
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:MCFD2-SRBD1 (FusionGDB2 ID:HG90411TG55133) |
Fusion Gene Summary for MCFD2-SRBD1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: MCFD2-SRBD1 | Fusion gene ID: hg90411tg55133 | Hgene | Tgene | Gene symbol | MCFD2 | SRBD1 | Gene ID | 90411 | 55133 |
| Gene name | multiple coagulation factor deficiency 2, ER cargo receptor complex subunit | S1 RNA binding domain 1 | |
| Synonyms | F5F8D|F5F8D2|LMAN1IP|SDNSF | - | |
| Cytomap | ('MCFD2')('SRBD1') 2p21 | 2p21 | |
| Type of gene | protein-coding | protein-coding | |
| Description | multiple coagulation factor deficiency protein 2multiple coagulation factor deficiency 2neural stem cell-derived neuronal survival protein | S1 RNA-binding domain-containing protein 1H_NH0576F01.1WUGSC:H_NH0576F01.1 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000319466, ENST00000409105, ENST00000409147, ENST00000409207, ENST00000409218, ENST00000409800, ENST00000409913, ENST00000409973, ENST00000444761, ENST00000493804, | ||
| Fusion gene scores | * DoF score | 5 X 7 X 4=140 | 3 X 3 X 3=27 |
| # samples | 6 | 4 | |
| ** MAII score | log2(6/140*10)=-1.22239242133645 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(4/27*10)=0.567040592723894 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | |
| Context | PubMed: MCFD2 [Title/Abstract] AND SRBD1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | MCFD2(47134949)-SRBD1(45647033), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | MCFD2-SRBD1 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. MCFD2-SRBD1 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. MCFD2-SRBD1 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. MCFD2-SRBD1 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. MCFD2-SRBD1 seems lost the major protein functional domain in Hgene partner, which is a cell metabolism gene due to the frame-shifted ORF. MCFD2-SRBD1 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. MCFD2-SRBD1 seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across MCFD2 (5'-gene) Fusion gene breakpoints across MCFD2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
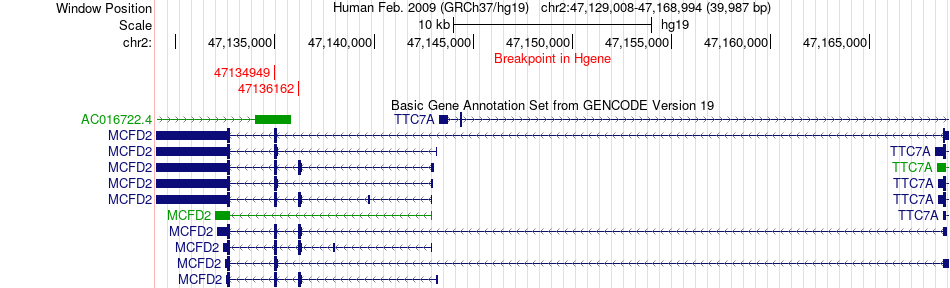 |
 Fusion gene breakpoints across SRBD1 (3'-gene) Fusion gene breakpoints across SRBD1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
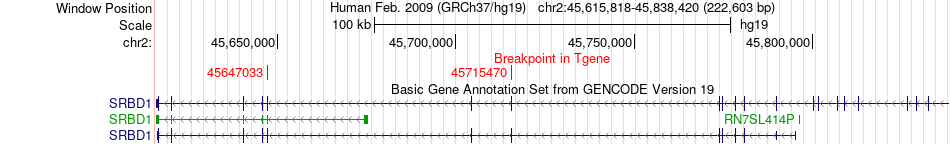 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-AR-A24H-01A | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| ChimerDB4 | BRCA | TCGA-AR-A24H-01A | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
Top |
Fusion Gene ORF analysis for MCFD2-SRBD1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000319466 | ENST00000490133 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| 5CDS-5UTR | ENST00000409105 | ENST00000490133 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| 5CDS-5UTR | ENST00000409147 | ENST00000490133 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| 5CDS-5UTR | ENST00000409207 | ENST00000490133 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| 5CDS-5UTR | ENST00000409218 | ENST00000490133 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| 5CDS-5UTR | ENST00000409800 | ENST00000490133 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| 5CDS-5UTR | ENST00000409913 | ENST00000490133 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| 5CDS-5UTR | ENST00000409973 | ENST00000490133 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| 5CDS-5UTR | ENST00000444761 | ENST00000490133 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| 5CDS-intron | ENST00000319466 | ENST00000490133 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| 5CDS-intron | ENST00000409105 | ENST00000490133 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| 5CDS-intron | ENST00000409207 | ENST00000490133 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| 5CDS-intron | ENST00000409218 | ENST00000490133 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| 5CDS-intron | ENST00000409973 | ENST00000490133 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| Frame-shift | ENST00000319466 | ENST00000263736 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| Frame-shift | ENST00000319466 | ENST00000263736 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| Frame-shift | ENST00000319466 | ENST00000535761 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| Frame-shift | ENST00000319466 | ENST00000535761 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| Frame-shift | ENST00000409147 | ENST00000263736 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| Frame-shift | ENST00000409147 | ENST00000535761 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| Frame-shift | ENST00000409218 | ENST00000263736 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| Frame-shift | ENST00000409218 | ENST00000263736 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| Frame-shift | ENST00000409218 | ENST00000535761 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| Frame-shift | ENST00000409218 | ENST00000535761 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| Frame-shift | ENST00000409800 | ENST00000263736 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| Frame-shift | ENST00000409800 | ENST00000535761 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| Frame-shift | ENST00000409913 | ENST00000263736 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| Frame-shift | ENST00000409913 | ENST00000535761 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| Frame-shift | ENST00000409973 | ENST00000263736 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| Frame-shift | ENST00000409973 | ENST00000263736 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| Frame-shift | ENST00000409973 | ENST00000535761 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| Frame-shift | ENST00000409973 | ENST00000535761 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| In-frame | ENST00000409105 | ENST00000263736 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| In-frame | ENST00000409105 | ENST00000263736 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| In-frame | ENST00000409105 | ENST00000535761 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| In-frame | ENST00000409105 | ENST00000535761 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| In-frame | ENST00000409207 | ENST00000263736 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| In-frame | ENST00000409207 | ENST00000263736 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| In-frame | ENST00000409207 | ENST00000535761 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| In-frame | ENST00000409207 | ENST00000535761 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| In-frame | ENST00000444761 | ENST00000263736 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| In-frame | ENST00000444761 | ENST00000535761 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| intron-3CDS | ENST00000409147 | ENST00000263736 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| intron-3CDS | ENST00000409147 | ENST00000535761 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| intron-3CDS | ENST00000409800 | ENST00000263736 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| intron-3CDS | ENST00000409800 | ENST00000535761 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| intron-3CDS | ENST00000409913 | ENST00000263736 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| intron-3CDS | ENST00000409913 | ENST00000535761 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| intron-3CDS | ENST00000444761 | ENST00000263736 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| intron-3CDS | ENST00000444761 | ENST00000535761 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| intron-3CDS | ENST00000493804 | ENST00000263736 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| intron-3CDS | ENST00000493804 | ENST00000263736 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| intron-3CDS | ENST00000493804 | ENST00000535761 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| intron-3CDS | ENST00000493804 | ENST00000535761 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| intron-5UTR | ENST00000493804 | ENST00000490133 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - |
| intron-intron | ENST00000409147 | ENST00000490133 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| intron-intron | ENST00000409800 | ENST00000490133 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| intron-intron | ENST00000409913 | ENST00000490133 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| intron-intron | ENST00000444761 | ENST00000490133 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
| intron-intron | ENST00000493804 | ENST00000490133 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000444761 | MCFD2 | chr2 | 47134949 | - | ENST00000263736 | SRBD1 | chr2 | 45647033 | - | 2013 | 444 | 51 | 1382 | 443 |
| ENST00000444761 | MCFD2 | chr2 | 47134949 | - | ENST00000535761 | SRBD1 | chr2 | 45647033 | - | 1600 | 444 | 51 | 1382 | 443 |
| ENST00000409105 | MCFD2 | chr2 | 47134949 | - | ENST00000263736 | SRBD1 | chr2 | 45647033 | - | 2058 | 489 | 99 | 1427 | 442 |
| ENST00000409105 | MCFD2 | chr2 | 47134949 | - | ENST00000535761 | SRBD1 | chr2 | 45647033 | - | 1645 | 489 | 99 | 1427 | 442 |
| ENST00000409207 | MCFD2 | chr2 | 47134949 | - | ENST00000263736 | SRBD1 | chr2 | 45647033 | - | 2045 | 476 | 80 | 1414 | 444 |
| ENST00000409207 | MCFD2 | chr2 | 47134949 | - | ENST00000535761 | SRBD1 | chr2 | 45647033 | - | 1632 | 476 | 80 | 1414 | 444 |
| ENST00000409105 | MCFD2 | chr2 | 47136162 | - | ENST00000263736 | SRBD1 | chr2 | 45715470 | - | 2073 | 329 | 99 | 1442 | 447 |
| ENST00000409105 | MCFD2 | chr2 | 47136162 | - | ENST00000535761 | SRBD1 | chr2 | 45715470 | - | 1660 | 329 | 99 | 1442 | 447 |
| ENST00000409207 | MCFD2 | chr2 | 47136162 | - | ENST00000263736 | SRBD1 | chr2 | 45715470 | - | 2060 | 316 | 80 | 1429 | 449 |
| ENST00000409207 | MCFD2 | chr2 | 47136162 | - | ENST00000535761 | SRBD1 | chr2 | 45715470 | - | 1647 | 316 | 80 | 1429 | 449 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000444761 | ENST00000263736 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - | 0.002529134 | 0.99747086 |
| ENST00000444761 | ENST00000535761 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - | 0.005305724 | 0.9946943 |
| ENST00000409105 | ENST00000263736 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - | 0.001296821 | 0.9987031 |
| ENST00000409105 | ENST00000535761 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - | 0.003077607 | 0.9969223 |
| ENST00000409207 | ENST00000263736 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - | 0.001582248 | 0.99841774 |
| ENST00000409207 | ENST00000535761 | MCFD2 | chr2 | 47134949 | - | SRBD1 | chr2 | 45647033 | - | 0.003332002 | 0.99666804 |
| ENST00000409105 | ENST00000263736 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - | 0.001655383 | 0.99834466 |
| ENST00000409105 | ENST00000535761 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - | 0.00488959 | 0.99511045 |
| ENST00000409207 | ENST00000263736 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - | 0.002118558 | 0.9978815 |
| ENST00000409207 | ENST00000535761 | MCFD2 | chr2 | 47136162 | - | SRBD1 | chr2 | 45715470 | - | 0.005056078 | 0.9949439 |
Top |
Fusion Genomic Features for MCFD2-SRBD1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
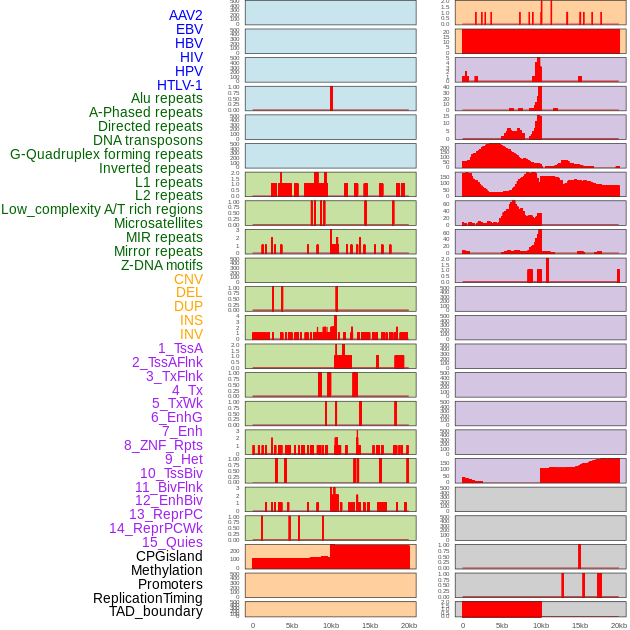 |
Top |
