
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:LMBR1-CHMP1A (FusionGDB2 ID:HG64327TG5119) |
Fusion Gene Summary for LMBR1-CHMP1A |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: LMBR1-CHMP1A | Fusion gene ID: hg64327tg5119 | Hgene | Tgene | Gene symbol | LMBR1 | CHMP1A | Gene ID | 64327 | 5119 |
| Gene name | limb development membrane protein 1 | charged multivesicular body protein 1A | |
| Synonyms | ACHP|C7orf2|DIF14|LSS|PPD2|THYP|TPT|ZRS | CHMP1|PCH8|PCOLN3|PRSM1|VPS46-1|VPS46A | |
| Cytomap | ('LMBR1')('CHMP1A') 7q36.3 | 16q24.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | limb region 1 protein homologdifferentiation-related gene 14 proteinlimb region 1 homolog | charged multivesicular body protein 1acharged multivesicular body protein 1/chromatin modifying protein 1chromatin modifying protein 1Aprocollagen (type III) N-endopeptidaseprotease, metallo, 1, 33kDvacuolar protein sorting-associated protein 46-1 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000353442, ENST00000354505, ENST00000359422, ENST00000540390, ENST00000430825, | ||
| Fusion gene scores | * DoF score | 14 X 11 X 12=1848 | 45 X 11 X 21=10395 |
| # samples | 20 | 48 | |
| ** MAII score | log2(20/1848*10)=-3.20789285164133 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(48/10395*10)=-4.43671154213721 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: LMBR1 [Title/Abstract] AND CHMP1A [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | LMBR1(156516806)-CHMP1A(89713739), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | LMBR1-CHMP1A seems lost the major protein functional domain in Hgene partner, which is a CGC by not retaining the major functional domain in the partially deleted in-frame ORF. LMBR1-CHMP1A seems lost the major protein functional domain in Hgene partner, which is a essential gene by not retaining the major functional domain in the partially deleted in-frame ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | CHMP1A | GO:0007076 | mitotic chromosome condensation | 11559747 |
| Tgene | CHMP1A | GO:0016192 | vesicle-mediated transport | 11559748 |
| Tgene | CHMP1A | GO:0016458 | gene silencing | 11559747 |
| Tgene | CHMP1A | GO:0045892 | negative regulation of transcription, DNA-templated | 11559747 |
 Fusion gene breakpoints across LMBR1 (5'-gene) Fusion gene breakpoints across LMBR1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
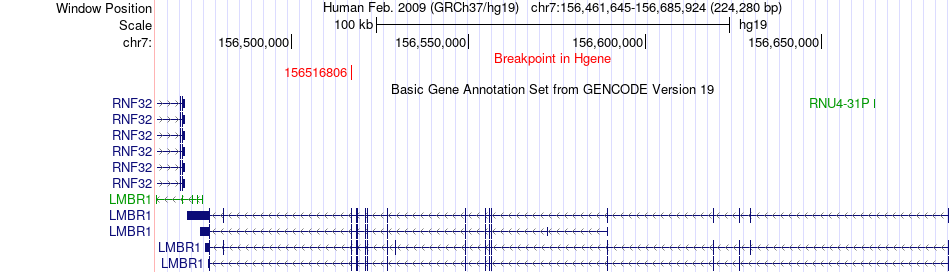 |
 Fusion gene breakpoints across CHMP1A (3'-gene) Fusion gene breakpoints across CHMP1A (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
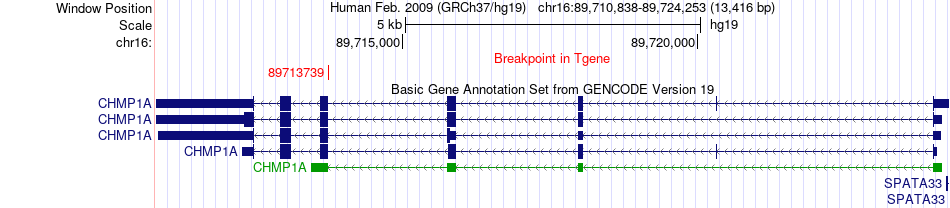 |
 Fusion gene information Fusion gene information * All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | ACC | TCGA-OR-A5LB-01A | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
Top |
Fusion Gene ORF analysis for LMBR1-CHMP1A |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000353442 | ENST00000547614 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| 5CDS-5UTR | ENST00000354505 | ENST00000547614 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| 5CDS-5UTR | ENST00000359422 | ENST00000547614 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| 5CDS-5UTR | ENST00000540390 | ENST00000547614 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| Frame-shift | ENST00000353442 | ENST00000397901 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| Frame-shift | ENST00000353442 | ENST00000535997 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| Frame-shift | ENST00000353442 | ENST00000550102 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| Frame-shift | ENST00000354505 | ENST00000397901 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| Frame-shift | ENST00000354505 | ENST00000535997 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| Frame-shift | ENST00000354505 | ENST00000550102 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| Frame-shift | ENST00000359422 | ENST00000253475 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| Frame-shift | ENST00000540390 | ENST00000397901 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| Frame-shift | ENST00000540390 | ENST00000535997 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| Frame-shift | ENST00000540390 | ENST00000550102 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| In-frame | ENST00000353442 | ENST00000253475 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| In-frame | ENST00000354505 | ENST00000253475 