
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:CHIC2-HSPA4 (FusionGDB2 ID:HG26511TG3308) |
Fusion Gene Summary for CHIC2-HSPA4 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: CHIC2-HSPA4 | Fusion gene ID: hg26511tg3308 | Hgene | Tgene | Gene symbol | CHIC2 | HSPA4 | Gene ID | 26511 | 3308 |
| Gene name | cysteine rich hydrophobic domain 2 | heat shock protein family A (Hsp70) member 4 | |
| Synonyms | BTL | APG-2|HEL-S-5a|HS24/P52|HSPH2|RY|hsp70|hsp70RY | |
| Cytomap | ('CHIC2')('HSPA4') 4q12 | 5q31.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | cysteine-rich hydrophobic domain-containing protein 2BRX-like translocated in leukemiacystein-rich hydrophobic domain 2cysteine-rich hydrophobic domain 2 protein | heat shock 70 kDa protein 4epididymis secretory sperm binding protein Li 5aheat shock 70-related protein APG-2heat shock 70kD protein 4heat shock 70kDa protein 4heat shock protein, 110 kDahsp70 RY | |
| Modification date | 20200313 | 20200327 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000263921, ENST00000512964, | ||
| Fusion gene scores | * DoF score | 11 X 7 X 9=693 | 10 X 11 X 4=440 |
| # samples | 12 | 12 | |
| ** MAII score | log2(12/693*10)=-2.5298209465287 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(12/440*10)=-1.87446911791614 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: CHIC2 [Title/Abstract] AND HSPA4 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | CHIC2(54930348)-HSPA4(132412345), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | CHIC2-HSPA4 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. CHIC2-HSPA4 seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. CHIC2-HSPA4 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. CHIC2-HSPA4 seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | HSPA4 | GO:0045040 | protein import into mitochondrial outer membrane | 15644312 |
| Tgene | HSPA4 | GO:0051131 | chaperone-mediated protein complex assembly | 15644312 |