Fusion Protein Features for MCFD2-SRBD1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr2:47134949/chr2:45647033) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000319466 | - | 3 | 4 | 81_92 | 103 | 147.0 | Calcium binding | Note=1 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409105 | - | 4 | 5 | 81_92 | 103 | 147.0 | Calcium binding | Note=1 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409207 | - | 3 | 4 | 81_92 | 103 | 147.0 | Calcium binding | Note=1 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409218 | - | 3 | 4 | 81_92 | 103 | 147.0 | Calcium binding | Note=1 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409973 | - | 4 | 5 | 81_92 | 103 | 147.0 | Calcium binding | Note=1 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000319466 | - | 3 | 4 | 68_103 | 103 | 147.0 | Domain | EF-hand 1 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409105 | - | 4 | 5 | 68_103 | 103 | 147.0 | Domain | EF-hand 1 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409207 | - | 3 | 4 | 68_103 | 103 | 147.0 | Domain | EF-hand 1 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409218 | - | 3 | 4 | 68_103 | 103 | 147.0 | Domain | EF-hand 1 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409973 | - | 4 | 5 | 68_103 | 103 | 147.0 | Domain | EF-hand 1 |
| Tgene | SRBD1 | chr2:47134949 | chr2:45647033 | ENST00000263736 | 15 | 21 | 919_992 | 683 | 996.0 | Domain | S1 motif | |
| Tgene | SRBD1 | chr2:47136162 | chr2:45715470 | ENST00000263736 | 13 | 21 | 919_992 | 624 | 996.0 | Domain | S1 motif |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000319466 | - | 3 | 4 | 129_140 | 103 | 147.0 | Calcium binding | Note=2 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409105 | - | 4 | 5 | 129_140 | 103 | 147.0 | Calcium binding | Note=2 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409147 | - | 2 | 3 | 129_140 | 51 | 95.0 | Calcium binding | Note=2 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409147 | - | 2 | 3 | 81_92 | 51 | 95.0 | Calcium binding | Note=1 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409207 | - | 3 | 4 | 129_140 | 103 | 147.0 | Calcium binding | Note=2 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409218 | - | 3 | 4 | 129_140 | 103 | 147.0 | Calcium binding | Note=2 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409800 | - | 2 | 3 | 129_140 | 51 | 95.0 | Calcium binding | Note=2 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409800 | - | 2 | 3 | 81_92 | 51 | 95.0 | Calcium binding | Note=1 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409913 | - | 2 | 3 | 129_140 | 51 | 95.0 | Calcium binding | Note=2 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409913 | - | 2 | 3 | 81_92 | 51 | 95.0 | Calcium binding | Note=1 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409973 | - | 4 | 5 | 129_140 | 103 | 147.0 | Calcium binding | Note=2 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000444761 | - | 2 | 3 | 129_140 | 84 | 128.0 | Calcium binding | Note=2 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000444761 | - | 2 | 3 | 81_92 | 84 | 128.0 | Calcium binding | Note=1 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000319466 | - | 2 | 4 | 129_140 | 49 | 147.0 | Calcium binding | Note=2 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000319466 | - | 2 | 4 | 81_92 | 49 | 147.0 | Calcium binding | Note=1 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409105 | - | 3 | 5 | 129_140 | 49 | 147.0 | Calcium binding | Note=2 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409105 | - | 3 | 5 | 81_92 | 49 | 147.0 | Calcium binding | Note=1 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409147 | - | 1 | 3 | 129_140 | 0 | 95.0 | Calcium binding | Note=2 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409147 | - | 1 | 3 | 81_92 | 0 | 95.0 | Calcium binding | Note=1 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409207 | - | 2 | 4 | 129_140 | 49 | 147.0 | Calcium binding | Note=2 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409207 | - | 2 | 4 | 81_92 | 49 | 147.0 | Calcium binding | Note=1 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409218 | - | 2 | 4 | 129_140 | 49 | 147.0 | Calcium binding | Note=2 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409218 | - | 2 | 4 | 81_92 | 49 | 147.0 | Calcium binding | Note=1 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409800 | - | 1 | 3 | 129_140 | 0 | 95.0 | Calcium binding | Note=2 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409800 | - | 1 | 3 | 81_92 | 0 | 95.0 | Calcium binding | Note=1 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409913 | - | 1 | 3 | 129_140 | 0 | 95.0 | Calcium binding | Note=2 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409913 | - | 1 | 3 | 81_92 | 0 | 95.0 | Calcium binding | Note=1 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409973 | - | 3 | 5 | 129_140 | 49 | 147.0 | Calcium binding | Note=2 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409973 | - | 3 | 5 | 81_92 | 49 | 147.0 | Calcium binding | Note=1 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000444761 | - | 1 | 3 | 129_140 | 0 | 128.0 | Calcium binding | Note=2 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000444761 | - | 1 | 3 | 81_92 | 0 | 128.0 | Calcium binding | Note=1 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000319466 | - | 3 | 4 | 116_146 | 103 | 147.0 | Domain | EF-hand 2 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409105 | - | 4 | 5 | 116_146 | 103 | 147.0 | Domain | EF-hand 2 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409147 | - | 2 | 3 | 116_146 | 51 | 95.0 | Domain | EF-hand 2 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409147 | - | 2 | 3 | 68_103 | 51 | 95.0 | Domain | EF-hand 1 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409207 | - | 3 | 4 | 116_146 | 103 | 147.0 | Domain | EF-hand 2 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409218 | - | 3 | 4 | 116_146 | 103 | 147.0 | Domain | EF-hand 2 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409800 | - | 2 | 3 | 116_146 | 51 | 95.0 | Domain | EF-hand 2 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409800 | - | 2 | 3 | 68_103 | 51 | 95.0 | Domain | EF-hand 1 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409913 | - | 2 | 3 | 116_146 | 51 | 95.0 | Domain | EF-hand 2 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409913 | - | 2 | 3 | 68_103 | 51 | 95.0 | Domain | EF-hand 1 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000409973 | - | 4 | 5 | 116_146 | 103 | 147.0 | Domain | EF-hand 2 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000444761 | - | 2 | 3 | 116_146 | 84 | 128.0 | Domain | EF-hand 2 |