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| In-frame | ENST00000359422 | ENST00000397901 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| In-frame | ENST00000359422 | ENST00000535997 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| In-frame | ENST00000359422 | ENST00000550102 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| In-frame | ENST00000540390 | ENST00000253475 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| intron-3CDS | ENST00000430825 | ENST00000253475 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| intron-3CDS | ENST00000430825 | ENST00000397901 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| intron-3CDS | ENST00000430825 | ENST00000535997 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| intron-3CDS | ENST00000430825 | ENST00000550102 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
| intron-5UTR | ENST00000430825 | ENST00000547614 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000353442 | LMBR1 | chr7 | 156516806 | - | ENST00000253475 | CHMP1A | chr16 | 89713739 | - | 3429 | 1462 | 237 | 1952 | 571 |
| ENST00000359422 | LMBR1 | chr7 | 156516806 | - | ENST00000397901 | CHMP1A | chr16 | 89713739 | - | 2959 | 985 | 210 | 1475 | 421 |
| ENST00000359422 | LMBR1 | chr7 | 156516806 | - | ENST00000535997 | CHMP1A | chr16 | 89713739 | - | 2927 | 985 | 210 | 1475 | 421 |
| ENST00000359422 | LMBR1 | chr7 | 156516806 | - | ENST00000550102 | CHMP1A | chr16 | 89713739 | - | 1501 | 985 | 210 | 1475 | 421 |
| ENST00000354505 | LMBR1 | chr7 | 156516806 | - | ENST00000253475 | CHMP1A | chr16 | 89713739 | - | 3469 | 1502 | 154 | 1992 | 612 |
| ENST00000540390 | LMBR1 | chr7 | 156516806 | - | ENST00000253475 | CHMP1A | chr16 | 89713739 | - | 3305 | 1338 | 104 | 1828 | 574 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000353442 | ENST00000253475 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - | 0.00466823 | 0.9953317 |
| ENST00000359422 | ENST00000397901 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - | 0.005775086 | 0.99422497 |
| ENST00000359422 | ENST00000535997 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - | 0.006270556 | 0.9937295 |
| ENST00000359422 | ENST00000550102 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - | 0.007768857 | 0.9922312 |
| ENST00000354505 | ENST00000253475 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - | 0.003293607 | 0.9967064 |
| ENST00000540390 | ENST00000253475 | LMBR1 | chr7 | 156516806 | - | CHMP1A | chr16 | 89713739 | - | 0.007331227 | 0.99266875 |
Top |
Fusion Genomic Features for LMBR1-CHMP1A |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
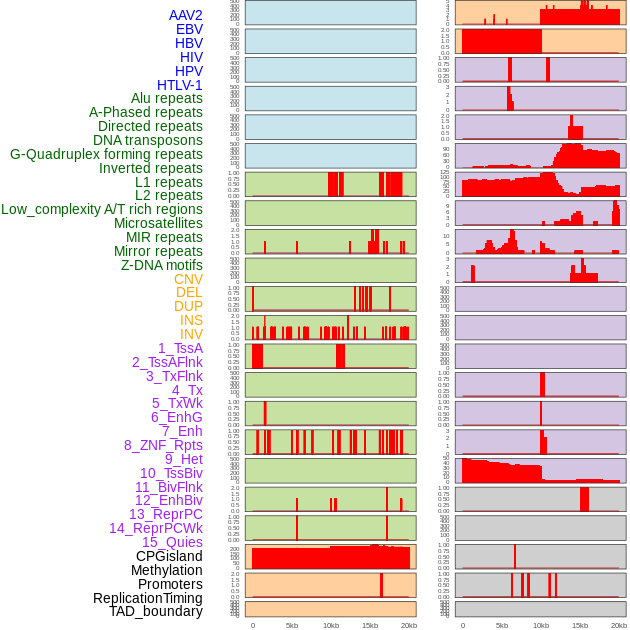 |
Top |
Fusion Protein Features for LMBR1-CHMP1A |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr7:156516806/chr16:89713739) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 250_287 | 408 | 491.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 250_287 | 449 | 532.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 132_151 | 408 | 491.0 | Topological domain | Cytoplasmic |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 173_187 | 408 | 491.0 | Topological domain | Extracellular |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 1_19 | 408 | 491.0 | Topological domain | Extracellular |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 209_291 | 408 | 491.0 | Topological domain | Cytoplasmic |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 313_339 | 408 | 491.0 | Topological domain | Extracellular |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 361_383 | 408 | 491.0 | Topological domain | Cytoplasmic |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 41_62 | 408 | 491.0 | Topological domain | Cytoplasmic |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 84_110 | 408 | 491.0 | Topological domain | Extracellular |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 132_151 | 449 | 532.0 | Topological domain | Cytoplasmic |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 173_187 | 449 | 532.0 | Topological domain | Extracellular |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 1_19 | 449 | 532.0 | Topological domain | Extracellular |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 209_291 | 449 | 532.0 | Topological domain | Cytoplasmic |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 313_339 | 449 | 532.0 | Topological domain | Extracellular |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 361_383 | 449 | 532.0 | Topological