 Fusion gene breakpoints across CHIC2 (5'-gene) Fusion gene breakpoints across CHIC2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
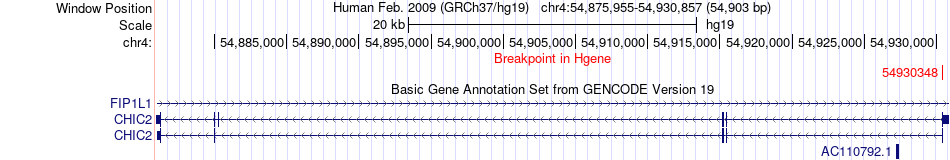 |
 Fusion gene breakpoints across HSPA4 (3'-gene) Fusion gene breakpoints across HSPA4 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
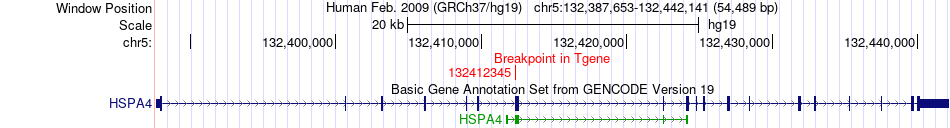 |
 Fusion gene information Fusion gene information * All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-E2-A572 | CHIC2 | chr4 | 54930348 | - | HSPA4 | chr5 | 132412345 | + |
Top |
Fusion Gene ORF analysis for CHIC2-HSPA4 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-3UTR | ENST00000263921 | ENST00000504328 | CHIC2 | chr4 | 54930348 | - | HSPA4 | chr5 | 132412345 | + |
| 5CDS-3UTR | ENST00000512964 | ENST00000504328 | CHIC2 | chr4 | 54930348 | - | HSPA4 | chr5 | 132412345 | + |
| In-frame | ENST00000263921 | ENST00000304858 | CHIC2 | chr4 | 54930348 | - | HSPA4 | chr5 | 132412345 | + |
| In-frame | ENST00000512964 | ENST00000304858 | CHIC2 | chr4 | 54930348 | - | HSPA4 | chr5 | 132412345 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000263921 | CHIC2 | chr4 | 54930348 | - | ENST00000304858 | HSPA4 | chr5 | 132412345 | + | 4382 | 509 | 512 | 2368 | 618 |
| ENST00000512964 | CHIC2 | chr4 | 54930348 | - | ENST00000304858 | HSPA4 | chr5 | 132412345 | + | 3992 | 119 | 122 | 1978 | 618 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000263921 | ENST00000304858 | CHIC2 | chr4 | 54930348 | - | HSPA4 | chr5 | 132412345 | + | 0.000180429 | 0.9998196 |
| ENST00000512964 | ENST00000304858 | CHIC2 | chr4 | 54930348 | - | HSPA4 | chr5 | 132412345 | + | 0.000149114 | 0.99985087 |
Top |
Fusion Genomic Features for CHIC2-HSPA4 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| CHIC2 | chr4 | 54930348 | - | HSPA4 | chr5 | 132412345 | + | 1.63E-09 | 1 |
| CHIC2 | chr4 | 54930348 | - | HSPA4 | chr5 | 132412345 | + | 1.63E-09 | 1 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