| Hgene | MCFD2 | chr2:47134949 | chr2:45647033 | ENST00000444761 | - | 2 | 3 | 68_103 | 84 | 128.0 | Domain | EF-hand 1 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000319466 | - | 2 | 4 | 116_146 | 49 | 147.0 | Domain | EF-hand 2 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000319466 | - | 2 | 4 | 68_103 | 49 | 147.0 | Domain | EF-hand 1 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409105 | - | 3 | 5 | 116_146 | 49 | 147.0 | Domain | EF-hand 2 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409105 | - | 3 | 5 | 68_103 | 49 | 147.0 | Domain | EF-hand 1 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409147 | - | 1 | 3 | 116_146 | 0 | 95.0 | Domain | EF-hand 2 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409147 | - | 1 | 3 | 68_103 | 0 | 95.0 | Domain | EF-hand 1 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409207 | - | 2 | 4 | 116_146 | 49 | 147.0 | Domain | EF-hand 2 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409207 | - | 2 | 4 | 68_103 | 49 | 147.0 | Domain | EF-hand 1 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409218 | - | 2 | 4 | 116_146 | 49 | 147.0 | Domain | EF-hand 2 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409218 | - | 2 | 4 | 68_103 | 49 | 147.0 | Domain | EF-hand 1 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409800 | - | 1 | 3 | 116_146 | 0 | 95.0 | Domain | EF-hand 2 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409800 | - | 1 | 3 | 68_103 | 0 | 95.0 | Domain | EF-hand 1 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409913 | - | 1 | 3 | 116_146 | 0 | 95.0 | Domain | EF-hand 2 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409913 | - | 1 | 3 | 68_103 | 0 | 95.0 | Domain | EF-hand 1 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409973 | - | 3 | 5 | 116_146 | 49 | 147.0 | Domain | EF-hand 2 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000409973 | - | 3 | 5 | 68_103 | 49 | 147.0 | Domain | EF-hand 1 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000444761 | - | 1 | 3 | 116_146 | 0 | 128.0 | Domain | EF-hand 2 |
| Hgene | MCFD2 | chr2:47136162 | chr2:45715470 | ENST00000444761 | - | 1 | 3 | 68_103 | 0 | 128.0 | Domain | EF-hand 1 |
| Tgene | SRBD1 | chr2:47134949 | chr2:45647033 | ENST00000263736 | 15 | 21 | 258_288 | 683 | 996.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | SRBD1 | chr2:47136162 | chr2:45715470 | ENST00000263736 | 13 | 21 | 258_288 | 624 | 996.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | SRBD1 | chr2:47134949 | chr2:45647033 | ENST00000263736 | 15 | 21 | 80_83 | 683 | 996.0 | Compositional bias | Note=Poly-Val | |
| Tgene | SRBD1 | chr2:47136162 | chr2:45715470 | ENST00000263736 | 13 | 21 | 80_83 | 624 | 996.0 | Compositional bias | Note=Poly-Val |
Top |
Fusion Gene Sequence for MCFD2-SRBD1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >52193_52193_1_MCFD2-SRBD1_MCFD2_chr2_47134949_ENST00000409105_SRBD1_chr2_45647033_ENST00000263736_length(transcript)=2058nt_BP=489nt ACGGGGGCTGGTGAGGCTCACGTTGGAGGGCTTCGCGTCTGCTTCGGAGACCGTAAGGCTGTCATCAGCTGTGGACGGGAGCCTTCTGAA GGGAAGCACTTGGGAGCCTGCAGGACGAATACCTACACCAGACTTGGAATTGAAAAGACCTCACTGGAGAAAGAGACATTTGATATATTG ATGACCATGAGATCCCTGCTCAGAACCCCCTTCCTGTGTGGCCTGCTCTGGGCCTTTTGTGCCCCAGGCGCCAGGGCTGAGGAGCCTGCA GCCAGCTTCTCCCAACCCGGCAGCATGGGCCTGGATAAGAACACAGTGCACGACCAAGAGCATATCATGGAGCATCTAGAAGGTGTCATC AACAAACCAGAGGCGGAGATGTCGCCACAAGAATTGCAGCTCCATTACTTCAAAATGCATGATTATGATGGCAATAATTTGCTTGATGGC TTAGAACTCTCCACAGCCATCACTCATGTCCATAAGGAGCATGACGTATCCCAGACTTTACTCAAGGCAACACTGGACAGTGTTGTAGAA GAATGTGTCAGCTTTGTGGGAGTGGATATTAACATCTGTTCAGAAGTTTTGTTAAGGCATATTGCAGGACTCAATGCCAACAGGGCCAAA AATATTATTGAATGGCGAGAGAAAAATGGACCCTTTATCAACCGAGAACAGCTGAAGAAAGTGAAAGGGCTGGGCCCAAAATCCTTCCAA CAGTGTGCTGGCTTCATCAGAATCAACCAGGATTATATCCGAACGTTTTGCAGTCAGCAAACTGAAACTTCAGGCCAAATTCAAGGAGTT GCTGTGACATCTTCAGCAGACGTTGAGGTCACAAATGAGAAGCAGGGCAAAAAGAAGAGCAAAACTGCAGTGAATGTTTTACTGAAGCCA AATCCTTTGGACCAAACTTGTATTCATCCAGAATCATATGACATAGCAATGAGGTTTTTGTCATCCATTGGAGGGACACTGTATGAGGTT GGAAAGCCTGAAATGCAACAAAAAATAAATTCATTCCTTGAAAAGGAAGGAATGGAGAAAATTGCAGAAAGATTGCAAACAACAGTACAC ACCTTACAGGTCATCATAGATGGTCTCAGCCAGCCTGAAAGCTTTGACTTTCGAACAGATTTTGATAAACCTGATTTCAAGAGAAGCATA GTATGCCTGGAAGATCTGCAGATTGGGACAGTTCTTACAGGCAAAGTTGAGAATGCCACTCTCTTTGGAATTTTTGTGGATATAGGAGTG GGGAAATCTGGGCTGATTCCCATACGAAATGTAACAGAAGCAAAACTTTCAAAAACAAAGAAGAGAAGAAGCCTTGGACTGGGCCCCGGA GAAAGAGTGGAAGTCCAAGTACTCAACATTGACATCCCCCGATCTAGGATTACTCTGGACCTCATTCGGGTGTTATGAGTATCCCACGAA GGCCAGACGCTGATTTTATTTTCTCATTTCCACAGATTGACAAGGATAAGTCAGTTGTTTGTAAACTCTAGGTAGCAGATGAGAAATAAT TCACTTAATATCAGAAATATTTTCCAAACACTTTCCTTTATTTTTTCTTCTGAATAAATAGAAAACCAACAGTTTGATTTCCTTTTCCCT TAAAGGAAACAACTAATACACATTCTTATATGGCTTTATGTAGTAATAGTTTTCTGACTAAAATTTTGTTTTTTATTTTTTGTAATTTAT CTTTAACTCCTTTTGCATTTTGTATAACAGATTGCTTAACTTCTACTTGCCAACATCTGCCTTGCTGGACTTGTATGGGATTGTCTTCTT GATTTGAATTGTACCGTCTTTGTTGACACAGTAGGGCTGGGCAGTGTTTAATCCTTCCATTTTATAGATTTTTTTTTAATCAGGCCTTTT GGACTTCATTCATAATTTTGCAATAATCTCTTTTCCCTTGTCATGCAAGCCAAAAATATACCAGTAAAACAGATTCTGACGTGTTTGTAG >52193_52193_1_MCFD2-SRBD1_MCFD2_chr2_47134949_ENST00000409105_SRBD1_chr2_45647033_ENST00000263736_length(amino acids)=442AA_BP=130 