domain | Cytoplasmic |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 405_426 | 449 | 532.0 | Topological domain | Extracellular |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 41_62 | 449 | 532.0 | Topological domain | Cytoplasmic |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 84_110 | 449 | 532.0 | Topological domain | Extracellular |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 111_131 | 408 | 491.0 | Transmembrane | Helical |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 152_172 | 408 | 491.0 | Transmembrane | Helical |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 188_208 | 408 | 491.0 | Transmembrane | Helical |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 20_40 | 408 | 491.0 | Transmembrane | Helical |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 292_312 | 408 | 491.0 | Transmembrane | Helical |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 340_360 | 408 | 491.0 | Transmembrane | Helical |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 384_404 | 408 | 491.0 | Transmembrane | Helical |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 63_83 | 408 | 491.0 | Transmembrane | Helical |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 111_131 | 449 | 532.0 | Transmembrane | Helical |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 152_172 | 449 | 532.0 | Transmembrane | Helical |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 188_208 | 449 | 532.0 | Transmembrane | Helical |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 20_40 | 449 | 532.0 | Transmembrane | Helical |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 292_312 | 449 | 532.0 | Transmembrane | Helical |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 340_360 | 449 | 532.0 | Transmembrane | Helical |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 384_404 | 449 | 532.0 | Transmembrane | Helical |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 427_447 | 449 | 532.0 | Transmembrane | Helical |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 63_83 | 449 | 532.0 | Transmembrane | Helical |
| Tgene | CHMP1A | chr7:156516806 | chr16:89713739 | ENST00000397901 | 3 | 7 | 102_124 | 84 | 197.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | CHMP1A | chr7:156516806 | chr16:89713739 | ENST00000397901 | 3 | 7 | 185_195 | 84 | 197.0 | Motif | Note=MIT-interacting motif |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 405_426 | 408 | 491.0 | Topological domain | Extracellular |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 448_490 | 408 | 491.0 | Topological domain | Cytoplasmic |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000354505 | - | 16 | 18 | 448_490 | 449 | 532.0 | Topological domain | Cytoplasmic |
| Hgene | LMBR1 | chr7:156516806 | chr16:89713739 | ENST00000353442 | - | 15 | 17 | 427_447 | 408 | 491.0 | Transmembrane | Helical |
| Tgene | CHMP1A | chr7:156516806 | chr16:89713739 | ENST00000397901 | 3 | 7 | 5_47 | 84 | 197.0 | Coiled coil | Ontology_term=ECO:0000255 |
Top |
Fusion Gene Sequence for LMBR1-CHMP1A |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >45733_45733_1_LMBR1-CHMP1A_LMBR1_chr7_156516806_ENST00000353442_CHMP1A_chr16_89713739_ENST00000253475_length(transcript)=3429nt_BP=1462nt GCCCCGCCCCCGCCCCGCCCCCGACTGCGCGGCGCCCGCCCTCCCTTAGTCAGAGCTGCTCGTCTGAGGCTGCTGAGGCGACGGCCGGTG TCGTGGTCGCGGTACCTGTTCCAACACGGCTCGCGGGCCCGTGCCGGCTCCGGTCCCCGGCGCGGCTGTCCGAGCCCCTGCGGCGGGCGG ACGATGGTGTGGCGGAGCACGCGGACGCGGGCGGCGCGGCGGCGGGCATGAAGGAGGATGGAAGGGCAGGACGAGGTGTCGGCGCGGGAG CAGCACTTCCACAGCCAAGTGCGGGAGTCCACGATATGTTTCCTTCTTTTTGCCATTCTCTACGTTGTTTCCTACTTCATCATCACAAGA TACAAGAGAAAATCAGATGAACAAGAAGATGAAGATGCCATCGTCAACAGGATTTCGTTGTTTTTGAGCACGTTCACTCTCGCAGTGTCA GCTGGGGCTGTTTTGCTTTTACCCTTCTCAATCATCAGCAATGAAATCCTGCTTTCTTTTCCTCAGAACTACTATATTCAGTGGCTAAAT GGCTCCCTGATTCATGGTTTGTGGAATCTTGCTTCCCTTTTTTCCAACCTTTGTTTATTTGTATTGATGCCCTTTGCCTTTTTCTTTCTG GAATCAGAAGGCTTTGCTGGCCTGAAAAAGGGAATCCGAGCCCGCATTTTAGAGACTTTGGTCATGCTTCTTCTTCTTGCGTTACTCATT CTTGGGATAGTGTGGGTAGCTTCAGCACTCATTGACAACGATGCCGCAAGCATGGAATCTTTATATGATCTCTGGGAGTTCTATCTACCC TATTTATATTCCTGTATATCATTGATGGGATGTTTGTTACTTCTCTTGTGTACACCAGTTGGCCTTTCTCGTATGTTCACAGTGATGGGT CAGTTGCTAGTGAAGCCAACAATTCTTGAAGACCTGGATGAACAAATTTATATCATTACCTTAGAGGAAGAAGCACTCCAGAGACGACTA AATGGGCTGTCTTCATCGGTGGAATACAACATAATGGAGTTGGAACAAGAACTTGAAAATGTAAAGACTCTTAAGACAAAATTAGAGAGG CGAAAAAAGGCTTCAGCATGGGAAAGAAATTTGGTGTATCCCGCTGTTATGGTTCTCCTTCTTATTGAGACATCCATCTCGGTCCTCTTG