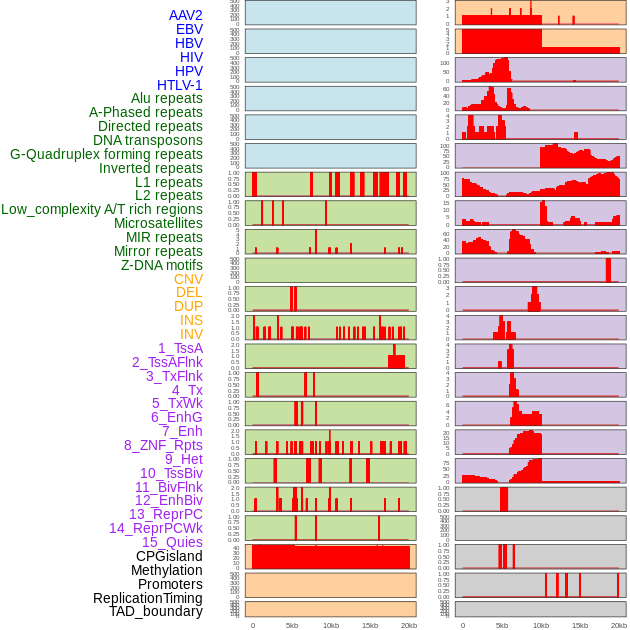 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
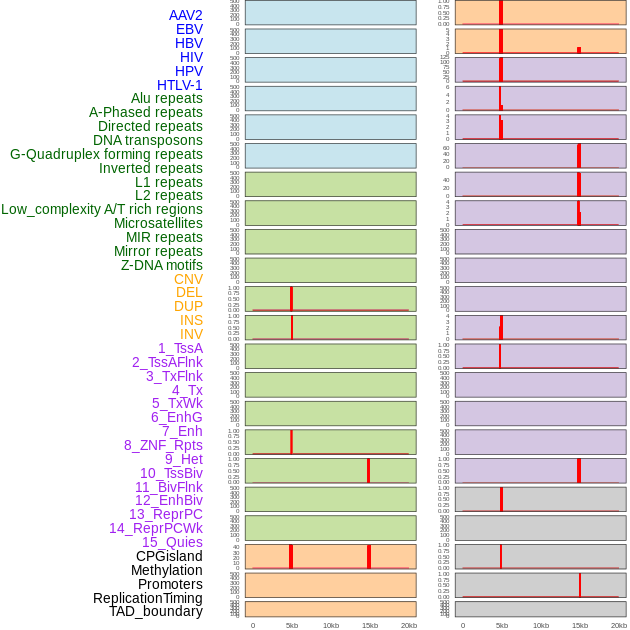 |
Top |
Fusion Protein Features for CHIC2-HSPA4 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr4:54930348/chr5:132412345) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CHIC2 | chr4:54930348 | chr5:132412345 | ENST00000263921 | - | 1 | 6 | 1_26 | 39 | 166.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | CHIC2 | chr4:54930348 | chr5:132412345 | ENST00000263921 | - | 1 | 6 | 9_15 | 39 | 166.0 | Compositional bias | Note=Poly-Glu |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CHIC2 | chr4:54930348 | chr5:132412345 | ENST00000263921 | - | 1 | 6 | 88_106 | 39 | 166.0 | Motif | Note=CHIC motif (Cys-rich) |
Top |
Fusion Gene Sequence for CHIC2-HSPA4 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >16480_16480_1_CHIC2-HSPA4_CHIC2_chr4_54930348_ENST00000263921_HSPA4_chr5_132412345_ENST00000304858_length(transcript)=4382nt_BP=509nt ACGCGCCCGCCCGCCCGTTGGCCCGGGAAGTCTCGCGATGCCGTAGCCGGCTGCTCCCGCTGCGGGTGGCTCGGGCTGTTGCTGTGGTTT CCTGAGTTGCTGCTGCTGCGGCGGCGGCAGCGGCGTCTGTGCTTGTGGAGGTGTCGGCCTCTGGGCGGATGTTGACATTGTGTTGTTGTT ATTGCTGATGGTAATGGCGGCGGCGGTGGCGGCGACGGTCCAGACCCCATCCCCTCTGTAGCCGGAGCCGAGACAGCCGACAGCGAACTC CGCGGCCTCGGAGCCGGCGGCAGCGGCGACTCCCCTCAGCCTCCGCCGCCTCGCCCGCCGGTACCCCGGCGCCAACCCCGGGAGTCAGGC CCTTTGGGCAGGGGAGCTCGGAGGCTCAGGATGGCGGATTTCGACGAAATCTATGAGGAAGAGGAGGACGAGGAGCGGGCCCTGGAGGAG CAGCTGCTCAAGTACTCGCCGGACCCGGTGGTCGTCCGCGGCTCCGGTCACGTCACCGTGTTCTGGCCACTGCATTTGACACGACATTGG GAGGTAGAAAATTTGATGAAGTGTTAGTAAATCACTTCTGTGAAGAATTTGGGAAGAAATACAAGCTAGACATTAAGTCCAAAATCCGTG CATTATTACGACTCTCTCAGGAGTGTGAGAAACTCAAGAAATTGATGAGTGCAAATGCTTCAGATCTCCCTTTGAGCATTGAATGTTTTA TGAATGATGTTGATGTATCTGGAACTATGAATAGAGGCAAATTTCTGGAGATGTGCAATGATCTCTTAGCTAGAGTGGAGCCACCACTTC GTAGTGTTTTGGAACAAACCAAGTTAAAGAAAGAAGATATTTATGCAGTGGAGATAGTTGGTGGTGCTACACGAATCCCTGCGGTAAAAG AGAAGATCAGCAAATTTTTCGGTAAAGAACTTAGTACAACATTAAATGCTGATGAAGCTGTCACTCGAGGCTGTGCATTGCAGTGTGCCA