
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:UBA1-UBA7 (FusionGDB2 ID:HG7317TG7318) |
Fusion Gene Summary for UBA1-UBA7 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: UBA1-UBA7 | Fusion gene ID: hg7317tg7318 | Hgene | Tgene | Gene symbol | UBA1 | UBA7 | Gene ID | 7317 | 7318 |
| Gene name | ubiquitin like modifier activating enzyme 1 | ubiquitin like modifier activating enzyme 7 | |
| Synonyms | A1S9|A1S9T|A1ST|AMCX1|CFAP124|GXP1|POC20|SMAX2|UBA1A|UBE1|UBE1X | D8|UBA1B|UBE1L|UBE2 | |
| Cytomap | ('UBA1')('UBA7') Xp11.3 | 3p21.31 | |
| Type of gene | protein-coding | protein-coding | |
| Description | ubiquitin-like modifier-activating enzyme 1A1S9T and BN75 temperature sensitivity complementingPOC20 centriolar protein homologUBA1, ubiquitin-activating enzyme E1 homolog Atesticular secretory protein Li 63 | ubiquitin-like modifier-activating enzyme 7UBA1, ubiquitin-activating enzyme E1 homolog BUBA7, ubiquitin-activating enzyme E1ubiquitin-activating enzyme 7ubiquitin-activating enzyme E1 homologubiquitin-activating enzyme E1-related proteinubiquitin-a | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000335972, ENST00000377269, ENST00000377351, ENST00000490869, | ||
| Fusion gene scores | * DoF score | 10 X 10 X 5=500 | 6 X 9 X 5=270 |
| # samples | 11 | 8 | |
| ** MAII score | log2(11/500*10)=-2.18442457113743 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(8/270*10)=-1.75488750216347 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: UBA1 [Title/Abstract] AND UBA7 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | UBA1(47071911)-UBA7(49845545), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | UBA1-UBA7 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. UBA1-UBA7 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. UBA1-UBA7 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. UBA1-UBA7 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | UBA1 | GO:0006974 | cellular response to DNA damage stimulus | 22456334 |
| Tgene | UBA7 | GO:0006464 | cellular protein modification process | 16428300 |
| Tgene | UBA7 | GO:0032020 | ISG15-protein conjugation | 16428300 |
 Fusion gene breakpoints across UBA1 (5'-gene) Fusion gene breakpoints across UBA1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
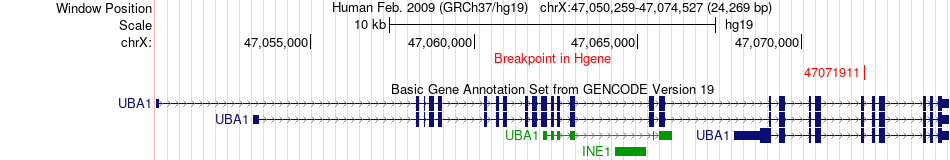 |
 Fusion gene breakpoints across UBA7 (3'-gene) Fusion gene breakpoints across UBA7 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