GTGGCTTGTAATATTCTTTGCCTATTGGTTGATGAAACAGCAATGCCAAAAGGAACAAGGGGGCCTGGAATAGGAAATGCCTCTCTTTCT ACGTTTGGTTTTGTGGGAGCTGCGCTTGAAATCATTTTGATTTTCTATCTTATGGTGTCCTCTGTTGTCGGCTTCTATAGCCTTCGATTT TTTGGAAACTTTACTCCCAAGAAAGATGACACAACTATGACAAAGATCATTGGAAATTGTGTGTCCATCTTGGTTTTGAGCTCTGCTCTG CCTGTGATGTCGAGAACACTGGGTGACCAAGAATATGGCCCAGGTGACCAAAGCCCTGGACAAGGCCCTGAGCACCATGGACCTGCAGAA GGTCTCCTCAGTGATGGACAGGTTCGAGCAGCAGGTGCAGAACCTGGACGTCCATACATCGGTGATGGAGGACTCCATGAGCTCGGCCAC CACCCTGACCACGCCGCAGGAGCAGGTGGACAGCCTCATCATGCAGATCGCCGAGGAGAATGGCCTGGAGGTGCTGGACCAGCTCAGCCA GCTGCCCGAGGGCGCCTCTGCCGTGGGCGAGAGCTCTGTGCGCAGCCAGGAGGACCAGCTGTCACGGAGGTTGGCCGCCTTGAGGAACTA GCCGTGCCCCGCCGGTGTGCACCGCCTCTGCCCCGTGATGTGCTGGAAGGCTCCTGTCCTCTCCCCACCGCGTCTTGCCTTTGTGCTGAC CCCGCGGGGCTGCGGCCGGCAGCCACTCTGCGTCTCTCACCTGCCAGGCCTGCGTGGCCTTAGGGTTGTTCCTGTTCTTTTAGGTTGGGC GGTGGGTCTGTGTCCTGGTGTTGAGTTTCTGCAAATTTCTGGGGGTGATTTCTGTGACTCTGGGCCCACAGCGGGGAGGCCAAGAGGGGC CCTGTGGACTTTCACCCAGCACTGTGGGGGCCTTCAGACTCTGGGGCAGCAGACATGCTGCTTCCCATCAGCCAGAGGGGGTCAGGGCTG CCCTGTTGCCAAACAACTCCCTGAGGCCTCTCCGCACCACCTCAGCGGGCAGGAGGTCCCACCATGTGGACAGACATAGCCCAAGGAGGC ACCACAGGTCTATGTGTGCTGGGGGATGTCAGGTGCCACCCAACGCTGTCCTGGTGGTATTTACAATGACATCCTCCTCCTCCATCACTC CAGGGGTGGTGTCTCGGCCGCCCCTACCAGCTGGCTGAGCCCCCTGGCCTCCTGCGCTCCCTCACTTCCCTCAGTTCCCAAAGCTGCCCA GTCCATGGGGACAGAACCGTCACTCAGATCCACATTCAAGTGTGCCCACCCTGCAGTCTTCATCCTCACTCAGCTGCTGCCTCTGGAGGT GCCTTTGGCCACATGTGCTGTGCTGTTTGTCTCCTCGACAGGGAGCCTGTCCACCAGCAGGCTGCGGTCCCAGCGGGTGCGTCTGCAGCT CCTCCCCTTGGGCAGCCTGGTTCTCCCGGAGGACCTTTCCTTGGGGCCCTGCTTCATGACGATGCTGCCTGTGTCACCCTCTACCATCTG TAAACAACTGGGTGCCTTCCCCGACCACACCCCAATGCCTTCCCAGCTTGGAAGCCAAGGCAGCTGATGAAGGGAGCTCAGGAGAGCCGT CTTCAGCTGGGCTGGGGTTGGGGCTGCTGTGAGGAAAACCTGCCATTGTGGCCCTGGAGAGTCACCAGCAGCTCTTGGGAAGGACTTGCT GGGAGGCTGAGAGAGGCTTTGGGCACAGCCTGCTGTCTTTTCCATTTCCTAAAGTTTACTTCATTGTCTTGAGGCTTCCAGGTTTTGTTT TTGTTTTTGCCAAAGTAGAAAAGGCAGGTGGTGGGCGGCTGGCAGGGAGTGCGGGTCCCCGCCCCTCTTCAGTCCTGCCCTCCCCTCCTC AGTCCTGCCCACCCCGTGCAGCCCATGCTGAGGCTGCAGTGGTGTCGTGGGTGTTACGTGCAGGAACGTGGAGACCCTGACGTGGGCTCA CTGCGTTTGGTTTTCTTTTCAGAACTTGGGAGCCCCCAGGGAGGGGCTAGTGTTGGTAGGTCCTAGACGTGGTTCCCTCCAGCCTCCCCA AAATCAACCCTGGTGTTGAGAGAACGTCCTTCTGTCCATCGTGGGTAACAGCCTTGGGGAGGGTGCAGAGCTCTGCAGAGCCATGGGCCA GGTGGGGCTGCCTCAGTCCTGTCCCCTTGGGCACTGAGGAGAGGGGCCCATTCACCTTTCTCCTAGAATGCTGTTGTAAATAAACAAATG GATCCCTGG >45733_45733_1_LMBR1-CHMP1A_LMBR1_chr7_156516806_ENST00000353442_CHMP1A_chr16_89713739_ENST00000253475_length(amino acids)=571AA_BP=408 MEGQDEVSAREQHFHSQVRESTICFLLFAILYVVSYFIITRYKRKSDEQEDEDAIVNRISLFLSTFTLAVSAGAVLLLPFSIISNEILLS FPQNYYIQWLNGSLIHGLWNLASLFSNLCLFVLMPFAFFFLESEGFAGLKKGIRARILETLVMLLLLALLILGIVWVASALIDNDAASME SLYDLWEFYLPYLYSCISLMGCLLLLLCTPVGLSRMFTVMGQLLVKPTILEDLDEQIYIITLEEEALQRRLNGLSSSVEYNIMELEQELE NVKTLKTKLERRKKASAWERNLVYPAVMVLLLIETSISVLLVACNILCLLVDETAMPKGTRGPGIGNASLSTFGFVGAALEIILIFYLMV SSVVGFYSLRFFGNFTPKKDDTTMTKIIGNCVSILVLSSALPVMSRTLGDQEYGPGDQSPGQGPEHHGPAEGLLSDGQVRAAGAEPGRPY IGDGGLHELGHHPDHAAGAGGQPHHADRRGEWPGGAGPAQPAARGRLCRGRELCAQPGGPAVTEVGRLEELAVPRRCAPPLPRDVLEGSC PLPTASCLCADPAGLRPAATLRLSPARPAWP -------------------------------------------------------------- >45733_45733_2_LMBR1-CHMP1A_LMBR1_chr7_156516806_ENST00000354505_CHMP1A_chr16_89713739_ENST00000253475_length(transcript)=3469nt_BP=1502nt GCCGGTGTCGTGGTCGCGGTACCTGTTCCAACACGGCTCGCGGGCCCGTGCCGGCTCCGGTCCCCGGCGCGGCTGTCCGAGCCCCTGCGG CGGGCGGACGATGGTGTGGCGGAGCACGCGGACGCGGGCGGCGCGGCGGCGGGCATGAAGGAGGATGGAAGGGCAGGACGAGGTGTCGGC GCGGGAGCAGCACTTCCACAGCCAAGTGCGGGAGTCCACGATATGTTTCCTTCTTTTTGCCATTCTCTACGTTGTTTCCTACTTCATCAT CACAAGATACAAGAGAAAATCAGATGAACAAGAAGATGAAGATGCCATCGTCAACAGGATTTCGTTGTTTTTGAGCACGTTCACTCTCGC AGTGTCAGCTGGGGCTGTTTTGCTTTTACCCTTCTCAATCATCAGCAATGAAATCCTGCTTTCTTTTCCTCAGAACTACTATATTCAGTG GCTAAATGGCTCCCTGATTCATGGTTTGTGGAATCTTGCTTCCCTTTTTTCCAACCTTTGTTTATTTGTATTGATGCCCTTTGCCTTTTT CTTTCTGGAATCAGAAGGCTTTGCTGGCCTGAAAAAGGGAATCCGAGCCCGCATTTTAGAGACTTTGGTCATGCTTCTTCTTCTTGCGTT ACTCATTCTTGGGATAGTGTGGGTAGCTTCAGCACTCATTGACAACGATGCCGCAAGCATGGAATCTTTATATGATCTCTGGGAGTTCTA TCTACCCTATTTATATTCCTGTATATCATTGATGGGATGTTTGTTACTTCTCTTGTGTACACCAGTTGGCCTTTCTCGTATGTTCACAGT GATGGGTCAGTTGCTAGTGAAGCCAACAATTCTTGAAGACCTGGATGAACAAATTTATATCATTACCTTAGAGGAAGAAGCACTCCAGAG ACGACTAAATGTGGGAATGCTTTGTGCGGGAAATCCAGAAGTTGCAACAGGAAGACAAGTGCCAGAAGAACAAGCTGGACAGCTACTTTT GGGAAAATGGATTCAGGAGGGAGGAGACATGATAGCAAACCCACGGCTGTCTTCATCGGTGGAATACAACATAATGGAGTTGGAACAAGA ACTTGAAAATGTAAAGACTCTTAAGACAAAATTAGAGAGGCGAAAAAAGGCTTCAGCATGGGAAAGAAATTTGGTGTATCCCGCTGTTAT GGTTCTCCTTCTTATTGAGACATCCATCTCGGTCCTCTTGGTGGCTTGTAATATTCTTTGCCTATTGGTTGATGAAACAGCAATGCCAAA AGGAACAAGGGGGCCTGGAATAGGAAATGCCTCTCTTTCTACGTTTGGTTTTGTGGGAGCTGCGCTTGAAATCATTTTGATTTTCTATCT