TCTTATCGCCTGCTTTCAAAGTCAGAGAATTTTCTATCACTGATGTAGTACCATATCCAATATCTCTGAGATGGAATTCTCCAGCTGAAG AAGGGTCAAGTGACTGTGAAGTCTTTTCCAAAAATCATGCTGCTCCTTTCTCTAAAGTTCTTACATTTTATAGAAAGGAACCTTTCACTC TTGAGGCCTACTACAGCTCTCCTCAGGATTTGCCCTATCCAGATCCTGCTATAGCTCAGTTTTCAGTTCAGAAAGTCACTCCTCAGTCTG ATGGCTCCAGTTCAAAAGTGAAAGTCAAAGTTCGAGTAAATGTCCATGGCATTTTCAGTGTGTCCAGTGCATCTTTAGTGGAGGTTCACA AGTCTGAGGAAAATGAGGAGCCAATGGAAACAGATCAGAATGCAAAGGAGGAAGAGAAGATGCAAGTGGACCAGGAGGAACCACATGTTG AAGAGCAACAGCAGCAGACACCAGCAGAAAATAAGGCAGAGTCTGAAGAAATGGAGACCTCTCAAGCTGGATCCAAGGATAAAAAGATGG ACCAACCACCCCAAGCCAAGAAGGCAAAAGTGAAGACCAGTACTGTGGACCTGCCAATCGAGAATCAGCTATTATGGCAGATAGACAGAG AGATGCTCAACTTGTACATTGAAAATGAGGGTAAGATGATCATGCAGGATAAACTGGAGAAGGAGCGGAATGATGCTAAGAACGCAGTGG AGGAATATGTGTATGAAATGAGAGACAAGCTTAGTGGTGAATATGAGAAGTTTGTGAGTGAAGATGATCGTAACAGTTTTACTTTGAAAC TGGAAGATACTGAAAATTGGTTGTATGAGGATGGAGAAGACCAGCCAAAGCAAGTTTATGTTGATAAGTTGGCTGAATTAAAAAATCTAG GTCAACCTATTAAGATACGTTTCCAGGAATCTGAAGAACGACCAAAATTATTTGAAGAACTAGGGAAACAGATCCAACAGTATATGAAAA TAATCAGCTCTTTCAAAAACAAGGAGGACCAGTATGATCATTTGGATGCTGCTGACATGACAAAGGTAGAAAAAAGCACAAATGAAGCAA TGGAGTGGATGAATAACAAGCTAAATCTGCAGAACAAGCAGAGTTTGACCATGGATCCAGTTGTCAAGTCAAAAGAGATTGAAGCTAAAA TTAAGGAGCTGACAAGTACTTGTAGCCCTATAATTTCAAAGCCCAAACCCAAAGTGGAACCTCCAAAAGAGGAACAAAAAAATGCAGAGC AGAATGGACCAGTGGATGGACAAGGAGACAACCCAGGCCCCCAGGCTGCTGAGCAGGGTACAGACACAGCTGTGCCTTCGGATTCAGACA AGAAGCTTCCTGAAATGGACATTGATTGATTCCAACACTTGTTTCTATTAAAACAGACTATTATAAAGCTTTAAGTTGTCAACTTTGTTC TAAATATCAACTAGCGCAAGTGAATACTGAAGATTTCTTAGTCAGTTTTTAGGGGATTTTCGGGGAGGGGAAATAGGTAATGTATGGAGC ATTTTCACTTCTAAATAGTTAGATACAGAAATTAAGTGCATTGTATCTTTTTCATAATGGTACTATTTAGAAGCCCAGTTAGTCTTACTG AGCTTATGCTTCACTCCTTTATGTTTAACCATGTGTCTACAAGAATAAGTTTGTTTTGGAAAGTTGAGCTATAGCTACAGCTCTAGCTAT CCAGCAGACTTTTCATTATGACTTACATGGCAGGAGCTCTAATTATGCTTTAAAAATCTGTTGTGGAGATTGCTTTAAATGCTCCCTGCC TGGTGTGGGGATGGGGTCCCCCTCTTTGTGAGGGCTGGAGCATGGCACGGCATGGATTAACACGGCAGAGGAACAAAGGTGTGCTCTGAG