
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:SS18-SSX2 (FusionGDB2 ID:HG6760TG6757) |
Fusion Gene Summary for SS18-SSX2 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: SS18-SSX2 | Fusion gene ID: hg6760tg6757 | Hgene | Tgene | Gene symbol | SS18 | SSX2 | Gene ID | 6760 | 6757 |
| Gene name | SS18 subunit of BAF chromatin remodeling complex | SSX family member 2 | |
| Synonyms | SSXT|SYT | CT5.2|CT5.2A|HD21|HOM-MEL-40|SSX | |
| Cytomap | ('SS18')('SSX2') 18q11.2 | Xp11.22 | |
| Type of gene | protein-coding | protein-coding | |
| Description | protein SSXTSS18, nBAF chromatin remodeling complex subunitsynovial sarcoma translocated to X chromosome proteinsynovial sarcoma translocation, chromosome 18synovial sarcoma, translocated to X chromosome | protein SSX2cancer/testis antigen 5.2cancer/testis antigen family 5, member 2asarcoma, synovial, X-chromosome-related 2synovial sarcoma, X breakpoint 2synovial sarcoma, X breakpoint 2Btumor antigen HOM-MEL-40 | |
| Modification date | 20200329 | 20200313 | |
| UniProtAcc | Q15532 | . | |
| Ensembl transtripts involved in fusion gene | ENST00000269137, ENST00000415083, ENST00000539849, ENST00000542420, ENST00000542743, ENST00000545952, ENST00000585241, | ENST00000269137, ENST00000415083, ENST00000539849, ENST00000542743, ENST00000545952, ENST00000585241, ENST00000542420, | |
| Fusion gene scores | * DoF score | 24 X 33 X 8=6336 | 3 X 17 X 5=255 |
| # samples | 38 | 13 | |
| ** MAII score | log2(38/6336*10)=-4.05950101174866 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(13/255*10)=-0.971985623830404 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: SS18 [Title/Abstract] AND SSX2 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | SS18(23612363)-SSX2(52729628), # samples:8 | ||
| Anticipated loss of major functional domain due to fusion event. | SS18-SSX2 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. SS18-SSX2 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. SS18-SSX2 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. SS18-SSX2 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. SS18-SSX2 seems lost the major protein functional domain in Hgene partner, which is a CGC due to the frame-shifted ORF. SS18-SSX2 seems lost the major protein functional domain in Tgene partner, which is a CGC due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | SS18 | GO:0045944 | positive regulation of transcription by RNA polymerase II | 15919756 |
 Fusion gene breakpoints across SS18 (5'-gene) Fusion gene breakpoints across SS18 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
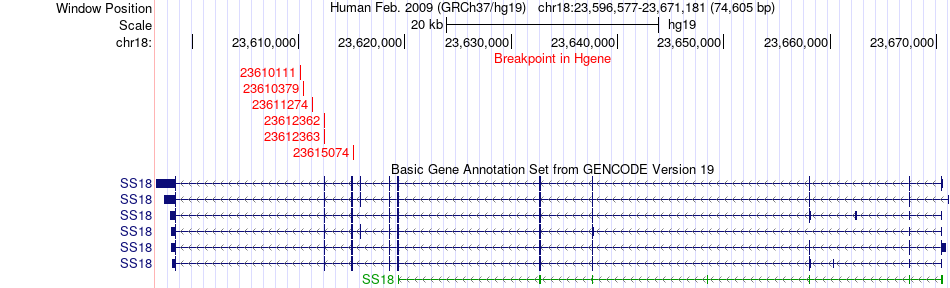 |
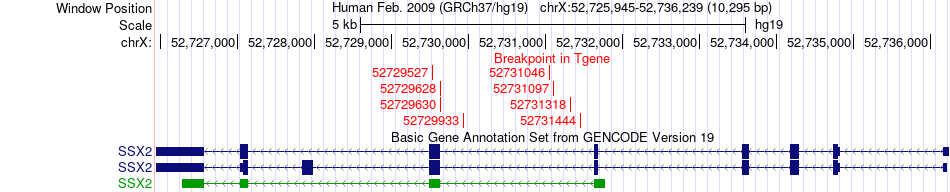
 Fusion gene breakpoints across SSX2 (3'-gene) Fusion gene breakpoints across SSX2 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | SARC | TCGA-DX-A7EO-01A | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| ChimerDB4 | SARC | TCGA-DX-A7ER-01A | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| ChimerDB4 | SARC | TCGA-DX-AB2Z-01A | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| ChimerDB4 | SARC | TCGA-DX-AB2Z-01A | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| ChimerDB4 | SARC | TCGA-DX-AB3C-01A | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| ChimerDB4 | SARC | TCGA-FX-A8OO-01A | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| ChimerDB4 | SARC | TCGA-FX-A8OO | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| ChimerDB4 | SARC | TCGA-WK-A8XT-01A | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| ChimerDB4 | SARC | TCGA-Z4-AAPF-01A | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| ChimerDB4 | synovial sarcoma | AB012575 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| ChimerKB3 | . | . | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| ChimerKB3 | . | . | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| ChimerKB3 | . | . | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| ChimerKB3 | . | . | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| ChimerKB3 | . | . | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| ChimerKB3 | . | . | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| ChimerKB3 | . | . | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| ChimerKB3 | . | . | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| ChimerKB3 | . | . | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| ChimerKB3 | . | . | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| ChimerKB3 | . | . | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| ChimerKB4 | . | . | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| ChiTaRS5.0 | N/A | AB300356 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| ChiTaRS5.0 | N/A | AF230662 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| ChiTaRS5.0 | N/A | AY138488 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| ChiTaRS5.0 | N/A | AY138489 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| ChiTaRS5.0 | N/A | AY138490 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| ChiTaRS5.0 | N/A | AY138491 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| ChiTaRS5.0 | N/A | AY138493 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| ChiTaRS5.0 | N/A | S79332 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| ChiTaRS5.0 | N/A | X79200 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
Top |
Fusion Gene ORF analysis for SS18-SSX2 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| 5CDS-5UTR | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| 5CDS-5UTR | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| 5CDS-5UTR | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| 5CDS-5UTR | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| 5CDS-5UTR | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| 5CDS-5UTR | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| 5CDS-5UTR | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| 5CDS-5UTR | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| 5CDS-5UTR | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| 5CDS-5UTR | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| 5CDS-5UTR | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| 5CDS-5UTR | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| 5CDS-5UTR | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| 5CDS-5UTR | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| 5CDS-5UTR | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| 5CDS-5UTR | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| 5CDS-5UTR | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| 5CDS-5UTR | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| 5CDS-5UTR | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| 5CDS-5UTR | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| 5CDS-5UTR | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| 5CDS-5UTR | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| 5CDS-5UTR | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| 5CDS-intron | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| 5CDS-intron | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| 5CDS-intron | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| 5CDS-intron | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| 5CDS-intron | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| 5CDS-intron | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| 5CDS-intron | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| 5CDS-intron | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| 5CDS-intron | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| 5CDS-intron | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| 5CDS-intron | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| 5CDS-intron | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| 5CDS-intron | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| 5CDS-intron | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| 5CDS-intron | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| 5CDS-intron | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| 5CDS-intron | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| 5CDS-intron | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| 5CDS-intron | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| 5CDS-intron | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| 5CDS-intron | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| 5CDS-intron | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| 5CDS-intron | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| 5CDS-intron | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| 5CDS-intron | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| 5CDS-intron | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| 5CDS-intron | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| 5CDS-intron | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| 5CDS-intron | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| 5CDS-intron | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| 5CDS-intron | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| 5CDS-intron | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| 5CDS-intron | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| 5CDS-intron | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| 5CDS-intron | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| 5CDS-intron | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| 5CDS-intron | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| 5CDS-intron | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| 5CDS-intron | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| 5CDS-intron | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| 5CDS-intron | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| 5CDS-intron | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| 5CDS-intron | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| 5CDS-intron | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| 5CDS-intron | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| 5CDS-intron | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| 5CDS-intron | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| 5CDS-intron | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| 5CDS-intron | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| 5CDS-intron | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| 5CDS-intron | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| 5CDS-intron | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| 5CDS-intron | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| 5CDS-intron | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| 5CDS-intron | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| 5CDS-intron | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| 5CDS-intron | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| 5CDS-intron | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| 5CDS-intron | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| 5CDS-intron | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| 5CDS-intron | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| 5CDS-intron | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| 5CDS-intron | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| 5CDS-intron | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| 5CDS-intron | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| 5CDS-intron | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| 5CDS-intron | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| 5CDS-intron | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| 5CDS-intron | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| 5CDS-intron | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| 5CDS-intron | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| 5CDS-intron | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| 5UTR-3CDS | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| 5UTR-3CDS | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| 5UTR-5UTR | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| Frame-shift | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| Frame-shift | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| Frame-shift | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| Frame-shift | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| Frame-shift | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| Frame-shift | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| Frame-shift | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| Frame-shift | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| Frame-shift | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| Frame-shift | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| Frame-shift | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| Frame-shift | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| Frame-shift | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| Frame-shift | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| Frame-shift | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| Frame-shift | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| Frame-shift | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| Frame-shift | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| Frame-shift | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| Frame-shift | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| Frame-shift | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| Frame-shift | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| Frame-shift | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| Frame-shift | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| In-frame | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| In-frame | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| In-frame | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| In-frame | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| In-frame | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| In-frame | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| In-frame | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| In-frame | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| In-frame | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| In-frame | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| In-frame | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| In-frame | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| In-frame | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| In-frame | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| In-frame | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| In-frame | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| In-frame | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| In-frame | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| In-frame | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| In-frame | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| In-frame | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| In-frame | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| In-frame | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23637552 | - | SSX2 | chrX | 52731680 | - |
| In-frame | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| intron-3CDS | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-3CDS | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-3CDS | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-3CDS | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-3CDS | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-3CDS | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-3CDS | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-3CDS | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-3CDS | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-3CDS | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-3CDS | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-3CDS | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-3CDS | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-3CDS | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-3CDS | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-3CDS | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-3CDS | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-3CDS | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-3CDS | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-3CDS | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-3CDS | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-3CDS | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-3CDS | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-3CDS | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-3CDS | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-3CDS | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-3CDS | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-3CDS | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-3CDS | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-3CDS | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-3CDS | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-3CDS | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-3CDS | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-3CDS | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-3CDS | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-3CDS | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-3CDS | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| intron-3CDS | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-3CDS | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| intron-3CDS | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-3CDS | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-3CDS | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| intron-3CDS | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| intron-3CDS | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-3CDS | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| intron-3CDS | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-3CDS | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-3CDS | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| intron-3UTR | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-3UTR | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-3UTR | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-3UTR | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-3UTR | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-3UTR | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-3UTR | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-3UTR | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-3UTR | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-3UTR | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-3UTR | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-3UTR | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-3UTR | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-3UTR | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-3UTR | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-3UTR | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-5UTR | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-5UTR | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-5UTR | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-5UTR | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-5UTR | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-5UTR | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-5UTR | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-5UTR | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-5UTR | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-5UTR | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-5UTR | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-5UTR | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-5UTR | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-5UTR | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-5UTR | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-5UTR | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-5UTR | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-5UTR | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-5UTR | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-5UTR | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-5UTR | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-5UTR | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-5UTR | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-5UTR | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-5UTR | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-5UTR | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-5UTR | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-5UTR | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-5UTR | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-5UTR | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-5UTR | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - |
| intron-5UTR | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 23615074 | SSX2 | chrX | 52729527 | ||
| intron-5UTR | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52729628 | - |
| intron-5UTR | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 23612390 | - | SSX2 | chrX | 52729539 | - |
| intron-5UTR | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52726465 | - |
| intron-5UTR | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 23615074 | - | SSX2 | chrX | 52729527 | - |
| intron-5UTR | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52726465 | - |
| intron-5UTR | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - |
| intron-intron | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000269137 | ENST00000336777 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000269137 | ENST00000337502 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-intron | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000269137 | ENST00000476392 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000415083 | ENST00000336777 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000415083 | ENST00000337502 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-intron | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000415083 | ENST00000476392 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000539849 | ENST00000336777 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000539849 | ENST00000337502 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-intron | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000539849 | ENST00000476392 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000542420 | ENST00000336777 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000542420 | ENST00000337502 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-intron | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000542420 | ENST00000476392 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000542743 | ENST00000336777 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000542743 | ENST00000337502 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-intron | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000542743 | ENST00000476392 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000545952 | ENST00000336777 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000545952 | ENST00000337502 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-intron | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000545952 | ENST00000476392 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| intron-intron | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| intron-intron | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| intron-intron | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| intron-intron | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000585241 | ENST00000336777 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| intron-intron | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| intron-intron | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| intron-intron | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| intron-intron | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000585241 | ENST00000337502 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
| intron-intron | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52813117 | - |
| intron-intron | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | + |
| intron-intron | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 23612362 | - | SSX2 | chrX | 52789420 | - |
| intron-intron | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52786931 | - |
| intron-intron | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 23612422 | - | SSX2 | chrX | 52726158 | - |
| intron-intron | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 23670542 | - | SSX2 | chrX | 52727038 | - |
| intron-intron | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 23610111 | - | SSX2 | chrX | 52729933 | - |
| intron-intron | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 23611274 | - | SSX2 | chrX | 52731046 | - |
| intron-intron | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 23610379 | - | SSX2 | chrX | 52731318 | - |
| intron-intron | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731444 | - |
| intron-intron | ENST00000585241 | ENST00000476392 | SS18 | chr18 | 22983419 | - | SSX2 | chrX | 52731097 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000415083 | SS18 | chr18 | 23612362 | - | ENST00000336777 | SSX2 | chrX | 52729628 | - | 2296 | 1286 | 56 | 1627 | 523 |
| ENST00000415083 | SS18 | chr18 | 23612362 | - | ENST00000337502 | SSX2 | chrX | 52729628 | - | 2150 | 1286 | 56 | 1522 | 488 |
| ENST00000269137 | SS18 | chr18 | 23612362 | - | ENST00000336777 | SSX2 | chrX | 52729628 | - | 2479 | 1469 | 161 | 1810 | 549 |
| ENST00000269137 | SS18 | chr18 | 23612362 | - | ENST00000337502 | SSX2 | chrX | 52729628 | - | 2333 | 1469 | 161 | 1705 | 514 |
| ENST00000545952 | SS18 | chr18 | 23612362 | - | ENST00000336777 | SSX2 | chrX | 52729628 | - | 2294 | 1284 | 303 | 1625 | 440 |
| ENST00000545952 | SS18 | chr18 | 23612362 | - | ENST00000337502 | SSX2 | chrX | 52729628 | - | 2148 | 1284 | 303 | 1520 | 405 |
| ENST00000415083 | SS18 | chr18 | 23612363 | - | ENST00000336777 | SSX2 | chrX | 52729628 | - | 2296 | 1286 | 56 | 1627 | 523 |