TATGGTGTCCTCTGTTGTCGGCTTCTATAGCCTTCGATTTTTTGGAAACTTTACTCCCAAGAAAGATGACACAACTATGACAAAGATCAT TGGAAATTGTGTGTCCATCTTGGTTTTGAGCTCTGCTCTGCCTGTGATGTCGAGAACACTGGGTGACCAAGAATATGGCCCAGGTGACCA AAGCCCTGGACAAGGCCCTGAGCACCATGGACCTGCAGAAGGTCTCCTCAGTGATGGACAGGTTCGAGCAGCAGGTGCAGAACCTGGACG TCCATACATCGGTGATGGAGGACTCCATGAGCTCGGCCACCACCCTGACCACGCCGCAGGAGCAGGTGGACAGCCTCATCATGCAGATCG CCGAGGAGAATGGCCTGGAGGTGCTGGACCAGCTCAGCCAGCTGCCCGAGGGCGCCTCTGCCGTGGGCGAGAGCTCTGTGCGCAGCCAGG AGGACCAGCTGTCACGGAGGTTGGCCGCCTTGAGGAACTAGCCGTGCCCCGCCGGTGTGCACCGCCTCTGCCCCGTGATGTGCTGGAAGG CTCCTGTCCTCTCCCCACCGCGTCTTGCCTTTGTGCTGACCCCGCGGGGCTGCGGCCGGCAGCCACTCTGCGTCTCTCACCTGCCAGGCC TGCGTGGCCTTAGGGTTGTTCCTGTTCTTTTAGGTTGGGCGGTGGGTCTGTGTCCTGGTGTTGAGTTTCTGCAAATTTCTGGGGGTGATT TCTGTGACTCTGGGCCCACAGCGGGGAGGCCAAGAGGGGCCCTGTGGACTTTCACCCAGCACTGTGGGGGCCTTCAGACTCTGGGGCAGC AGACATGCTGCTTCCCATCAGCCAGAGGGGGTCAGGGCTGCCCTGTTGCCAAACAACTCCCTGAGGCCTCTCCGCACCACCTCAGCGGGC AGGAGGTCCCACCATGTGGACAGACATAGCCCAAGGAGGCACCACAGGTCTATGTGTGCTGGGGGATGTCAGGTGCCACCCAACGCTGTC CTGGTGGTATTTACAATGACATCCTCCTCCTCCATCACTCCAGGGGTGGTGTCTCGGCCGCCCCTACCAGCTGGCTGAGCCCCCTGGCCT CCTGCGCTCCCTCACTTCCCTCAGTTCCCAAAGCTGCCCAGTCCATGGGGACAGAACCGTCACTCAGATCCACATTCAAGTGTGCCCACC CTGCAGTCTTCATCCTCACTCAGCTGCTGCCTCTGGAGGTGCCTTTGGCCACATGTGCTGTGCTGTTTGTCTCCTCGACAGGGAGCCTGT CCACCAGCAGGCTGCGGTCCCAGCGGGTGCGTCTGCAGCTCCTCCCCTTGGGCAGCCTGGTTCTCCCGGAGGACCTTTCCTTGGGGCCCT GCTTCATGACGATGCTGCCTGTGTCACCCTCTACCATCTGTAAACAACTGGGTGCCTTCCCCGACCACACCCCAATGCCTTCCCAGCTTG GAAGCCAAGGCAGCTGATGAAGGGAGCTCAGGAGAGCCGTCTTCAGCTGGGCTGGGGTTGGGGCTGCTGTGAGGAAAACCTGCCATTGTG GCCCTGGAGAGTCACCAGCAGCTCTTGGGAAGGACTTGCTGGGAGGCTGAGAGAGGCTTTGGGCACAGCCTGCTGTCTTTTCCATTTCCT AAAGTTTACTTCATTGTCTTGAGGCTTCCAGGTTTTGTTTTTGTTTTTGCCAAAGTAGAAAAGGCAGGTGGTGGGCGGCTGGCAGGGAGT GCGGGTCCCCGCCCCTCTTCAGTCCTGCCCTCCCCTCCTCAGTCCTGCCCACCCCGTGCAGCCCATGCTGAGGCTGCAGTGGTGTCGTGG GTGTTACGTGCAGGAACGTGGAGACCCTGACGTGGGCTCACTGCGTTTGGTTTTCTTTTCAGAACTTGGGAGCCCCCAGGGAGGGGCTAG TGTTGGTAGGTCCTAGACGTGGTTCCCTCCAGCCTCCCCAAAATCAACCCTGGTGTTGAGAGAACGTCCTTCTGTCCATCGTGGGTAACA GCCTTGGGGAGGGTGCAGAGCTCTGCAGAGCCATGGGCCAGGTGGGGCTGCCTCAGTCCTGTCCCCTTGGGCACTGAGGAGAGGGGCCCA TTCACCTTTCTCCTAGAATGCTGTTGTAAATAAACAAATGGATCCCTGG >45733_45733_2_LMBR1-CHMP1A_LMBR1_chr7_156516806_ENST00000354505_CHMP1A_chr16_89713739_ENST00000253475_length(amino acids)=612AA_BP=449 MEGQDEVSAREQHFHSQVRESTICFLLFAILYVVSYFIITRYKRKSDEQEDEDAIVNRISLFLSTFTLAVSAGAVLLLPFSIISNEILLS FPQNYYIQWLNGSLIHGLWNLASLFSNLCLFVLMPFAFFFLESEGFAGLKKGIRARILETLVMLLLLALLILGIVWVASALIDNDAASME SLYDLWEFYLPYLYSCISLMGCLLLLLCTPVGLSRMFTVMGQLLVKPTILEDLDEQIYIITLEEEALQRRLNVGMLCAGNPEVATGRQVP EEQAGQLLLGKWIQEGGDMIANPRLSSSVEYNIMELEQELENVKTLKTKLERRKKASAWERNLVYPAVMVLLLIETSISVLLVACNILCL LVDETAMPKGTRGPGIGNASLSTFGFVGAALEIILIFYLMVSSVVGFYSLRFFGNFTPKKDDTTMTKIIGNCVSILVLSSALPVMSRTLG DQEYGPGDQSPGQGPEHHGPAEGLLSDGQVRAAGAEPGRPYIGDGGLHELGHHPDHAAGAGGQPHHADRRGEWPGGAGPAQPAARGRLCR GRELCAQPGGPAVTEVGRLEELAVPRRCAPPLPRDVLEGSCPLPTASCLCADPAGLRPAATLRLSPARPAWP -------------------------------------------------------------- >45733_45733_3_LMBR1-CHMP1A_LMBR1_chr7_156516806_ENST00000359422_CHMP1A_chr16_89713739_ENST00000397901_length(transcript)=2959nt_BP=985nt GGTTTGTGGAATCTTGCTTCCCTTTTTTCCAACCTTTGTTTATTTGTATTGATGCCCTTTGCCTTTTTCTTTCTGGAATCAGAAGGCTTT GCTGGCCTGAAAAAGGAGGGCAAGATGTCCAGGCAAAACCTTGTTTAAATGTTCATCACCATAGAAACATGACAGCTTCTCCATAGAGCT TTTGGAATCCGAGCCCGCATTTTAGAGACTTTGGTCATGCTTCTTCTTCTTGCGTTACTCATTCTTGGGATAGTGTGGGTAGCTTCAGCA CTCATTGACAACGATGCCGCAAGCATGGAATCTTTATATGATCTCTGGGAGTTCTATCTACCCTATTTATATTCCTGTATATCATTGATG GGATGTTTGTTACTTCTCTTGTGTACACCAGTTGGCCTTTCTCGTATGTTCACAGTGATGGGTCAGTTGCTAGTGAAGCCAACAATTCTT GAAGACCTGGATGAACAAATTTATATCATTACCTTAGAGGAAGAAGCACTCCAGAGACGACTAAATGGGCTGTCTTCATCGGTGGAATAC AACATAATGGAGTTGGAACAAGAACTTGAAAATGTAAAGACTCTTAAGACAAAATTAGAGAGGCGAAAAAAGGCTTCAGCATGGGAAAGA AATTTGGTGTATCCCGCTGTTATGGTTCTCCTTCTTATTGAGACATCCATCTCGGTCCTCTTGGTGGCTTGTAATATTCTTTGCCTATTG GTTGATGAAACAGCAATGCCAAAAGGAACAAGGGGGCCTGGAATAGGAAATGCCTCTCTTTCTACGTTTGGTTTTGTGGGAGCTGCGCTT GAAATCATTTTGATTTTCTATCTTATGGTGTCCTCTGTTGTCGGCTTCTATAGCCTTCGATTTTTTGGAAACTTTACTCCCAAGAAAGAT GACACAACTATGACAAAGATCATTGGAAATTGTGTGTCCATCTTGGTTTTGAGCTCTGCTCTGCCTGTGATGTCGAGAACACTGGGTGAC CAAGAATATGGCCCAGGTGACCAAAGCCCTGGACAAGGCCCTGAGCACCATGGACCTGCAGAAGGTCTCCTCAGTGATGGACAGGTTCGA GCAGCAGGTGCAGAACCTGGACGTCCATACATCGGTGATGGAGGACTCCATGAGCTCGGCCACCACCCTGACCACGCCGCAGGAGCAGGT GGACAGCCTCATCATGCAGATCGCCGAGGAGAATGGCCTGGAGGTGCTGGACCAGCTCAGCCAGCTGCCCGAGGGCGCCTCTGCCGTGGG CGAGAGCTCTGTGCGCAGCCAGGAGGACCAGCTGTCACGGAGGTTGGCCGCCTTGAGGAACTAGCCGTGCCCCGCCGGTGTGCACCGCCT