CTTCTTCATATTTCACCTTCACCCTCACCTGTGTTCTCTTCCCTCTCTCCCAATAAAAGGGCTCCCATTATAAATGCCATGTACTTCTCT TGGGAAAATAGACCCCCTTGCCTAGAGTAAGTTGTTAACTGAGGGCTTTAAACCTGGAGGCTCTTCCTGAAAGTATGTTCATGAATACCC CAAGCATCAAGGTCTAAATAATTTTCAGAAGATTAGAATTGGGTAGATATACTGTTGGATATAGCCATGGTAAATTTAACTGAGGAATTA AATCCTTGTTAATTTTGGTGTAAAGATTTGTATCCTGGCCTGTTTATTCAGGTGGGAGATGTTCTGTATAATTTTGAGGTATAGCTTGAA CTCCTAGAACAAGAGTTAAAAGTTCTTTTTAGCTCATTCACAATCACTTGTGTGTTAAGTGTTAATATCTGTCATGAACCTGCCAATTTG TACGTGTTAAGCAGCTGACAGATGATGCAAAAAAAACTAAGATACACTGGATTGTACTGGATGAGTCTGAAATTAGTGTTAAAGAAGTGT GGTGCCTGATGTTATAACAACACCTCATTCTTAACTGTGTGGCCAATGTCAGGTATACCAAAGCATTCCTCTGACTGCTTTTTGGGGCAG TGTTGACAGGAGTGAGGCATTTTGGAAACTCCAAGGGAAGCCTGTGTTTGAGGCCAGAGTTGGTACCTGCTTTAAGAAACTGTTACAGGG CTTGTTGGTAATGTTTTGAATCTTTGTAAAAATTGATATTCTGTATAACAGAGTGCCTCTCTGTTACTTTTGGCCTATGTTGTTAGAAAT AAGATGCTATCTTGCATAGTTAGAATTAAGCGTTTGTCCATCTCTAGATAACATTGAAAAGTTTGAGTGTTACAGGCTCTAAAGTGCAAT GGAGAACAATTGCTTAGGAGGAGGAAGGGGAAAGTATACTTATGTTGGTGTGTAAGCATGCATGGAGGCCTCGAGGCACCACAACAAGGC CAGGTTCTTAGCACATCCTCTGTTCTCTACCTGTCATGCTTCAAGAGTTCCAGCTGGCCTGCTGTGAAGCCAGTAGCTATTTTAAAAGTG CTGGCTGCAGGATTAAGCCAACCCTTTACAACTGGCCCAGACCGTAATGGCCATTTCTTCTTAAAAAAAAAAAAATTTTTTTTTTTTTTT TTTTTTTTTTTTTTTTTTTTTTTGGTGTGTGTGTGTGTGTGTGTGGGGAAGGGTGTGGGTGCTGGGATGGGGGTGAATTTCAAGTTGAAA CACGATGATGTGGTCTGCTTGCTCTCTGCTTAGACAGGCTTTAAGACCAATTGGATTTTCATGGAGGCCAGGACAGAGAGTGGTAATTGA TGGCTTGTGTACAGACAAGCATTTATCCAGGTTGCGTTATCCAAACTGATTTGTTAAAGCTCCAGGTCACGTTCTTACACTAAGAAAATC ATTCAGTAAACATGAGACAAGTTTAGGAAAAACTAATAAAATAAACCTGTCTTAAACTTGAA >16480_16480_1_CHIC2-HSPA4_CHIC2_chr4_54930348_ENST00000263921_HSPA4_chr5_132412345_ENST00000304858_length(amino acids)=618AA_BP=1 MATAFDTTLGGRKFDEVLVNHFCEEFGKKYKLDIKSKIRALLRLSQECEKLKKLMSANASDLPLSIECFMNDVDVSGTMNRGKFLEMCND LLARVEPPLRSVLEQTKLKKEDIYAVEIVGGATRIPAVKEKISKFFGKELSTTLNADEAVTRGCALQCAILSPAFKVREFSITDVVPYPI SLRWNSPAEEGSSDCEVFSKNHAAPFSKVLTFYRKEPFTLEAYYSSPQDLPYPDPAIAQFSVQKVTPQSDGSSSKVKVKVRVNVHGIFSV SSASLVEVHKSEENEEPMETDQNAKEEEKMQVDQEEPHVEEQQQQTPAENKAESEEMETSQAGSKDKKMDQPPQAKKAKVKTSTVDLPIE