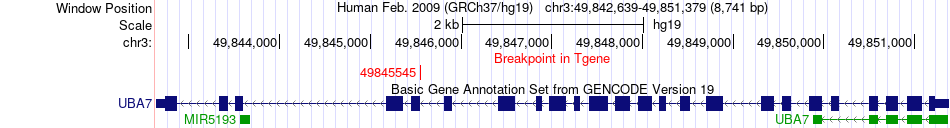 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | Non-Cancer | 201N | UBA1 | chrX | 47071911 | + | UBA7 | chr3 | 49845545 | - |
| ChimerDB4 | Non-Cancer | 5759N | UBA1 | chrX | 47071911 | + | UBA7 | chr3 | 49845545 | - |
| ChimerDB4 | Non-Cancer | NT15 | UBA1 | chrX | 47071911 | + | UBA7 | chr3 | 49845545 | - |
Top |
Fusion Gene ORF analysis for UBA1-UBA7 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000335972 | ENST00000494212 | UBA1 | chrX | 47071911 | + | UBA7 | chr3 | 49845545 | - |
| 5CDS-intron | ENST00000377269 | ENST00000494212 | UBA1 | chrX | 47071911 | + | UBA7 | chr3 | 49845545 | - |
| 5CDS-intron | ENST00000377351 | ENST00000494212 | UBA1 | chrX | 47071911 | + | UBA7 | chr3 | 49845545 | - |
| In-frame | ENST00000335972 | ENST00000333486 | UBA1 | chrX | 47071911 | + | UBA7 | chr3 | 49845545 | - |
| In-frame | ENST00000377269 | ENST00000333486 | UBA1 | chrX | 47071911 | + | UBA7 | chr3 | 49845545 | - |
| In-frame | ENST00000377351 | ENST00000333486 | UBA1 | chrX | 47071911 | + | UBA7 | chr3 | 49845545 | - |
| intron-3CDS | ENST00000490869 | ENST00000333486 | UBA1 | chrX | 47071911 | + | UBA7 | chr3 | 49845545 | - |
| intron-intron | ENST00000490869 | ENST00000494212 | UBA1 | chrX | 47071911 | + | UBA7 | chr3 | 49845545 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000377351 | UBA1 | chrX | 47071911 | + | ENST00000333486 | UBA7 | chr3 | 49845545 | - | 3353 | 2643 | 30 | 3251 | 1073 |
| ENST00000335972 | UBA1 | chrX | 47071911 | + | ENST00000333486 | UBA7 | chr3 | 49845545 | - | 3446 | 2736 | 150 | 3344 | 1064 |
| ENST00000377269 | UBA1 | chrX | 47071911 | + | ENST00000333486 | UBA7 | chr3 | 49845545 | - | 2386 | 1676 | 779 | 2284 | 501 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000377351 | ENST00000333486 | UBA1 | chrX | 47071911 | + | UBA7 | chr3 | 49845545 | - | 0.002621877 | 0.9973781 |
| ENST00000335972 | ENST00000333486 | UBA1 | chrX | 47071911 | + | UBA7 | chr3 | 49845545 | - | 0.003174356 | 0.9968257 |
| ENST00000377269 | ENST00000333486 | UBA1 | chrX | 47071911 | + | UBA7 | chr3 | 49845545 | - | 0.03270269 | 0.96729726 |
Top |
Fusion Genomic Features for UBA1-UBA7 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
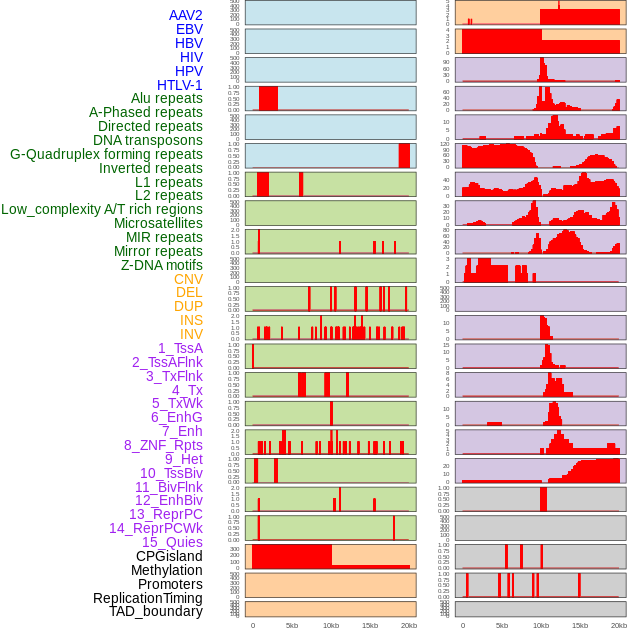 |
Top |
Fusion Protein Features for UBA1-UBA7 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chrX:47071911/chr3:49845545) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | UBA1 | chrX:47071911 | chr3:49845545 | ENST00000335972 | + | 21 | 26 | 5_11 | 851 | 1059.0 | Motif | Note=Nuclear localization signal |