| ENST00000415083 | SS18 | chr18 | 23612363 | - | ENST00000337502 | SSX2 | chrX | 52729628 | - | 2150 | 1286 | 56 | 1522 | 488 |
| ENST00000269137 | SS18 | chr18 | 23612363 | - | ENST00000336777 | SSX2 | chrX | 52729628 | - | 2479 | 1469 | 161 | 1810 | 549 |
| ENST00000269137 | SS18 | chr18 | 23612363 | - | ENST00000337502 | SSX2 | chrX | 52729628 | - | 2333 | 1469 | 161 | 1705 | 514 |
| ENST00000545952 | SS18 | chr18 | 23612363 | - | ENST00000336777 | SSX2 | chrX | 52729628 | - | 2294 | 1284 | 303 | 1625 | 440 |
| ENST00000545952 | SS18 | chr18 | 23612363 | - | ENST00000337502 | SSX2 | chrX | 52729628 | - | 2148 | 1284 | 303 | 1520 | 405 |
| ENST00000415083 | SS18 | chr18 | 23637552 | - | ENST00000336777 | SSX2 | chrX | 52731680 | - | 1501 | 441 | 56 | 832 | 258 |
| ENST00000415083 | SS18 | chr18 | 23637552 | - | ENST00000337502 | SSX2 | chrX | 52731680 | - | 1355 | 441 | 56 | 727 | 223 |
| ENST00000269137 | SS18 | chr18 | 23637552 | - | ENST00000336777 | SSX2 | chrX | 52731680 | - | 1777 | 717 | 161 | 1108 | 315 |
| ENST00000269137 | SS18 | chr18 | 23637552 | - | ENST00000337502 | SSX2 | chrX | 52731680 | - | 1631 | 717 | 161 | 1003 | 280 |
| ENST00000545952 | SS18 | chr18 | 23637552 | - | ENST00000336777 | SSX2 | chrX | 52731680 | - | 1592 | 532 | 303 | 923 | 206 |
| ENST00000545952 | SS18 | chr18 | 23637552 | - | ENST00000337502 | SSX2 | chrX | 52731680 | - | 1446 | 532 | 303 | 818 | 171 |
| ENST00000415083 | SS18 | chr18 | 23612363 | - | ENST00000336777 | SSX2 | chrX | 52729628 | - | 2296 | 1286 | 56 | 1627 | 523 |
| ENST00000415083 | SS18 | chr18 | 23612363 | - | ENST00000337502 | SSX2 | chrX | 52729628 | - | 2150 | 1286 | 56 | 1522 | 488 |
| ENST00000269137 | SS18 | chr18 | 23612363 | - | ENST00000336777 | SSX2 | chrX | 52729628 | - | 2479 | 1469 | 161 | 1810 | 549 |
| ENST00000269137 | SS18 | chr18 | 23612363 | - | ENST00000337502 | SSX2 | chrX | 52729628 | - | 2333 | 1469 | 161 | 1705 | 514 |
| ENST00000545952 | SS18 | chr18 | 23612363 | - | ENST00000336777 | SSX2 | chrX | 52729628 | - | 2294 | 1284 | 303 | 1625 | 440 |
| ENST00000545952 | SS18 | chr18 | 23612363 | - | ENST00000337502 | SSX2 | chrX | 52729628 | - | 2148 | 1284 | 303 | 1520 | 405 |
| ENST00000415083 | SS18 | chr18 | 23612363 | - | ENST00000336777 | SSX2 | chrX | 52729630 | - | 2296 | 1286 | 56 | 1627 | 523 |
| ENST00000415083 | SS18 | chr18 | 23612363 | - | ENST00000337502 | SSX2 | chrX | 52729630 | - | 2150 | 1286 | 56 | 1522 | 488 |
| ENST00000269137 | SS18 | chr18 | 23612363 | - | ENST00000336777 | SSX2 | chrX | 52729630 | - | 2479 | 1469 | 161 | 1810 | 549 |
| ENST00000269137 | SS18 | chr18 | 23612363 | - | ENST00000337502 | SSX2 | chrX | 52729630 | - | 2333 | 1469 | 161 | 1705 | 514 |
| ENST00000545952 | SS18 | chr18 | 23612363 | - | ENST00000336777 | SSX2 | chrX | 52729630 | - | 2294 | 1284 | 303 | 1625 | 440 |
| ENST00000545952 | SS18 | chr18 | 23612363 | - | ENST00000337502 | SSX2 | chrX | 52729630 | - | 2148 | 1284 | 303 | 1520 | 405 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - | 0.006816891 | 0.9931831 |
| ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - | 0.003513192 | 0.9964868 |
| ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - | 0.006717681 | 0.9932823 |
| ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - | 0.00366823 | 0.99633175 |
| ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - | 0.003093663 | 0.99690634 |
| ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729628 | - | 0.001545619 | 0.99845433 |
| ENST00000415083 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - | 0.006816891 | 0.9931831 |
| ENST00000415083 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - | 0.003513192 | 0.9964868 |
| ENST00000269137 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - | 0.006717681 | 0.9932823 |
| ENST00000269137 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - | 0.00366823 | 0.99633175 |
| ENST00000545952 | ENST00000336777 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - | 0.003093663 | 0.99690634 |
| ENST00000545952 | ENST00000337502 | SS18 | chr18 | 23612363 | - | SSX2 | chrX | 52729630 | - | 0.001545619 | 0.99845433 |
Top |
Fusion Genomic Features for SS18-SSX2 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
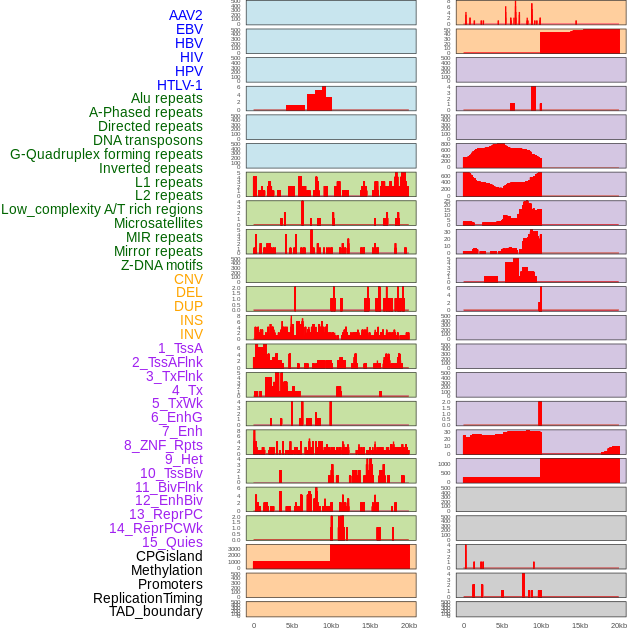 |
Top |
Fusion Protein Features for SS18-SSX2 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr18:23612363/chrX:52729628) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| SS18 | . |
| FUNCTION: Appears to function synergistically with RBM14 as a transcriptional coactivator. Isoform 1 and isoform 2 function in nuclear receptor coactivation. Isoform 1 and isoform 2 function in general transcriptional coactivation. Component of SWI/SNF chromatin remodeling subcomplex GBAF that carries out key enzymatic activities, changing chromatin structure by altering DNA-histone contacts within a nucleosome in an ATP-dependent manner (PubMed:29374058). {ECO:0000269|PubMed:15919756, ECO:0000269|PubMed:29374058}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000269137 | - | 9 | 10 | 95_99 | 379 | 388.0 | Compositional bias | Note=Poly-Pro |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000415083 | - | 10 | 11 | 95_99 | 410 | 419.0 | Compositional bias | Note=Poly-Pro |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000269137 | - | 9 | 10 | 95_99 | 379 | 388.0 | Compositional bias | Note=Poly-Pro |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000415083 | - | 10 | 11 | 95_99 | 410 | 419.0 | Compositional bias | Note=Poly-Pro |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000269137 | - | 9 | 10 | 374_377 | 379 | 388.0 | Motif | SH2-binding |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000269137 | - | 9 | 10 | 50_53 | 379 | 388.0 | Motif | SH2-binding |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000415083 | - | 10 | 11 | 374_377 | 410 | 419.0 | Motif | SH2-binding |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000415083 | - | 10 | 11 | 392_401 | 410 | 419.0 | Motif | SH3-binding |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000415083 | - | 10 | 11 | 50_53 | 410 | 419.0 | Motif | SH2-binding |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000269137 | - | 9 | 10 | 374_377 | 379 | 388.0 | Motif | SH2-binding |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000269137 | - | 9 | 10 | 50_53 | 379 | 388.0 | Motif | SH2-binding |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000415083 | - | 10 | 11 | 374_377 | 410 | 419.0 | Motif | SH2-binding |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000415083 | - | 10 | 11 | 392_401 | 410 | 419.0 | Motif | SH3-binding |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000415083 | - | 10 | 11 | 50_53 | 410 | 419.0 | Motif | SH2-binding |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000269137 | - | 9 | 10 | 2_186 | 379 | 388.0 | Region | Note=Transcriptional activation |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000269137 | - | 9 | 10 | 344_369 | 379 | 388.0 | Region | Note=2 X 13 AA imperfect tandem repeats |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000415083 | - | 10 | 11 | 2_186 | 410 | 419.0 | Region | Note=Transcriptional activation |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000415083 | - | 10 | 11 | 344_369 | 410 | 419.0 | Region | Note=2 X 13 AA imperfect tandem repeats |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000269137 | - | 9 | 10 | 2_186 | 379 | 388.0 | Region | Note=Transcriptional activation |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000269137 | - | 9 | 10 | 344_369 | 379 | 388.0 | Region | Note=2 X 13 AA imperfect tandem repeats |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000415083 | - | 10 | 11 | 2_186 | 410 | 419.0 | Region | Note=Transcriptional activation |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000415083 | - | 10 | 11 | 344_369 | 410 | 419.0 | Region | Note=2 X 13 AA imperfect tandem repeats |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000269137 | - | 9 | 10 | 344_356 | 379 | 388.0 | Repeat | Note=1 |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000269137 | - | 9 | 10 | 357_369 | 379 | 388.0 | Repeat | Note=2 |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000415083 | - | 10 | 11 | 344_356 | 410 | 419.0 | Repeat | Note=1 |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000415083 | - | 10 | 11 | 357_369 | 410 | 419.0 | Repeat | Note=2 |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000269137 | - | 9 | 10 | 344_356 | 379 | 388.0 | Repeat | Note=1 |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000269137 | - | 9 | 10 | 357_369 | 379 | 388.0 | Repeat | Note=2 |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000415083 | - | 10 | 11 | 344_356 | 410 | 419.0 | Repeat | Note=1 |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000415083 | - | 10 | 11 | 357_369 | 410 | 419.0 | Repeat | Note=2 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000269137 | - | 9 | 10 | 175_418 | 379 | 388.0 | Compositional bias | Note=Gln-rich |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000415083 | - | 10 | 11 | 175_418 | 410 | 419.0 | Compositional bias | Note=Gln-rich |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000269137 | - | 9 | 10 | 175_418 | 379 | 388.0 | Compositional bias | Note=Gln-rich |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000415083 | - | 10 | 11 | 175_418 | 410 | 419.0 | Compositional bias | Note=Gln-rich |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000269137 | - | 9 | 10 | 392_401 | 379 | 388.0 | Motif | SH3-binding |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000269137 | - | 9 | 10 | 413_416 | 379 | 388.0 | Motif | SH2-binding |
| Hgene | SS18 | chr18:23612363 | chrX:52729628 | ENST00000415083 | - | 10 | 11 | 413_416 | 410 | 419.0 | Motif | SH2-binding |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000269137 | - | 9 | 10 | 392_401 | 379 | 388.0 | Motif | SH3-binding |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000269137 | - | 9 | 10 | 413_416 | 379 | 388.0 | Motif | SH2-binding |
| Hgene | SS18 | chr18:23612363 | chrX:52729630 | ENST00000415083 | - | 10 | 11 | 413_416 | 410 | 419.0 | Motif | SH2-binding |
| Tgene | SSX2 | chr18:23612363 | chrX:52729628 | ENST00000336777 | 4 | 9 | 20_83 | 110 | 420.3333333333333 | Domain | KRAB-related | |
| Tgene | SSX2 | chr18:23612363 | chrX:52729628 | ENST00000337502 | 4 | 8 | 20_83 | 110 | 376.0 | Domain | KRAB-related | |
| Tgene | SSX2 | chr18:23612363 | chrX:52729630 | ENST00000336777 | 4 | 9 | 20_83 | 110 | 420.3333333333333 | Domain | KRAB-related | |
| Tgene | SSX2 | chr18:23612363 | chrX:52729630 | ENST00000337502 | 4 | 8 | 20_83 | 110 | 376.0 | Domain | KRAB-related |
Top |
Fusion Gene Sequence for SS18-SSX2 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >86683_86683_1_SS18-SSX2_SS18_chr18_23612362_ENST00000269137_SSX2_chrX_52729628_ENST00000336777_length(transcript)=2479nt_BP=1469nt TTTCCTGAATCGCTGAAGCGGAGGGAGGACTAGGGCGCGACTCTACCATTAGCGCAAAGGGCCAGGGGGTGCCCATAAGGAGAGCTCGGG GGATGCAGGGACGGTCAAGAAAGTTCGGCCAGGGCTAGGGGTCCCCCAGAGTGCGCAGAAGCGAGACATCCCTGGCCAGAGTGCCTCCTT CAGGCCTGCCCCGCCGCCCGTCCGCTTGGCCCCGGAGCATCCCCCCTCCCCGCATTCCCGAGGTGGTTCAGCCCGAGAGGCCGGCGTCTC TCCCCCAGTTTGCCGTTCACCCGGAGCGCTCGGGACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGC AGCGAGGCAAGGGGGAGATCACTCCCGCTGCGATTCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGA ATAAAGGAAAGACCTCAGAGTGTTCTCAGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAA ATATGCAGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCAC GCTCTCACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGGGC CTAACCATATGCCTATGCAGGGACCTGGACCCAATCAACTCAATATGACAAACAGTTCCATGAATATGCCTTCAAGTAGCCATGGATCCA TGGGAGGTTACAACCATTCTGTGCCATCATCACAGAGCATGCCAGTACAGAATCAGATGACAATGAGTCAGGGACAACCAATGGGAAACT ATGGTCCCAGACCAAATATGAGTATGCAGCCAAACCAAGGTCCAATGATGCATCAGCAGCCTCCTTCTCAGCAATACAATATGCCACAGG GAGGCGGACAGCATTACCAAGGACAGCAGCCACCTATGGGAATGATGGGTCAAGTTAACCAAGGCAATCATATGATGGGTCAGAGACAGA TTCCTCCCTATAGACCTCCTCAACAGGGCCCACCACAGCAGTACTCAGGCCAGGAAGACTATTACGGGGACCAATACAGTCATGGTGGAC AAGGTCCTCCAGAAGGCATGAACCAGCAATATTACCCTGATGGAAATTCACAGTATGGCCAACAGCAAGATGCATACCAGGGACCACCTC CACAACAGGGATATCCACCCCAGCAGCAGCAGTACCCAGGGCAGCAAGGTTACCCAGGACAGCAGCAGGGCTACGGTCCTTCACAGGGTG GTCCAGGTCCTCAGTATCCTAACTACCCACAGGGACAAGGTCAGCAGTATGGAGGATATAGACCAACACAGCCTGGACCACCACAGCCAC CCCAGCAGAGGCCTTATGGATATGACCAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGCCAGAAGCATCTG GCCCACAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGAAATAGGGAGGCCC AAGAAAAGGAAGAGAGACGCGGAACAGCTCATCGGTGGAGCAGTCAGAACACACACAACATTGGTCGATTCAGTTTGTCAACTTCTATGG GTGCAGTTCATGGTACCCCCAAAACAATTACACACAACAGGGACCCAAAAGGGGGGAACATGCCTGGACCCACAGACTGCGTGAGAGAAA ACAGCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCCCCTCAGGGATACGACACATGCCCATGATGAGAAG CAGAACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAGGTGCATAGCAAGTGAAAGCAAGTGTTCACAACAG TGAAAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTACGTTGTATTTTCTTACACTGTGTCATTCTGTTAGA TACTAACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTATCCATGCACCTACCTTAGAAAACAAGTATTGTCG GTTACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAGGGCAGTTCTGAGTGTTTAATTTCAGATTTTTTCC