NQLLWQIDREMLNLYIENEGKMIMQDKLEKERNDAKNAVEEYVYEMRDKLSGEYEKFVSEDDRNSFTLKLEDTENWLYEDGEDQPKQVYV DKLAELKNLGQPIKIRFQESEERPKLFEELGKQIQQYMKIISSFKNKEDQYDHLDAADMTKVEKSTNEAMEWMNNKLNLQNKQSLTMDPV VKSKEIEAKIKELTSTCSPIISKPKPKVEPPKEEQKNAEQNGPVDGQGDNPGPQAAEQGTDTAVPSDSDKKLPEMDID -------------------------------------------------------------- >16480_16480_2_CHIC2-HSPA4_CHIC2_chr4_54930348_ENST00000512964_HSPA4_chr5_132412345_ENST00000304858_length(transcript)=3992nt_BP=119nt ATGGCGGATTTCGACGAAATCTATGAGGAAGAGGAGGACGAGGAGCGGGCCCTGGAGGAGCAGCTGCTCAAGTACTCGCCGGACCCGGTG GTCGTCCGCGGCTCCGGTCACGTCACCGTGTTCTGGCCACTGCATTTGACACGACATTGGGAGGTAGAAAATTTGATGAAGTGTTAGTAA ATCACTTCTGTGAAGAATTTGGGAAGAAATACAAGCTAGACATTAAGTCCAAAATCCGTGCATTATTACGACTCTCTCAGGAGTGTGAGA AACTCAAGAAATTGATGAGTGCAAATGCTTCAGATCTCCCTTTGAGCATTGAATGTTTTATGAATGATGTTGATGTATCTGGAACTATGA ATAGAGGCAAATTTCTGGAGATGTGCAATGATCTCTTAGCTAGAGTGGAGCCACCACTTCGTAGTGTTTTGGAACAAACCAAGTTAAAGA AAGAAGATATTTATGCAGTGGAGATAGTTGGTGGTGCTACACGAATCCCTGCGGTAAAAGAGAAGATCAGCAAATTTTTCGGTAAAGAAC TTAGTACAACATTAAATGCTGATGAAGCTGTCACTCGAGGCTGTGCATTGCAGTGTGCCATCTTATCGCCTGCTTTCAAAGTCAGAGAAT TTTCTATCACTGATGTAGTACCATATCCAATATCTCTGAGATGGAATTCTCCAGCTGAAGAAGGGTCAAGTGACTGTGAAGTCTTTTCCA AAAATCATGCTGCTCCTTTCTCTAAAGTTCTTACATTTTATAGAAAGGAACCTTTCACTCTTGAGGCCTACTACAGCTCTCCTCAGGATT TGCCCTATCCAGATCCTGCTATAGCTCAGTTTTCAGTTCAGAAAGTCACTCCTCAGTCTGATGGCTCCAGTTCAAAAGTGAAAGTCAAAG TTCGAGTAAATGTCCATGGCATTTTCAGTGTGTCCAGTGCATCTTTAGTGGAGGTTCACAAGTCTGAGGAAAATGAGGAGCCAATGGAAA CAGATCAGAATGCAAAGGAGGAAGAGAAGATGCAAGTGGACCAGGAGGAACCACATGTTGAAGAGCAACAGCAGCAGACACCAGCAGAAA ATAAGGCAGAGTCTGAAGAAATGGAGACCTCTCAAGCTGGATCCAAGGATAAAAAGATGGACCAACCACCCCAAGCCAAGAAGGCAAAAG TGAAGACCAGTACTGTGGACCTGCCAATCGAGAATCAGCTATTATGGCAGATAGACAGAGAGATGCTCAACTTGTACATTGAAAATGAGG GTAAGATGATCATGCAGGATAAACTGGAGAAGGAGCGGAATGATGCTAAGAACGCAGTGGAGGAATATGTGTATGAAATGAGAGACAAGC TTAGTGGTGAATATGAGAAGTTTGTGAGTGAAGATGATCGTAACAGTTTTACTTTGAAACTGGAAGATACTGAAAATTGGTTGTATGAGG ATGGAGAAGACCAGCCAAAGCAAGTTTATGTTGATAAGTTGGCTGAATTAAAAAATCTAGGTCAACCTATTAAGATACGTTTCCAGGAAT