CTGCCCCGTGATGTGCTGGAAGGCTCCTGTCCTCTCCCCACCGCGTCTTGCCTTTGTGCTGACCCCGCGGGGCTGCGGCCGGCAGCCACT CTGCGTCTCTCACCTGCCAGGCCTGCGTGGCCTTAGGGTTGTTCCTGTTCTTTTAGGTTGGGCGGTGGGTCTGTGTCCTGGTGTTGAGTT TCTGCAAATTTCTGGGGGTGATTTCTGTGACTCTGGGCCCACAGCGGGGAGGCCAAGAGGGGCCCTGTGGACTTTCACCCAGCACTGTGG GGGCCTTCAGACTCTGGGGCAGCAGACATGCTGCTTCCCATCAGCCAGAGGGGGTCAGGGCTGCCCTGTTGCCAAACAACTCCCTGAGGC CTCTCCGCACCACCTCAGCGGGCAGGAGGTCCCACCATGTGGACAGACATAGCCCAAGGAGGCACCACAGGTCTATGTGTGCTGGGGGAT GTCAGGTGCCACCCAACGCTGTCCTGGTGGTATTTACAATGACATCCTCCTCCTCCATCACTCCAGGGGTGGTGTCTCGGCCGCCCCTAC CAGCTGGCTGAGCCCCCTGGCCTCCTGCGCTCCCTCACTTCCCTCAGTTCCCAAAGCTGCCCAGTCCATGGGGACAGAACCGTCACTCAG ATCCACATTCAAGTGTGCCCACCCTGCAGTCTTCATCCTCACTCAGCTGCTGCCTCTGGAGGTGCCTTTGGCCACATGTGCTGTGCTGTT TGTCTCCTCGACAGGGAGCCTGTCCACCAGCAGGCTGCGGTCCCAGCGGGTGCGTCTGCAGCTCCTCCCCTTGGGCAGCCTGGTTCTCCC GGAGGACCTTTCCTTGGGGCCCTGCTTCATGACGATGCTGCCTGTGTCACCCTCTACCATCTGTAAACAACTGGGTGCCTTCCCCGACCA CACCCCAATGCCTTCCCAGCTTGGAAGCCAAGGCAGCTGATGAAGGGAGCTCAGGAGAGCCGTCTTCAGCTGGGCTGGGGTTGGGGCTGC TGTGAGGAAAACCTGCCATTGTGGCCCTGGAGAGTCACCAGCAGCTCTTGGGAAGGACTTGCTGGGAGGCTGAGAGAGGCTTTGGGCACA GCCTGCTGTCTTTTCCATTTCCTAAAGTTTACTTCATTGTCTTGAGGCTTCCAGGTTTTGTTTTTGTTTTTGCCAAAGTAGAAAAGGCAG GTGGTGGGCGGCTGGCAGGGAGTGCGGGTCCCCGCCCCTCTTCAGTCCTGCCCTCCCCTCCTCAGTCCTGCCCACCCCGTGCAGCCCATG CTGAGGCTGCAGTGGTGTCGTGGGTGTTACGTGCAGGAACGTGGAGACCCTGACGTGGGCTCACTGCGTTTGGTTTTCTTTTCAGAACTT GGGAGCCCCCAGGGAGGGGCTAGTGTTGGTAGGTCCTAGACGTGGTTCCCTCCAGCCTCCCCAAAATCAACCCTGGTGTTGAGAGAACGT CCTTCTGTCCATCGTGGGTAACAGCCTTGGGGAGGGTGCAGAGCTCTGCAGAGCCATGGGCCAGGTGGGGCTGCCTCAGTCCTGTCCCCT TGGGCACTGAGGAGAGGGGCCCATTCACCTTTCTCCTAGAATGCTGTTGTAAATAAACAAATGGATCCCTGGAAACTTT >45733_45733_3_LMBR1-CHMP1A_LMBR1_chr7_156516806_ENST00000359422_CHMP1A_chr16_89713739_ENST00000397901_length(amino acids)=421AA_BP=258 MVMLLLLALLILGIVWVASALIDNDAASMESLYDLWEFYLPYLYSCISLMGCLLLLLCTPVGLSRMFTVMGQLLVKPTILEDLDEQIYII TLEEEALQRRLNGLSSSVEYNIMELEQELENVKTLKTKLERRKKASAWERNLVYPAVMVLLLIETSISVLLVACNILCLLVDETAMPKGT RGPGIGNASLSTFGFVGAALEIILIFYLMVSSVVGFYSLRFFGNFTPKKDDTTMTKIIGNCVSILVLSSALPVMSRTLGDQEYGPGDQSP GQGPEHHGPAEGLLSDGQVRAAGAEPGRPYIGDGGLHELGHHPDHAAGAGGQPHHADRRGEWPGGAGPAQPAARGRLCRGRELCAQPGGP AVTEVGRLEELAVPRRCAPPLPRDVLEGSCPLPTASCLCADPAGLRPAATLRLSPARPAWP -------------------------------------------------------------- >45733_45733_4_LMBR1-CHMP1A_LMBR1_chr7_156516806_ENST00000359422_CHMP1A_chr16_89713739_ENST00000535997_length(transcript)=2927nt_BP=985nt GGTTTGTGGAATCTTGCTTCCCTTTTTTCCAACCTTTGTTTATTTGTATTGATGCCCTTTGCCTTTTTCTTTCTGGAATCAGAAGGCTTT GCTGGCCTGAAAAAGGAGGGCAAGATGTCCAGGCAAAACCTTGTTTAAATGTTCATCACCATAGAAACATGACAGCTTCTCCATAGAGCT TTTGGAATCCGAGCCCGCATTTTAGAGACTTTGGTCATGCTTCTTCTTCTTGCGTTACTCATTCTTGGGATAGTGTGGGTAGCTTCAGCA CTCATTGACAACGATGCCGCAAGCATGGAATCTTTATATGATCTCTGGGAGTTCTATCTACCCTATTTATATTCCTGTATATCATTGATG GGATGTTTGTTACTTCTCTTGTGTACACCAGTTGGCCTTTCTCGTATGTTCACAGTGATGGGTCAGTTGCTAGTGAAGCCAACAATTCTT GAAGACCTGGATGAACAAATTTATATCATTACCTTAGAGGAAGAAGCACTCCAGAGACGACTAAATGGGCTGTCTTCATCGGTGGAATAC AACATAATGGAGTTGGAACAAGAACTTGAAAATGTAAAGACTCTTAAGACAAAATTAGAGAGGCGAAAAAAGGCTTCAGCATGGGAAAGA AATTTGGTGTATCCCGCTGTTATGGTTCTCCTTCTTATTGAGACATCCATCTCGGTCCTCTTGGTGGCTTGTAATATTCTTTGCCTATTG GTTGATGAAACAGCAATGCCAAAAGGAACAAGGGGGCCTGGAATAGGAAATGCCTCTCTTTCTACGTTTGGTTTTGTGGGAGCTGCGCTT GAAATCATTTTGATTTTCTATCTTATGGTGTCCTCTGTTGTCGGCTTCTATAGCCTTCGATTTTTTGGAAACTTTACTCCCAAGAAAGAT GACACAACTATGACAAAGATCATTGGAAATTGTGTGTCCATCTTGGTTTTGAGCTCTGCTCTGCCTGTGATGTCGAGAACACTGGGTGAC CAAGAATATGGCCCAGGTGACCAAAGCCCTGGACAAGGCCCTGAGCACCATGGACCTGCAGAAGGTCTCCTCAGTGATGGACAGGTTCGA GCAGCAGGTGCAGAACCTGGACGTCCATACATCGGTGATGGAGGACTCCATGAGCTCGGCCACCACCCTGACCACGCCGCAGGAGCAGGT GGACAGCCTCATCATGCAGATCGCCGAGGAGAATGGCCTGGAGGTGCTGGACCAGCTCAGCCAGCTGCCCGAGGGCGCCTCTGCCGTGGG CGAGAGCTCTGTGCGCAGCCAGGAGGACCAGCTGTCACGGAGGTTGGCCGCCTTGAGGAACTAGCCGTGCCCCGCCGGTGTGCACCGCCT CTGCCCCGTGATGTGCTGGAAGGCTCCTGTCCTCTCCCCACCGCGTCTTGCCTTTGTGCTGACCCCGCGGGGCTGCGGCCGGCAGCCACT CTGCGTCTCTCACCTGCCAGGCCTGCGTGGCCTTAGGGTTGTTCCTGTTCTTTTAGGTTGGGCGGTGGGTCTGTGTCCTGGTGTTGAGTT TCTGCAAATTTCTGGGGGTGATTTCTGTGACTCTGGGCCCACAGCGGGGAGGCCAAGAGGGGCCCTGTGGACTTTCACCCAGCACTGTGG GGGCCTTCAGACTCTGGGGCAGCAGACATGCTGCTTCCCATCAGCCAGAGGGGGTCAGGGCTGCCCTGTTGCCAAACAACTCCCTGAGGC CTCTCCGCACCACCTCAGCGGGCAGGAGGTCCCACCATGTGGACAGACATAGCCCAAGGAGGCACCACAGGTCTATGTGTGCTGGGGGAT GTCAGGTGCCACCCAACGCTGTCCTGGTGGTATTTACAATGACATCCTCCTCCTCCATCACTCCAGGGGTGGTGTCTCGGCCGCCCCTAC CAGCTGGCTGAGCCCCCTGGCCTCCTGCGCTCCCTCACTTCCCTCAGTTCCCAAAGCTGCCCAGTCCATGGGGACAGAACCGTCACTCAG ATCCACATTCAAGTGTGCCCACCCTGCAGTCTTCATCCTCACTCAGCTGCTGCCTCTGGAGGTGCCTTTGGCCACATGTGCTGTGCTGTT