| Hgene | UBA1 | chrX:47071911 | chr3:49845545 | ENST00000377351 | + | 21 | 26 | 5_11 | 851 | 1059.0 | Motif | Note=Nuclear localization signal |
| Hgene | UBA1 | chrX:47071911 | chr3:49845545 | ENST00000335972 | + | 21 | 26 | 576_577 | 851 | 1059.0 | Nucleotide binding | ATP |
| Hgene | UBA1 | chrX:47071911 | chr3:49845545 | ENST00000377351 | + | 21 | 26 | 576_577 | 851 | 1059.0 | Nucleotide binding | ATP |
| Hgene | UBA1 | chrX:47071911 | chr3:49845545 | ENST00000335972 | + | 21 | 26 | 63_611 | 851 | 1059.0 | Region | Note=2 approximate repeats |
| Hgene | UBA1 | chrX:47071911 | chr3:49845545 | ENST00000377351 | + | 21 | 26 | 63_611 | 851 | 1059.0 | Region | Note=2 approximate repeats |
| Hgene | UBA1 | chrX:47071911 | chr3:49845545 | ENST00000335972 | + | 21 | 26 | 459_611 | 851 | 1059.0 | Repeat | Note=1-2 |
| Hgene | UBA1 | chrX:47071911 | chr3:49845545 | ENST00000335972 | + | 21 | 26 | 63_199 | 851 | 1059.0 | Repeat | Note=1-1 |
| Hgene | UBA1 | chrX:47071911 | chr3:49845545 | ENST00000377351 | + | 21 | 26 | 459_611 | 851 | 1059.0 | Repeat | Note=1-2 |
| Hgene | UBA1 | chrX:47071911 | chr3:49845545 | ENST00000377351 | + | 21 | 26 | 63_199 | 851 | 1059.0 | Repeat | Note=1-1 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | UBA7 | chrX:47071911 | chr3:49845545 | ENST00000333486 | 18 | 24 | 442_471 | 810 | 1013.0 | Nucleotide binding | ATP | |
| Tgene | UBA7 | chrX:47071911 | chr3:49845545 | ENST00000333486 | 18 | 24 | 23_575 | 810 | 1013.0 | Region | Note=2 approximate repeats | |
| Tgene | UBA7 | chrX:47071911 | chr3:49845545 | ENST00000333486 | 18 | 24 | 23_159 | 810 | 1013.0 | Repeat | Note=1-1 | |
| Tgene | UBA7 | chrX:47071911 | chr3:49845545 | ENST00000333486 | 18 | 24 | 423_575 | 810 | 1013.0 | Repeat | Note=1-2 |
Top |
Fusion Gene Sequence for UBA1-UBA7 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >95805_95805_1_UBA1-UBA7_UBA1_chrX_47071911_ENST00000335972_UBA7_chr3_49845545_ENST00000333486_length(transcript)=3446nt_BP=2736nt GGCTCGGCTGTAAGGAGGTGGCAGGGACAACCACAACCACAACGGCCGGGGGAGGAGAAGGCGGCAGCGGCGATTCTAGGCGGCCCAGGC GGCGGGGAGGAGGAGAAGGAGGAGGGTGGCGGCCGGGCTTGGCTTCGGCTCCTTGAGGAGTTGGCGGCGGCGCGACCCGGGGAACCGGCA TTGATGTCCAGCTCGCCGCTGTCCAAGAAACGTCGCGTGTCCGGGCCTGATCCAAAGCCGGGTTCTAACTGCTCCCCTGCCCAGTCCGTG TTGTCCGAAGTGCCCTCGGTGCCAACCAACGGAATGGCCAAGAACGGCAGTGAAGCAGACATAGACGAGGGCCTTTACTCCCGGCAGCTG TATGTGTTGGGCCATGAGGCAATGAAGCGGCTCCAGACATCCAGTGTCCTGGTATCAGGCCTGCGGGGCCTGGGCGTGGAGATCGCTAAG