CTGAAGAACGACCAAAATTATTTGAAGAACTAGGGAAACAGATCCAACAGTATATGAAAATAATCAGCTCTTTCAAAAACAAGGAGGACC AGTATGATCATTTGGATGCTGCTGACATGACAAAGGTAGAAAAAAGCACAAATGAAGCAATGGAGTGGATGAATAACAAGCTAAATCTGC AGAACAAGCAGAGTTTGACCATGGATCCAGTTGTCAAGTCAAAAGAGATTGAAGCTAAAATTAAGGAGCTGACAAGTACTTGTAGCCCTA TAATTTCAAAGCCCAAACCCAAAGTGGAACCTCCAAAAGAGGAACAAAAAAATGCAGAGCAGAATGGACCAGTGGATGGACAAGGAGACA ACCCAGGCCCCCAGGCTGCTGAGCAGGGTACAGACACAGCTGTGCCTTCGGATTCAGACAAGAAGCTTCCTGAAATGGACATTGATTGAT TCCAACACTTGTTTCTATTAAAACAGACTATTATAAAGCTTTAAGTTGTCAACTTTGTTCTAAATATCAACTAGCGCAAGTGAATACTGA AGATTTCTTAGTCAGTTTTTAGGGGATTTTCGGGGAGGGGAAATAGGTAATGTATGGAGCATTTTCACTTCTAAATAGTTAGATACAGAA ATTAAGTGCATTGTATCTTTTTCATAATGGTACTATTTAGAAGCCCAGTTAGTCTTACTGAGCTTATGCTTCACTCCTTTATGTTTAACC ATGTGTCTACAAGAATAAGTTTGTTTTGGAAAGTTGAGCTATAGCTACAGCTCTAGCTATCCAGCAGACTTTTCATTATGACTTACATGG CAGGAGCTCTAATTATGCTTTAAAAATCTGTTGTGGAGATTGCTTTAAATGCTCCCTGCCTGGTGTGGGGATGGGGTCCCCCTCTTTGTG AGGGCTGGAGCATGGCACGGCATGGATTAACACGGCAGAGGAACAAAGGTGTGCTCTGAGCTTCTTCATATTTCACCTTCACCCTCACCT GTGTTCTCTTCCCTCTCTCCCAATAAAAGGGCTCCCATTATAAATGCCATGTACTTCTCTTGGGAAAATAGACCCCCTTGCCTAGAGTAA GTTGTTAACTGAGGGCTTTAAACCTGGAGGCTCTTCCTGAAAGTATGTTCATGAATACCCCAAGCATCAAGGTCTAAATAATTTTCAGAA GATTAGAATTGGGTAGATATACTGTTGGATATAGCCATGGTAAATTTAACTGAGGAATTAAATCCTTGTTAATTTTGGTGTAAAGATTTG TATCCTGGCCTGTTTATTCAGGTGGGAGATGTTCTGTATAATTTTGAGGTATAGCTTGAACTCCTAGAACAAGAGTTAAAAGTTCTTTTT AGCTCATTCACAATCACTTGTGTGTTAAGTGTTAATATCTGTCATGAACCTGCCAATTTGTACGTGTTAAGCAGCTGACAGATGATGCAA AAAAAACTAAGATACACTGGATTGTACTGGATGAGTCTGAAATTAGTGTTAAAGAAGTGTGGTGCCTGATGTTATAACAACACCTCATTC TTAACTGTGTGGCCAATGTCAGGTATACCAAAGCATTCCTCTGACTGCTTTTTGGGGCAGTGTTGACAGGAGTGAGGCATTTTGGAAACT CCAAGGGAAGCCTGTGTTTGAGGCCAGAGTTGGTACCTGCTTTAAGAAACTGTTACAGGGCTTGTTGGTAATGTTTTGAATCTTTGTAAA AATTGATATTCTGTATAACAGAGTGCCTCTCTGTTACTTTTGGCCTATGTTGTTAGAAATAAGATGCTATCTTGCATAGTTAGAATTAAG CGTTTGTCCATCTCTAGATAACATTGAAAAGTTTGAGTGTTACAGGCTCTAAAGTGCAATGGAGAACAATTGCTTAGGAGGAGGAAGGGG