TGTCTCCTCGACAGGGAGCCTGTCCACCAGCAGGCTGCGGTCCCAGCGGGTGCGTCTGCAGCTCCTCCCCTTGGGCAGCCTGGTTCTCCC GGAGGACCTTTCCTTGGGGCCCTGCTTCATGACGATGCTGCCTGTGTCACCCTCTACCATCTGTAAACAACTGGGTGCCTTCCCCGACCA CACCCCAATGCCTTCCCAGCTTGGAAGCCAAGGCAGCTGATGAAGGGAGCTCAGGAGAGCCGTCTTCAGCTGGGCTGGGGTTGGGGCTGC TGTGAGGAAAACCTGCCATTGTGGCCCTGGAGAGTCACCAGCAGCTCTTGGGAAGGACTTGCTGGGAGGCTGAGAGAGGCTTTGGGCACA GCCTGCTGTCTTTTCCATTTCCTAAAGTTTACTTCATTGTCTTGAGGCTTCCAGGTTTTGTTTTTGTTTTTGCCAAAGTAGAAAAGGCAG GTGGTGGGCGGCTGGCAGGGAGTGCGGGTCCCCGCCCCTCTTCAGTCCTGCCCTCCCCTCCTCAGTCCTGCCCACCCCGTGCAGCCCATG CTGAGGCTGCAGTGGTGTCGTGGGTGTTACGTGCAGGAACGTGGAGACCCTGACGTGGGCTCACTGCGTTTGGTTTTCTTTTCAGAACTT GGGAGCCCCCAGGGAGGGGCTAGTGTTGGTAGGTCCTAGACGTGGTTCCCTCCAGCCTCCCCAAAATCAACCCTGGTGTTGAGAGAACGT CCTTCTGTCCATCGTGGGTAACAGCCTTGGGGAGGGTGCAGAGCTCTGCAGAGCCATGGGCCAGGTGGGGCTGCCTCAGTCCTGTCCCCT TGGGCACTGAGGAGAGGGGCCCATTCACCTTTCTCCTAGAATGCTGT >45733_45733_4_LMBR1-CHMP1A_LMBR1_chr7_156516806_ENST00000359422_CHMP1A_chr16_89713739_ENST00000535997_length(amino acids)=421AA_BP=258 MVMLLLLALLILGIVWVASALIDNDAASMESLYDLWEFYLPYLYSCISLMGCLLLLLCTPVGLSRMFTVMGQLLVKPTILEDLDEQIYII TLEEEALQRRLNGLSSSVEYNIMELEQELENVKTLKTKLERRKKASAWERNLVYPAVMVLLLIETSISVLLVACNILCLLVDETAMPKGT RGPGIGNASLSTFGFVGAALEIILIFYLMVSSVVGFYSLRFFGNFTPKKDDTTMTKIIGNCVSILVLSSALPVMSRTLGDQEYGPGDQSP GQGPEHHGPAEGLLSDGQVRAAGAEPGRPYIGDGGLHELGHHPDHAAGAGGQPHHADRRGEWPGGAGPAQPAARGRLCRGRELCAQPGGP AVTEVGRLEELAVPRRCAPPLPRDVLEGSCPLPTASCLCADPAGLRPAATLRLSPARPAWP -------------------------------------------------------------- >45733_45733_5_LMBR1-CHMP1A_LMBR1_chr7_156516806_ENST00000359422_CHMP1A_chr16_89713739_ENST00000550102_length(transcript)=1501nt_BP=985nt GGTTTGTGGAATCTTGCTTCCCTTTTTTCCAACCTTTGTTTATTTGTATTGATGCCCTTTGCCTTTTTCTTTCTGGAATCAGAAGGCTTT GCTGGCCTGAAAAAGGAGGGCAAGATGTCCAGGCAAAACCTTGTTTAAATGTTCATCACCATAGAAACATGACAGCTTCTCCATAGAGCT TTTGGAATCCGAGCCCGCATTTTAGAGACTTTGGTCATGCTTCTTCTTCTTGCGTTACTCATTCTTGGGATAGTGTGGGTAGCTTCAGCA CTCATTGACAACGATGCCGCAAGCATGGAATCTTTATATGATCTCTGGGAGTTCTATCTACCCTATTTATATTCCTGTATATCATTGATG GGATGTTTGTTACTTCTCTTGTGTACACCAGTTGGCCTTTCTCGTATGTTCACAGTGATGGGTCAGTTGCTAGTGAAGCCAACAATTCTT GAAGACCTGGATGAACAAATTTATATCATTACCTTAGAGGAAGAAGCACTCCAGAGACGACTAAATGGGCTGTCTTCATCGGTGGAATAC AACATAATGGAGTTGGAACAAGAACTTGAAAATGTAAAGACTCTTAAGACAAAATTAGAGAGGCGAAAAAAGGCTTCAGCATGGGAAAGA AATTTGGTGTATCCCGCTGTTATGGTTCTCCTTCTTATTGAGACATCCATCTCGGTCCTCTTGGTGGCTTGTAATATTCTTTGCCTATTG GTTGATGAAACAGCAATGCCAAAAGGAACAAGGGGGCCTGGAATAGGAAATGCCTCTCTTTCTACGTTTGGTTTTGTGGGAGCTGCGCTT GAAATCATTTTGATTTTCTATCTTATGGTGTCCTCTGTTGTCGGCTTCTATAGCCTTCGATTTTTTGGAAACTTTACTCCCAAGAAAGAT GACACAACTATGACAAAGATCATTGGAAATTGTGTGTCCATCTTGGTTTTGAGCTCTGCTCTGCCTGTGATGTCGAGAACACTGGGTGAC CAAGAATATGGCCCAGGTGACCAAAGCCCTGGACAAGGCCCTGAGCACCATGGACCTGCAGAAGGTCTCCTCAGTGATGGACAGGTTCGA GCAGCAGGTGCAGAACCTGGACGTCCATACATCGGTGATGGAGGACTCCATGAGCTCGGCCACCACCCTGACCACGCCGCAGGAGCAGGT GGACAGCCTCATCATGCAGATCGCCGAGGAGAATGGCCTGGAGGTGCTGGACCAGCTCAGCCAGCTGCCCGAGGGCGCCTCTGCCGTGGG CGAGAGCTCTGTGCGCAGCCAGGAGGACCAGCTGTCACGGAGGTTGGCCGCCTTGAGGAACTAGCCGTGCCCCGCCGGTGTGCACCGCCT CTGCCCCGTGATGTGCTGGAAGGCTCCTGTCCTCTCCCCACCGCGTCTTGCCTTTGTGCTGACCCCGCGGGGCTGCGGCCGGCAGCCACT CTGCGTCTCTCACCTGCCAGGCCTGCGTGGCCTTAGGGTTGTTCCTGTTCTTTTAGGTTGG >45733_45733_5_LMBR1-CHMP1A_LMBR1_chr7_156516806_ENST00000359422_CHMP1A_chr16_89713739_ENST00000550102_length(amino acids)=421AA_BP=258 MVMLLLLALLILGIVWVASALIDNDAASMESLYDLWEFYLPYLYSCISLMGCLLLLLCTPVGLSRMFTVMGQLLVKPTILEDLDEQIYII TLEEEALQRRLNGLSSSVEYNIMELEQELENVKTLKTKLERRKKASAWERNLVYPAVMVLLLIETSISVLLVACNILCLLVDETAMPKGT RGPGIGNASLSTFGFVGAALEIILIFYLMVSSVVGFYSLRFFGNFTPKKDDTTMTKIIGNCVSILVLSSALPVMSRTLGDQEYGPGDQSP GQGPEHHGPAEGLLSDGQVRAAGAEPGRPYIGDGGLHELGHHPDHAAGAGGQPHHADRRGEWPGGAGPAQPAARGRLCRGRELCAQPGGP AVTEVGRLEELAVPRRCAPPLPRDVLEGSCPLPTASCLCADPAGLRPAATLRLSPARPAWP -------------------------------------------------------------- >45733_45733_6_LMBR1-CHMP1A_LMBR1_chr7_156516806_ENST00000540390_CHMP1A_chr16_89713739_ENST00000253475_length(transcript)=3305nt_BP=1338nt AGAGCTGCTCGTCTGAGGCTGCTGAGGCGACGGCCGGTGTCGTGGTCGCGGTACCTGTTCCAACACGGCTCGCGGGCCCGTGCCGGCTCC GGTCCCCGGCGCGGCTGTCCGAGCCCCTGCGGCGGGCGGACGATGGTGTGGCGGAGCACGCGGACGCGGGCGGCGCGGCGGCGGGCATGA AGGAGGATGGAAGGGCAGGACGAGGTGTCGGCGCGGGAGCAGCACTTCCACAGCCAAGTGCGGGAGTCCACGATGAACAAGAAGATGAAG ATGCCATCGTCAACAGGATTTCGTTGTTTTTGAGCACGTTCACTCTCGCAGTGTCAGCTGGGGCTGTTTTGCTTTTACCCTTCTCAATCA TCAGCAATGAAATCCTGCTTTCTTTTCCTCAGAACTACTATATTCAGTGGCTAAATGGCTCCCTGATTCATGGTTTGTGGAATCTTGCTT CCCTTTTTTCCAACCTTTGTTTATTTGTATTGATGCCCTTTGCCTTTTTCTTTCTGGAATCAGAAGGCTTTGCTGGCCTGAAAAAGGGAA