AACATCATCCTTGGTGGGGTCAAGGCTGTTACCCTACATGACCAGGGCACTGCCCAGTGGGCTGATCTTTCCTCCCAGTTCTACCTGCGG GAGGAGGACATCGGTAAAAACCGGGCCGAGGTATCACAGCCCCGCCTCGCTGAGCTCAACAGCTATGTGCCTGTCACTGCCTACACTGGA CCCCTCGTTGAGGACTTCCTTAGTGGTTTCCAGGTGGTGGTGCTCACCAACACCCCCCTGGAGGACCAGCTGCGAGTGGGTGAGTTCTGT CACAACCGTGGCATCAAGCTGGTGGTGGCAGACACGCGGGGCCTGTTTGGGCAGCTCTTCTGTGACTTTGGAGAGGAAATGATCCTCACA GATTCCAATGGGGAGCAGCCACTCAGTGCTATGGTTTCTATGGTTACCAAGGACAACCCCGGTGTGGTTACCTGCCTGGATGAGGCCCGA CACGGGTTTGAGAGCGGGGACTTTGTCTCCTTTTCAGAAGTACAGGGCATGGTTGAACTCAACGGAAATCAGCCCATGGAGATCAAAGTC CTGGGTCCTTATACCTTTAGCATCTGTGACACCTCCAACTTCTCCGACTACATCCGTGGAGGCATCGTCAGTCAGGTCAAAGTACCTAAG AAGATTAGCTTTAAATCCTTGGTGGCCTCACTGGCAGAACCTGACTTTGTGGTGACGGACTTCGCCAAGTTTTCTCGCCCTGCCCAGCTG CACATTGGCTTCCAGGCCCTGCACCAGTTCTGTGCTCAGCATGGCCGGCCACCTCGGCCCCGCAATGAGGAGGATGCAGCAGAACTGGTA GCCTTAGCACAGGCTGTGAATGCTCGAGCCCTGCCAGCAGTGCAGCAAAATAACCTGGACGAGGACCTCATCCGGAAGCTGGCATATGTG GCTGCTGGGGATCTGGCACCCATAAACGCCTTCATTGGGGGCCTGGCTGCCCAGGAAGTCATGAAGGCCTGCTCCGGGAAGTTCATGCCC ATCATGCAGTGGCTATACTTTGATGCCCTTGAGTGTCTCCCTGAGGACAAAGAGGTCCTCACAGAGGACAAGTGCCTCCAGCGCCAGAAC CGTTATGACGGGCAAGTGGCTGTGTTTGGCTCAGACCTGCAAGAGAAGCTGGGCAAGCAGAAGTATTTCCTGGTGGGTGCGGGGGCCATT GGCTGTGAGCTGCTCAAGAACTTTGCCATGATTGGGCTGGGCTGCGGGGAGGGTGGAGAAATCATCGTTACAGACATGGACACCATTGAG AAGTCAAATCTGAATCGACAGTTTCTTTTCCGGCCCTGGGATGTCACGAAGTTAAAGTCTGACACGGCTGCTGCAGCTGTGCGCCAAATG AATCCACATATCCGGGTGACAAGCCACCAGAACCGTGTGGGTCCTGACACGGAGCGCATCTATGATGACGATTTTTTCCAAAACCTAGAT GGCGTGGCCAATGCCCTGGACAACGTGGATGCCCGCATGTACATGGACCGCCGCTGTGTCTACTACCGGAAGCCACTGCTGGAGTCAGGC ACACTGGGCACCAAAGGCAATGTGCAGGTGGTGATCCCCTTCCTGACAGAGTCGTACAGTTCCAGCCAGGACCCACCTGAGAAGTCCATC CCCATCTGTACCCTGAAGAACTTCCCTAATGCCATCGAGCACACCCTGCAGTGGGCTCGGGATGAGTTTGAAGGCCTCTTCAAGCAGCCA GCAGAAAATGTCAACCAGTACCTCACAGACCCCAAGTTTGTGGAGCGAACACTGCGGCTGGCAGGCACTCAGCCCTTGGAGGTGCTGGAG GCTGTGCAGCGCAGCCTGGTGCTGCAGCGACCACAGACCTGGGCTGACTGCGTGACCTGGGCCTGCCACCACTGGCACACCCAGTACTCG AACAACATCCGGCAGCTGCTGCACAACTTCCCTCCTGACCAGCTCACAAGCTCAGGAGCGCCGTTCTGGTCTGGGCCCAAACGCTGTCCA CACCCGCTCACCTTTGATGTCAACAATCCCCTGCATCTGGACTATGTGATGGCTGCTGCCAACCTGTTTGCCCAGACCTACGGGCTGACA GGCTCTCAGGACCGAGCTGCTGTGGCCACATTCCTGCAGTCTGTGCAGGTCCCCGAATTCACCCCCAAGTCTGGCGTCAAGATCCATGTT TCTGACCAGGAGCTGCAGAGCGCCAATGCCTCTGTTGATGACAGTCGTCTAGAGGAGCTCAAAGCCACTCTGCCCAGCCCAGACAAGCTC CCTGGATTCAAGATGTACCCCATTGACTTTGAGAAGGATGATGACAGCAACTTCCATGTGGACTTTGTGGTAGCGGCAGCTAGCCTGAGA TGTCAGAACTACGGGATTCCACCGGTCAACCGTGCCCAGAGCAAGCGAATTGTGGGCCAGATTATCCCAGCCATTGCCACCACTACAGCA