AAAGTATACTTATGTTGGTGTGTAAGCATGCATGGAGGCCTCGAGGCACCACAACAAGGCCAGGTTCTTAGCACATCCTCTGTTCTCTAC CTGTCATGCTTCAAGAGTTCCAGCTGGCCTGCTGTGAAGCCAGTAGCTATTTTAAAAGTGCTGGCTGCAGGATTAAGCCAACCCTTTACA ACTGGCCCAGACCGTAATGGCCATTTCTTCTTAAAAAAAAAAAAATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTGGTGTGT GTGTGTGTGTGTGTGGGGAAGGGTGTGGGTGCTGGGATGGGGGTGAATTTCAAGTTGAAACACGATGATGTGGTCTGCTTGCTCTCTGCT TAGACAGGCTTTAAGACCAATTGGATTTTCATGGAGGCCAGGACAGAGAGTGGTAATTGATGGCTTGTGTACAGACAAGCATTTATCCAG GTTGCGTTATCCAAACTGATTTGTTAAAGCTCCAGGTCACGTTCTTACACTAAGAAAATCATTCAGTAAACATGAGACAAGTTTAGGAAA AACTAATAAAATAAACCTGTCTTAAACTTGAA >16480_16480_2_CHIC2-HSPA4_CHIC2_chr4_54930348_ENST00000512964_HSPA4_chr5_132412345_ENST00000304858_length(amino acids)=618AA_BP=1 MATAFDTTLGGRKFDEVLVNHFCEEFGKKYKLDIKSKIRALLRLSQECEKLKKLMSANASDLPLSIECFMNDVDVSGTMNRGKFLEMCND LLARVEPPLRSVLEQTKLKKEDIYAVEIVGGATRIPAVKEKISKFFGKELSTTLNADEAVTRGCALQCAILSPAFKVREFSITDVVPYPI SLRWNSPAEEGSSDCEVFSKNHAAPFSKVLTFYRKEPFTLEAYYSSPQDLPYPDPAIAQFSVQKVTPQSDGSSSKVKVKVRVNVHGIFSV SSASLVEVHKSEENEEPMETDQNAKEEEKMQVDQEEPHVEEQQQQTPAENKAESEEMETSQAGSKDKKMDQPPQAKKAKVKTSTVDLPIE NQLLWQIDREMLNLYIENEGKMIMQDKLEKERNDAKNAVEEYVYEMRDKLSGEYEKFVSEDDRNSFTLKLEDTENWLYEDGEDQPKQVYV DKLAELKNLGQPIKIRFQESEERPKLFEELGKQIQQYMKIISSFKNKEDQYDHLDAADMTKVEKSTNEAMEWMNNKLNLQNKQSLTMDPV VKSKEIEAKIKELTSTCSPIISKPKPKVEPPKEEQKNAEQNGPVDGQGDNPGPQAAEQGTDTAVPSDSDKKLPEMDID -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for CHIC2-HSPA4 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for CHIC2-HSPA4 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for CHIC2-HSPA4 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
| Tgene | C0005586 | Bipolar Disorder | 2 | PSYGENET | |
| Tgene | C0001969 | Alcoholic Intoxication | 1 | PSYGENET | |
| Tgene | C0238288 | Muscular Dystrophy, Facioscapulohumeral | 1 | CTD_human | |
| Tgene | C0338831 | Manic | 1 | PSYGENET | |
| Tgene | C0525045 | Mood Disorders | 1 | PSYGENET | |
| Tgene | C0564408 | Manic mood | 1 | PSYGENET | |
| Tgene | C3496069 | cocaine use | 1 | PSYGENET |