TCTGCATTTACACACACACGCACACAAACCACACCACACACACACACACACACACACACACACACACACACACACACCAAGTACCAGTAT AAGCATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCACTCTGTTTCCTGGTCAGCATGTACTCCCCTCATCC >86683_86683_1_SS18-SSX2_SS18_chr18_23612362_ENST00000269137_SSX2_chrX_52729628_ENST00000336777_length(amino acids)=549AA_BP=436 MARVPPSGLPRRPSAWPRSIPPPRIPEVVQPERPASLPQFAVHPERSGLADSGDGGNMSVAFAAPRQRGKGEITPAAIQKMLDDNNHLIQ CIMDSQNKGKTSECSQYQQMLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGMNQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQM NGQMPGPNHMPMQGPGPNQLNMTNSSMNMPSSSHGSMGGYNHSVPSSQSMPVQNQMTMSQGQPMGNYGPRPNMSMQPNQGPMMHQQPPSQ QYNMPQGGGQHYQGQQPPMGMMGQVNQGNHMMGQRQIPPYRPPQQGPPQQYSGQEDYYGDQYSHGGQGPPEGMNQQYYPDGNSQYGQQQD AYQGPPPQQGYPPQQQQYPGQQGYPGQQQGYGPSQGGPGPQYPNYPQGQGQQYGGYRPTQPGPPQPPQQRPYGYDQIMPKKPAEEGNDSE EVPEASGPQNDGKELCPPGKPTTSEKIHERSGNREAQEKEERRGTAHRWSSQNTHNIGRFSLSTSMGAVHGTPKTITHNRDPKGGNMPGP -------------------------------------------------------------- >86683_86683_2_SS18-SSX2_SS18_chr18_23612362_ENST00000269137_SSX2_chrX_52729628_ENST00000337502_length(transcript)=2333nt_BP=1469nt TTTCCTGAATCGCTGAAGCGGAGGGAGGACTAGGGCGCGACTCTACCATTAGCGCAAAGGGCCAGGGGGTGCCCATAAGGAGAGCTCGGG GGATGCAGGGACGGTCAAGAAAGTTCGGCCAGGGCTAGGGGTCCCCCAGAGTGCGCAGAAGCGAGACATCCCTGGCCAGAGTGCCTCCTT CAGGCCTGCCCCGCCGCCCGTCCGCTTGGCCCCGGAGCATCCCCCCTCCCCGCATTCCCGAGGTGGTTCAGCCCGAGAGGCCGGCGTCTC TCCCCCAGTTTGCCGTTCACCCGGAGCGCTCGGGACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGC AGCGAGGCAAGGGGGAGATCACTCCCGCTGCGATTCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGA ATAAAGGAAAGACCTCAGAGTGTTCTCAGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAA ATATGCAGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCAC GCTCTCACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGGGC CTAACCATATGCCTATGCAGGGACCTGGACCCAATCAACTCAATATGACAAACAGTTCCATGAATATGCCTTCAAGTAGCCATGGATCCA TGGGAGGTTACAACCATTCTGTGCCATCATCACAGAGCATGCCAGTACAGAATCAGATGACAATGAGTCAGGGACAACCAATGGGAAACT ATGGTCCCAGACCAAATATGAGTATGCAGCCAAACCAAGGTCCAATGATGCATCAGCAGCCTCCTTCTCAGCAATACAATATGCCACAGG GAGGCGGACAGCATTACCAAGGACAGCAGCCACCTATGGGAATGATGGGTCAAGTTAACCAAGGCAATCATATGATGGGTCAGAGACAGA TTCCTCCCTATAGACCTCCTCAACAGGGCCCACCACAGCAGTACTCAGGCCAGGAAGACTATTACGGGGACCAATACAGTCATGGTGGAC AAGGTCCTCCAGAAGGCATGAACCAGCAATATTACCCTGATGGAAATTCACAGTATGGCCAACAGCAAGATGCATACCAGGGACCACCTC CACAACAGGGATATCCACCCCAGCAGCAGCAGTACCCAGGGCAGCAAGGTTACCCAGGACAGCAGCAGGGCTACGGTCCTTCACAGGGTG GTCCAGGTCCTCAGTATCCTAACTACCCACAGGGACAAGGTCAGCAGTATGGAGGATATAGACCAACACAGCCTGGACCACCACAGCCAC CCCAGCAGAGGCCTTATGGATATGACCAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGCCAGAAGCATCTG GCCCACAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGACCCAAAAGGGGGG AACATGCCTGGACCCACAGACTGCGTGAGAGAAAACAGCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCC CCTCAGGGATACGACACATGCCCATGATGAGAAGCAGAACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAG GTGCATAGCAAGTGAAAGCAAGTGTTCACAACAGTGAAAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTAC GTTGTATTTTCTTACACTGTGTCATTCTGTTAGATACTAACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTA TCCATGCACCTACCTTAGAAAACAAGTATTGTCGGTTACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAG GGCAGTTCTGAGTGTTTAATTTCAGATTTTTTCCTCTGCATTTACACACACACGCACACAAACCACACCACACACACACACACACACACA CACACACACACACACACACACCAAGTACCAGTATAAGCATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCAC >86683_86683_2_SS18-SSX2_SS18_chr18_23612362_ENST00000269137_SSX2_chrX_52729628_ENST00000337502_length(amino acids)=514AA_BP=436 MARVPPSGLPRRPSAWPRSIPPPRIPEVVQPERPASLPQFAVHPERSGLADSGDGGNMSVAFAAPRQRGKGEITPAAIQKMLDDNNHLIQ CIMDSQNKGKTSECSQYQQMLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGMNQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQM NGQMPGPNHMPMQGPGPNQLNMTNSSMNMPSSSHGSMGGYNHSVPSSQSMPVQNQMTMSQGQPMGNYGPRPNMSMQPNQGPMMHQQPPSQ QYNMPQGGGQHYQGQQPPMGMMGQVNQGNHMMGQRQIPPYRPPQQGPPQQYSGQEDYYGDQYSHGGQGPPEGMNQQYYPDGNSQYGQQQD AYQGPPPQQGYPPQQQQYPGQQGYPGQQQGYGPSQGGPGPQYPNYPQGQGQQYGGYRPTQPGPPQPPQQRPYGYDQIMPKKPAEEGNDSE -------------------------------------------------------------- >86683_86683_3_SS18-SSX2_SS18_chr18_23612362_ENST00000415083_SSX2_chrX_52729628_ENST00000336777_length(transcript)=2296nt_BP=1286nt AGTTTGCCGTTCACCCGGAGCGCTCGGGACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGCAGCGAG GCAAGGGGGAGATCACTCCCGCTGCGATTCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGAATAAAG GAAAGACCTCAGAGTGTTCTCAGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAAATATGC AGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCACGCTCTC ACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGGGCCTAACC ATATGCCTATGCAGGGACCTGGACCCAATCAACTCAATATGACAAACAGTTCCATGAATATGCCTTCAAGTAGCCATGGATCCATGGGAG GTTACAACCATTCTGTGCCATCATCACAGAGCATGCCAGTACAGAATCAGATGACAATGAGTCAGGGACAACCAATGGGAAACTATGGTC CCAGACCAAATATGAGTATGCAGCCAAACCAAGGTCCAATGATGCATCAGCAGCCTCCTTCTCAGCAATACAATATGCCACAGGGAGGCG GACAGCATTACCAAGGACAGCAGCCACCTATGGGAATGATGGGTCAAGTTAACCAAGGCAATCATATGATGGGTCAGAGACAGATTCCTC CCTATAGACCTCCTCAACAGGGCCCACCACAGCAGTACTCAGGCCAGGAAGACTATTACGGGGACCAATACAGTCATGGTGGACAAGGTC CTCCAGAAGGCATGAACCAGCAATATTACCCTGATGGTCATAATGATTACGGTTATCAGCAACCGTCGTATCCTGAACAAGGCTACGATA GGCCTTATGAGGATTCCTCACAACATTACTACGAAGGAGGAAATTCACAGTATGGCCAACAGCAAGATGCATACCAGGGACCACCTCCAC AACAGGGATATCCACCCCAGCAGCAGCAGTACCCAGGGCAGCAAGGTTACCCAGGACAGCAGCAGGGCTACGGTCCTTCACAGGGTGGTC CAGGTCCTCAGTATCCTAACTACCCACAGGGACAAGGTCAGCAGTATGGAGGATATAGACCAACACAGCCTGGACCACCACAGCCACCCC AGCAGAGGCCTTATGGATATGACCAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGCCAGAAGCATCTGGCC CACAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGAAATAGGGAGGCCCAAG AAAAGGAAGAGAGACGCGGAACAGCTCATCGGTGGAGCAGTCAGAACACACACAACATTGGTCGATTCAGTTTGTCAACTTCTATGGGTG CAGTTCATGGTACCCCCAAAACAATTACACACAACAGGGACCCAAAAGGGGGGAACATGCCTGGACCCACAGACTGCGTGAGAGAAAACA GCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCCCCTCAGGGATACGACACATGCCCATGATGAGAAGCAG AACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAGGTGCATAGCAAGTGAAAGCAAGTGTTCACAACAGTGA AAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTACGTTGTATTTTCTTACACTGTGTCATTCTGTTAGATAC TAACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTATCCATGCACCTACCTTAGAAAACAAGTATTGTCGGTT ACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAGGGCAGTTCTGAGTGTTTAATTTCAGATTTTTTCCTCT GCATTTACACACACACGCACACAAACCACACCACACACACACACACACACACACACACACACACACACACACACCAAGTACCAGTATAAG CATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCACTCTGTTTCCTGGTCAGCATGTACTCCCCTCATCCGAT >86683_86683_3_SS18-SSX2_SS18_chr18_23612362_ENST00000415083_SSX2_chrX_52729628_ENST00000336777_length(amino acids)=523AA_BP=410 MSVAFAAPRQRGKGEITPAAIQKMLDDNNHLIQCIMDSQNKGKTSECSQYQQMLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGM NQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQMNGQMPGPNHMPMQGPGPNQLNMTNSSMNMPSSSHGSMGGYNHSVPSSQSMPVQNQMT MSQGQPMGNYGPRPNMSMQPNQGPMMHQQPPSQQYNMPQGGGQHYQGQQPPMGMMGQVNQGNHMMGQRQIPPYRPPQQGPPQQYSGQEDY YGDQYSHGGQGPPEGMNQQYYPDGHNDYGYQQPSYPEQGYDRPYEDSSQHYYEGGNSQYGQQQDAYQGPPPQQGYPPQQQQYPGQQGYPG QQQGYGPSQGGPGPQYPNYPQGQGQQYGGYRPTQPGPPQPPQQRPYGYDQIMPKKPAEEGNDSEEVPEASGPQNDGKELCPPGKPTTSEK -------------------------------------------------------------- >86683_86683_4_SS18-SSX2_SS18_chr18_23612362_ENST00000415083_SSX2_chrX_52729628_ENST00000337502_length(transcript)=2150nt_BP=1286nt AGTTTGCCGTTCACCCGGAGCGCTCGGGACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGCAGCGAG GCAAGGGGGAGATCACTCCCGCTGCGATTCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGAATAAAG GAAAGACCTCAGAGTGTTCTCAGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAAATATGC AGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCACGCTCTC ACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGGGCCTAACC ATATGCCTATGCAGGGACCTGGACCCAATCAACTCAATATGACAAACAGTTCCATGAATATGCCTTCAAGTAGCCATGGATCCATGGGAG GTTACAACCATTCTGTGCCATCATCACAGAGCATGCCAGTACAGAATCAGATGACAATGAGTCAGGGACAACCAATGGGAAACTATGGTC CCAGACCAAATATGAGTATGCAGCCAAACCAAGGTCCAATGATGCATCAGCAGCCTCCTTCTCAGCAATACAATATGCCACAGGGAGGCG GACAGCATTACCAAGGACAGCAGCCACCTATGGGAATGATGGGTCAAGTTAACCAAGGCAATCATATGATGGGTCAGAGACAGATTCCTC CCTATAGACCTCCTCAACAGGGCCCACCACAGCAGTACTCAGGCCAGGAAGACTATTACGGGGACCAATACAGTCATGGTGGACAAGGTC CTCCAGAAGGCATGAACCAGCAATATTACCCTGATGGTCATAATGATTACGGTTATCAGCAACCGTCGTATCCTGAACAAGGCTACGATA GGCCTTATGAGGATTCCTCACAACATTACTACGAAGGAGGAAATTCACAGTATGGCCAACAGCAAGATGCATACCAGGGACCACCTCCAC AACAGGGATATCCACCCCAGCAGCAGCAGTACCCAGGGCAGCAAGGTTACCCAGGACAGCAGCAGGGCTACGGTCCTTCACAGGGTGGTC CAGGTCCTCAGTATCCTAACTACCCACAGGGACAAGGTCAGCAGTATGGAGGATATAGACCAACACAGCCTGGACCACCACAGCCACCCC AGCAGAGGCCTTATGGATATGACCAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGCCAGAAGCATCTGGCC CACAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGACCCAAAAGGGGGGAAC ATGCCTGGACCCACAGACTGCGTGAGAGAAAACAGCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCCCCT CAGGGATACGACACATGCCCATGATGAGAAGCAGAACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAGGTG CATAGCAAGTGAAAGCAAGTGTTCACAACAGTGAAAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTACGTT GTATTTTCTTACACTGTGTCATTCTGTTAGATACTAACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTATCC ATGCACCTACCTTAGAAAACAAGTATTGTCGGTTACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAGGGC AGTTCTGAGTGTTTAATTTCAGATTTTTTCCTCTGCATTTACACACACACGCACACAAACCACACCACACACACACACACACACACACAC ACACACACACACACACACCAAGTACCAGTATAAGCATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCACTCT >86683_86683_4_SS18-SSX2_SS18_chr18_23612362_ENST00000415083_SSX2_chrX_52729628_ENST00000337502_length(amino acids)=488AA_BP=410 MSVAFAAPRQRGKGEITPAAIQKMLDDNNHLIQCIMDSQNKGKTSECSQYQQMLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGM NQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQMNGQMPGPNHMPMQGPGPNQLNMTNSSMNMPSSSHGSMGGYNHSVPSSQSMPVQNQMT MSQGQPMGNYGPRPNMSMQPNQGPMMHQQPPSQQYNMPQGGGQHYQGQQPPMGMMGQVNQGNHMMGQRQIPPYRPPQQGPPQQYSGQEDY YGDQYSHGGQGPPEGMNQQYYPDGHNDYGYQQPSYPEQGYDRPYEDSSQHYYEGGNSQYGQQQDAYQGPPPQQGYPPQQQQYPGQQGYPG QQQGYGPSQGGPGPQYPNYPQGQGQQYGGYRPTQPGPPQPPQQRPYGYDQIMPKKPAEEGNDSEEVPEASGPQNDGKELCPPGKPTTSEK -------------------------------------------------------------- >86683_86683_5_SS18-SSX2_SS18_chr18_23612362_ENST00000545952_SSX2_chrX_52729628_ENST00000336777_length(transcript)=2294nt_BP=1284nt ACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGCAGCGAGGCAAGGGGGAGATCACTCCCGCTGCGAT TCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGAATAAAGGAAAGACCTCAGAGTGTTCTCAGGTCAT GAATAAACCACATCTGTTAGTATGGTTTGGTCAGTGGTCCTCAGAAAATCTCATGTCGTGCAACCATACTAATACTCAAGCAAAGGAAAA CTTGAAGGCCTAGACTGAAACAGGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAAATATG CAGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCACGCTCT CACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGGGCCTAAC CATATGCCTATGCAGGGACCTGGACCCAATCAACTCAATATGACAAACAGTTCCATGAATATGCCTTCAAGTAGCCATGGATCCATGGGA GGTTACAACCATTCTGTGCCATCATCACAGAGCATGCCAGTACAGAATCAGATGACAATGAGTCAGGGACAACCAATGGGAAACTATGGT CCCAGACCAAATATGAGTATGCAGCCAAACCAAGGTCCAATGATGCATCAGCAGCCTCCTTCTCAGCAATACAATATGCCACAGGGAGGC GGACAGCATTACCAAGGACAGCAGCCACCTATGGGAATGATGGGTCAAGTTAACCAAGGCAATCATATGATGGGTCAGAGACAGATTCCT CCCTATAGACCTCCTCAACAGGGCCCACCACAGCAGTACTCAGGCCAGGAAGACTATTACGGGGACCAATACAGTCATGGTGGACAAGGT CCTCCAGAAGGCATGAACCAGCAATATTACCCTGATGGAAATTCACAGTATGGCCAACAGCAAGATGCATACCAGGGACCACCTCCACAA CAGGGATATCCACCCCAGCAGCAGCAGTACCCAGGGCAGCAAGGTTACCCAGGACAGCAGCAGGGCTACGGTCCTTCACAGGGTGGTCCA GGTCCTCAGTATCCTAACTACCCACAGGGACAAGGTCAGCAGTATGGAGGATATAGACCAACACAGCCTGGACCACCACAGCCACCCCAG CAGAGGCCTTATGGATATGACCAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGCCAGAAGCATCTGGCCCA CAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGAAATAGGGAGGCCCAAGAA AAGGAAGAGAGACGCGGAACAGCTCATCGGTGGAGCAGTCAGAACACACACAACATTGGTCGATTCAGTTTGTCAACTTCTATGGGTGCA GTTCATGGTACCCCCAAAACAATTACACACAACAGGGACCCAAAAGGGGGGAACATGCCTGGACCCACAGACTGCGTGAGAGAAAACAGC TGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCCCCTCAGGGATACGACACATGCCCATGATGAGAAGCAGAA CGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAGGTGCATAGCAAGTGAAAGCAAGTGTTCACAACAGTGAAA AGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTACGTTGTATTTTCTTACACTGTGTCATTCTGTTAGATACTA ACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTATCCATGCACCTACCTTAGAAAACAAGTATTGTCGGTTAC CTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAGGGCAGTTCTGAGTGTTTAATTTCAGATTTTTTCCTCTGC ATTTACACACACACGCACACAAACCACACCACACACACACACACACACACACACACACACACACACACACACCAAGTACCAGTATAAGCA TCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCACTCTGTTTCCTGGTCAGCATGTACTCCCCTCATCCGATTC >86683_86683_5_SS18-SSX2_SS18_chr18_23612362_ENST00000545952_SSX2_chrX_52729628_ENST00000336777_length(amino acids)=440AA_BP=327 MLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGMNQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQMNGQMPGPNHMPMQGPGPNQ LNMTNSSMNMPSSSHGSMGGYNHSVPSSQSMPVQNQMTMSQGQPMGNYGPRPNMSMQPNQGPMMHQQPPSQQYNMPQGGGQHYQGQQPPM GMMGQVNQGNHMMGQRQIPPYRPPQQGPPQQYSGQEDYYGDQYSHGGQGPPEGMNQQYYPDGNSQYGQQQDAYQGPPPQQGYPPQQQQYP GQQGYPGQQQGYGPSQGGPGPQYPNYPQGQGQQYGGYRPTQPGPPQPPQQRPYGYDQIMPKKPAEEGNDSEEVPEASGPQNDGKELCPPG -------------------------------------------------------------- >86683_86683_6_SS18-SSX2_SS18_chr18_23612362_ENST00000545952_SSX2_chrX_52729628_ENST00000337502_length(transcript)=2148nt_BP=1284nt ACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGCAGCGAGGCAAGGGGGAGATCACTCCCGCTGCGAT TCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGAATAAAGGAAAGACCTCAGAGTGTTCTCAGGTCAT GAATAAACCACATCTGTTAGTATGGTTTGGTCAGTGGTCCTCAGAAAATCTCATGTCGTGCAACCATACTAATACTCAAGCAAAGGAAAA CTTGAAGGCCTAGACTGAAACAGGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAAATATG CAGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCACGCTCT CACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGGGCCTAAC CATATGCCTATGCAGGGACCTGGACCCAATCAACTCAATATGACAAACAGTTCCATGAATATGCCTTCAAGTAGCCATGGATCCATGGGA GGTTACAACCATTCTGTGCCATCATCACAGAGCATGCCAGTACAGAATCAGATGACAATGAGTCAGGGACAACCAATGGGAAACTATGGT CCCAGACCAAATATGAGTATGCAGCCAAACCAAGGTCCAATGATGCATCAGCAGCCTCCTTCTCAGCAATACAATATGCCACAGGGAGGC GGACAGCATTACCAAGGACAGCAGCCACCTATGGGAATGATGGGTCAAGTTAACCAAGGCAATCATATGATGGGTCAGAGACAGATTCCT CCCTATAGACCTCCTCAACAGGGCCCACCACAGCAGTACTCAGGCCAGGAAGACTATTACGGGGACCAATACAGTCATGGTGGACAAGGT CCTCCAGAAGGCATGAACCAGCAATATTACCCTGATGGAAATTCACAGTATGGCCAACAGCAAGATGCATACCAGGGACCACCTCCACAA CAGGGATATCCACCCCAGCAGCAGCAGTACCCAGGGCAGCAAGGTTACCCAGGACAGCAGCAGGGCTACGGTCCTTCACAGGGTGGTCCA GGTCCTCAGTATCCTAACTACCCACAGGGACAAGGTCAGCAGTATGGAGGATATAGACCAACACAGCCTGGACCACCACAGCCACCCCAG CAGAGGCCTTATGGATATGACCAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGCCAGAAGCATCTGGCCCA CAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGACCCAAAAGGGGGGAACAT GCCTGGACCCACAGACTGCGTGAGAGAAAACAGCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCCCCTCA GGGATACGACACATGCCCATGATGAGAAGCAGAACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAGGTGCA TAGCAAGTGAAAGCAAGTGTTCACAACAGTGAAAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTACGTTGT ATTTTCTTACACTGTGTCATTCTGTTAGATACTAACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTATCCAT GCACCTACCTTAGAAAACAAGTATTGTCGGTTACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAGGGCAG TTCTGAGTGTTTAATTTCAGATTTTTTCCTCTGCATTTACACACACACGCACACAAACCACACCACACACACACACACACACACACACAC ACACACACACACACACCAAGTACCAGTATAAGCATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCACTCTGT >86683_86683_6_SS18-SSX2_SS18_chr18_23612362_ENST00000545952_SSX2_chrX_52729628_ENST00000337502_length(amino acids)=405AA_BP=327 MLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGMNQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQMNGQMPGPNHMPMQGPGPNQ LNMTNSSMNMPSSSHGSMGGYNHSVPSSQSMPVQNQMTMSQGQPMGNYGPRPNMSMQPNQGPMMHQQPPSQQYNMPQGGGQHYQGQQPPM GMMGQVNQGNHMMGQRQIPPYRPPQQGPPQQYSGQEDYYGDQYSHGGQGPPEGMNQQYYPDGNSQYGQQQDAYQGPPPQQGYPPQQQQYP GQQGYPGQQQGYGPSQGGPGPQYPNYPQGQGQQYGGYRPTQPGPPQPPQQRPYGYDQIMPKKPAEEGNDSEEVPEASGPQNDGKELCPPG -------------------------------------------------------------- >86683_86683_7_SS18-SSX2_SS18_chr18_23612363_ENST00000269137_SSX2_chrX_52729628_ENST00000336777_length(transcript)=2479nt_BP=1469nt TTTCCTGAATCGCTGAAGCGGAGGGAGGACTAGGGCGCGACTCTACCATTAGCGCAAAGGGCCAGGGGGTGCCCATAAGGAGAGCTCGGG GGATGCAGGGACGGTCAAGAAAGTTCGGCCAGGGCTAGGGGTCCCCCAGAGTGCGCAGAAGCGAGACATCCCTGGCCAGAGTGCCTCCTT CAGGCCTGCCCCGCCGCCCGTCCGCTTGGCCCCGGAGCATCCCCCCTCCCCGCATTCCCGAGGTGGTTCAGCCCGAGAGGCCGGCGTCTC TCCCCCAGTTTGCCGTTCACCCGGAGCGCTCGGGACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGC AGCGAGGCAAGGGGGAGATCACTCCCGCTGCGATTCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGA ATAAAGGAAAGACCTCAGAGTGTTCTCAGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAA ATATGCAGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCAC GCTCTCACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGGGC CTAACCATATGCCTATGCAGGGACCTGGACCCAATCAACTCAATATGACAAACAGTTCCATGAATATGCCTTCAAGTAGCCATGGATCCA TGGGAGGTTACAACCATTCTGTGCCATCATCACAGAGCATGCCAGTACAGAATCAGATGACAATGAGTCAGGGACAACCAATGGGAAACT ATGGTCCCAGACCAAATATGAGTATGCAGCCAAACCAAGGTCCAATGATGCATCAGCAGCCTCCTTCTCAGCAATACAATATGCCACAGG GAGGCGGACAGCATTACCAAGGACAGCAGCCACCTATGGGAATGATGGGTCAAGTTAACCAAGGCAATCATATGATGGGTCAGAGACAGA TTCCTCCCTATAGACCTCCTCAACAGGGCCCACCACAGCAGTACTCAGGCCAGGAAGACTATTACGGGGACCAATACAGTCATGGTGGAC AAGGTCCTCCAGAAGGCATGAACCAGCAATATTACCCTGATGGAAATTCACAGTATGGCCAACAGCAAGATGCATACCAGGGACCACCTC CACAACAGGGATATCCACCCCAGCAGCAGCAGTACCCAGGGCAGCAAGGTTACCCAGGACAGCAGCAGGGCTACGGTCCTTCACAGGGTG GTCCAGGTCCTCAGTATCCTAACTACCCACAGGGACAAGGTCAGCAGTATGGAGGATATAGACCAACACAGCCTGGACCACCACAGCCAC CCCAGCAGAGGCCTTATGGATATGACCAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGCCAGAAGCATCTG GCCCACAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGAAATAGGGAGGCCC AAGAAAAGGAAGAGAGACGCGGAACAGCTCATCGGTGGAGCAGTCAGAACACACACAACATTGGTCGATTCAGTTTGTCAACTTCTATGG GTGCAGTTCATGGTACCCCCAAAACAATTACACACAACAGGGACCCAAAAGGGGGGAACATGCCTGGACCCACAGACTGCGTGAGAGAAA ACAGCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCCCCTCAGGGATACGACACATGCCCATGATGAGAAG CAGAACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAGGTGCATAGCAAGTGAAAGCAAGTGTTCACAACAG TGAAAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTACGTTGTATTTTCTTACACTGTGTCATTCTGTTAGA TACTAACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTATCCATGCACCTACCTTAGAAAACAAGTATTGTCG GTTACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAGGGCAGTTCTGAGTGTTTAATTTCAGATTTTTTCC TCTGCATTTACACACACACGCACACAAACCACACCACACACACACACACACACACACACACACACACACACACACACCAAGTACCAGTAT AAGCATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCACTCTGTTTCCTGGTCAGCATGTACTCCCCTCATCC >86683_86683_7_SS18-SSX2_SS18_chr18_23612363_ENST00000269137_SSX2_chrX_52729628_ENST00000336777_length(amino acids)=549AA_BP=436 MARVPPSGLPRRPSAWPRSIPPPRIPEVVQPERPASLPQFAVHPERSGLADSGDGGNMSVAFAAPRQRGKGEITPAAIQKMLDDNNHLIQ CIMDSQNKGKTSECSQYQQMLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGMNQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQM NGQMPGPNHMPMQGPGPNQLNMTNSSMNMPSSSHGSMGGYNHSVPSSQSMPVQNQMTMSQGQPMGNYGPRPNMSMQPNQGPMMHQQPPSQ QYNMPQGGGQHYQGQQPPMGMMGQVNQGNHMMGQRQIPPYRPPQQGPPQQYSGQEDYYGDQYSHGGQGPPEGMNQQYYPDGNSQYGQQQD AYQGPPPQQGYPPQQQQYPGQQGYPGQQQGYGPSQGGPGPQYPNYPQGQGQQYGGYRPTQPGPPQPPQQRPYGYDQIMPKKPAEEGNDSE EVPEASGPQNDGKELCPPGKPTTSEKIHERSGNREAQEKEERRGTAHRWSSQNTHNIGRFSLSTSMGAVHGTPKTITHNRDPKGGNMPGP -------------------------------------------------------------- >86683_86683_8_SS18-SSX2_SS18_chr18_23612363_ENST00000269137_SSX2_chrX_52729628_ENST00000337502_length(transcript)=2333nt_BP=1469nt TTTCCTGAATCGCTGAAGCGGAGGGAGGACTAGGGCGCGACTCTACCATTAGCGCAAAGGGCCAGGGGGTGCCCATAAGGAGAGCTCGGG GGATGCAGGGACGGTCAAGAAAGTTCGGCCAGGGCTAGGGGTCCCCCAGAGTGCGCAGAAGCGAGACATCCCTGGCCAGAGTGCCTCCTT CAGGCCTGCCCCGCCGCCCGTCCGCTTGGCCCCGGAGCATCCCCCCTCCCCGCATTCCCGAGGTGGTTCAGCCCGAGAGGCCGGCGTCTC TCCCCCAGTTTGCCGTTCACCCGGAGCGCTCGGGACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGC AGCGAGGCAAGGGGGAGATCACTCCCGCTGCGATTCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGA ATAAAGGAAAGACCTCAGAGTGTTCTCAGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAA ATATGCAGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCAC GCTCTCACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGGGC CTAACCATATGCCTATGCAGGGACCTGGACCCAATCAACTCAATATGACAAACAGTTCCATGAATATGCCTTCAAGTAGCCATGGATCCA TGGGAGGTTACAACCATTCTGTGCCATCATCACAGAGCATGCCAGTACAGAATCAGATGACAATGAGTCAGGGACAACCAATGGGAAACT ATGGTCCCAGACCAAATATGAGTATGCAGCCAAACCAAGGTCCAATGATGCATCAGCAGCCTCCTTCTCAGCAATACAATATGCCACAGG GAGGCGGACAGCATTACCAAGGACAGCAGCCACCTATGGGAATGATGGGTCAAGTTAACCAAGGCAATCATATGATGGGTCAGAGACAGA TTCCTCCCTATAGACCTCCTCAACAGGGCCCACCACAGCAGTACTCAGGCCAGGAAGACTATTACGGGGACCAATACAGTCATGGTGGAC AAGGTCCTCCAGAAGGCATGAACCAGCAATATTACCCTGATGGAAATTCACAGTATGGCCAACAGCAAGATGCATACCAGGGACCACCTC CACAACAGGGATATCCACCCCAGCAGCAGCAGTACCCAGGGCAGCAAGGTTACCCAGGACAGCAGCAGGGCTACGGTCCTTCACAGGGTG GTCCAGGTCCTCAGTATCCTAACTACCCACAGGGACAAGGTCAGCAGTATGGAGGATATAGACCAACACAGCCTGGACCACCACAGCCAC CCCAGCAGAGGCCTTATGGATATGACCAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGCCAGAAGCATCTG GCCCACAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGACCCAAAAGGGGGG AACATGCCTGGACCCACAGACTGCGTGAGAGAAAACAGCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCC CCTCAGGGATACGACACATGCCCATGATGAGAAGCAGAACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAG GTGCATAGCAAGTGAAAGCAAGTGTTCACAACAGTGAAAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTAC GTTGTATTTTCTTACACTGTGTCATTCTGTTAGATACTAACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTA TCCATGCACCTACCTTAGAAAACAAGTATTGTCGGTTACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAG GGCAGTTCTGAGTGTTTAATTTCAGATTTTTTCCTCTGCATTTACACACACACGCACACAAACCACACCACACACACACACACACACACA CACACACACACACACACACACCAAGTACCAGTATAAGCATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCAC >86683_86683_8_SS18-SSX2_SS18_chr18_23612363_ENST00000269137_SSX2_chrX_52729628_ENST00000337502_length(amino acids)=514AA_BP=436 MARVPPSGLPRRPSAWPRSIPPPRIPEVVQPERPASLPQFAVHPERSGLADSGDGGNMSVAFAAPRQRGKGEITPAAIQKMLDDNNHLIQ CIMDSQNKGKTSECSQYQQMLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGMNQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQM NGQMPGPNHMPMQGPGPNQLNMTNSSMNMPSSSHGSMGGYNHSVPSSQSMPVQNQMTMSQGQPMGNYGPRPNMSMQPNQGPMMHQQPPSQ QYNMPQGGGQHYQGQQPPMGMMGQVNQGNHMMGQRQIPPYRPPQQGPPQQYSGQEDYYGDQYSHGGQGPPEGMNQQYYPDGNSQYGQQQD AYQGPPPQQGYPPQQQQYPGQQGYPGQQQGYGPSQGGPGPQYPNYPQGQGQQYGGYRPTQPGPPQPPQQRPYGYDQIMPKKPAEEGNDSE -------------------------------------------------------------- >86683_86683_9_SS18-SSX2_SS18_chr18_23612363_ENST00000269137_SSX2_chrX_52729630_ENST00000336777_length(transcript)=2479nt_BP=1469nt TTTCCTGAATCGCTGAAGCGGAGGGAGGACTAGGGCGCGACTCTACCATTAGCGCAAAGGGCCAGGGGGTGCCCATAAGGAGAGCTCGGG GGATGCAGGGACGGTCAAGAAAGTTCGGCCAGGGCTAGGGGTCCCCCAGAGTGCGCAGAAGCGAGACATCCCTGGCCAGAGTGCCTCCTT CAGGCCTGCCCCGCCGCCCGTCCGCTTGGCCCCGGAGCATCCCCCCTCCCCGCATTCCCGAGGTGGTTCAGCCCGAGAGGCCGGCGTCTC TCCCCCAGTTTGCCGTTCACCCGGAGCGCTCGGGACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGC AGCGAGGCAAGGGGGAGATCACTCCCGCTGCGATTCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGA ATAAAGGAAAGACCTCAGAGTGTTCTCAGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAA ATATGCAGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCAC GCTCTCACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGGGC CTAACCATATGCCTATGCAGGGACCTGGACCCAATCAACTCAATATGACAAACAGTTCCATGAATATGCCTTCAAGTAGCCATGGATCCA TGGGAGGTTACAACCATTCTGTGCCATCATCACAGAGCATGCCAGTACAGAATCAGATGACAATGAGTCAGGGACAACCAATGGGAAACT ATGGTCCCAGACCAAATATGAGTATGCAGCCAAACCAAGGTCCAATGATGCATCAGCAGCCTCCTTCTCAGCAATACAATATGCCACAGG GAGGCGGACAGCATTACCAAGGACAGCAGCCACCTATGGGAATGATGGGTCAAGTTAACCAAGGCAATCATATGATGGGTCAGAGACAGA TTCCTCCCTATAGACCTCCTCAACAGGGCCCACCACAGCAGTACTCAGGCCAGGAAGACTATTACGGGGACCAATACAGTCATGGTGGAC AAGGTCCTCCAGAAGGCATGAACCAGCAATATTACCCTGATGGAAATTCACAGTATGGCCAACAGCAAGATGCATACCAGGGACCACCTC CACAACAGGGATATCCACCCCAGCAGCAGCAGTACCCAGGGCAGCAAGGTTACCCAGGACAGCAGCAGGGCTACGGTCCTTCACAGGGTG GTCCAGGTCCTCAGTATCCTAACTACCCACAGGGACAAGGTCAGCAGTATGGAGGATATAGACCAACACAGCCTGGACCACCACAGCCAC CCCAGCAGAGGCCTTATGGATATGACCAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGCCAGAAGCATCTG GCCCACAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGAAATAGGGAGGCCC AAGAAAAGGAAGAGAGACGCGGAACAGCTCATCGGTGGAGCAGTCAGAACACACACAACATTGGTCGATTCAGTTTGTCAACTTCTATGG GTGCAGTTCATGGTACCCCCAAAACAATTACACACAACAGGGACCCAAAAGGGGGGAACATGCCTGGACCCACAGACTGCGTGAGAGAAA ACAGCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCCCCTCAGGGATACGACACATGCCCATGATGAGAAG CAGAACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAGGTGCATAGCAAGTGAAAGCAAGTGTTCACAACAG TGAAAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTACGTTGTATTTTCTTACACTGTGTCATTCTGTTAGA TACTAACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTATCCATGCACCTACCTTAGAAAACAAGTATTGTCG GTTACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAGGGCAGTTCTGAGTGTTTAATTTCAGATTTTTTCC TCTGCATTTACACACACACGCACACAAACCACACCACACACACACACACACACACACACACACACACACACACACACCAAGTACCAGTAT AAGCATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCACTCTGTTTCCTGGTCAGCATGTACTCCCCTCATCC >86683_86683_9_SS18-SSX2_SS18_chr18_23612363_ENST00000269137_SSX2_chrX_52729630_ENST00000336777_length(amino acids)=549AA_BP=436 MARVPPSGLPRRPSAWPRSIPPPRIPEVVQPERPASLPQFAVHPERSGLADSGDGGNMSVAFAAPRQRGKGEITPAAIQKMLDDNNHLIQ CIMDSQNKGKTSECSQYQQMLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGMNQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQM NGQMPGPNHMPMQGPGPNQLNMTNSSMNMPSSSHGSMGGYNHSVPSSQSMPVQNQMTMSQGQPMGNYGPRPNMSMQPNQGPMMHQQPPSQ QYNMPQGGGQHYQGQQPPMGMMGQVNQGNHMMGQRQIPPYRPPQQGPPQQYSGQEDYYGDQYSHGGQGPPEGMNQQYYPDGNSQYGQQQD AYQGPPPQQGYPPQQQQYPGQQGYPGQQQGYGPSQGGPGPQYPNYPQGQGQQYGGYRPTQPGPPQPPQQRPYGYDQIMPKKPAEEGNDSE EVPEASGPQNDGKELCPPGKPTTSEKIHERSGNREAQEKEERRGTAHRWSSQNTHNIGRFSLSTSMGAVHGTPKTITHNRDPKGGNMPGP -------------------------------------------------------------- >86683_86683_10_SS18-SSX2_SS18_chr18_23612363_ENST00000269137_SSX2_chrX_52729630_ENST00000337502_length(transcript)=2333nt_BP=1469nt TTTCCTGAATCGCTGAAGCGGAGGGAGGACTAGGGCGCGACTCTACCATTAGCGCAAAGGGCCAGGGGGTGCCCATAAGGAGAGCTCGGG GGATGCAGGGACGGTCAAGAAAGTTCGGCCAGGGCTAGGGGTCCCCCAGAGTGCGCAGAAGCGAGACATCCCTGGCCAGAGTGCCTCCTT CAGGCCTGCCCCGCCGCCCGTCCGCTTGGCCCCGGAGCATCCCCCCTCCCCGCATTCCCGAGGTGGTTCAGCCCGAGAGGCCGGCGTCTC TCCCCCAGTTTGCCGTTCACCCGGAGCGCTCGGGACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGC AGCGAGGCAAGGGGGAGATCACTCCCGCTGCGATTCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGA ATAAAGGAAAGACCTCAGAGTGTTCTCAGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAA ATATGCAGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCAC GCTCTCACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGGGC CTAACCATATGCCTATGCAGGGACCTGGACCCAATCAACTCAATATGACAAACAGTTCCATGAATATGCCTTCAAGTAGCCATGGATCCA