MGACRTNTYTRLGIEKTSLEKETFDILMTMRSLLRTPFLCGLLWAFCAPGARAEEPAASFSQPGSMGLDKNTVHDQEHIMEHLEGVINKP EAEMSPQELQLHYFKMHDYDGNNLLDGLELSTAITHVHKEHDVSQTLLKATLDSVVEECVSFVGVDINICSEVLLRHIAGLNANRAKNII EWREKNGPFINREQLKKVKGLGPKSFQQCAGFIRINQDYIRTFCSQQTETSGQIQGVAVTSSADVEVTNEKQGKKKSKTAVNVLLKPNPL DQTCIHPESYDIAMRFLSSIGGTLYEVGKPEMQQKINSFLEKEGMEKIAERLQTTVHTLQVIIDGLSQPESFDFRTDFDKPDFKRSIVCL -------------------------------------------------------------- >52193_52193_2_MCFD2-SRBD1_MCFD2_chr2_47134949_ENST00000409105_SRBD1_chr2_45647033_ENST00000535761_length(transcript)=1645nt_BP=489nt ACGGGGGCTGGTGAGGCTCACGTTGGAGGGCTTCGCGTCTGCTTCGGAGACCGTAAGGCTGTCATCAGCTGTGGACGGGAGCCTTCTGAA GGGAAGCACTTGGGAGCCTGCAGGACGAATACCTACACCAGACTTGGAATTGAAAAGACCTCACTGGAGAAAGAGACATTTGATATATTG ATGACCATGAGATCCCTGCTCAGAACCCCCTTCCTGTGTGGCCTGCTCTGGGCCTTTTGTGCCCCAGGCGCCAGGGCTGAGGAGCCTGCA GCCAGCTTCTCCCAACCCGGCAGCATGGGCCTGGATAAGAACACAGTGCACGACCAAGAGCATATCATGGAGCATCTAGAAGGTGTCATC AACAAACCAGAGGCGGAGATGTCGCCACAAGAATTGCAGCTCCATTACTTCAAAATGCATGATTATGATGGCAATAATTTGCTTGATGGC TTAGAACTCTCCACAGCCATCACTCATGTCCATAAGGAGCATGACGTATCCCAGACTTTACTCAAGGCAACACTGGACAGTGTTGTAGAA GAATGTGTCAGCTTTGTGGGAGTGGATATTAACATCTGTTCAGAAGTTTTGTTAAGGCATATTGCAGGACTCAATGCCAACAGGGCCAAA AATATTATTGAATGGCGAGAGAAAAATGGACCCTTTATCAACCGAGAACAGCTGAAGAAAGTGAAAGGGCTGGGCCCAAAATCCTTCCAA CAGTGTGCTGGCTTCATCAGAATCAACCAGGATTATATCCGAACGTTTTGCAGTCAGCAAACTGAAACTTCAGGCCAAATTCAAGGAGTT GCTGTGACATCTTCAGCAGACGTTGAGGTCACAAATGAGAAGCAGGGCAAAAAGAAGAGCAAAACTGCAGTGAATGTTTTACTGAAGCCA AATCCTTTGGACCAAACTTGTATTCATCCAGAATCATATGACATAGCAATGAGGTTTTTGTCATCCATTGGAGGGACACTGTATGAGGTT GGAAAGCCTGAAATGCAACAAAAAATAAATTCATTCCTTGAAAAGGAAGGAATGGAGAAAATTGCAGAAAGATTGCAAACAACAGTACAC ACCTTACAGGTCATCATAGATGGTCTCAGCCAGCCTGAAAGCTTTGACTTTCGAACAGATTTTGATAAACCTGATTTCAAGAGAAGCATA GTATGCCTGGAAGATCTGCAGATTGGGACAGTTCTTACAGGCAAAGTTGAGAATGCCACTCTCTTTGGAATTTTTGTGGATATAGGAGTG GGGAAATCTGGGCTGATTCCCATACGAAATGTAACAGAAGCAAAACTTTCAAAAACAAAGAAGAGAAGAAGCCTTGGACTGGGCCCCGGA GAAAGAGTGGAAGTCCAAGTACTCAACATTGACATCCCCCGATCTAGGATTACTCTGGACCTCATTCGGGTGTTATGAGTATCCCACGAA GGCCAGACGCTGATTTTATTTTCTCATTTCCACAGATTGACAAGGATAAGTCAGTTGTTTGTAAACTCTAGGTAGCAGATGAGAAATAAT TCACTTAATATCAGAAATATTTTCCAAACACTTTCCTTTATTTTTTCTTCTGAATAAATAGAAAACCAACAGTTTGATTTCCTTTTCCCT >52193_52193_2_MCFD2-SRBD1_MCFD2_chr2_47134949_ENST00000409105_SRBD1_chr2_45647033_ENST00000535761_length(amino acids)=442AA_BP=130 MGACRTNTYTRLGIEKTSLEKETFDILMTMRSLLRTPFLCGLLWAFCAPGARAEEPAASFSQPGSMGLDKNTVHDQEHIMEHLEGVINKP EAEMSPQELQLHYFKMHDYDGNNLLDGLELSTAITHVHKEHDVSQTLLKATLDSVVEECVSFVGVDINICSEVLLRHIAGLNANRAKNII EWREKNGPFINREQLKKVKGLGPKSFQQCAGFIRINQDYIRTFCSQQTETSGQIQGVAVTSSADVEVTNEKQGKKKSKTAVNVLLKPNPL DQTCIHPESYDIAMRFLSSIGGTLYEVGKPEMQQKINSFLEKEGMEKIAERLQTTVHTLQVIIDGLSQPESFDFRTDFDKPDFKRSIVCL -------------------------------------------------------------- >52193_52193_3_MCFD2-SRBD1_MCFD2_chr2_47134949_ENST00000409207_SRBD1_chr2_45647033_ENST00000263736_length(transcript)=2045nt_BP=476nt TACTCACCGGTGTCCGGAAAGGTGAACGCTGCGCTCGGGCTGCCTCGCCTGTTACCTCCGCCGCCGGGCATGCTCAGCGTCTGCAGCTGC CGGACCAGCTCCGGCATGCGGTCCCAGTGGCCCTCGGCGCGGCAGCGCTCCAGCTCGCTCTCCACCTTCAGATATTGATGACCATGAGAT CCCTGCTCAGAACCCCCTTCCTGTGTGGCCTGCTCTGGGCCTTTTGTGCCCCAGGCGCCAGGGCTGAGGAGCCTGCAGCCAGCTTCTCCC AACCCGGCAGCATGGGCCTGGATAAGAACACAGTGCACGACCAAGAGCATATCATGGAGCATCTAGAAGGTGTCATCAACAAACCAGAGG CGGAGATGTCGCCACAAGAATTGCAGCTCCATTACTTCAAAATGCATGATTATGATGGCAATAATTTGCTTGATGGCTTAGAACTCTCCA CAGCCATCACTCATGTCCATAAGGAGCATGACGTATCCCAGACTTTACTCAAGGCAACACTGGACAGTGTTGTAGAAGAATGTGTCAGCT TTGTGGGAGTGGATATTAACATCTGTTCAGAAGTTTTGTTAAGGCATATTGCAGGACTCAATGCCAACAGGGCCAAAAATATTATTGAAT GGCGAGAGAAAAATGGACCCTTTATCAACCGAGAACAGCTGAAGAAAGTGAAAGGGCTGGGCCCAAAATCCTTCCAACAGTGTGCTGGCT TCATCAGAATCAACCAGGATTATATCCGAACGTTTTGCAGTCAGCAAACTGAAACTTCAGGCCAAATTCAAGGAGTTGCTGTGACATCTT CAGCAGACGTTGAGGTCACAAATGAGAAGCAGGGCAAAAAGAAGAGCAAAACTGCAGTGAATGTTTTACTGAAGCCAAATCCTTTGGACC AAACTTGTATTCATCCAGAATCATATGACATAGCAATGAGGTTTTTGTCATCCATTGGAGGGACACTGTATGAGGTTGGAAAGCCTGAAA TGCAACAAAAAATAAATTCATTCCTTGAAAAGGAAGGAATGGAGAAAATTGCAGAAAGATTGCAAACAACAGTACACACCTTACAGGTCA TCATAGATGGTCTCAGCCAGCCTGAAAGCTTTGACTTTCGAACAGATTTTGATAAACCTGATTTCAAGAGAAGCATAGTATGCCTGGAAG ATCTGCAGATTGGGACAGTTCTTACAGGCAAAGTTGAGAATGCCACTCTCTTTGGAATTTTTGTGGATATAGGAGTGGGGAAATCTGGGC TGATTCCCATACGAAATGTAACAGAAGCAAAACTTTCAAAAACAAAGAAGAGAAGAAGCCTTGGACTGGGCCCCGGAGAAAGAGTGGAAG TCCAAGTACTCAACATTGACATCCCCCGATCTAGGATTACTCTGGACCTCATTCGGGTGTTATGAGTATCCCACGAAGGCCAGACGCTGA TTTTATTTTCTCATTTCCACAGATTGACAAGGATAAGTCAGTTGTTTGTAAACTCTAGGTAGCAGATGAGAAATAATTCACTTAATATCA GAAATATTTTCCAAACACTTTCCTTTATTTTTTCTTCTGAATAAATAGAAAACCAACAGTTTGATTTCCTTTTCCCTTAAAGGAAACAAC TAATACACATTCTTATATGGCTTTATGTAGTAATAGTTTTCTGACTAAAATTTTGTTTTTTATTTTTTGTAATTTATCTTTAACTCCTTT TGCATTTTGTATAACAGATTGCTTAACTTCTACTTGCCAACATCTGCCTTGCTGGACTTGTATGGGATTGTCTTCTTGATTTGAATTGTA CCGTCTTTGTTGACACAGTAGGGCTGGGCAGTGTTTAATCCTTCCATTTTATAGATTTTTTTTTAATCAGGCCTTTTGGACTTCATTCAT AATTTTGCAATAATCTCTTTTCCCTTGTCATGCAAGCCAAAAATATACCAGTAAAACAGATTCTGACGTGTTTGTAGTTATCAAATGAAT >52193_52193_3_MCFD2-SRBD1_MCFD2_chr2_47134949_ENST00000409207_SRBD1_chr2_45647033_ENST00000263736_length(amino acids)=444AA_BP=132 MQLPDQLRHAVPVALGAAALQLALHLQILMTMRSLLRTPFLCGLLWAFCAPGARAEEPAASFSQPGSMGLDKNTVHDQEHIMEHLEGVIN KPEAEMSPQELQLHYFKMHDYDGNNLLDGLELSTAITHVHKEHDVSQTLLKATLDSVVEECVSFVGVDINICSEVLLRHIAGLNANRAKN IIEWREKNGPFINREQLKKVKGLGPKSFQQCAGFIRINQDYIRTFCSQQTETSGQIQGVAVTSSADVEVTNEKQGKKKSKTAVNVLLKPN PLDQTCIHPESYDIAMRFLSSIGGTLYEVGKPEMQQKINSFLEKEGMEKIAERLQTTVHTLQVIIDGLSQPESFDFRTDFDKPDFKRSIV -------------------------------------------------------------- >52193_52193_4_MCFD2-SRBD1_MCFD2_chr2_47134949_ENST00000409207_SRBD1_chr2_45647033_ENST00000535761_length(transcript)=1632nt_BP=476nt TACTCACCGGTGTCCGGAAAGGTGAACGCTGCGCTCGGGCTGCCTCGCCTGTTACCTCCGCCGCCGGGCATGCTCAGCGTCTGCAGCTGC CGGACCAGCTCCGGCATGCGGTCCCAGTGGCCCTCGGCGCGGCAGCGCTCCAGCTCGCTCTCCACCTTCAGATATTGATGACCATGAGAT CCCTGCTCAGAACCCCCTTCCTGTGTGGCCTGCTCTGGGCCTTTTGTGCCCCAGGCGCCAGGGCTGAGGAGCCTGCAGCCAGCTTCTCCC