCAGAGTAGTTTGCATTGACACCTGTAGTAATCGCCAAACCTCTTTTTCTGCTCATAAAGG AGGTAGAATCACATTAGGAAGATACGTAATGTTTCATCTGTGGATTTGATGACAACCTCTGATTGAGGGAATGGTAGTAAACTCCTCATG GGCTTGTGACCCATGGCTTTGACTTAGATAATTGACATTAATATACTTTGTTTTTTTGATAAGGTAGAGAAAGAAATTCAATAGAAAGTA AGGAGCCACTCAGTCAAATAATATAATTCGTATAAAACTTGACAAGTCTTTTTATTTTTGTGACTTGTGAGATAGTTTAAGATATTTTTA TGTGACCAATTTGTAGAATGAATACATATTTTCAAAGTTATCTTTTCAAAATTCAACTTTTAGAGTATTTTAGCAAAATATATGTATAGT TTATAATATAGAAGTATAGCTTTATAAAGATATTAAGGATTTATTTTTTAAAATAATTCTCTGAATAGTTTGCTGTAGTAGTTGCCTTCA GCTTAATAAAACCATAGTACTTCTACATACATCTTTATAAGTTAGGCTATGCCTACAAAGGTCTTAAAAAGTAGATAATTTACCGGGTGT AGTGGCTCACACCTGTAATCCCAGCACTTTGGGAGGCCGAGACGGGTGGATCACCTGAGGTCAGGGGTTCGAGACCAGCCTGGCCAACAT GACGAAACCCCATCTCTACTAAAAATACAAAAAAAGTAGCCAGGCATGGTGGCAAGCACCTGTAGTCCCAGTTACTCGGGTGGCTGAGGC AGGAGAATCACTTGAACCTGGGAGGTGGAGACTGCAGTGAGCCGAGATTGTGCCACTGCACTCGAGCCTGGGTGACAGAGTGAGACTCCG TCTCGGGAAAAGAAAAAAAAGAAGTAGATAATTAAAAAAATCTTGCATAAAAAAAAATCTTCCATAAAGGGCAAAAAGGCTTTAACTTCC ATTATTCAGATTTAATCCAACTAAGATAGTCTTTGGCTTCATTCACTGTTCAGTTTCACAAGTTTTTTACAACTACTAGTAACAGAGCTA ATACAGAATAGTAATCCACATTATAAACATAGATCAAAATACCACATCATACCCCATAAATACATGCAGTTATTATTTGTCAATTAAAAA TAAAATTTTTTTAAAAGAAAAGTACAGTTCGACCCTAGGCAAGAAGAAATTTCAAATTGTATATGCATATTTTGAATTTGGTTTATATTT TGTGGTATCTGAAATATTCCCTGGACTAGAAGCAAAACCACAGAAACAAAACTTTACATTTAGAAAAAAATTTCAAGACTTCTATTGCTA ACTTTATTACATTAGTGGTTTTTAGAAGTTATTTAGAGCAGTCTTGTCTGATAGAACTTTCCACCATGATGGAAATGTAAATAAATATAC CACATTAAAAATATAAATTTGCAATGTCCAATAGGCTCGTATGGCTATTGAGCATTTAAGATGTAATTAATTAGACAGAGGAAGTAAATT TAACATTTTCTCTCATTTTAGCTAATTAGAATTTAAAGTGAAATAGCCACAAGTGGATAGTGGCTACTGTGTTGGATGGCACAGTTCAAA ATGCCTCTTATTTATAAAAAAAAGTGTTTAAAAGTGGTTCCCATGGCTTGAAGAGGAATAATATACACATAACTTCTTTAAAATCACATT TTGTTTGAGATGGAGATCTTGCTATATTGCCCAGGCTTATCTGGAACTCCTGGGCTCAAGAGGGAGGATCACATTTGACTGGACAAAATA CATTTTATAAAGGTGTTTTGTATTTTATGTTAAAGTGGAATTGTGTCTTCAAAATTGTGTCTTCAAAATTTCACAAGTTAATACTATTAC TGTATTAGTCCAATGTTAAACATTTTAAATGCCTGCTCATAAGACTCATTTTTGATAAATAAATGTATCTTTAATGTTTTGTTTCCCAAA >83869_83869_4_SMARCA2-BEND2_SMARCA2_chr9_2039900_ENST00000382203_BEND2_chrX_18213562_ENST00000380033_length(amino acids)=718AA_BP=261 MSTPTDPGAMPHPGPSPGPGPSPGPILGPSPGPGPSPGSVHSMMGPSPGPPSVSHPMPTMGSTDFPQEGMHQMHKPIDGIHDKGIVEDIH CGSMKGTGMRPPHPGMGPPQSPMDQHSQGYMSPHPSPLGAPEHVSSPMSGGGPTPPQMPPSQPGALIPGDPQAMSQPNRGPSPFSPVQLH QLRAQILAYKMLARGQPLPETLQLAVQGKRTLPGLQQQQQQQQQQQQQQQQQQQQQQQPQQQPPQPQTQQQQQPALVNYNRPSAEKIILT EMPGTTETNVENNSQTVYYPALSGNTSAPYPASSYLPITSNFESGPQMSYGTMSYSTEMKNNCDQDDASASACLTPDFALLPLNILVKVD TNTENSVNTMNRSTLLDSDSGQDSSSSSVCIPPKYGYLGDPKRNVRVLKIHLLAVQNMAKPKQAACYLVRILFSKEILISSSVDIHLKDS QSLDPNKMAALREYLATTFPTCDLHEHGKDWQDCISGINSMIYCLCSEGKSTPKTVRKNKKRTNRVASASADRNDQRGRDGGEGCSWMFQ PMNNSKMREKRNLQPNSNAIPEGMREPSTDNPEEPGEAWSYFGRPWRNIRMPCSVLTLAKTKSCASLSARYLIQKLFTKDVLVQSNVYGN -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for SMARCA2-BEND2 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for SMARCA2-BEND2 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for SMARCA2-BEND2 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
| Hgene | SMARCA2 | C0036341 | Schizophrenia | 4 | PSYGENET |
| Hgene | SMARCA2 | C1303073 | Nicolaides Baraitser syndrome | 4 | CTD_human;GENOMICS_ENGLAND;ORPHANET;UNIPROT |
| Hgene | SMARCA2 | C0010606 | Adenoid Cystic Carcinoma | 1 | CTD_human |
| Hgene | SMARCA2 | C0265338 | Coffin-Siris syndrome | 1 | CTD_human;GENOMICS_ENGLAND |
| Hgene | SMARCA2 | C0345967 | Malignant mesothelioma | 1 | CTD_human |
| Hgene | SMARCA2 | C3281201 | MENTAL RETARDATION, AUTOSOMAL DOMINANT 12 | 1 | GENOMICS_ENGLAND |
| Tgene | C0087031 | Juvenile-Onset Still Disease | 1 | CTD_human | |
| Tgene | C3495559 | Juvenile arthritis | 1 | CTD_human | |
| Tgene | C3714758 | Juvenile psoriatic arthritis | 1 | CTD_human | |
| Tgene | C4552091 | Polyarthritis, Juvenile, Rheumatoid Factor Negative | 1 | CTD_human | |
| Tgene | C4704862 | Polyarthritis, Juvenile, Rheumatoid Factor Positive | 1 | CTD_human |