TGGGAGGTTACAACCATTCTGTGCCATCATCACAGAGCATGCCAGTACAGAATCAGATGACAATGAGTCAGGGACAACCAATGGGAAACT ATGGTCCCAGACCAAATATGAGTATGCAGCCAAACCAAGGTCCAATGATGCATCAGCAGCCTCCTTCTCAGCAATACAATATGCCACAGG GAGGCGGACAGCATTACCAAGGACAGCAGCCACCTATGGGAATGATGGGTCAAGTTAACCAAGGCAATCATATGATGGGTCAGAGACAGA TTCCTCCCTATAGACCTCCTCAACAGGGCCCACCACAGCAGTACTCAGGCCAGGAAGACTATTACGGGGACCAATACAGTCATGGTGGAC AAGGTCCTCCAGAAGGCATGAACCAGCAATATTACCCTGATGGAAATTCACAGTATGGCCAACAGCAAGATGCATACCAGGGACCACCTC CACAACAGGGATATCCACCCCAGCAGCAGCAGTACCCAGGGCAGCAAGGTTACCCAGGACAGCAGCAGGGCTACGGTCCTTCACAGGGTG GTCCAGGTCCTCAGTATCCTAACTACCCACAGGGACAAGGTCAGCAGTATGGAGGATATAGACCAACACAGCCTGGACCACCACAGCCAC CCCAGCAGAGGCCTTATGGATATGACCAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGCCAGAAGCATCTG GCCCACAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGACCCAAAAGGGGGG AACATGCCTGGACCCACAGACTGCGTGAGAGAAAACAGCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCC CCTCAGGGATACGACACATGCCCATGATGAGAAGCAGAACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAG GTGCATAGCAAGTGAAAGCAAGTGTTCACAACAGTGAAAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTAC GTTGTATTTTCTTACACTGTGTCATTCTGTTAGATACTAACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTA TCCATGCACCTACCTTAGAAAACAAGTATTGTCGGTTACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAG GGCAGTTCTGAGTGTTTAATTTCAGATTTTTTCCTCTGCATTTACACACACACGCACACAAACCACACCACACACACACACACACACACA CACACACACACACACACACACCAAGTACCAGTATAAGCATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCAC >86683_86683_10_SS18-SSX2_SS18_chr18_23612363_ENST00000269137_SSX2_chrX_52729630_ENST00000337502_length(amino acids)=514AA_BP=436 MARVPPSGLPRRPSAWPRSIPPPRIPEVVQPERPASLPQFAVHPERSGLADSGDGGNMSVAFAAPRQRGKGEITPAAIQKMLDDNNHLIQ CIMDSQNKGKTSECSQYQQMLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGMNQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQM NGQMPGPNHMPMQGPGPNQLNMTNSSMNMPSSSHGSMGGYNHSVPSSQSMPVQNQMTMSQGQPMGNYGPRPNMSMQPNQGPMMHQQPPSQ QYNMPQGGGQHYQGQQPPMGMMGQVNQGNHMMGQRQIPPYRPPQQGPPQQYSGQEDYYGDQYSHGGQGPPEGMNQQYYPDGNSQYGQQQD AYQGPPPQQGYPPQQQQYPGQQGYPGQQQGYGPSQGGPGPQYPNYPQGQGQQYGGYRPTQPGPPQPPQQRPYGYDQIMPKKPAEEGNDSE -------------------------------------------------------------- >86683_86683_11_SS18-SSX2_SS18_chr18_23612363_ENST00000415083_SSX2_chrX_52729628_ENST00000336777_length(transcript)=2296nt_BP=1286nt AGTTTGCCGTTCACCCGGAGCGCTCGGGACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGCAGCGAG GCAAGGGGGAGATCACTCCCGCTGCGATTCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGAATAAAG GAAAGACCTCAGAGTGTTCTCAGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAAATATGC AGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCACGCTCTC ACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGGGCCTAACC ATATGCCTATGCAGGGACCTGGACCCAATCAACTCAATATGACAAACAGTTCCATGAATATGCCTTCAAGTAGCCATGGATCCATGGGAG GTTACAACCATTCTGTGCCATCATCACAGAGCATGCCAGTACAGAATCAGATGACAATGAGTCAGGGACAACCAATGGGAAACTATGGTC CCAGACCAAATATGAGTATGCAGCCAAACCAAGGTCCAATGATGCATCAGCAGCCTCCTTCTCAGCAATACAATATGCCACAGGGAGGCG GACAGCATTACCAAGGACAGCAGCCACCTATGGGAATGATGGGTCAAGTTAACCAAGGCAATCATATGATGGGTCAGAGACAGATTCCTC CCTATAGACCTCCTCAACAGGGCCCACCACAGCAGTACTCAGGCCAGGAAGACTATTACGGGGACCAATACAGTCATGGTGGACAAGGTC CTCCAGAAGGCATGAACCAGCAATATTACCCTGATGGTCATAATGATTACGGTTATCAGCAACCGTCGTATCCTGAACAAGGCTACGATA GGCCTTATGAGGATTCCTCACAACATTACTACGAAGGAGGAAATTCACAGTATGGCCAACAGCAAGATGCATACCAGGGACCACCTCCAC AACAGGGATATCCACCCCAGCAGCAGCAGTACCCAGGGCAGCAAGGTTACCCAGGACAGCAGCAGGGCTACGGTCCTTCACAGGGTGGTC CAGGTCCTCAGTATCCTAACTACCCACAGGGACAAGGTCAGCAGTATGGAGGATATAGACCAACACAGCCTGGACCACCACAGCCACCCC AGCAGAGGCCTTATGGATATGACCAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGCCAGAAGCATCTGGCC CACAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGAAATAGGGAGGCCCAAG AAAAGGAAGAGAGACGCGGAACAGCTCATCGGTGGAGCAGTCAGAACACACACAACATTGGTCGATTCAGTTTGTCAACTTCTATGGGTG CAGTTCATGGTACCCCCAAAACAATTACACACAACAGGGACCCAAAAGGGGGGAACATGCCTGGACCCACAGACTGCGTGAGAGAAAACA GCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCCCCTCAGGGATACGACACATGCCCATGATGAGAAGCAG AACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAGGTGCATAGCAAGTGAAAGCAAGTGTTCACAACAGTGA AAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTACGTTGTATTTTCTTACACTGTGTCATTCTGTTAGATAC TAACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTATCCATGCACCTACCTTAGAAAACAAGTATTGTCGGTT ACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAGGGCAGTTCTGAGTGTTTAATTTCAGATTTTTTCCTCT GCATTTACACACACACGCACACAAACCACACCACACACACACACACACACACACACACACACACACACACACACCAAGTACCAGTATAAG CATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCACTCTGTTTCCTGGTCAGCATGTACTCCCCTCATCCGAT >86683_86683_11_SS18-SSX2_SS18_chr18_23612363_ENST00000415083_SSX2_chrX_52729628_ENST00000336777_length(amino acids)=523AA_BP=410 MSVAFAAPRQRGKGEITPAAIQKMLDDNNHLIQCIMDSQNKGKTSECSQYQQMLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGM NQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQMNGQMPGPNHMPMQGPGPNQLNMTNSSMNMPSSSHGSMGGYNHSVPSSQSMPVQNQMT MSQGQPMGNYGPRPNMSMQPNQGPMMHQQPPSQQYNMPQGGGQHYQGQQPPMGMMGQVNQGNHMMGQRQIPPYRPPQQGPPQQYSGQEDY YGDQYSHGGQGPPEGMNQQYYPDGHNDYGYQQPSYPEQGYDRPYEDSSQHYYEGGNSQYGQQQDAYQGPPPQQGYPPQQQQYPGQQGYPG QQQGYGPSQGGPGPQYPNYPQGQGQQYGGYRPTQPGPPQPPQQRPYGYDQIMPKKPAEEGNDSEEVPEASGPQNDGKELCPPGKPTTSEK -------------------------------------------------------------- >86683_86683_12_SS18-SSX2_SS18_chr18_23612363_ENST00000415083_SSX2_chrX_52729628_ENST00000337502_length(transcript)=2150nt_BP=1286nt AGTTTGCCGTTCACCCGGAGCGCTCGGGACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGCAGCGAG GCAAGGGGGAGATCACTCCCGCTGCGATTCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGAATAAAG GAAAGACCTCAGAGTGTTCTCAGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAAATATGC AGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCACGCTCTC ACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGGGCCTAACC ATATGCCTATGCAGGGACCTGGACCCAATCAACTCAATATGACAAACAGTTCCATGAATATGCCTTCAAGTAGCCATGGATCCATGGGAG GTTACAACCATTCTGTGCCATCATCACAGAGCATGCCAGTACAGAATCAGATGACAATGAGTCAGGGACAACCAATGGGAAACTATGGTC CCAGACCAAATATGAGTATGCAGCCAAACCAAGGTCCAATGATGCATCAGCAGCCTCCTTCTCAGCAATACAATATGCCACAGGGAGGCG GACAGCATTACCAAGGACAGCAGCCACCTATGGGAATGATGGGTCAAGTTAACCAAGGCAATCATATGATGGGTCAGAGACAGATTCCTC CCTATAGACCTCCTCAACAGGGCCCACCACAGCAGTACTCAGGCCAGGAAGACTATTACGGGGACCAATACAGTCATGGTGGACAAGGTC CTCCAGAAGGCATGAACCAGCAATATTACCCTGATGGTCATAATGATTACGGTTATCAGCAACCGTCGTATCCTGAACAAGGCTACGATA GGCCTTATGAGGATTCCTCACAACATTACTACGAAGGAGGAAATTCACAGTATGGCCAACAGCAAGATGCATACCAGGGACCACCTCCAC AACAGGGATATCCACCCCAGCAGCAGCAGTACCCAGGGCAGCAAGGTTACCCAGGACAGCAGCAGGGCTACGGTCCTTCACAGGGTGGTC CAGGTCCTCAGTATCCTAACTACCCACAGGGACAAGGTCAGCAGTATGGAGGATATAGACCAACACAGCCTGGACCACCACAGCCACCCC AGCAGAGGCCTTATGGATATGACCAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGCCAGAAGCATCTGGCC CACAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGACCCAAAAGGGGGGAAC ATGCCTGGACCCACAGACTGCGTGAGAGAAAACAGCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCCCCT CAGGGATACGACACATGCCCATGATGAGAAGCAGAACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAGGTG CATAGCAAGTGAAAGCAAGTGTTCACAACAGTGAAAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTACGTT GTATTTTCTTACACTGTGTCATTCTGTTAGATACTAACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTATCC ATGCACCTACCTTAGAAAACAAGTATTGTCGGTTACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAGGGC AGTTCTGAGTGTTTAATTTCAGATTTTTTCCTCTGCATTTACACACACACGCACACAAACCACACCACACACACACACACACACACACAC ACACACACACACACACACCAAGTACCAGTATAAGCATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCACTCT >86683_86683_12_SS18-SSX2_SS18_chr18_23612363_ENST00000415083_SSX2_chrX_52729628_ENST00000337502_length(amino acids)=488AA_BP=410 MSVAFAAPRQRGKGEITPAAIQKMLDDNNHLIQCIMDSQNKGKTSECSQYQQMLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGM NQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQMNGQMPGPNHMPMQGPGPNQLNMTNSSMNMPSSSHGSMGGYNHSVPSSQSMPVQNQMT MSQGQPMGNYGPRPNMSMQPNQGPMMHQQPPSQQYNMPQGGGQHYQGQQPPMGMMGQVNQGNHMMGQRQIPPYRPPQQGPPQQYSGQEDY YGDQYSHGGQGPPEGMNQQYYPDGHNDYGYQQPSYPEQGYDRPYEDSSQHYYEGGNSQYGQQQDAYQGPPPQQGYPPQQQQYPGQQGYPG QQQGYGPSQGGPGPQYPNYPQGQGQQYGGYRPTQPGPPQPPQQRPYGYDQIMPKKPAEEGNDSEEVPEASGPQNDGKELCPPGKPTTSEK -------------------------------------------------------------- >86683_86683_13_SS18-SSX2_SS18_chr18_23612363_ENST00000415083_SSX2_chrX_52729630_ENST00000336777_length(transcript)=2296nt_BP=1286nt AGTTTGCCGTTCACCCGGAGCGCTCGGGACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGCAGCGAG GCAAGGGGGAGATCACTCCCGCTGCGATTCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGAATAAAG GAAAGACCTCAGAGTGTTCTCAGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAAATATGC AGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCACGCTCTC ACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGGGCCTAACC ATATGCCTATGCAGGGACCTGGACCCAATCAACTCAATATGACAAACAGTTCCATGAATATGCCTTCAAGTAGCCATGGATCCATGGGAG GTTACAACCATTCTGTGCCATCATCACAGAGCATGCCAGTACAGAATCAGATGACAATGAGTCAGGGACAACCAATGGGAAACTATGGTC CCAGACCAAATATGAGTATGCAGCCAAACCAAGGTCCAATGATGCATCAGCAGCCTCCTTCTCAGCAATACAATATGCCACAGGGAGGCG GACAGCATTACCAAGGACAGCAGCCACCTATGGGAATGATGGGTCAAGTTAACCAAGGCAATCATATGATGGGTCAGAGACAGATTCCTC CCTATAGACCTCCTCAACAGGGCCCACCACAGCAGTACTCAGGCCAGGAAGACTATTACGGGGACCAATACAGTCATGGTGGACAAGGTC CTCCAGAAGGCATGAACCAGCAATATTACCCTGATGGTCATAATGATTACGGTTATCAGCAACCGTCGTATCCTGAACAAGGCTACGATA GGCCTTATGAGGATTCCTCACAACATTACTACGAAGGAGGAAATTCACAGTATGGCCAACAGCAAGATGCATACCAGGGACCACCTCCAC AACAGGGATATCCACCCCAGCAGCAGCAGTACCCAGGGCAGCAAGGTTACCCAGGACAGCAGCAGGGCTACGGTCCTTCACAGGGTGGTC CAGGTCCTCAGTATCCTAACTACCCACAGGGACAAGGTCAGCAGTATGGAGGATATAGACCAACACAGCCTGGACCACCACAGCCACCCC AGCAGAGGCCTTATGGATATGACCAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGCCAGAAGCATCTGGCC CACAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGAAATAGGGAGGCCCAAG AAAAGGAAGAGAGACGCGGAACAGCTCATCGGTGGAGCAGTCAGAACACACACAACATTGGTCGATTCAGTTTGTCAACTTCTATGGGTG CAGTTCATGGTACCCCCAAAACAATTACACACAACAGGGACCCAAAAGGGGGGAACATGCCTGGACCCACAGACTGCGTGAGAGAAAACA GCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCCCCTCAGGGATACGACACATGCCCATGATGAGAAGCAG AACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAGGTGCATAGCAAGTGAAAGCAAGTGTTCACAACAGTGA AAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTACGTTGTATTTTCTTACACTGTGTCATTCTGTTAGATAC TAACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTATCCATGCACCTACCTTAGAAAACAAGTATTGTCGGTT ACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAGGGCAGTTCTGAGTGTTTAATTTCAGATTTTTTCCTCT GCATTTACACACACACGCACACAAACCACACCACACACACACACACACACACACACACACACACACACACACACCAAGTACCAGTATAAG CATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCACTCTGTTTCCTGGTCAGCATGTACTCCCCTCATCCGAT >86683_86683_13_SS18-SSX2_SS18_chr18_23612363_ENST00000415083_SSX2_chrX_52729630_ENST00000336777_length(amino acids)=523AA_BP=410 MSVAFAAPRQRGKGEITPAAIQKMLDDNNHLIQCIMDSQNKGKTSECSQYQQMLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGM NQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQMNGQMPGPNHMPMQGPGPNQLNMTNSSMNMPSSSHGSMGGYNHSVPSSQSMPVQNQMT MSQGQPMGNYGPRPNMSMQPNQGPMMHQQPPSQQYNMPQGGGQHYQGQQPPMGMMGQVNQGNHMMGQRQIPPYRPPQQGPPQQYSGQEDY YGDQYSHGGQGPPEGMNQQYYPDGHNDYGYQQPSYPEQGYDRPYEDSSQHYYEGGNSQYGQQQDAYQGPPPQQGYPPQQQQYPGQQGYPG QQQGYGPSQGGPGPQYPNYPQGQGQQYGGYRPTQPGPPQPPQQRPYGYDQIMPKKPAEEGNDSEEVPEASGPQNDGKELCPPGKPTTSEK -------------------------------------------------------------- >86683_86683_14_SS18-SSX2_SS18_chr18_23612363_ENST00000415083_SSX2_chrX_52729630_ENST00000337502_length(transcript)=2150nt_BP=1286nt AGTTTGCCGTTCACCCGGAGCGCTCGGGACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGCAGCGAG GCAAGGGGGAGATCACTCCCGCTGCGATTCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGAATAAAG GAAAGACCTCAGAGTGTTCTCAGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAAATATGC AGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCACGCTCTC ACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGGGCCTAACC ATATGCCTATGCAGGGACCTGGACCCAATCAACTCAATATGACAAACAGTTCCATGAATATGCCTTCAAGTAGCCATGGATCCATGGGAG GTTACAACCATTCTGTGCCATCATCACAGAGCATGCCAGTACAGAATCAGATGACAATGAGTCAGGGACAACCAATGGGAAACTATGGTC CCAGACCAAATATGAGTATGCAGCCAAACCAAGGTCCAATGATGCATCAGCAGCCTCCTTCTCAGCAATACAATATGCCACAGGGAGGCG GACAGCATTACCAAGGACAGCAGCCACCTATGGGAATGATGGGTCAAGTTAACCAAGGCAATCATATGATGGGTCAGAGACAGATTCCTC CCTATAGACCTCCTCAACAGGGCCCACCACAGCAGTACTCAGGCCAGGAAGACTATTACGGGGACCAATACAGTCATGGTGGACAAGGTC CTCCAGAAGGCATGAACCAGCAATATTACCCTGATGGTCATAATGATTACGGTTATCAGCAACCGTCGTATCCTGAACAAGGCTACGATA GGCCTTATGAGGATTCCTCACAACATTACTACGAAGGAGGAAATTCACAGTATGGCCAACAGCAAGATGCATACCAGGGACCACCTCCAC AACAGGGATATCCACCCCAGCAGCAGCAGTACCCAGGGCAGCAAGGTTACCCAGGACAGCAGCAGGGCTACGGTCCTTCACAGGGTGGTC CAGGTCCTCAGTATCCTAACTACCCACAGGGACAAGGTCAGCAGTATGGAGGATATAGACCAACACAGCCTGGACCACCACAGCCACCCC AGCAGAGGCCTTATGGATATGACCAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGCCAGAAGCATCTGGCC CACAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGACCCAAAAGGGGGGAAC ATGCCTGGACCCACAGACTGCGTGAGAGAAAACAGCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCCCCT CAGGGATACGACACATGCCCATGATGAGAAGCAGAACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAGGTG CATAGCAAGTGAAAGCAAGTGTTCACAACAGTGAAAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTACGTT GTATTTTCTTACACTGTGTCATTCTGTTAGATACTAACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTATCC ATGCACCTACCTTAGAAAACAAGTATTGTCGGTTACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAGGGC AGTTCTGAGTGTTTAATTTCAGATTTTTTCCTCTGCATTTACACACACACGCACACAAACCACACCACACACACACACACACACACACAC ACACACACACACACACACCAAGTACCAGTATAAGCATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCACTCT >86683_86683_14_SS18-SSX2_SS18_chr18_23612363_ENST00000415083_SSX2_chrX_52729630_ENST00000337502_length(amino acids)=488AA_BP=410 MSVAFAAPRQRGKGEITPAAIQKMLDDNNHLIQCIMDSQNKGKTSECSQYQQMLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGM NQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQMNGQMPGPNHMPMQGPGPNQLNMTNSSMNMPSSSHGSMGGYNHSVPSSQSMPVQNQMT MSQGQPMGNYGPRPNMSMQPNQGPMMHQQPPSQQYNMPQGGGQHYQGQQPPMGMMGQVNQGNHMMGQRQIPPYRPPQQGPPQQYSGQEDY YGDQYSHGGQGPPEGMNQQYYPDGHNDYGYQQPSYPEQGYDRPYEDSSQHYYEGGNSQYGQQQDAYQGPPPQQGYPPQQQQYPGQQGYPG QQQGYGPSQGGPGPQYPNYPQGQGQQYGGYRPTQPGPPQPPQQRPYGYDQIMPKKPAEEGNDSEEVPEASGPQNDGKELCPPGKPTTSEK -------------------------------------------------------------- >86683_86683_15_SS18-SSX2_SS18_chr18_23612363_ENST00000545952_SSX2_chrX_52729628_ENST00000336777_length(transcript)=2294nt_BP=1284nt ACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGCAGCGAGGCAAGGGGGAGATCACTCCCGCTGCGAT TCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGAATAAAGGAAAGACCTCAGAGTGTTCTCAGGTCAT GAATAAACCACATCTGTTAGTATGGTTTGGTCAGTGGTCCTCAGAAAATCTCATGTCGTGCAACCATACTAATACTCAAGCAAAGGAAAA CTTGAAGGCCTAGACTGAAACAGGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAAATATG CAGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCACGCTCT CACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGGGCCTAAC CATATGCCTATGCAGGGACCTGGACCCAATCAACTCAATATGACAAACAGTTCCATGAATATGCCTTCAAGTAGCCATGGATCCATGGGA GGTTACAACCATTCTGTGCCATCATCACAGAGCATGCCAGTACAGAATCAGATGACAATGAGTCAGGGACAACCAATGGGAAACTATGGT CCCAGACCAAATATGAGTATGCAGCCAAACCAAGGTCCAATGATGCATCAGCAGCCTCCTTCTCAGCAATACAATATGCCACAGGGAGGC GGACAGCATTACCAAGGACAGCAGCCACCTATGGGAATGATGGGTCAAGTTAACCAAGGCAATCATATGATGGGTCAGAGACAGATTCCT CCCTATAGACCTCCTCAACAGGGCCCACCACAGCAGTACTCAGGCCAGGAAGACTATTACGGGGACCAATACAGTCATGGTGGACAAGGT CCTCCAGAAGGCATGAACCAGCAATATTACCCTGATGGAAATTCACAGTATGGCCAACAGCAAGATGCATACCAGGGACCACCTCCACAA CAGGGATATCCACCCCAGCAGCAGCAGTACCCAGGGCAGCAAGGTTACCCAGGACAGCAGCAGGGCTACGGTCCTTCACAGGGTGGTCCA GGTCCTCAGTATCCTAACTACCCACAGGGACAAGGTCAGCAGTATGGAGGATATAGACCAACACAGCCTGGACCACCACAGCCACCCCAG CAGAGGCCTTATGGATATGACCAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGCCAGAAGCATCTGGCCCA CAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGAAATAGGGAGGCCCAAGAA AAGGAAGAGAGACGCGGAACAGCTCATCGGTGGAGCAGTCAGAACACACACAACATTGGTCGATTCAGTTTGTCAACTTCTATGGGTGCA GTTCATGGTACCCCCAAAACAATTACACACAACAGGGACCCAAAAGGGGGGAACATGCCTGGACCCACAGACTGCGTGAGAGAAAACAGC TGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCCCCTCAGGGATACGACACATGCCCATGATGAGAAGCAGAA CGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAGGTGCATAGCAAGTGAAAGCAAGTGTTCACAACAGTGAAA AGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTACGTTGTATTTTCTTACACTGTGTCATTCTGTTAGATACTA ACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTATCCATGCACCTACCTTAGAAAACAAGTATTGTCGGTTAC CTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAGGGCAGTTCTGAGTGTTTAATTTCAGATTTTTTCCTCTGC ATTTACACACACACGCACACAAACCACACCACACACACACACACACACACACACACACACACACACACACACCAAGTACCAGTATAAGCA TCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCACTCTGTTTCCTGGTCAGCATGTACTCCCCTCATCCGATTC >86683_86683_15_SS18-SSX2_SS18_chr18_23612363_ENST00000545952_SSX2_chrX_52729628_ENST00000336777_length(amino acids)=440AA_BP=327 MLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGMNQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQMNGQMPGPNHMPMQGPGPNQ LNMTNSSMNMPSSSHGSMGGYNHSVPSSQSMPVQNQMTMSQGQPMGNYGPRPNMSMQPNQGPMMHQQPPSQQYNMPQGGGQHYQGQQPPM GMMGQVNQGNHMMGQRQIPPYRPPQQGPPQQYSGQEDYYGDQYSHGGQGPPEGMNQQYYPDGNSQYGQQQDAYQGPPPQQGYPPQQQQYP GQQGYPGQQQGYGPSQGGPGPQYPNYPQGQGQQYGGYRPTQPGPPQPPQQRPYGYDQIMPKKPAEEGNDSEEVPEASGPQNDGKELCPPG -------------------------------------------------------------- >86683_86683_16_SS18-SSX2_SS18_chr18_23612363_ENST00000545952_SSX2_chrX_52729628_ENST00000337502_length(transcript)=2148nt_BP=1284nt ACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGCAGCGAGGCAAGGGGGAGATCACTCCCGCTGCGAT TCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGAATAAAGGAAAGACCTCAGAGTGTTCTCAGGTCAT GAATAAACCACATCTGTTAGTATGGTTTGGTCAGTGGTCCTCAGAAAATCTCATGTCGTGCAACCATACTAATACTCAAGCAAAGGAAAA CTTGAAGGCCTAGACTGAAACAGGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAAATATG CAGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCACGCTCT CACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGGGCCTAAC CATATGCCTATGCAGGGACCTGGACCCAATCAACTCAATATGACAAACAGTTCCATGAATATGCCTTCAAGTAGCCATGGATCCATGGGA GGTTACAACCATTCTGTGCCATCATCACAGAGCATGCCAGTACAGAATCAGATGACAATGAGTCAGGGACAACCAATGGGAAACTATGGT CCCAGACCAAATATGAGTATGCAGCCAAACCAAGGTCCAATGATGCATCAGCAGCCTCCTTCTCAGCAATACAATATGCCACAGGGAGGC GGACAGCATTACCAAGGACAGCAGCCACCTATGGGAATGATGGGTCAAGTTAACCAAGGCAATCATATGATGGGTCAGAGACAGATTCCT CCCTATAGACCTCCTCAACAGGGCCCACCACAGCAGTACTCAGGCCAGGAAGACTATTACGGGGACCAATACAGTCATGGTGGACAAGGT CCTCCAGAAGGCATGAACCAGCAATATTACCCTGATGGAAATTCACAGTATGGCCAACAGCAAGATGCATACCAGGGACCACCTCCACAA CAGGGATATCCACCCCAGCAGCAGCAGTACCCAGGGCAGCAAGGTTACCCAGGACAGCAGCAGGGCTACGGTCCTTCACAGGGTGGTCCA GGTCCTCAGTATCCTAACTACCCACAGGGACAAGGTCAGCAGTATGGAGGATATAGACCAACACAGCCTGGACCACCACAGCCACCCCAG CAGAGGCCTTATGGATATGACCAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGCCAGAAGCATCTGGCCCA CAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGACCCAAAAGGGGGGAACAT GCCTGGACCCACAGACTGCGTGAGAGAAAACAGCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCCCCTCA GGGATACGACACATGCCCATGATGAGAAGCAGAACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAGGTGCA TAGCAAGTGAAAGCAAGTGTTCACAACAGTGAAAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTACGTTGT ATTTTCTTACACTGTGTCATTCTGTTAGATACTAACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTATCCAT GCACCTACCTTAGAAAACAAGTATTGTCGGTTACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAGGGCAG TTCTGAGTGTTTAATTTCAGATTTTTTCCTCTGCATTTACACACACACGCACACAAACCACACCACACACACACACACACACACACACAC ACACACACACACACACCAAGTACCAGTATAAGCATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCACTCTGT >86683_86683_16_SS18-SSX2_SS18_chr18_23612363_ENST00000545952_SSX2_chrX_52729628_ENST00000337502_length(amino acids)=405AA_BP=327 MLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGMNQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQMNGQMPGPNHMPMQGPGPNQ LNMTNSSMNMPSSSHGSMGGYNHSVPSSQSMPVQNQMTMSQGQPMGNYGPRPNMSMQPNQGPMMHQQPPSQQYNMPQGGGQHYQGQQPPM GMMGQVNQGNHMMGQRQIPPYRPPQQGPPQQYSGQEDYYGDQYSHGGQGPPEGMNQQYYPDGNSQYGQQQDAYQGPPPQQGYPPQQQQYP GQQGYPGQQQGYGPSQGGPGPQYPNYPQGQGQQYGGYRPTQPGPPQPPQQRPYGYDQIMPKKPAEEGNDSEEVPEASGPQNDGKELCPPG -------------------------------------------------------------- >86683_86683_17_SS18-SSX2_SS18_chr18_23612363_ENST00000545952_SSX2_chrX_52729630_ENST00000336777_length(transcript)=2294nt_BP=1284nt ACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGCAGCGAGGCAAGGGGGAGATCACTCCCGCTGCGAT TCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGAATAAAGGAAAGACCTCAGAGTGTTCTCAGGTCAT GAATAAACCACATCTGTTAGTATGGTTTGGTCAGTGGTCCTCAGAAAATCTCATGTCGTGCAACCATACTAATACTCAAGCAAAGGAAAA CTTGAAGGCCTAGACTGAAACAGGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAAATATG CAGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCACGCTCT CACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGGGCCTAAC CATATGCCTATGCAGGGACCTGGACCCAATCAACTCAATATGACAAACAGTTCCATGAATATGCCTTCAAGTAGCCATGGATCCATGGGA GGTTACAACCATTCTGTGCCATCATCACAGAGCATGCCAGTACAGAATCAGATGACAATGAGTCAGGGACAACCAATGGGAAACTATGGT CCCAGACCAAATATGAGTATGCAGCCAAACCAAGGTCCAATGATGCATCAGCAGCCTCCTTCTCAGCAATACAATATGCCACAGGGAGGC GGACAGCATTACCAAGGACAGCAGCCACCTATGGGAATGATGGGTCAAGTTAACCAAGGCAATCATATGATGGGTCAGAGACAGATTCCT CCCTATAGACCTCCTCAACAGGGCCCACCACAGCAGTACTCAGGCCAGGAAGACTATTACGGGGACCAATACAGTCATGGTGGACAAGGT CCTCCAGAAGGCATGAACCAGCAATATTACCCTGATGGAAATTCACAGTATGGCCAACAGCAAGATGCATACCAGGGACCACCTCCACAA CAGGGATATCCACCCCAGCAGCAGCAGTACCCAGGGCAGCAAGGTTACCCAGGACAGCAGCAGGGCTACGGTCCTTCACAGGGTGGTCCA GGTCCTCAGTATCCTAACTACCCACAGGGACAAGGTCAGCAGTATGGAGGATATAGACCAACACAGCCTGGACCACCACAGCCACCCCAG CAGAGGCCTTATGGATATGACCAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGCCAGAAGCATCTGGCCCA CAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGAAATAGGGAGGCCCAAGAA AAGGAAGAGAGACGCGGAACAGCTCATCGGTGGAGCAGTCAGAACACACACAACATTGGTCGATTCAGTTTGTCAACTTCTATGGGTGCA GTTCATGGTACCCCCAAAACAATTACACACAACAGGGACCCAAAAGGGGGGAACATGCCTGGACCCACAGACTGCGTGAGAGAAAACAGC TGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCCCCTCAGGGATACGACACATGCCCATGATGAGAAGCAGAA CGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAGGTGCATAGCAAGTGAAAGCAAGTGTTCACAACAGTGAAA AGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTACGTTGTATTTTCTTACACTGTGTCATTCTGTTAGATACTA ACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTATCCATGCACCTACCTTAGAAAACAAGTATTGTCGGTTAC CTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAGGGCAGTTCTGAGTGTTTAATTTCAGATTTTTTCCTCTGC ATTTACACACACACGCACACAAACCACACCACACACACACACACACACACACACACACACACACACACACACCAAGTACCAGTATAAGCA TCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCACTCTGTTTCCTGGTCAGCATGTACTCCCCTCATCCGATTC >86683_86683_17_SS18-SSX2_SS18_chr18_23612363_ENST00000545952_SSX2_chrX_52729630_ENST00000336777_length(amino acids)=440AA_BP=327 MLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGMNQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQMNGQMPGPNHMPMQGPGPNQ LNMTNSSMNMPSSSHGSMGGYNHSVPSSQSMPVQNQMTMSQGQPMGNYGPRPNMSMQPNQGPMMHQQPPSQQYNMPQGGGQHYQGQQPPM GMMGQVNQGNHMMGQRQIPPYRPPQQGPPQQYSGQEDYYGDQYSHGGQGPPEGMNQQYYPDGNSQYGQQQDAYQGPPPQQGYPPQQQQYP GQQGYPGQQQGYGPSQGGPGPQYPNYPQGQGQQYGGYRPTQPGPPQPPQQRPYGYDQIMPKKPAEEGNDSEEVPEASGPQNDGKELCPPG -------------------------------------------------------------- >86683_86683_18_SS18-SSX2_SS18_chr18_23612363_ENST00000545952_SSX2_chrX_52729630_ENST00000337502_length(transcript)=2148nt_BP=1284nt ACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGCAGCGAGGCAAGGGGGAGATCACTCCCGCTGCGAT TCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGAATAAAGGAAAGACCTCAGAGTGTTCTCAGGTCAT GAATAAACCACATCTGTTAGTATGGTTTGGTCAGTGGTCCTCAGAAAATCTCATGTCGTGCAACCATACTAATACTCAAGCAAAGGAAAA CTTGAAGGCCTAGACTGAAACAGGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAAATATG CAGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCACGCTCT CACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGGGCCTAAC CATATGCCTATGCAGGGACCTGGACCCAATCAACTCAATATGACAAACAGTTCCATGAATATGCCTTCAAGTAGCCATGGATCCATGGGA GGTTACAACCATTCTGTGCCATCATCACAGAGCATGCCAGTACAGAATCAGATGACAATGAGTCAGGGACAACCAATGGGAAACTATGGT CCCAGACCAAATATGAGTATGCAGCCAAACCAAGGTCCAATGATGCATCAGCAGCCTCCTTCTCAGCAATACAATATGCCACAGGGAGGC GGACAGCATTACCAAGGACAGCAGCCACCTATGGGAATGATGGGTCAAGTTAACCAAGGCAATCATATGATGGGTCAGAGACAGATTCCT CCCTATAGACCTCCTCAACAGGGCCCACCACAGCAGTACTCAGGCCAGGAAGACTATTACGGGGACCAATACAGTCATGGTGGACAAGGT CCTCCAGAAGGCATGAACCAGCAATATTACCCTGATGGAAATTCACAGTATGGCCAACAGCAAGATGCATACCAGGGACCACCTCCACAA CAGGGATATCCACCCCAGCAGCAGCAGTACCCAGGGCAGCAAGGTTACCCAGGACAGCAGCAGGGCTACGGTCCTTCACAGGGTGGTCCA GGTCCTCAGTATCCTAACTACCCACAGGGACAAGGTCAGCAGTATGGAGGATATAGACCAACACAGCCTGGACCACCACAGCCACCCCAG CAGAGGCCTTATGGATATGACCAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGCCAGAAGCATCTGGCCCA CAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGACCCAAAAGGGGGGAACAT GCCTGGACCCACAGACTGCGTGAGAGAAAACAGCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCCCCTCA GGGATACGACACATGCCCATGATGAGAAGCAGAACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAGGTGCA TAGCAAGTGAAAGCAAGTGTTCACAACAGTGAAAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTACGTTGT ATTTTCTTACACTGTGTCATTCTGTTAGATACTAACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTATCCAT GCACCTACCTTAGAAAACAAGTATTGTCGGTTACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAGGGCAG TTCTGAGTGTTTAATTTCAGATTTTTTCCTCTGCATTTACACACACACGCACACAAACCACACCACACACACACACACACACACACACAC ACACACACACACACACCAAGTACCAGTATAAGCATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCACTCTGT >86683_86683_18_SS18-SSX2_SS18_chr18_23612363_ENST00000545952_SSX2_chrX_52729630_ENST00000337502_length(amino acids)=405AA_BP=327 MLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGMNQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQMNGQMPGPNHMPMQGPGPNQ LNMTNSSMNMPSSSHGSMGGYNHSVPSSQSMPVQNQMTMSQGQPMGNYGPRPNMSMQPNQGPMMHQQPPSQQYNMPQGGGQHYQGQQPPM GMMGQVNQGNHMMGQRQIPPYRPPQQGPPQQYSGQEDYYGDQYSHGGQGPPEGMNQQYYPDGNSQYGQQQDAYQGPPPQQGYPPQQQQYP GQQGYPGQQQGYGPSQGGPGPQYPNYPQGQGQQYGGYRPTQPGPPQPPQQRPYGYDQIMPKKPAEEGNDSEEVPEASGPQNDGKELCPPG -------------------------------------------------------------- >86683_86683_19_SS18-SSX2_SS18_chr18_23637552_ENST00000269137_SSX2_chrX_52731680_ENST00000336777_length(transcript)=1777nt_BP=717nt TTTCCTGAATCGCTGAAGCGGAGGGAGGACTAGGGCGCGACTCTACCATTAGCGCAAAGGGCCAGGGGGTGCCCATAAGGAGAGCTCGGG GGATGCAGGGACGGTCAAGAAAGTTCGGCCAGGGCTAGGGGTCCCCCAGAGTGCGCAGAAGCGAGACATCCCTGGCCAGAGTGCCTCCTT CAGGCCTGCCCCGCCGCCCGTCCGCTTGGCCCCGGAGCATCCCCCCTCCCCGCATTCCCGAGGTGGTTCAGCCCGAGAGGCCGGCGTCTC TCCCCCAGTTTGCCGTTCACCCGGAGCGCTCGGGACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGC AGCGAGGCAAGGGGGAGATCACTCCCGCTGCGATTCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGA ATAAAGGAAAGACCTCAGAGTGTTCTCAGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAA ATATGCAGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCAC GCTCTCACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGTTG AACGTCCTCAGATGACTTTCGGCAGGCTCCAGGGAATCTCCCCGAAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGG AAGTGCCAGAAGCATCTGGCCCACAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGAT CTGGAAATAGGGAGGCCCAAGAAAAGGAAGAGAGACGCGGAACAGCTCATCGGTGGAGCAGTCAGAACACACACAACATTGGTCGATTCA GTTTGTCAACTTCTATGGGTGCAGTTCATGGTACCCCCAAAACAATTACACACAACAGGGACCCAAAAGGGGGGAACATGCCTGGACCCA CAGACTGCGTGAGAGAAAACAGCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCCCCTCAGGGATACGACA CATGCCCATGATGAGAAGCAGAACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAGGTGCATAGCAAGTGAA AGCAAGTGTTCACAACAGTGAAAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTACGTTGTATTTTCTTACA CTGTGTCATTCTGTTAGATACTAACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTATCCATGCACCTACCTT AGAAAACAAGTATTGTCGGTTACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAGGGCAGTTCTGAGTGTT TAATTTCAGATTTTTTCCTCTGCATTTACACACACACGCACACAAACCACACCACACACACACACACACACACACACACACACACACACA CACACCAAGTACCAGTATAAGCATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCACTCTGTTTCCTGGTCAG >86683_86683_19_SS18-SSX2_SS18_chr18_23637552_ENST00000269137_SSX2_chrX_52731680_ENST00000336777_length(amino acids)=315AA_BP=185 MARVPPSGLPRRPSAWPRSIPPPRIPEVVQPERPASLPQFAVHPERSGLADSGDGGNMSVAFAAPRQRGKGEITPAAIQKMLDDNNHLIQ CIMDSQNKGKTSECSQYQQMLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGMNQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQM NGQMPVERPQMTFGRLQGISPKIMPKKPAEEGNDSEEVPEASGPQNDGKELCPPGKPTTSEKIHERSGNREAQEKEERRGTAHRWSSQNT -------------------------------------------------------------- >86683_86683_20_SS18-SSX2_SS18_chr18_23637552_ENST00000269137_SSX2_chrX_52731680_ENST00000337502_length(transcript)=1631nt_BP=717nt TTTCCTGAATCGCTGAAGCGGAGGGAGGACTAGGGCGCGACTCTACCATTAGCGCAAAGGGCCAGGGGGTGCCCATAAGGAGAGCTCGGG GGATGCAGGGACGGTCAAGAAAGTTCGGCCAGGGCTAGGGGTCCCCCAGAGTGCGCAGAAGCGAGACATCCCTGGCCAGAGTGCCTCCTT CAGGCCTGCCCCGCCGCCCGTCCGCTTGGCCCCGGAGCATCCCCCCTCCCCGCATTCCCGAGGTGGTTCAGCCCGAGAGGCCGGCGTCTC TCCCCCAGTTTGCCGTTCACCCGGAGCGCTCGGGACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGC AGCGAGGCAAGGGGGAGATCACTCCCGCTGCGATTCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGA ATAAAGGAAAGACCTCAGAGTGTTCTCAGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAA ATATGCAGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCAC GCTCTCACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGTTG AACGTCCTCAGATGACTTTCGGCAGGCTCCAGGGAATCTCCCCGAAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGG AAGTGCCAGAAGCATCTGGCCCACAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGAT CTGGACCCAAAAGGGGGGAACATGCCTGGACCCACAGACTGCGTGAGAGAAAACAGCTGGTGATTTATGAAGAGATCAGCGACCCTGAGG AAGATGACGAGTAACTCCCCTCAGGGATACGACACATGCCCATGATGAGAAGCAGAACGTGGTGACCTTTCACGAACATGGGCATGGCTG CGGACCCCTCGTCATCAGGTGCATAGCAAGTGAAAGCAAGTGTTCACAACAGTGAAAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAG TTCGATGTTAGCGTTTACGTTGTATTTTCTTACACTGTGTCATTCTGTTAGATACTAACATTTTCATTGATGAGCAAGACATACTTAATG CATATTTTGGTTTGTGTATCCATGCACCTACCTTAGAAAACAAGTATTGTCGGTTACCTCTGCATGGAACAGCATTACCCTCCTCTCTCC CCAGATGTGACTACTGAGGGCAGTTCTGAGTGTTTAATTTCAGATTTTTTCCTCTGCATTTACACACACACGCACACAAACCACACCACA CACACACACACACACACACACACACACACACACACACACCAAGTACCAGTATAAGCATCTGCCATCTGCTTTTCCCATTGCCATGCGTCC TGGTCAAGCTCCCCTCACTCTGTTTCCTGGTCAGCATGTACTCCCCTCATCCGATTCCCCTGTAGCAGTCACTGACAGTTAATAAACCTT >86683_86683_20_SS18-SSX2_SS18_chr18_23637552_ENST00000269137_SSX2_chrX_52731680_ENST00000337502_length(amino acids)=280AA_BP=185 MARVPPSGLPRRPSAWPRSIPPPRIPEVVQPERPASLPQFAVHPERSGLADSGDGGNMSVAFAAPRQRGKGEITPAAIQKMLDDNNHLIQ CIMDSQNKGKTSECSQYQQMLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGMNQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQM NGQMPVERPQMTFGRLQGISPKIMPKKPAEEGNDSEEVPEASGPQNDGKELCPPGKPTTSEKIHERSGPKRGEHAWTHRLRERKQLVIYE -------------------------------------------------------------- >86683_86683_21_SS18-SSX2_SS18_chr18_23637552_ENST00000415083_SSX2_chrX_52731680_ENST00000336777_length(transcript)=1501nt_BP=441nt AGTTTGCCGTTCACCCGGAGCGCTCGGGACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGCAGCGAG GCAAGGGGGAGATCACTCCCGCTGCGATTCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGAATAAAG GAAAGACCTCAGAGTGTTCTCAGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAAATATGC AGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCACGCTCTC ACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGTTGAACGTC CTCAGATGACTTTCGGCAGGCTCCAGGGAATCTCCCCGAAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGC CAGAAGCATCTGGCCCACAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGAA ATAGGGAGGCCCAAGAAAAGGAAGAGAGACGCGGAACAGCTCATCGGTGGAGCAGTCAGAACACACACAACATTGGTCGATTCAGTTTGT CAACTTCTATGGGTGCAGTTCATGGTACCCCCAAAACAATTACACACAACAGGGACCCAAAAGGGGGGAACATGCCTGGACCCACAGACT GCGTGAGAGAAAACAGCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCCCCTCAGGGATACGACACATGCC CATGATGAGAAGCAGAACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAGGTGCATAGCAAGTGAAAGCAAG TGTTCACAACAGTGAAAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTACGTTGTATTTTCTTACACTGTGT CATTCTGTTAGATACTAACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTATCCATGCACCTACCTTAGAAAA CAAGTATTGTCGGTTACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAGGGCAGTTCTGAGTGTTTAATTT CAGATTTTTTCCTCTGCATTTACACACACACGCACACAAACCACACCACACACACACACACACACACACACACACACACACACACACACC AAGTACCAGTATAAGCATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCACTCTGTTTCCTGGTCAGCATGTA >86683_86683_21_SS18-SSX2_SS18_chr18_23637552_ENST00000415083_SSX2_chrX_52731680_ENST00000336777_length(amino acids)=258AA_BP=128 MSVAFAAPRQRGKGEITPAAIQKMLDDNNHLIQCIMDSQNKGKTSECSQYQQMLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGM NQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQMNGQMPVERPQMTFGRLQGISPKIMPKKPAEEGNDSEEVPEASGPQNDGKELCPPGKP -------------------------------------------------------------- >86683_86683_22_SS18-SSX2_SS18_chr18_23637552_ENST00000415083_SSX2_chrX_52731680_ENST00000337502_length(transcript)=1355nt_BP=441nt AGTTTGCCGTTCACCCGGAGCGCTCGGGACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGCAGCGAG GCAAGGGGGAGATCACTCCCGCTGCGATTCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGAATAAAG GAAAGACCTCAGAGTGTTCTCAGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAAATATGC AGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCACGCTCTC ACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGTTGAACGTC CTCAGATGACTTTCGGCAGGCTCCAGGGAATCTCCCCGAAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTGC CAGAAGCATCTGGCCCACAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGAC CCAAAAGGGGGGAACATGCCTGGACCCACAGACTGCGTGAGAGAAAACAGCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATG ACGAGTAACTCCCCTCAGGGATACGACACATGCCCATGATGAGAAGCAGAACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACC CCTCGTCATCAGGTGCATAGCAAGTGAAAGCAAGTGTTCACAACAGTGAAAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGAT GTTAGCGTTTACGTTGTATTTTCTTACACTGTGTCATTCTGTTAGATACTAACATTTTCATTGATGAGCAAGACATACTTAATGCATATT TTGGTTTGTGTATCCATGCACCTACCTTAGAAAACAAGTATTGTCGGTTACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGAT GTGACTACTGAGGGCAGTTCTGAGTGTTTAATTTCAGATTTTTTCCTCTGCATTTACACACACACGCACACAAACCACACCACACACACA CACACACACACACACACACACACACACACACACCAAGTACCAGTATAAGCATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCA AGCTCCCCTCACTCTGTTTCCTGGTCAGCATGTACTCCCCTCATCCGATTCCCCTGTAGCAGTCACTGACAGTTAATAAACCTTTGCAAA >86683_86683_22_SS18-SSX2_SS18_chr18_23637552_ENST00000415083_SSX2_chrX_52731680_ENST00000337502_length(amino acids)=223AA_BP=128 MSVAFAAPRQRGKGEITPAAIQKMLDDNNHLIQCIMDSQNKGKTSECSQYQQMLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGM NQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQMNGQMPVERPQMTFGRLQGISPKIMPKKPAEEGNDSEEVPEASGPQNDGKELCPPGKP -------------------------------------------------------------- >86683_86683_23_SS18-SSX2_SS18_chr18_23637552_ENST00000545952_SSX2_chrX_52731680_ENST00000336777_length(transcript)=1592nt_BP=532nt ACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGCAGCGAGGCAAGGGGGAGATCACTCCCGCTGCGAT TCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGAATAAAGGAAAGACCTCAGAGTGTTCTCAGGTCAT GAATAAACCACATCTGTTAGTATGGTTTGGTCAGTGGTCCTCAGAAAATCTCATGTCGTGCAACCATACTAATACTCAAGCAAAGGAAAA CTTGAAGGCCTAGACTGAAACAGGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAAATATG CAGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCACGCTCT CACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGTTGAACGT CCTCAGATGACTTTCGGCAGGCTCCAGGGAATCTCCCCGAAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTG CCAGAAGCATCTGGCCCACAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGA AATAGGGAGGCCCAAGAAAAGGAAGAGAGACGCGGAACAGCTCATCGGTGGAGCAGTCAGAACACACACAACATTGGTCGATTCAGTTTG TCAACTTCTATGGGTGCAGTTCATGGTACCCCCAAAACAATTACACACAACAGGGACCCAAAAGGGGGGAACATGCCTGGACCCACAGAC TGCGTGAGAGAAAACAGCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGATGACGAGTAACTCCCCTCAGGGATACGACACATGC CCATGATGAGAAGCAGAACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGACCCCTCGTCATCAGGTGCATAGCAAGTGAAAGCAA GTGTTCACAACAGTGAAAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGATGTTAGCGTTTACGTTGTATTTTCTTACACTGTG TCATTCTGTTAGATACTAACATTTTCATTGATGAGCAAGACATACTTAATGCATATTTTGGTTTGTGTATCCATGCACCTACCTTAGAAA ACAAGTATTGTCGGTTACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGATGTGACTACTGAGGGCAGTTCTGAGTGTTTAATT TCAGATTTTTTCCTCTGCATTTACACACACACGCACACAAACCACACCACACACACACACACACACACACACACACACACACACACACAC CAAGTACCAGTATAAGCATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTCAAGCTCCCCTCACTCTGTTTCCTGGTCAGCATGT >86683_86683_23_SS18-SSX2_SS18_chr18_23637552_ENST00000545952_SSX2_chrX_52731680_ENST00000336777_length(amino acids)=206AA_BP=76 MLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGMNQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQMNGQMPVERPQMTFGRLQGI SPKIMPKKPAEEGNDSEEVPEASGPQNDGKELCPPGKPTTSEKIHERSGNREAQEKEERRGTAHRWSSQNTHNIGRFSLSTSMGAVHGTP -------------------------------------------------------------- >86683_86683_24_SS18-SSX2_SS18_chr18_23637552_ENST00000545952_SSX2_chrX_52731680_ENST00000337502_length(transcript)=1446nt_BP=532nt ACTTGCCGATAGTGGTGACGGCGGCAACATGTCTGTGGCTTTCGCGGCCCCGAGGCAGCGAGGCAAGGGGGAGATCACTCCCGCTGCGAT TCAGAAGATGTTGGATGACAATAACCATCTTATTCAGTGTATAATGGACTCTCAGAATAAAGGAAAGACCTCAGAGTGTTCTCAGGTCAT GAATAAACCACATCTGTTAGTATGGTTTGGTCAGTGGTCCTCAGAAAATCTCATGTCGTGCAACCATACTAATACTCAAGCAAAGGAAAA CTTGAAGGCCTAGACTGAAACAGGTATCAGCAGATGTTGCACACAAACTTGGTATACCTTGCTACAATAGCAGATTCTAATCAAAATATG CAGTCTCTTTTACCAGCACCACCCACACAGAATATGCCTATGGGTCCTGGAGGGATGAATCAGAGCGGCCCTCCCCCACCTCCACGCTCT CACAACATGCCTTCAGATGGAATGGTAGGTGGGGGTCCTCCTGCACCGCACATGCAGAACCAGATGAACGGCCAGATGCCTGTTGAACGT CCTCAGATGACTTTCGGCAGGCTCCAGGGAATCTCCCCGAAGATCATGCCCAAGAAGCCAGCAGAGGAAGGAAATGATTCGGAGGAAGTG CCAGAAGCATCTGGCCCACAAAATGATGGGAAAGAGCTGTGCCCCCCGGGAAAACCAACTACCTCTGAGAAGATTCACGAGAGATCTGGA CCCAAAAGGGGGGAACATGCCTGGACCCACAGACTGCGTGAGAGAAAACAGCTGGTGATTTATGAAGAGATCAGCGACCCTGAGGAAGAT GACGAGTAACTCCCCTCAGGGATACGACACATGCCCATGATGAGAAGCAGAACGTGGTGACCTTTCACGAACATGGGCATGGCTGCGGAC CCCTCGTCATCAGGTGCATAGCAAGTGAAAGCAAGTGTTCACAACAGTGAAAAGTTGAGCGTCATTTTTCTTAGTGTGCCAAGAGTTCGA TGTTAGCGTTTACGTTGTATTTTCTTACACTGTGTCATTCTGTTAGATACTAACATTTTCATTGATGAGCAAGACATACTTAATGCATAT TTTGGTTTGTGTATCCATGCACCTACCTTAGAAAACAAGTATTGTCGGTTACCTCTGCATGGAACAGCATTACCCTCCTCTCTCCCCAGA TGTGACTACTGAGGGCAGTTCTGAGTGTTTAATTTCAGATTTTTTCCTCTGCATTTACACACACACGCACACAAACCACACCACACACAC ACACACACACACACACACACACACACACACACACCAAGTACCAGTATAAGCATCTGCCATCTGCTTTTCCCATTGCCATGCGTCCTGGTC AAGCTCCCCTCACTCTGTTTCCTGGTCAGCATGTACTCCCCTCATCCGATTCCCCTGTAGCAGTCACTGACAGTTAATAAACCTTTGCAA >86683_86683_24_SS18-SSX2_SS18_chr18_23637552_ENST00000545952_SSX2_chrX_52731680_ENST00000337502_length(amino acids)=171AA_BP=76 MLHTNLVYLATIADSNQNMQSLLPAPPTQNMPMGPGGMNQSGPPPPPRSHNMPSDGMVGGGPPAPHMQNQMNGQMPVERPQMTFGRLQGI -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for SS18-SSX2 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Tgene | SSX2 | chr18:23612363 | chrX:52729628 | ENST00000336777 | 4 | 9 | 25_80 | 110.0 | 420.3333333333333 | RAB3IP | |
| Tgene | SSX2 | chr18:23612363 | chrX:52729628 | ENST00000337502 | 4 | 8 | 25_80 | 110.0 | 376.0 | RAB3IP | |
| Tgene | SSX2 | chr18:23612363 | chrX:52729630 | ENST00000336777 | 4 | 9 | 25_80 | 110.0 | 420.3333333333333 | RAB3IP | |
| Tgene | SSX2 | chr18:23612363 | chrX:52729630 | ENST00000337502 | 4 | 8 | 25_80 | 110.0 | 376.0 | RAB3IP | |
| Tgene | SSX2 | chr18:23612363 | chrX:52729628 | ENST00000336777 | 4 | 9 | 1_80 | 110.0 | 420.3333333333333 | SSX2IP | |
| Tgene | SSX2 | chr18:23612363 | chrX:52729628 | ENST00000337502 | 4 | 8 | 1_80 | 110.0 | 376.0 | SSX2IP | |
| Tgene | SSX2 | chr18:23612363 | chrX:52729630 | ENST00000336777 | 4 | 9 | 1_80 | 110.0 | 420.3333333333333 | SSX2IP | |
| Tgene | SSX2 | chr18:23612363 | chrX:52729630 | ENST00000337502 | 4 | 8 | 1_80 | 110.0 | 376.0 | SSX2IP |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for SS18-SSX2 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for SS18-SSX2 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
| Hgene | SS18 | C0039101 | synovial sarcoma | 2 | CTD_human;ORPHANET |
| Tgene | C0033578 | Prostatic Neoplasms | 1 | CTD_human | |
| Tgene | C0039101 | synovial sarcoma | 1 | CTD_human;ORPHANET | |
| Tgene | C0376358 | Malignant neoplasm of prostate | 1 | CTD_human |