AACCCGGCAGCATGGGCCTGGATAAGAACACAGTGCACGACCAAGAGCATATCATGGAGCATCTAGAAGGTGTCATCAACAAACCAGAGG CGGAGATGTCGCCACAAGAATTGCAGCTCCATTACTTCAAAATGCATGATTATGATGGCAATAATTTGCTTGATGGCTTAGAACTCTCCA CAGCCATCACTCATGTCCATAAGGAGCATGACGTATCCCAGACTTTACTCAAGGCAACACTGGACAGTGTTGTAGAAGAATGTGTCAGCT TTGTGGGAGTGGATATTAACATCTGTTCAGAAGTTTTGTTAAGGCATATTGCAGGACTCAATGCCAACAGGGCCAAAAATATTATTGAAT GGCGAGAGAAAAATGGACCCTTTATCAACCGAGAACAGCTGAAGAAAGTGAAAGGGCTGGGCCCAAAATCCTTCCAACAGTGTGCTGGCT TCATCAGAATCAACCAGGATTATATCCGAACGTTTTGCAGTCAGCAAACTGAAACTTCAGGCCAAATTCAAGGAGTTGCTGTGACATCTT CAGCAGACGTTGAGGTCACAAATGAGAAGCAGGGCAAAAAGAAGAGCAAAACTGCAGTGAATGTTTTACTGAAGCCAAATCCTTTGGACC AAACTTGTATTCATCCAGAATCATATGACATAGCAATGAGGTTTTTGTCATCCATTGGAGGGACACTGTATGAGGTTGGAAAGCCTGAAA TGCAACAAAAAATAAATTCATTCCTTGAAAAGGAAGGAATGGAGAAAATTGCAGAAAGATTGCAAACAACAGTACACACCTTACAGGTCA TCATAGATGGTCTCAGCCAGCCTGAAAGCTTTGACTTTCGAACAGATTTTGATAAACCTGATTTCAAGAGAAGCATAGTATGCCTGGAAG ATCTGCAGATTGGGACAGTTCTTACAGGCAAAGTTGAGAATGCCACTCTCTTTGGAATTTTTGTGGATATAGGAGTGGGGAAATCTGGGC TGATTCCCATACGAAATGTAACAGAAGCAAAACTTTCAAAAACAAAGAAGAGAAGAAGCCTTGGACTGGGCCCCGGAGAAAGAGTGGAAG TCCAAGTACTCAACATTGACATCCCCCGATCTAGGATTACTCTGGACCTCATTCGGGTGTTATGAGTATCCCACGAAGGCCAGACGCTGA TTTTATTTTCTCATTTCCACAGATTGACAAGGATAAGTCAGTTGTTTGTAAACTCTAGGTAGCAGATGAGAAATAATTCACTTAATATCA GAAATATTTTCCAAACACTTTCCTTTATTTTTTCTTCTGAATAAATAGAAAACCAACAGTTTGATTTCCTTTTCCCTTAAAGGAAACAAC >52193_52193_4_MCFD2-SRBD1_MCFD2_chr2_47134949_ENST00000409207_SRBD1_chr2_45647033_ENST00000535761_length(amino acids)=444AA_BP=132 MQLPDQLRHAVPVALGAAALQLALHLQILMTMRSLLRTPFLCGLLWAFCAPGARAEEPAASFSQPGSMGLDKNTVHDQEHIMEHLEGVIN KPEAEMSPQELQLHYFKMHDYDGNNLLDGLELSTAITHVHKEHDVSQTLLKATLDSVVEECVSFVGVDINICSEVLLRHIAGLNANRAKN IIEWREKNGPFINREQLKKVKGLGPKSFQQCAGFIRINQDYIRTFCSQQTETSGQIQGVAVTSSADVEVTNEKQGKKKSKTAVNVLLKPN PLDQTCIHPESYDIAMRFLSSIGGTLYEVGKPEMQQKINSFLEKEGMEKIAERLQTTVHTLQVIIDGLSQPESFDFRTDFDKPDFKRSIV -------------------------------------------------------------- >52193_52193_5_MCFD2-SRBD1_MCFD2_chr2_47134949_ENST00000444761_SRBD1_chr2_45647033_ENST00000263736_length(transcript)=2013nt_BP=444nt GCTCTTAAGACTAGCACGGTGGGCTGGCAGGCGCCCGCTGTCGGCCGAGGACTGTCTGGGCGCGCTTCCTGGAGGAGGCGGTGCGTTTCG CGCGCTCGCTGCCGGCGAGCCAGCCTCTTCCCTTACTCACCGGTGTCCGGAAAGGTGAACGCTGCGCTCGGGCTGCCTCGCCTGTTACCT CCGCCGCCGGGCATGCTCAGCGTCTGCAGCTGCCGGACCAGCTCCGGCATGCGGTCCCAGTGGCCCTCGGCGCGGCAGCGCTCCAGCTCG CTCTCCACCTTCAGGCATATCATGGAGCATCTAGAAGGTGTCATCAACAAACCAGAGGCGGAGATGTCGCCACAAGAATTGCAGCTCCAT TACTTCAAAATGCATGATTATGATGGCAATAATTTGCTTGATGGCTTAGAACTCTCCACAGCCATCACTCATGTCCATAAGGAGCATGAC GTATCCCAGACTTTACTCAAGGCAACACTGGACAGTGTTGTAGAAGAATGTGTCAGCTTTGTGGGAGTGGATATTAACATCTGTTCAGAA GTTTTGTTAAGGCATATTGCAGGACTCAATGCCAACAGGGCCAAAAATATTATTGAATGGCGAGAGAAAAATGGACCCTTTATCAACCGA GAACAGCTGAAGAAAGTGAAAGGGCTGGGCCCAAAATCCTTCCAACAGTGTGCTGGCTTCATCAGAATCAACCAGGATTATATCCGAACG TTTTGCAGTCAGCAAACTGAAACTTCAGGCCAAATTCAAGGAGTTGCTGTGACATCTTCAGCAGACGTTGAGGTCACAAATGAGAAGCAG GGCAAAAAGAAGAGCAAAACTGCAGTGAATGTTTTACTGAAGCCAAATCCTTTGGACCAAACTTGTATTCATCCAGAATCATATGACATA GCAATGAGGTTTTTGTCATCCATTGGAGGGACACTGTATGAGGTTGGAAAGCCTGAAATGCAACAAAAAATAAATTCATTCCTTGAAAAG GAAGGAATGGAGAAAATTGCAGAAAGATTGCAAACAACAGTACACACCTTACAGGTCATCATAGATGGTCTCAGCCAGCCTGAAAGCTTT GACTTTCGAACAGATTTTGATAAACCTGATTTCAAGAGAAGCATAGTATGCCTGGAAGATCTGCAGATTGGGACAGTTCTTACAGGCAAA GTTGAGAATGCCACTCTCTTTGGAATTTTTGTGGATATAGGAGTGGGGAAATCTGGGCTGATTCCCATACGAAATGTAACAGAAGCAAAA CTTTCAAAAACAAAGAAGAGAAGAAGCCTTGGACTGGGCCCCGGAGAAAGAGTGGAAGTCCAAGTACTCAACATTGACATCCCCCGATCT AGGATTACTCTGGACCTCATTCGGGTGTTATGAGTATCCCACGAAGGCCAGACGCTGATTTTATTTTCTCATTTCCACAGATTGACAAGG ATAAGTCAGTTGTTTGTAAACTCTAGGTAGCAGATGAGAAATAATTCACTTAATATCAGAAATATTTTCCAAACACTTTCCTTTATTTTT TCTTCTGAATAAATAGAAAACCAACAGTTTGATTTCCTTTTCCCTTAAAGGAAACAACTAATACACATTCTTATATGGCTTTATGTAGTA ATAGTTTTCTGACTAAAATTTTGTTTTTTATTTTTTGTAATTTATCTTTAACTCCTTTTGCATTTTGTATAACAGATTGCTTAACTTCTA CTTGCCAACATCTGCCTTGCTGGACTTGTATGGGATTGTCTTCTTGATTTGAATTGTACCGTCTTTGTTGACACAGTAGGGCTGGGCAGT GTTTAATCCTTCCATTTTATAGATTTTTTTTTAATCAGGCCTTTTGGACTTCATTCATAATTTTGCAATAATCTCTTTTCCCTTGTCATG CAAGCCAAAAATATACCAGTAAAACAGATTCTGACGTGTTTGTAGTTATCAAATGAATGGCTCGAAACACTTCTCAAAAGGATATACGTA >52193_52193_5_MCFD2-SRBD1_MCFD2_chr2_47134949_ENST00000444761_SRBD1_chr2_45647033_ENST00000263736_length(amino acids)=443AA_BP=131 MSGRASWRRRCVSRARCRRASLFPYSPVSGKVNAALGLPRLLPPPPGMLSVCSCRTSSGMRSQWPSARQRSSSLSTFRHIMEHLEGVINK PEAEMSPQELQLHYFKMHDYDGNNLLDGLELSTAITHVHKEHDVSQTLLKATLDSVVEECVSFVGVDINICSEVLLRHIAGLNANRAKNI IEWREKNGPFINREQLKKVKGLGPKSFQQCAGFIRINQDYIRTFCSQQTETSGQIQGVAVTSSADVEVTNEKQGKKKSKTAVNVLLKPNP LDQTCIHPESYDIAMRFLSSIGGTLYEVGKPEMQQKINSFLEKEGMEKIAERLQTTVHTLQVIIDGLSQPESFDFRTDFDKPDFKRSIVC -------------------------------------------------------------- >52193_52193_6_MCFD2-SRBD1_MCFD2_chr2_47134949_ENST00000444761_SRBD1_chr2_45647033_ENST00000535761_length(transcript)=1600nt_BP=444nt GCTCTTAAGACTAGCACGGTGGGCTGGCAGGCGCCCGCTGTCGGCCGAGGACTGTCTGGGCGCGCTTCCTGGAGGAGGCGGTGCGTTTCG CGCGCTCGCTGCCGGCGAGCCAGCCTCTTCCCTTACTCACCGGTGTCCGGAAAGGTGAACGCTGCGCTCGGGCTGCCTCGCCTGTTACCT CCGCCGCCGGGCATGCTCAGCGTCTGCAGCTGCCGGACCAGCTCCGGCATGCGGTCCCAGTGGCCCTCGGCGCGGCAGCGCTCCAGCTCG CTCTCCACCTTCAGGCATATCATGGAGCATCTAGAAGGTGTCATCAACAAACCAGAGGCGGAGATGTCGCCACAAGAATTGCAGCTCCAT TACTTCAAAATGCATGATTATGATGGCAATAATTTGCTTGATGGCTTAGAACTCTCCACAGCCATCACTCATGTCCATAAGGAGCATGAC GTATCCCAGACTTTACTCAAGGCAACACTGGACAGTGTTGTAGAAGAATGTGTCAGCTTTGTGGGAGTGGATATTAACATCTGTTCAGAA GTTTTGTTAAGGCATATTGCAGGACTCAATGCCAACAGGGCCAAAAATATTATTGAATGGCGAGAGAAAAATGGACCCTTTATCAACCGA GAACAGCTGAAGAAAGTGAAAGGGCTGGGCCCAAAATCCTTCCAACAGTGTGCTGGCTTCATCAGAATCAACCAGGATTATATCCGAACG TTTTGCAGTCAGCAAACTGAAACTTCAGGCCAAATTCAAGGAGTTGCTGTGACATCTTCAGCAGACGTTGAGGTCACAAATGAGAAGCAG GGCAAAAAGAAGAGCAAAACTGCAGTGAATGTTTTACTGAAGCCAAATCCTTTGGACCAAACTTGTATTCATCCAGAATCATATGACATA GCAATGAGGTTTTTGTCATCCATTGGAGGGACACTGTATGAGGTTGGAAAGCCTGAAATGCAACAAAAAATAAATTCATTCCTTGAAAAG GAAGGAATGGAGAAAATTGCAGAAAGATTGCAAACAACAGTACACACCTTACAGGTCATCATAGATGGTCTCAGCCAGCCTGAAAGCTTT