GCTGTGGCAGGCCTGTTGGGCCTGGAGCTGTATAAGGTGGTGAGTGGGCCACGGCCTCGTAGTGCCTTTCGCCACAGCTACCTACATCTG GCTGAAAACTACCTCATCCGCTATATGCCTTTTGCCCCAGCCATCCAGACGTTCCATCACCTGAAGTGGACCTCTTGGGACCGTCTGAAG GTACCAGCTGGGCAGCCTGAGAGGACCCTGGAGTCGCTGCTGGCTCATCTTCAGGAGCAGCACGGGTTGAGGGTGAGGATCCTGCTGCAC GGCTCAGCCCTGCTCTATGCGGCCGGATGGTCACCTGAAAAGCAGGCCCAGCACCTGCCCCTCAGGGTGACAGAACTGGTTCAGCAGCTG ACAGGCCAGGCACCTGCTCCTGGGCAGCGGGTGTTGGTGCTAGAGCTGAGCTGTGAGGGTGACGACGAGGACACTGCCTTCCCACCTCTG CACTATGAGCTGTGACAAGGCAGCCACCCTGTCACCTAGCTCAATGGAGCCCCGGATCCCAAGCCCTGCATTGTAAGCCCACAGTAGGCA >95805_95805_1_UBA1-UBA7_UBA1_chrX_47071911_ENST00000335972_UBA7_chr3_49845545_ENST00000333486_length(amino acids)=1064AA_BP=70 MAAARPGEPALMSSSPLSKKRRVSGPDPKPGSNCSPAQSVLSEVPSVPTNGMAKNGSEADIDEGLYSRQLYVLGHEAMKRLQTSSVLVSG LRGLGVEIAKNIILGGVKAVTLHDQGTAQWADLSSQFYLREEDIGKNRAEVSQPRLAELNSYVPVTAYTGPLVEDFLSGFQVVVLTNTPL EDQLRVGEFCHNRGIKLVVADTRGLFGQLFCDFGEEMILTDSNGEQPLSAMVSMVTKDNPGVVTCLDEARHGFESGDFVSFSEVQGMVEL NGNQPMEIKVLGPYTFSICDTSNFSDYIRGGIVSQVKVPKKISFKSLVASLAEPDFVVTDFAKFSRPAQLHIGFQALHQFCAQHGRPPRP RNEEDAAELVALAQAVNARALPAVQQNNLDEDLIRKLAYVAAGDLAPINAFIGGLAAQEVMKACSGKFMPIMQWLYFDALECLPEDKEVL TEDKCLQRQNRYDGQVAVFGSDLQEKLGKQKYFLVGAGAIGCELLKNFAMIGLGCGEGGEIIVTDMDTIEKSNLNRQFLFRPWDVTKLKS DTAAAAVRQMNPHIRVTSHQNRVGPDTERIYDDDFFQNLDGVANALDNVDARMYMDRRCVYYRKPLLESGTLGTKGNVQVVIPFLTESYS SSQDPPEKSIPICTLKNFPNAIEHTLQWARDEFEGLFKQPAENVNQYLTDPKFVERTLRLAGTQPLEVLEAVQRSLVLQRPQTWADCVTW ACHHWHTQYSNNIRQLLHNFPPDQLTSSGAPFWSGPKRCPHPLTFDVNNPLHLDYVMAAANLFAQTYGLTGSQDRAAVATFLQSVQVPEF TPKSGVKIHVSDQELQSANASVDDSRLEELKATLPSPDKLPGFKMYPIDFEKDDDSNFHVDFVVAAASLRCQNYGIPPVNRAQSKRIVGQ IIPAIATTTAAVAGLLGLELYKVVSGPRPRSAFRHSYLHLAENYLIRYMPFAPAIQTFHHLKWTSWDRLKVPAGQPERTLESLLAHLQEQ -------------------------------------------------------------- >95805_95805_2_UBA1-UBA7_UBA1_chrX_47071911_ENST00000377269_UBA7_chr3_49845545_ENST00000333486_length(transcript)=2386nt_BP=1676nt CTAATTGAAGTTTTTATTTTTTATGGAGGCATAACATATCTAGAATAAGGTACACAGATTTTTTTTTTTTTTTTTTTGAGGTGGAGTTTT GCTCATGTTGCCCAGGTTGGAGTGCAGTGGCGCAGTCTCGGCTCACTGCAGCCTCCGCCTCCCAGGTTCAAGCGATTCTCCTGCCTCAGC CTTCCAAAGTGCTGGGAATACAGGCGTGAGCCACCTCGCCTGGCACTCAGATCTTAATACATTTTTAGGCTGGGCATGGCGGCTGACACC TATAATCTCAGCACTCAGGAGGCCAAGGAGTGAGGTTTGTTTGAGCCCAGGAGTTCAAGACCAGCCTAGGCAACATAGCAAGACCCTGTC TCTACAACAAATCTTAAAAATTAGCCGGGTGTGGTGGCATGTGCCTGCTGTCCCAGCTATTCGGGAGGCTGAGGTGGGAGGATCGTTTCA ACCTGGGAGGTCAAGGCTGCTGTGAGCCGTGATTGCATCGCTGCACTCCAGCTCGGGTGACAGTGTGAGACTTTGTCTCAAACAACAAAA CAAAACAGTTTTCCACATACACAGCACTATTGTAACCACTGCCCAAATCAACTTCATTGATTTTTCTTTTTTTGAGACAGTCTTCCTCTG