TCCGAGCCCGCATTTTAGAGACTTTGGTCATGCTTCTTCTTCTTGCGTTACTCATTCTTGGGATAGTGTGGGTAGCTTCAGCACTCATTG ACAACGATGCCGCAAGCATGGAATCTTTATATGATCTCTGGGAGTTCTATCTACCCTATTTATATTCCTGTATATCATTGATGGGATGTT TGTTACTTCTCTTGTGTACACCAGTTGGCCTTTCTCGTATGTTCACAGTGATGGGTCAGTTGCTAGTGAAGCCAACAATTCTTGAAGACC TGGATGAACAAATTTATATCATTACCTTAGAGGAAGAAGCACTCCAGAGACGACTAAATGGGCTGTCTTCATCGGTGGAATACAACATAA TGGAGTTGGAACAAGAACTTGAAAATGTAAAGACTCTTAAGACAAAATTAGAGAGGCGAAAAAAGGCTTCAGCATGGGAAAGAAATTTGG TGTATCCCGCTGTTATGGTTCTCCTTCTTATTGAGACATCCATCTCGGTCCTCTTGGTGGCTTGTAATATTCTTTGCCTATTGGTTGATG AAACAGCAATGCCAAAAGGAACAAGGGGGCCTGGAATAGGAAATGCCTCTCTTTCTACGTTTGGTTTTGTGGGAGCTGCGCTTGAAATCA TTTTGATTTTCTATCTTATGGTGTCCTCTGTTGTCGGCTTCTATAGCCTTCGATTTTTTGGAAACTTTACTCCCAAGAAAGATGACACAA CTATGACAAAGATCATTGGAAATTGTGTGTCCATCTTGGTTTTGAGCTCTGCTCTGCCTGTGATGTCGAGAACACTGGGTGACCAAGAAT ATGGCCCAGGTGACCAAAGCCCTGGACAAGGCCCTGAGCACCATGGACCTGCAGAAGGTCTCCTCAGTGATGGACAGGTTCGAGCAGCAG GTGCAGAACCTGGACGTCCATACATCGGTGATGGAGGACTCCATGAGCTCGGCCACCACCCTGACCACGCCGCAGGAGCAGGTGGACAGC CTCATCATGCAGATCGCCGAGGAGAATGGCCTGGAGGTGCTGGACCAGCTCAGCCAGCTGCCCGAGGGCGCCTCTGCCGTGGGCGAGAGC TCTGTGCGCAGCCAGGAGGACCAGCTGTCACGGAGGTTGGCCGCCTTGAGGAACTAGCCGTGCCCCGCCGGTGTGCACCGCCTCTGCCCC GTGATGTGCTGGAAGGCTCCTGTCCTCTCCCCACCGCGTCTTGCCTTTGTGCTGACCCCGCGGGGCTGCGGCCGGCAGCCACTCTGCGTC TCTCACCTGCCAGGCCTGCGTGGCCTTAGGGTTGTTCCTGTTCTTTTAGGTTGGGCGGTGGGTCTGTGTCCTGGTGTTGAGTTTCTGCAA ATTTCTGGGGGTGATTTCTGTGACTCTGGGCCCACAGCGGGGAGGCCAAGAGGGGCCCTGTGGACTTTCACCCAGCACTGTGGGGGCCTT CAGACTCTGGGGCAGCAGACATGCTGCTTCCCATCAGCCAGAGGGGGTCAGGGCTGCCCTGTTGCCAAACAACTCCCTGAGGCCTCTCCG CACCACCTCAGCGGGCAGGAGGTCCCACCATGTGGACAGACATAGCCCAAGGAGGCACCACAGGTCTATGTGTGCTGGGGGATGTCAGGT GCCACCCAACGCTGTCCTGGTGGTATTTACAATGACATCCTCCTCCTCCATCACTCCAGGGGTGGTGTCTCGGCCGCCCCTACCAGCTGG CTGAGCCCCCTGGCCTCCTGCGCTCCCTCACTTCCCTCAGTTCCCAAAGCTGCCCAGTCCATGGGGACAGAACCGTCACTCAGATCCACA TTCAAGTGTGCCCACCCTGCAGTCTTCATCCTCACTCAGCTGCTGCCTCTGGAGGTGCCTTTGGCCACATGTGCTGTGCTGTTTGTCTCC TCGACAGGGAGCCTGTCCACCAGCAGGCTGCGGTCCCAGCGGGTGCGTCTGCAGCTCCTCCCCTTGGGCAGCCTGGTTCTCCCGGAGGAC CTTTCCTTGGGGCCCTGCTTCATGACGATGCTGCCTGTGTCACCCTCTACCATCTGTAAACAACTGGGTGCCTTCCCCGACCACACCCCA ATGCCTTCCCAGCTTGGAAGCCAAGGCAGCTGATGAAGGGAGCTCAGGAGAGCCGTCTTCAGCTGGGCTGGGGTTGGGGCTGCTGTGAGG AAAACCTGCCATTGTGGCCCTGGAGAGTCACCAGCAGCTCTTGGGAAGGACTTGCTGGGAGGCTGAGAGAGGCTTTGGGCACAGCCTGCT GTCTTTTCCATTTCCTAAAGTTTACTTCATTGTCTTGAGGCTTCCAGGTTTTGTTTTTGTTTTTGCCAAAGTAGAAAAGGCAGGTGGTGG GCGGCTGGCAGGGAGTGCGGGTCCCCGCCCCTCTTCAGTCCTGCCCTCCCCTCCTCAGTCCTGCCCACCCCGTGCAGCCCATGCTGAGGC TGCAGTGGTGTCGTGGGTGTTACGTGCAGGAACGTGGAGACCCTGACGTGGGCTCACTGCGTTTGGTTTTCTTTTCAGAACTTGGGAGCC CCCAGGGAGGGGCTAGTGTTGGTAGGTCCTAGACGTGGTTCCCTCCAGCCTCCCCAAAATCAACCCTGGTGTTGAGAGAACGTCCTTCTG TCCATCGTGGGTAACAGCCTTGGGGAGGGTGCAGAGCTCTGCAGAGCCATGGGCCAGGTGGGGCTGCCTCAGTCCTGTCCCCTTGGGCAC TGAGGAGAGGGGCCCATTCACCTTTCTCCTAGAATGCTGTTGTAAATAAACAAATGGATCCCTGG >45733_45733_6_LMBR1-CHMP1A_LMBR1_chr7_156516806_ENST00000540390_CHMP1A_chr16_89713739_ENST00000253475_length(amino acids)=574AA_BP=411 MSEPLRRADDGVAEHADAGGAAAGMKEDGRAGRGVGAGAALPQPSAGVHDEQEDEDAIVNRISLFLSTFTLAVSAGAVLLLPFSIISNEI LLSFPQNYYIQWLNGSLIHGLWNLASLFSNLCLFVLMPFAFFFLESEGFAGLKKGIRARILETLVMLLLLALLILGIVWVASALIDNDAA SMESLYDLWEFYLPYLYSCISLMGCLLLLLCTPVGLSRMFTVMGQLLVKPTILEDLDEQIYIITLEEEALQRRLNGLSSSVEYNIMELEQ ELENVKTLKTKLERRKKASAWERNLVYPAVMVLLLIETSISVLLVACNILCLLVDETAMPKGTRGPGIGNASLSTFGFVGAALEIILIFY LMVSSVVGFYSLRFFGNFTPKKDDTTMTKIIGNCVSILVLSSALPVMSRTLGDQEYGPGDQSPGQGPEHHGPAEGLLSDGQVRAAGAEPG RPYIGDGGLHELGHHPDHAAGAGGQPHHADRRGEWPGGAGPAQPAARGRLCRGRELCAQPGGPAVTEVGRLEELAVPRRCAPPLPRDVLE GSCPLPTASCLCADPAGLRPAATLRLSPARPAWP -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for LMBR1-CHMP1A |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for LMBR1-CHMP1A |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for LMBR1-CHMP1A |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
| Hgene | LMBR1 | C1868114 | POLYDACTYLY, PREAXIAL II (disorder) | 4 | CTD_human;GENOMICS_ENGLAND;ORPHANET |
| Hgene | LMBR1 | C1851100 | LAURIN-SANDROW SYNDROME | 2 | CTD_human;GENOMICS_ENGLAND;ORPHANET |
| Hgene | LMBR1 | C1861355 | Syndactyly, Type IV | 2 | CTD_human;GENOMICS_ENGLAND;ORPHANET |
| Hgene | LMBR1 | C0265559 | Acheiropodia | 1 | CTD_human;GENOMICS_ENGLAND;ORPHANET |
| Hgene | LMBR1 | C0265581 | Longitudinal deficiency of radius | 1 | ORPHANET |
| Hgene | LMBR1 | C1861098 | Tibia, Hypoplasia of, with Polydactyly | 1 | GENOMICS_ENGLAND |
| Tgene | C0266468 | Congenital pontocerebellar hypoplasia | 1 | GENOMICS_ENGLAND | |
| Tgene | C3554209 | Congenital pontocerebellar hypoplasia type 8 | 1 | CTD_human;GENOMICS_ENGLAND;ORPHANET |