GACTTTCGAACAGATTTTGATAAACCTGATTTCAAGAGAAGCATAGTATGCCTGGAAGATCTGCAGATTGGGACAGTTCTTACAGGCAAA GTTGAGAATGCCACTCTCTTTGGAATTTTTGTGGATATAGGAGTGGGGAAATCTGGGCTGATTCCCATACGAAATGTAACAGAAGCAAAA CTTTCAAAAACAAAGAAGAGAAGAAGCCTTGGACTGGGCCCCGGAGAAAGAGTGGAAGTCCAAGTACTCAACATTGACATCCCCCGATCT AGGATTACTCTGGACCTCATTCGGGTGTTATGAGTATCCCACGAAGGCCAGACGCTGATTTTATTTTCTCATTTCCACAGATTGACAAGG ATAAGTCAGTTGTTTGTAAACTCTAGGTAGCAGATGAGAAATAATTCACTTAATATCAGAAATATTTTCCAAACACTTTCCTTTATTTTT >52193_52193_6_MCFD2-SRBD1_MCFD2_chr2_47134949_ENST00000444761_SRBD1_chr2_45647033_ENST00000535761_length(amino acids)=443AA_BP=131 MSGRASWRRRCVSRARCRRASLFPYSPVSGKVNAALGLPRLLPPPPGMLSVCSCRTSSGMRSQWPSARQRSSSLSTFRHIMEHLEGVINK PEAEMSPQELQLHYFKMHDYDGNNLLDGLELSTAITHVHKEHDVSQTLLKATLDSVVEECVSFVGVDINICSEVLLRHIAGLNANRAKNI IEWREKNGPFINREQLKKVKGLGPKSFQQCAGFIRINQDYIRTFCSQQTETSGQIQGVAVTSSADVEVTNEKQGKKKSKTAVNVLLKPNP LDQTCIHPESYDIAMRFLSSIGGTLYEVGKPEMQQKINSFLEKEGMEKIAERLQTTVHTLQVIIDGLSQPESFDFRTDFDKPDFKRSIVC -------------------------------------------------------------- >52193_52193_7_MCFD2-SRBD1_MCFD2_chr2_47136162_ENST00000409105_SRBD1_chr2_45715470_ENST00000263736_length(transcript)=2073nt_BP=329nt ACGGGGGCTGGTGAGGCTCACGTTGGAGGGCTTCGCGTCTGCTTCGGAGACCGTAAGGCTGTCATCAGCTGTGGACGGGAGCCTTCTGAA GGGAAGCACTTGGGAGCCTGCAGGACGAATACCTACACCAGACTTGGAATTGAAAAGACCTCACTGGAGAAAGAGACATTTGATATATTG ATGACCATGAGATCCCTGCTCAGAACCCCCTTCCTGTGTGGCCTGCTCTGGGCCTTTTGTGCCCCAGGCGCCAGGGCTGAGGAGCCTGCA GCCAGCTTCTCCCAACCCGGCAGCATGGGCCTGGATAAGAACACAGTGCACGACCAAGATATCGTCAGTGAAGCAGGAGCATCAATCTAC AGTGTCAGCCCTGAAGCTAACAAAGAGATGCCAGGGCTGGACCCTAATTTGAGAAGTGCAGTTTCCATAGCAAGGCGTGTACAAGATCCA TTAGCTGAGCTAGTGAAAATTGAGCCAAAGCACATTGGAGTTGGAATGTATCAGCATGACGTATCCCAGACTTTACTCAAGGCAACACTG GACAGTGTTGTAGAAGAATGTGTCAGCTTTGTGGGAGTGGATATTAACATCTGTTCAGAAGTTTTGTTAAGGCATATTGCAGGACTCAAT GCCAACAGGGCCAAAAATATTATTGAATGGCGAGAGAAAAATGGACCCTTTATCAACCGAGAACAGCTGAAGAAAGTGAAAGGGCTGGGC CCAAAATCCTTCCAACAGTGTGCTGGCTTCATCAGAATCAACCAGGATTATATCCGAACGTTTTGCAGTCAGCAAACTGAAACTTCAGGC CAAATTCAAGGAGTTGCTGTGACATCTTCAGCAGACGTTGAGGTCACAAATGAGAAGCAGGGCAAAAAGAAGAGCAAAACTGCAGTGAAT GTTTTACTGAAGCCAAATCCTTTGGACCAAACTTGTATTCATCCAGAATCATATGACATAGCAATGAGGTTTTTGTCATCCATTGGAGGG ACACTGTATGAGGTTGGAAAGCCTGAAATGCAACAAAAAATAAATTCATTCCTTGAAAAGGAAGGAATGGAGAAAATTGCAGAAAGATTG CAAACAACAGTACACACCTTACAGGTCATCATAGATGGTCTCAGCCAGCCTGAAAGCTTTGACTTTCGAACAGATTTTGATAAACCTGAT TTCAAGAGAAGCATAGTATGCCTGGAAGATCTGCAGATTGGGACAGTTCTTACAGGCAAAGTTGAGAATGCCACTCTCTTTGGAATTTTT GTGGATATAGGAGTGGGGAAATCTGGGCTGATTCCCATACGAAATGTAACAGAAGCAAAACTTTCAAAAACAAAGAAGAGAAGAAGCCTT GGACTGGGCCCCGGAGAAAGAGTGGAAGTCCAAGTACTCAACATTGACATCCCCCGATCTAGGATTACTCTGGACCTCATTCGGGTGTTA TGAGTATCCCACGAAGGCCAGACGCTGATTTTATTTTCTCATTTCCACAGATTGACAAGGATAAGTCAGTTGTTTGTAAACTCTAGGTAG CAGATGAGAAATAATTCACTTAATATCAGAAATATTTTCCAAACACTTTCCTTTATTTTTTCTTCTGAATAAATAGAAAACCAACAGTTT GATTTCCTTTTCCCTTAAAGGAAACAACTAATACACATTCTTATATGGCTTTATGTAGTAATAGTTTTCTGACTAAAATTTTGTTTTTTA TTTTTTGTAATTTATCTTTAACTCCTTTTGCATTTTGTATAACAGATTGCTTAACTTCTACTTGCCAACATCTGCCTTGCTGGACTTGTA TGGGATTGTCTTCTTGATTTGAATTGTACCGTCTTTGTTGACACAGTAGGGCTGGGCAGTGTTTAATCCTTCCATTTTATAGATTTTTTT TTAATCAGGCCTTTTGGACTTCATTCATAATTTTGCAATAATCTCTTTTCCCTTGTCATGCAAGCCAAAAATATACCAGTAAAACAGATT CTGACGTGTTTGTAGTTATCAAATGAATGGCTCGAAACACTTCTCAAAAGGATATACGTATTGACCCCAACAATAAATGTTTGTGGCTAG >52193_52193_7_MCFD2-SRBD1_MCFD2_chr2_47136162_ENST00000409105_SRBD1_chr2_45715470_ENST00000263736_length(amino acids)=447AA_BP=77 MGACRTNTYTRLGIEKTSLEKETFDILMTMRSLLRTPFLCGLLWAFCAPGARAEEPAASFSQPGSMGLDKNTVHDQDIVSEAGASIYSVS PEANKEMPGLDPNLRSAVSIARRVQDPLAELVKIEPKHIGVGMYQHDVSQTLLKATLDSVVEECVSFVGVDINICSEVLLRHIAGLNANR AKNIIEWREKNGPFINREQLKKVKGLGPKSFQQCAGFIRINQDYIRTFCSQQTETSGQIQGVAVTSSADVEVTNEKQGKKKSKTAVNVLL KPNPLDQTCIHPESYDIAMRFLSSIGGTLYEVGKPEMQQKINSFLEKEGMEKIAERLQTTVHTLQVIIDGLSQPESFDFRTDFDKPDFKR -------------------------------------------------------------- >52193_52193_8_MCFD2-SRBD1_MCFD2_chr2_47136162_ENST00000409105_SRBD1_chr2_45715470_ENST00000535761_length(transcript)=1660nt_BP=329nt ACGGGGGCTGGTGAGGCTCACGTTGGAGGGCTTCGCGTCTGCTTCGGAGACCGTAAGGCTGTCATCAGCTGTGGACGGGAGCCTTCTGAA GGGAAGCACTTGGGAGCCTGCAGGACGAATACCTACACCAGACTTGGAATTGAAAAGACCTCACTGGAGAAAGAGACATTTGATATATTG ATGACCATGAGATCCCTGCTCAGAACCCCCTTCCTGTGTGGCCTGCTCTGGGCCTTTTGTGCCCCAGGCGCCAGGGCTGAGGAGCCTGCA GCCAGCTTCTCCCAACCCGGCAGCATGGGCCTGGATAAGAACACAGTGCACGACCAAGATATCGTCAGTGAAGCAGGAGCATCAATCTAC AGTGTCAGCCCTGAAGCTAACAAAGAGATGCCAGGGCTGGACCCTAATTTGAGAAGTGCAGTTTCCATAGCAAGGCGTGTACAAGATCCA TTAGCTGAGCTAGTGAAAATTGAGCCAAAGCACATTGGAGTTGGAATGTATCAGCATGACGTATCCCAGACTTTACTCAAGGCAACACTG GACAGTGTTGTAGAAGAATGTGTCAGCTTTGTGGGAGTGGATATTAACATCTGTTCAGAAGTTTTGTTAAGGCATATTGCAGGACTCAAT GCCAACAGGGCCAAAAATATTATTGAATGGCGAGAGAAAAATGGACCCTTTATCAACCGAGAACAGCTGAAGAAAGTGAAAGGGCTGGGC CCAAAATCCTTCCAACAGTGTGCTGGCTTCATCAGAATCAACCAGGATTATATCCGAACGTTTTGCAGTCAGCAAACTGAAACTTCAGGC CAAATTCAAGGAGTTGCTGTGACATCTTCAGCAGACGTTGAGGTCACAAATGAGAAGCAGGGCAAAAAGAAGAGCAAAACTGCAGTGAAT GTTTTACTGAAGCCAAATCCTTTGGACCAAACTTGTATTCATCCAGAATCATATGACATAGCAATGAGGTTTTTGTCATCCATTGGAGGG ACACTGTATGAGGTTGGAAAGCCTGAAATGCAACAAAAAATAAATTCATTCCTTGAAAAGGAAGGAATGGAGAAAATTGCAGAAAGATTG CAAACAACAGTACACACCTTACAGGTCATCATAGATGGTCTCAGCCAGCCTGAAAGCTTTGACTTTCGAACAGATTTTGATAAACCTGAT TTCAAGAGAAGCATAGTATGCCTGGAAGATCTGCAGATTGGGACAGTTCTTACAGGCAAAGTTGAGAATGCCACTCTCTTTGGAATTTTT GTGGATATAGGAGTGGGGAAATCTGGGCTGATTCCCATACGAAATGTAACAGAAGCAAAACTTTCAAAAACAAAGAAGAGAAGAAGCCTT GGACTGGGCCCCGGAGAAAGAGTGGAAGTCCAAGTACTCAACATTGACATCCCCCGATCTAGGATTACTCTGGACCTCATTCGGGTGTTA TGAGTATCCCACGAAGGCCAGACGCTGATTTTATTTTCTCATTTCCACAGATTGACAAGGATAAGTCAGTTGTTTGTAAACTCTAGGTAG CAGATGAGAAATAATTCACTTAATATCAGAAATATTTTCCAAACACTTTCCTTTATTTTTTCTTCTGAATAAATAGAAAACCAACAGTTT >52193_52193_8_MCFD2-SRBD1_MCFD2_chr2_47136162_ENST00000409105_SRBD1_chr2_45715470_ENST00000535761_length(amino acids)=447AA_BP=77 MGACRTNTYTRLGIEKTSLEKETFDILMTMRSLLRTPFLCGLLWAFCAPGARAEEPAASFSQPGSMGLDKNTVHDQDIVSEAGASIYSVS