TCACCCAGGCTGGAGTGCAATGGCATGATCTCGGCTCACTGCAACCTCCGCCTCCCGGGTTCAAGAGATTTCCTGCCTCAGCCTCCTGAG TAGCTGGGATTACAGGCATCTGCCACCACGCCCAGCTACTTTTTGTATTTTTAGTAGAGATGGGGTTTCACCATGTTGGTCAGGCTGGTC TCGAACCCTTGACCTCAGGTGATCCATCTGCCTTAGCCTCCCAAAGTGCTGGGATTACAGGTGTGAGCTACTGTGCCTGGCCTGGTTTTT CAATTGAGGTGGGGGAATGGGGAAGGCTAGAGGGCTTCCCCACTTCCAGAGTAGCTGTGCGCCTTGTACTTGCTTTATTCACTGATGGTC CTTCTAATAATGCCTGCGGAAACCCATTTTGTCAACTGTGGCCACATGTAATCTGTTTGCTCTGTCTGCAGTGGGCTCGGGATGAGTTTG AAGGCCTCTTCAAGCAGCCAGCAGAAAATGTCAACCAGTACCTCACAGACCCCAAGTTTGTGGAGCGAACACTGCGGCTGGCAGGCACTC AGCCCTTGGAGGTGCTGGAGGCTGTGCAGCGCAGCCTGGTGCTGCAGCGACCACAGACCTGGGCTGACTGCGTGACCTGGGCCTGCCACC ACTGGCACACCCAGTACTCGAACAACATCCGGCAGCTGCTGCACAACTTCCCTCCTGACCAGCTCACAAGCTCAGGAGCGCCGTTCTGGT CTGGGCCCAAACGCTGTCCACACCCGCTCACCTTTGATGTCAACAATCCCCTGCATCTGGACTATGTGATGGCTGCTGCCAACCTGTTTG CCCAGACCTACGGGCTGACAGGCTCTCAGGACCGAGCTGCTGTGGCCACATTCCTGCAGTCTGTGCAGGTCCCCGAATTCACCCCCAAGT CTGGCGTCAAGATCCATGTTTCTGACCAGGAGCTGCAGAGCGCCAATGCCTCTGTTGATGACAGTCGTCTAGAGGAGCTCAAAGCCACTC TGCCCAGCCCAGACAAGCTCCCTGGATTCAAGATGTACCCCATTGACTTTGAGAAGGATGATGACAGCAACTTCCATGTGGACTTTGTGG TAGCGGCAGCTAGCCTGAGATGTCAGAACTACGGGATTCCACCGGTCAACCGTGCCCAGAGCAAGCGAATTGTGGGCCAGATTATCCCAG CCATTGCCACCACTACAGCAGCTGTGGCAGGCCTGTTGGGCCTGGAGCTGTATAAGGTGGTGAGTGGGCCACGGCCTCGTAGTGCCTTTC GCCACAGCTACCTACATCTGGCTGAAAACTACCTCATCCGCTATATGCCTTTTGCCCCAGCCATCCAGACGTTCCATCACCTGAAGTGGA CCTCTTGGGACCGTCTGAAGGTACCAGCTGGGCAGCCTGAGAGGACCCTGGAGTCGCTGCTGGCTCATCTTCAGGAGCAGCACGGGTTGA GGGTGAGGATCCTGCTGCACGGCTCAGCCCTGCTCTATGCGGCCGGATGGTCACCTGAAAAGCAGGCCCAGCACCTGCCCCTCAGGGTGA CAGAACTGGTTCAGCAGCTGACAGGCCAGGCACCTGCTCCTGGGCAGCGGGTGTTGGTGCTAGAGCTGAGCTGTGAGGGTGACGACGAGG ACACTGCCTTCCCACCTCTGCACTATGAGCTGTGACAAGGCAGCCACCCTGTCACCTAGCTCAATGGAGCCCCGGATCCCAAGCCCTGCA >95805_95805_2_UBA1-UBA7_UBA1_chrX_47071911_ENST00000377269_UBA7_chr3_49845545_ENST00000333486_length(amino acids)=501AA_BP=1 MGFHHVGQAGLEPLTSGDPSALASQSAGITGVSYCAWPGFSIEVGEWGRLEGFPTSRVAVRLVLALFTDGPSNNACGNPFCQLWPHVICL LCLQWARDEFEGLFKQPAENVNQYLTDPKFVERTLRLAGTQPLEVLEAVQRSLVLQRPQTWADCVTWACHHWHTQYSNNIRQLLHNFPPD QLTSSGAPFWSGPKRCPHPLTFDVNNPLHLDYVMAAANLFAQTYGLTGSQDRAAVATFLQSVQVPEFTPKSGVKIHVSDQELQSANASVD DSRLEELKATLPSPDKLPGFKMYPIDFEKDDDSNFHVDFVVAAASLRCQNYGIPPVNRAQSKRIVGQIIPAIATTTAAVAGLLGLELYKV VSGPRPRSAFRHSYLHLAENYLIRYMPFAPAIQTFHHLKWTSWDRLKVPAGQPERTLESLLAHLQEQHGLRVRILLHGSALLYAAGWSPE -------------------------------------------------------------- >95805_95805_3_UBA1-UBA7_UBA1_chrX_47071911_ENST00000377351_UBA7_chr3_49845545_ENST00000333486_length(transcript)=3353nt_BP=2643nt