PEANKEMPGLDPNLRSAVSIARRVQDPLAELVKIEPKHIGVGMYQHDVSQTLLKATLDSVVEECVSFVGVDINICSEVLLRHIAGLNANR AKNIIEWREKNGPFINREQLKKVKGLGPKSFQQCAGFIRINQDYIRTFCSQQTETSGQIQGVAVTSSADVEVTNEKQGKKKSKTAVNVLL KPNPLDQTCIHPESYDIAMRFLSSIGGTLYEVGKPEMQQKINSFLEKEGMEKIAERLQTTVHTLQVIIDGLSQPESFDFRTDFDKPDFKR -------------------------------------------------------------- >52193_52193_9_MCFD2-SRBD1_MCFD2_chr2_47136162_ENST00000409207_SRBD1_chr2_45715470_ENST00000263736_length(transcript)=2060nt_BP=316nt TACTCACCGGTGTCCGGAAAGGTGAACGCTGCGCTCGGGCTGCCTCGCCTGTTACCTCCGCCGCCGGGCATGCTCAGCGTCTGCAGCTGC CGGACCAGCTCCGGCATGCGGTCCCAGTGGCCCTCGGCGCGGCAGCGCTCCAGCTCGCTCTCCACCTTCAGATATTGATGACCATGAGAT CCCTGCTCAGAACCCCCTTCCTGTGTGGCCTGCTCTGGGCCTTTTGTGCCCCAGGCGCCAGGGCTGAGGAGCCTGCAGCCAGCTTCTCCC AACCCGGCAGCATGGGCCTGGATAAGAACACAGTGCACGACCAAGATATCGTCAGTGAAGCAGGAGCATCAATCTACAGTGTCAGCCCTG AAGCTAACAAAGAGATGCCAGGGCTGGACCCTAATTTGAGAAGTGCAGTTTCCATAGCAAGGCGTGTACAAGATCCATTAGCTGAGCTAG TGAAAATTGAGCCAAAGCACATTGGAGTTGGAATGTATCAGCATGACGTATCCCAGACTTTACTCAAGGCAACACTGGACAGTGTTGTAG AAGAATGTGTCAGCTTTGTGGGAGTGGATATTAACATCTGTTCAGAAGTTTTGTTAAGGCATATTGCAGGACTCAATGCCAACAGGGCCA AAAATATTATTGAATGGCGAGAGAAAAATGGACCCTTTATCAACCGAGAACAGCTGAAGAAAGTGAAAGGGCTGGGCCCAAAATCCTTCC AACAGTGTGCTGGCTTCATCAGAATCAACCAGGATTATATCCGAACGTTTTGCAGTCAGCAAACTGAAACTTCAGGCCAAATTCAAGGAG TTGCTGTGACATCTTCAGCAGACGTTGAGGTCACAAATGAGAAGCAGGGCAAAAAGAAGAGCAAAACTGCAGTGAATGTTTTACTGAAGC CAAATCCTTTGGACCAAACTTGTATTCATCCAGAATCATATGACATAGCAATGAGGTTTTTGTCATCCATTGGAGGGACACTGTATGAGG TTGGAAAGCCTGAAATGCAACAAAAAATAAATTCATTCCTTGAAAAGGAAGGAATGGAGAAAATTGCAGAAAGATTGCAAACAACAGTAC ACACCTTACAGGTCATCATAGATGGTCTCAGCCAGCCTGAAAGCTTTGACTTTCGAACAGATTTTGATAAACCTGATTTCAAGAGAAGCA TAGTATGCCTGGAAGATCTGCAGATTGGGACAGTTCTTACAGGCAAAGTTGAGAATGCCACTCTCTTTGGAATTTTTGTGGATATAGGAG TGGGGAAATCTGGGCTGATTCCCATACGAAATGTAACAGAAGCAAAACTTTCAAAAACAAAGAAGAGAAGAAGCCTTGGACTGGGCCCCG GAGAAAGAGTGGAAGTCCAAGTACTCAACATTGACATCCCCCGATCTAGGATTACTCTGGACCTCATTCGGGTGTTATGAGTATCCCACG AAGGCCAGACGCTGATTTTATTTTCTCATTTCCACAGATTGACAAGGATAAGTCAGTTGTTTGTAAACTCTAGGTAGCAGATGAGAAATA ATTCACTTAATATCAGAAATATTTTCCAAACACTTTCCTTTATTTTTTCTTCTGAATAAATAGAAAACCAACAGTTTGATTTCCTTTTCC CTTAAAGGAAACAACTAATACACATTCTTATATGGCTTTATGTAGTAATAGTTTTCTGACTAAAATTTTGTTTTTTATTTTTTGTAATTT ATCTTTAACTCCTTTTGCATTTTGTATAACAGATTGCTTAACTTCTACTTGCCAACATCTGCCTTGCTGGACTTGTATGGGATTGTCTTC TTGATTTGAATTGTACCGTCTTTGTTGACACAGTAGGGCTGGGCAGTGTTTAATCCTTCCATTTTATAGATTTTTTTTTAATCAGGCCTT TTGGACTTCATTCATAATTTTGCAATAATCTCTTTTCCCTTGTCATGCAAGCCAAAAATATACCAGTAAAACAGATTCTGACGTGTTTGT >52193_52193_9_MCFD2-SRBD1_MCFD2_chr2_47136162_ENST00000409207_SRBD1_chr2_45715470_ENST00000263736_length(amino acids)=449AA_BP=79 MQLPDQLRHAVPVALGAAALQLALHLQILMTMRSLLRTPFLCGLLWAFCAPGARAEEPAASFSQPGSMGLDKNTVHDQDIVSEAGASIYS VSPEANKEMPGLDPNLRSAVSIARRVQDPLAELVKIEPKHIGVGMYQHDVSQTLLKATLDSVVEECVSFVGVDINICSEVLLRHIAGLNA NRAKNIIEWREKNGPFINREQLKKVKGLGPKSFQQCAGFIRINQDYIRTFCSQQTETSGQIQGVAVTSSADVEVTNEKQGKKKSKTAVNV LLKPNPLDQTCIHPESYDIAMRFLSSIGGTLYEVGKPEMQQKINSFLEKEGMEKIAERLQTTVHTLQVIIDGLSQPESFDFRTDFDKPDF -------------------------------------------------------------- >52193_52193_10_MCFD2-SRBD1_MCFD2_chr2_47136162_ENST00000409207_SRBD1_chr2_45715470_ENST00000535761_length(transcript)=1647nt_BP=316nt TACTCACCGGTGTCCGGAAAGGTGAACGCTGCGCTCGGGCTGCCTCGCCTGTTACCTCCGCCGCCGGGCATGCTCAGCGTCTGCAGCTGC CGGACCAGCTCCGGCATGCGGTCCCAGTGGCCCTCGGCGCGGCAGCGCTCCAGCTCGCTCTCCACCTTCAGATATTGATGACCATGAGAT CCCTGCTCAGAACCCCCTTCCTGTGTGGCCTGCTCTGGGCCTTTTGTGCCCCAGGCGCCAGGGCTGAGGAGCCTGCAGCCAGCTTCTCCC AACCCGGCAGCATGGGCCTGGATAAGAACACAGTGCACGACCAAGATATCGTCAGTGAAGCAGGAGCATCAATCTACAGTGTCAGCCCTG AAGCTAACAAAGAGATGCCAGGGCTGGACCCTAATTTGAGAAGTGCAGTTTCCATAGCAAGGCGTGTACAAGATCCATTAGCTGAGCTAG TGAAAATTGAGCCAAAGCACATTGGAGTTGGAATGTATCAGCATGACGTATCCCAGACTTTACTCAAGGCAACACTGGACAGTGTTGTAG AAGAATGTGTCAGCTTTGTGGGAGTGGATATTAACATCTGTTCAGAAGTTTTGTTAAGGCATATTGCAGGACTCAATGCCAACAGGGCCA AAAATATTATTGAATGGCGAGAGAAAAATGGACCCTTTATCAACCGAGAACAGCTGAAGAAAGTGAAAGGGCTGGGCCCAAAATCCTTCC AACAGTGTGCTGGCTTCATCAGAATCAACCAGGATTATATCCGAACGTTTTGCAGTCAGCAAACTGAAACTTCAGGCCAAATTCAAGGAG TTGCTGTGACATCTTCAGCAGACGTTGAGGTCACAAATGAGAAGCAGGGCAAAAAGAAGAGCAAAACTGCAGTGAATGTTTTACTGAAGC CAAATCCTTTGGACCAAACTTGTATTCATCCAGAATCATATGACATAGCAATGAGGTTTTTGTCATCCATTGGAGGGACACTGTATGAGG TTGGAAAGCCTGAAATGCAACAAAAAATAAATTCATTCCTTGAAAAGGAAGGAATGGAGAAAATTGCAGAAAGATTGCAAACAACAGTAC ACACCTTACAGGTCATCATAGATGGTCTCAGCCAGCCTGAAAGCTTTGACTTTCGAACAGATTTTGATAAACCTGATTTCAAGAGAAGCA TAGTATGCCTGGAAGATCTGCAGATTGGGACAGTTCTTACAGGCAAAGTTGAGAATGCCACTCTCTTTGGAATTTTTGTGGATATAGGAG TGGGGAAATCTGGGCTGATTCCCATACGAAATGTAACAGAAGCAAAACTTTCAAAAACAAAGAAGAGAAGAAGCCTTGGACTGGGCCCCG GAGAAAGAGTGGAAGTCCAAGTACTCAACATTGACATCCCCCGATCTAGGATTACTCTGGACCTCATTCGGGTGTTATGAGTATCCCACG AAGGCCAGACGCTGATTTTATTTTCTCATTTCCACAGATTGACAAGGATAAGTCAGTTGTTTGTAAACTCTAGGTAGCAGATGAGAAATA ATTCACTTAATATCAGAAATATTTTCCAAACACTTTCCTTTATTTTTTCTTCTGAATAAATAGAAAACCAACAGTTTGATTTCCTTTTCC >52193_52193_10_MCFD2-SRBD1_MCFD2_chr2_47136162_ENST00000409207_SRBD1_chr2_45715470_ENST00000535761_length(amino acids)=449AA_BP=79 MQLPDQLRHAVPVALGAAALQLALHLQILMTMRSLLRTPFLCGLLWAFCAPGARAEEPAASFSQPGSMGLDKNTVHDQDIVSEAGASIYS VSPEANKEMPGLDPNLRSAVSIARRVQDPLAELVKIEPKHIGVGMYQHDVSQTLLKATLDSVVEECVSFVGVDINICSEVLLRHIAGLNA NRAKNIIEWREKNGPFINREQLKKVKGLGPKSFQQCAGFIRINQDYIRTFCSQQTETSGQIQGVAVTSSADVEVTNEKQGKKKSKTAVNV LLKPNPLDQTCIHPESYDIAMRFLSSIGGTLYEVGKPEMQQKINSFLEKEGMEKIAERLQTTVHTLQVIIDGLSQPESFDFRTDFDKPDF -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for MCFD2-SRBD1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for MCFD2-SRBD1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for MCFD2-SRBD1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
| Hgene | MCFD2 | C3150889 | FACTOR V AND FACTOR VIII, COMBINED DEFICIENCY OF, 2 | 5 | CTD_human;GENOMICS_ENGLAND;UNIPROT |
| Hgene | MCFD2 | C1856883 | FACTOR V AND FACTOR VIII, COMBINED DEFICIENCY OF | 2 | ORPHANET |
| Hgene | MCFD2 | C4551981 | Familial Multiple Coagulation Factor Deficiency I | 1 | GENOMICS_ENGLAND |