CTTGTACGACAGAGGTGGTTTGCTCTTCCGTTGCCCCGTGGCTTCAGCTCATCTTTGGCAGGAAGGCGAGGCTTCCGCCCGGCACAGGGG ATGTCCAGCTCGCCGCTGTCCAAGAAACGTCGCGTGTCCGGGCCTGATCCAAAGCCGGGTTCTAACTGCTCCCCTGCCCAGTCCGTGTTG TCCGAAGTGCCCTCGGTGCCAACCAACGGAATGGCCAAGAACGGCAGTGAAGCAGACATAGACGAGGGCCTTTACTCCCGGCAGCTGTAT GTGTTGGGCCATGAGGCAATGAAGCGGCTCCAGACATCCAGTGTCCTGGTATCAGGCCTGCGGGGCCTGGGCGTGGAGATCGCTAAGAAC ATCATCCTTGGTGGGGTCAAGGCTGTTACCCTACATGACCAGGGCACTGCCCAGTGGGCTGATCTTTCCTCCCAGTTCTACCTGCGGGAG GAGGACATCGGTAAAAACCGGGCCGAGGTATCACAGCCCCGCCTCGCTGAGCTCAACAGCTATGTGCCTGTCACTGCCTACACTGGACCC CTCGTTGAGGACTTCCTTAGTGGTTTCCAGGTGGTGGTGCTCACCAACACCCCCCTGGAGGACCAGCTGCGAGTGGGTGAGTTCTGTCAC AACCGTGGCATCAAGCTGGTGGTGGCAGACACGCGGGGCCTGTTTGGGCAGCTCTTCTGTGACTTTGGAGAGGAAATGATCCTCACAGAT TCCAATGGGGAGCAGCCACTCAGTGCTATGGTTTCTATGGTTACCAAGGACAACCCCGGTGTGGTTACCTGCCTGGATGAGGCCCGACAC GGGTTTGAGAGCGGGGACTTTGTCTCCTTTTCAGAAGTACAGGGCATGGTTGAACTCAACGGAAATCAGCCCATGGAGATCAAAGTCCTG GGTCCTTATACCTTTAGCATCTGTGACACCTCCAACTTCTCCGACTACATCCGTGGAGGCATCGTCAGTCAGGTCAAAGTACCTAAGAAG ATTAGCTTTAAATCCTTGGTGGCCTCACTGGCAGAACCTGACTTTGTGGTGACGGACTTCGCCAAGTTTTCTCGCCCTGCCCAGCTGCAC ATTGGCTTCCAGGCCCTGCACCAGTTCTGTGCTCAGCATGGCCGGCCACCTCGGCCCCGCAATGAGGAGGATGCAGCAGAACTGGTAGCC TTAGCACAGGCTGTGAATGCTCGAGCCCTGCCAGCAGTGCAGCAAAATAACCTGGACGAGGACCTCATCCGGAAGCTGGCATATGTGGCT GCTGGGGATCTGGCACCCATAAACGCCTTCATTGGGGGCCTGGCTGCCCAGGAAGTCATGAAGGCCTGCTCCGGGAAGTTCATGCCCATC ATGCAGTGGCTATACTTTGATGCCCTTGAGTGTCTCCCTGAGGACAAAGAGGTCCTCACAGAGGACAAGTGCCTCCAGCGCCAGAACCGT TATGACGGGCAAGTGGCTGTGTTTGGCTCAGACCTGCAAGAGAAGCTGGGCAAGCAGAAGTATTTCCTGGTGGGTGCGGGGGCCATTGGC TGTGAGCTGCTCAAGAACTTTGCCATGATTGGGCTGGGCTGCGGGGAGGGTGGAGAAATCATCGTTACAGACATGGACACCATTGAGAAG TCAAATCTGAATCGACAGTTTCTTTTCCGGCCCTGGGATGTCACGAAGTTAAAGTCTGACACGGCTGCTGCAGCTGTGCGCCAAATGAAT CCACATATCCGGGTGACAAGCCACCAGAACCGTGTGGGTCCTGACACGGAGCGCATCTATGATGACGATTTTTTCCAAAACCTAGATGGC GTGGCCAATGCCCTGGACAACGTGGATGCCCGCATGTACATGGACCGCCGCTGTGTCTACTACCGGAAGCCACTGCTGGAGTCAGGCACA CTGGGCACCAAAGGCAATGTGCAGGTGGTGATCCCCTTCCTGACAGAGTCGTACAGTTCCAGCCAGGACCCACCTGAGAAGTCCATCCCC ATCTGTACCCTGAAGAACTTCCCTAATGCCATCGAGCACACCCTGCAGTGGGCTCGGGATGAGTTTGAAGGCCTCTTCAAGCAGCCAGCA GAAAATGTCAACCAGTACCTCACAGACCCCAAGTTTGTGGAGCGAACACTGCGGCTGGCAGGCACTCAGCCCTTGGAGGTGCTGGAGGCT GTGCAGCGCAGCCTGGTGCTGCAGCGACCACAGACCTGGGCTGACTGCGTGACCTGGGCCTGCCACCACTGGCACACCCAGTACTCGAAC AACATCCGGCAGCTGCTGCACAACTTCCCTCCTGACCAGCTCACAAGCTCAGGAGCGCCGTTCTGGTCTGGGCCCAAACGCTGTCCACAC CCGCTCACCTTTGATGTCAACAATCCCCTGCATCTGGACTATGTGATGGCTGCTGCCAACCTGTTTGCCCAGACCTACGGGCTGACAGGC TCTCAGGACCGAGCTGCTGTGGCCACATTCCTGCAGTCTGTGCAGGTCCCCGAATTCACCCCCAAGTCTGGCGTCAAGATCCATGTTTCT GACCAGGAGCTGCAGAGCGCCAATGCCTCTGTTGATGACAGTCGTCTAGAGGAGCTCAAAGCCACTCTGCCCAGCCCAGACAAGCTCCCT GGATTCAAGATGTACCCCATTGACTTTGAGAAGGATGATGACAGCAACTTCCATGTGGACTTTGTGGTAGCGGCAGCTAGCCTGAGATGT CAGAACTACGGGATTCCACCGGTCAACCGTGCCCAGAGCAAGCGAATTGTGGGCCAGATTATCCCAGCCATTGCCACCACTACAGCAGCT GTGGCAGGCCTGTTGGGCCTGGAGCTGTATAAGGTGGTGAGTGGGCCACGGCCTCGTAGTGCCTTTCGCCACAGCTACCTACATCTGGCT GAAAACTACCTCATCCGCTATATGCCTTTTGCCCCAGCCATCCAGACGTTCCATCACCTGAAGTGGACCTCTTGGGACCGTCTGAAGGTA CCAGCTGGGCAGCCTGAGAGGACCCTGGAGTCGCTGCTGGCTCATCTTCAGGAGCAGCACGGGTTGAGGGTGAGGATCCTGCTGCACGGC TCAGCCCTGCTCTATGCGGCCGGATGGTCACCTGAAAAGCAGGCCCAGCACCTGCCCCTCAGGGTGACAGAACTGGTTCAGCAGCTGACA GGCCAGGCACCTGCTCCTGGGCAGCGGGTGTTGGTGCTAGAGCTGAGCTGTGAGGGTGACGACGAGGACACTGCCTTCCCACCTCTGCAC TATGAGCTGTGACAAGGCAGCCACCCTGTCACCTAGCTCAATGGAGCCCCGGATCCCAAGCCCTGCATTGTAAGCCCACAGTAGGCACTC >95805_95805_3_UBA1-UBA7_UBA1_chrX_47071911_ENST00000377351_UBA7_chr3_49845545_ENST00000333486_length(amino acids)=1073AA_BP=79 MPRGFSSSLAGRRGFRPAQGMSSSPLSKKRRVSGPDPKPGSNCSPAQSVLSEVPSVPTNGMAKNGSEADIDEGLYSRQLYVLGHEAMKRL QTSSVLVSGLRGLGVEIAKNIILGGVKAVTLHDQGTAQWADLSSQFYLREEDIGKNRAEVSQPRLAELNSYVPVTAYTGPLVEDFLSGFQ VVVLTNTPLEDQLRVGEFCHNRGIKLVVADTRGLFGQLFCDFGEEMILTDSNGEQPLSAMVSMVTKDNPGVVTCLDEARHGFESGDFVSF SEVQGMVELNGNQPMEIKVLGPYTFSICDTSNFSDYIRGGIVSQVKVPKKISFKSLVASLAEPDFVVTDFAKFSRPAQLHIGFQALHQFC AQHGRPPRPRNEEDAAELVALAQAVNARALPAVQQNNLDEDLIRKLAYVAAGDLAPINAFIGGLAAQEVMKACSGKFMPIMQWLYFDALE CLPEDKEVLTEDKCLQRQNRYDGQVAVFGSDLQEKLGKQKYFLVGAGAIGCELLKNFAMIGLGCGEGGEIIVTDMDTIEKSNLNRQFLFR PWDVTKLKSDTAAAAVRQMNPHIRVTSHQNRVGPDTERIYDDDFFQNLDGVANALDNVDARMYMDRRCVYYRKPLLESGTLGTKGNVQVV IPFLTESYSSSQDPPEKSIPICTLKNFPNAIEHTLQWARDEFEGLFKQPAENVNQYLTDPKFVERTLRLAGTQPLEVLEAVQRSLVLQRP QTWADCVTWACHHWHTQYSNNIRQLLHNFPPDQLTSSGAPFWSGPKRCPHPLTFDVNNPLHLDYVMAAANLFAQTYGLTGSQDRAAVATF LQSVQVPEFTPKSGVKIHVSDQELQSANASVDDSRLEELKATLPSPDKLPGFKMYPIDFEKDDDSNFHVDFVVAAASLRCQNYGIPPVNR AQSKRIVGQIIPAIATTTAAVAGLLGLELYKVVSGPRPRSAFRHSYLHLAENYLIRYMPFAPAIQTFHHLKWTSWDRLKVPAGQPERTLE -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for UBA1-UBA7 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for UBA1-UBA7 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for UBA1-UBA7 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
| Hgene | UBA1 | C1844934 | Arthrogryposis multiplex congenita, distal, X-linked | 2 | CTD_human;GENOMICS_ENGLAND;ORPHANET;UNIPROT |
| Hgene | UBA1 | C0007137 | Squamous cell carcinoma | 1 | CTD_human |
| Tgene | C0007137 | Squamous cell carcinoma | 1 | CTD_human |
